Abstract
N2O is an important greenhouse gas influencing global warming, and agricultural land is the predominant (anthropogenic) source of N2O emissions. Here, we report the high N2O-reducing activity of Bradyrhizobium ottawaense, suggesting the potential for efficiently mitigating N2O emission from agricultural lands. Among the 15 B. ottawaense isolates examined, the N2O-reducing activities of most (13) strains were approximately five-fold higher than that of Bradyrhizobium diazoefficiens USDA110T under anaerobic conditions. This robust N2O-reducing activity of B. ottawaense was confirmed by N2O reductase (NosZ) protein levels and by mitigation of N2O emitted by nodule decomposition in laboratory system. While the NosZ of B. ottawaense and B. diazoefficiens showed high homology, nosZ gene expression in B. ottawaense was over 150-fold higher than that in B. diazoefficiens USDA110T, suggesting the high N2O-reducing activity of B. ottawaense is achieved by high nos expression. Furthermore, we examined the nos operon transcription start sites and found that, unlike B. diazoefficiens, B. ottawaense has two transcription start sites under N2O-respiring conditions, which may contribute to the high nosZ expression. Our study indicates the potential of B. ottawaense for effective N2O reduction and unique regulation of nos gene expression towards the high performance of N2O mitigation in the soil.
Similar content being viewed by others
Introduction
The expansion of human activities is triggering irreversible environmental damage, including global warming and stratospheric ozone depletion. N2O is a long-lived greenhouse gas (GHG) whose atmospheric lifetime is an estimated 116 ± 9 years1. Moreover, N2O has a stratospheric ozone-depleting effect. Although N2O concentration in the atmosphere is still low compared with other GHG such as CO2 and CH4, N2O is an alarming GHG due to its high global warming potential per unit2. Agricultural land is the primary source of N2O, accounting for 52% of anthropogenic origin emissions3. N2O is markedly emitted from nitrogen-rich environments, such as agricultural fields in which excess N fertilizers are applied and crop residues, including nodulated legume roots4,5. Biochemically, microbial nitrification and denitrification are the two major processes of N2O generation6,7. During nitrification, N2O is produced as a byproduct when ammonia is oxidized to nitrite via hydroxylamine. N2O is also generated from NO during incomplete denitrification, which intricately involves diverse soil bacteria, fungi, and archaea8. However, to date, only one microbial enzyme, N2O reductase (encoded by the nosZ gene), reportedly reduces N2O to N27.
Since some rhizobial species possess the nosZ gene, strategies to reduce N2O emissions from agricultural fields using rhizobia have been studied. In particular, soybeans are grown globally, and the amount of N2O emitted from nodulated soybeans is higher than that from corn or wheat. For example, N2O emissions from soybean fields in Argentina are estimated to reach 5.1 kg N ha−1 year−19. The use of rhizobia is, therefore, an effective approach to reducing global GHG emissions. Bradyrhizobium nodulates various legumes, including soybean, and has been studied as a model denitrification microorganism. Soybean roots nodulated with Bradyrhizobium diazoefficiens USDA110T scavenges exogenous N2O, even in ambient air containing a low concentration of N2O (0.34 ppm)10. Moreover, N2O fluxes from soybean fields have been mitigated by inoculation with B. diazoefficiens mutants with high N2O reductase activity (nos++ mutants)11. The utility of B. diazoefficiens in N2O mitigation has also been verified in soybean ecosystems in Japan12, France13, and South America14.
On the other hand, rhizobial strains carrying nos genes are uncommon; nos genes and N2O-reducing activity have been observed only in B. diazoefficiens, soybean rhizobia10, and Ensifer meliloti, an alfalfa endosymbiont15. Several soybean rhizobia species, including B. diazoefficiens, B. japonicum, B. elkanii, and Ensifer fredii, have been identified, but most soybean rhizobia in Japan and the world are non-nos-possessing (nos−) species16. However, nos gene clusters have been recently found in Bradyrhizobium ottawaense17 and Rhizobium leguminosarum13, suggesting that the strategy for mitigating N2O emissions from the legume rhizosphere using rhizobia could be expanded to various legume and rhizobial species.
Soybean rhizosphere is a hotspot for complicated nitrogen transformations including production and reduction of N2O. Nodule decomposition is a major source of N2O through ammonification, nitrification and denitrification in soil organisms and rhizobia18. N2O is only emitted by decomposed nodules at the late growth period of soybean growth, but not by fresh nodules or roots5,11,19,20,21. N2O formed by nodule decomposition is either emitted into the atmosphere or is further reduced to N2 by N2O reductase of soybean bradyrhizobia possessing nosZ gene (nos+ strains)10,21,22. Due to the balance between the production and reduction of N2O during nodule decomposition, N2O emission occurs in root systems with nodules harboring nos– and even nos+ strains. Therefore, to effectively prevent N2O release, using rhizobia with high N2O-reducing activity is necessary.
Denitrification reactions involving N2O reduction occur under anaerobic conditions. In bradyrhizobia, N2O reductase (nos) genes are regulated by three different two-component regulatory systems18. The FixLJK2 cascade is the primary oxygen-sensing regulator for nos operons. Under moderate low oxygen concentration conditions (< 5%), FixLJK2 recognizes the FixK box [TTG(A/C)-N6-(T/G)CAA] located upstream of nosR and promotes nos operon expression23,24. It has also been shown that the NasST two-component regulatory system, which senses NO3- concentrations and regulates the NO3- assimilation gene (nas) operon, is also responsible for regulating the nos operon25. NasT act as activators of the nas/nos operons and NasS acts on NasT, inhibiting its function: in the absence of NO3-/NO2-, NasS and NasT bind to each other, and transcription is arrested by the terminator structure upstream of the nas/nos operon. On the other hand, in the presence of NO3-/NO2-, NasT is released from NasS and binds to the mRNA upstream of the nos operon, resulting in a conformational change in the hairpin termination structure of the mRNA and read-through transcription of the nos genes18. In nasS deletion mutants, transcription of the nos operon is activated independently of NO3-. Itakura et al.11 developed nos++ strains from naturally occurring nasS mutants and verified their utility in N2O reduction in laboratory and field experiments. Additionally, the RegSR two-component regulatory system presumably controls nosR expression via the NifA protein26.
The catalytic unit of N2O reductase requires auxiliary proteins which are encoded in nos gene cluster (nosRZDFYLX)27. The flavoproteins NosR and NosX form an electron transport pathway from the quinone pool to NosZ. NosR is also required for the transcription of nos genes. NosD, NosF, NosY, and NosL are involved in maturation of the CuZ site of NosZ27,28.
In this study, we characterized nos-possessing B. ottawaense strains isolated from sorghum roots based on their genome sequence and activity. B. ottawaense has been reported to form effective nitrogen fixing nodules on soybeans29 and nitrogen fixation activity was comparable to that of B. diazoefficiens at the plant level30. Most B. ottawaense strains analyzed in this study showed significantly higher N2O-reducing activity than that of B. diazoefficiens USDA110T. Gene expression and promoter analyses showed that B. ottawaense strongly expressed the nosZ gene under both N2O- and NO3--reducing conditions, and its high-level expression is thought to be achieved by different nos operon transcription start sites and not by already known regulation systems. Our study indicates the potential of B. ottawaense in N2O mitigation and the unique regulation of nos gene expression that contributes to the high performance of N2O reduction.
Results
N 2 O- reducing activity in B. ottawaense
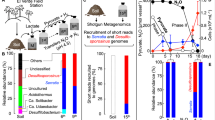
The B. ottawaense strains used in this study are listed in Supplementary Table 1. Among them, the phylogenetic relationships and gene conservation of the denitrification pathway of four strains (SG09, TM102, TM233, and TM239) have been reported17. To confirm species classification and gene organization, we determined the draft genome sequence of 10 strains, including 3 strains reported by Wasai-Hara et al.17 (see Supplementary Table 1). All isolates showed more than 95.0% average nucleotide identity (ANI) values with the type strain B. ottawaense OO99T31, indicating that the isolates were classified into B. ottawaense (see Supplementary Table 2). Furthermore, phylogenetic analysis based on multiple housekeeping genes (AMPHORA32) supported this classification, as shown in Supplementary Fig. 1.
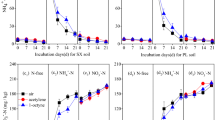
The N2O-reducing activity of the B. ottawaense strains was determined under free-living, N2O-respiring conditions (Fig. 1a, see also Supplementary Fig. 2). Almost all isolates and the type strain B. ottawaense OO99T showed activity in the range of 11.5–15.8 nmol h−1 protein−1, which was 5.5–7.4-fold higher than that of B. diazoefficiens USDA110T (2.0 nmol h−1 protein−1). Also, growth under N2O-respiring conditions was better in B. ottawaense strains than in B. diazoefficiens USDA110T (Supplementary Fig. 3). Conversely, two strains (SF12 and SF19) showed relatively low activity, with values of 2.4 and 3.4 nmol h−1 protein−1, respectively, comparable to that of B. diazoefficiens USDA110T (Fig. 1a). We also analyzed the N2O-reducing activity of TM102, TM233, and TM239, which lack nodulation and nitrogen-fixing ability on soybeans17, but no significant difference was observed from the other nodulating strains of B. ottawaense (p < 0.05, Tukey’s test). Monitoring N2O concentrations over time showed a rapid decrease in B. ottawaense (SG09, OO99T, TM102, and TM233), while B. diazoefficiens USDA110T exhibited a slow decrease (Fig. 1b).
N2O-reducing activity of B. ottawaense. (a) N2O-reducing activity of B. ottawaense isolates, type strain OO99T, B. diazoefficiens stain USDA110T, and the nosZ-deficient strain (ΔnosZ). Different letters above the bars represent significant differences between inoculation treatments analyzed using Tukey’s test after analysis of variance (ANOVA; p < 0.05). Parentheses after the strain name indicate nodule-forming ability. R = nodule forming strain (rhizobia), N = non-nodulation and non-diazotroph. (b) N2O-reducing activity in representative strains of B. ottawaense and B. diazoefficiens. The graph shows the changes in N2O concentration over time in the gas phase in the test tube.
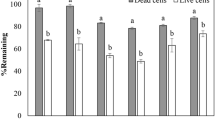
Next, we examined the effects of B. ottawaense inoculation on the N2O flux associated with nodule degradation in a soybean rhizosphere in a laboratory system (Fig. 2, Supplementary Fig. 4). After soybean seeds (cv Enrei) were inoculated with B. japonicum USDA6T (nos-), B. diazoefficiens USDA110T (nos+) and B. ottawaense SG09 (nos+), the root system of 36 days-old soybean plants (the nodule fresh weights were similar (1.9–2.1 g fresh weight plant-1) irrespective to USDA6T, USDA110T and SG09 inoculation treatments) were decapitated and transferred into open bottles containing a field soil. After 20 days incubation, we determined N2O flux from the nodulating root-soil system containing decomposing nodules in closed bottles under an aerobic condition, which simulates the rhizosphere of field-grown soybeans. Under atmospheric conditions (approximately 340 ppb of N2O in air), N2O flux from soybean rhizospheres inoculated with B. japonicum USDA 6T, B. diazoefficiens USDA110T and B. ottawaense SG09 was 29.2, 7.1 and 2.3 nmol h−1 plant−1, respectively (Fig. 2a). N2O flux following SG09 inoculation significantly decreased relative to that after B. japonicum USDA 6T (nos−) and B. diazoefficiens USDA110T (nos+) inoculation. N2O flux of USDA110T inoculation also significantly decreased relative to that B. japonicum USDA 6T (nos−) that is similar to the previous results11. In N2O-supplemented air (50 ppm N2O), B. ottawaense SG09 inoculation exclusively showed negative N2O flux. However, such negative flux was not observed with B. diazoefficiens USDA 110T and USDA 6T inoculation (Fig. 2b). These results demonstrate the effectiveness of B. ottawaense inoculation in reducing N2O emissions from the soybean rhizosphere.
N2O flux in rhizosphere inoculated with either Bradyrhizobium ottawaense, B. diazoefficiens and B. japonicum. N2O flux from the rhizosphere of soybean plants inoculated with B. ottawaense SG09 (nos +), B. diazoefficiens USDA110T (nos+), and B. japonicum USDA 6T (nos−) under (a) an atmospheric concentration (approximately 340 ppb) of N2O and (b) N2O-supplemented air (50 ppm). Asterisks represent significant differences at p < 0.05 by the t-test.
nosZ gene expression and protein activity of wild-type B. ottawaense strains
nosZ expression in B. ottawaense strains was evaluated by RT-qPCR under both N2O- and NO3--respiring conditions based on B. diazoefficiens USDA110T (Table 1). Under N2O-respiring conditions, wild-type (WT) B. ottawaense SG09 and OO99T strains showed 211.5- and 163.5-fold higher expression levels than WT B. diazoefficiens USDA110T, respectively. Under NO3--respiring conditions, the nosZ expression of WT strains was upregulated in both B. ottawaense and B. diazoefficiens being 19.3-fold (USDA110T), 8.9-fold (SG09), and 12.6-fold (OO99T) higher than that under N2O-respiring conditions. In the comparison among strains, B. ottawaense SG09 and OO99T showed 109- and 119-fold higher nosZ expression than that of USDA110T respectively, even under NO3--respiring conditions. On the other hand, the two strains with low N2O-reducing activity (SF12 and SF19; Fig. 1a) showed low expression levels that were 31.2- and 40.6-fold higher than those of USDA110T, respectively, and less than 1/5 those of SG09 under N2O-respiring conditions.
We next analyzed NosZ protein activity in B. ottawaense and B. diazoefficiens by specific activity staining with methyl viologen after sodium deoxycholate polyacrylamide gel electrophoresis (DOC-PAGE). Equal amounts of B. ottawaense and B. diazoefficiens total protein were loaded and confirmed by Coomassie brilliant blue (CBB) staining (Fig. 3a, see also Supplementary Fig. 5). On the same gel, the intensity of activity staining was clearly higher for SG09 than that for USDA110T; the intensity of the eightfold diluted SG09 lane was comparable to that of the non-diluted USDA110T lane, indicating that NosZ protein activity in B. ottawaense was approximately eightfold higher than that in B. diazoefficiens (Fig. 3b, c).
Activity of the NosZ protein of Bradyrhizobium ottawaense and B. diazoefficiens. Coomassie brilliant blue staining (a) and NosZ-specific activity staining (DOC-PAGE, b) protein extracted from B. ottawaense SG09 and B. diazoefficiens USDA110T. The numbers in each lane indicate the concentration (x) rate of extracted protein samples. ‘M’ indicates the protein size marker (60, 120, and 240 kDa were indicated). The gel images are cropped; full images are shown in Supplemental Fig. 5. Panel (c) shows the signal intensity of the NosZ-specific activity staining.
nosZ expression in nasS deletion mutants
To investigate the high nosZ expression in B. ottawaense, we compared the sequences of the genes involved in the expression of the nos operon, nasST, fixLJK2, and regSR, in B. ottawaense SG09 and B. diazoefficiens USDA110T. As shown in Supplementary Table 3, all genes showed > 90% identity in amino acid sequence. However, the upstream sequence of the nos operon, which is recognized by NasT to suppress transcription, showed only 48% identity in the nucleotide sequence. Therefore, we examined whether the NasST regulatory system is also functional in B. ottawaense, similar to B. diazoefficiens USDA110T25. To this end, we analyzed nosZ gene expression in nasS deletion mutants of B. ottawaense SG09 and OO99T (Table 2). In B. diazoefficiens USDA110T, nosZ expression was significantly increased (3.4-fold) in the ΔnasS mutant. Similarly, nosZ expression was significantly increased in the B. ottawaense SG09 (2.03-fold) and OO99T (2.06-fold) ΔnasS mutants under N2O-respiring conditions. This significant expression increase in nasS deletion mutants was not observed in the presence of NO3-. The effect of nasS deletion was also observed in N2O-reducing activity, although the N2O-reducing activities of ΔnasS mutants of B. ottawaense SG09 and OO99T were not significantly higher than those of their parent strains SG09 and OO99T, respectively (See Supplementary Fig. 6).
Nos operon transcription system in B. ottawaense
To investigate the effect of the transcriptional regulation of the nos operon on nosZ expression, we determined the transcriptional start site of nosR, which is located upstream of the operon (see Supplementary Fig. 7). The transcription start site was investigated under anaerobic conditions with NO3- or N2O as the sole electron acceptor by 5′ rapid amplification of cDNA ends (5′ RACE).
Under N2O-respiring conditions, two start sites were detected in B. ottawaense SG09 at C and G, 212 and 79 nucleotides (nt) upstream of the nosR start codon, respectively (Pd1 and Pd2, respectively; Fig. 4, Supplementary Fig. 8). The -35/-10 consensus sequence was predicted upstream of the two transcription start sites. In addition, the putative FixK box was predicted upstream of Pd1. On the other hand, under NO3--respiring conditions, a single transcription start site, Pd1, was observed in B. ottawaense SG09 (Fig. 4 and Supplementary Fig. 8). We also confirmed that B. ottawaense OO99T has identical promoter sequences and two transcription start sites (Pd1 and Pd2; Fig. 4 and Supplementary Fig. 8). In B. ottawaense OO99T, two transcription start points (Pd1 and Pd2) were detected under NO3--respiring conditions, but comparing the band intensities, it was clear that Pd1, which was also detected in SG09, was strongly transcribed. We additionally examined the transcription start site in USDA110T under N2O-respiring conditions and detected a single site (G, 84 nucleotides upstream of nosR) that was identical to the previously reported site of USDA110T under NO3--respiring conditions24,33 (see Supplementary Fig. 8). Our results indicate that, unlike B. diazoefficiens USDA110T, the nosR of B. ottawaense has two transcription start sites under N2O-respiring conditions.
Transcriptional organization of nosR in Bradyrhizobium ottawaense SG09 and OO99T. Pd1(c) is the transcription start site under N2O- and NO3--respiring conditions. Pd2(g) is the transcription start site under N2O-respiring conditions. -35/-10 consensus sequences preceding each transcription start site are indicated by underlining. Putative FixK box located upstream of Pd1 is shown in the box. The translational start codon (ATG) of nosR is shown in bold case. The promoter sequences of B. ottawaense SG09 and OO99T are completely identical. The dotted underlined region indicates the 56 bp deleted in strains SF12 and SF19 (see Supplementary Fig. 7 for details).
Two B. ottawaense strains with low N2O-reducing activity
Among the B. ottawaense analyzed in this study, two strains SF12 and SF19 showed low N2O-reducing activity and nosZ expression (Fig. 1a, Table 1). When compared nos gene clusters (see Supplementary Fig. 7a, b), we could not identify the differences responsible for the N2O-reducing activity as their nos genes are identical in amino acid sequence. However, when compared the genomic sequence of the upstream region of nosR, 56 bp deletion was commonly observed in the low N2O-reducing activity strains SF12 and SF19; the deleted region includes the start codon (ATG) of nosR (Fig. 4 and Supplementary Fig. 7c). To confirm that 56 bp deletion is the cause of low activity, the deletion mutants of SG09 and OO99T(SG09Δ56, OO99TΔ56) were generated. In the 56 bp deletion mutants, both N2O-reducing activity and nosZ expression levels were decreased to levels comparable to those of SF12 and SF19 (Table 3), confirming that 56 bp deletion is the cause of the low activity of SF12 and SF19.
Discussion
In this study, we demonstrated that B. ottawaense has higher N2O-reducing activity than that of B. diazoefficiens. In a previous study, B. diazoefficiens mutants with high N2O-reducing activity (nos++ mutants) were generated, and the mutants mitigated N2O emission at the laboratory and field levels11,34. One of the nos++ mutants (5M09) was established as a non-genetically modified organism; however, this strain has 66 mutations in the genome, raising concerns for actual agricultural use. The B. ottawaense described in this study is a WT strain that exhibits high N2O reduction activity comparable to that of the artificially generated USDA110T nos++ mutant strains; both the nos++ mutants (5M09) and B. ottawaense exhibited approximately fivefold higher N2O reduction activity than the WT USDA110T (Fig. 1 and25). Furthermore, we demonstrated that SG09 inoculation resulted in almost no N2O release in the rhizosphere. Notably, negative N2O flux was observed under a 50 ppm N2O gas phase in the laboratory experiment, suggesting a system is in place to reduce high N2O concentrations (Fig. 2). Given that GHG reduction is a current key issue, B. ottawaense is quite beneficial as it can contribute to the mitigation of N2O in agricultural fields.
High N2O-reducing ability is considered adaptive in environments with high N2O concentrations. B. ottawaense was first isolated from a soybean field in Canada in 2012 as a novel species29,35. Other B. ottawaense isolates have been reported from soybean and peanut fields in China and Japan and woody legumes in Ethiopia36,37,38,39. B. ottawaense can form nodules in soybeans, but it is rarely detected in soybean fields in Japan37, suggesting that it is adapted to different environments than those of conventional soybean rhizobacteria such as B. diazoefficiens, B. japonicum, and B. elkanii. N2O reduction occurs preferentially over NO3- reduction40, and B. ottawaense can grow better than B. diazoefficiens under N2O-respiring conditions (see Supplementary Fig. 3). Therefore, it is possible that the ability to reduce N2O, as in B. ottawaense, may have been important to survive in specific environments.
In the current study, the N2O-reducing activity of bradyrhizobia showed a similar behavior to the expression of the nosZ gene. B. ottawaense strains with high N2O-reducing activity (SG09 and OO99T) strongly express the nosZ gene under both N2O- and NO3--respiring conditions (Figs. 1 and 3, Table 1). In addition, Bradyrhizobium with low N2O-reducing activity (USDA110T, SF12, SF19) showed relatively low nosZ expression compared to that of high N2O-reducing strains (Fig. 1, Table 1). Given the relatively high homology of nosZ between B. diazoefficiens and B. ottawaense (92% identity in amino acid sequence, see Supplementary Fig. 7a), our results suggest that the N2O-reducing activity of Bradyrhizobium is determined by the expression of the nosZ gene rather than NosZ protein activity. However, further experiments, such as swapping of promotor or coding regions of nosZ gene, are required to prove this idea.
To investigate the cause of high nosZ expression in B. ottawaense, we first focused on the NasST regulatory system and examined whether it is functional in B. ottawaense. In the nasS deletion mutants (OO99TΔnasS and SG09ΔnasS), nosZ expression levels increased under N2O-respiring condition but not increased under NO3--respiring condition (Table 1), indicating that the NasST regulatory system is functional in B. ottawaense as in B. diazoefficiens25,33. In addition, it seems that the NasST regulatory system is not a main factor for the high expression in B. ottawaense because the nosZ expression of WT B. ottawaense (211 in SG09, 163 in OO99T) was higher than that of USDA110 ΔnasS (3.4) under N2O-respiring conditions (Table 2).
Analysis of ΔnasS mutants also showed that nosZ gene expression levels are not directly reflected in N2O-reducing activity. B. ottawaense ΔnasS mutants exhibited higher nosZ gene expression than that of WT (Table 2), but N2O-reducing activity did not significantly differ between WT and ΔnasS mutants (see Supplementary Fig. 6). In addition, when comparing nosZ expression in ΔnasS mutants, B. ottawaense demonstrated a 100-fold higher expression than that of B. diazoefficiens (Table 2), but N2O-reducing activity only slightly differed (see Supplementary Fig. 6). The lack of linearity between gene expression and N2O-reducing activity may indicate upper limits for NosZ protein activity. This may be due to translation efficiency or depletion of the components required for NosZ activity, such as copper and electrons, during N2O reduction. Moreover, NosZ protein activation requires highly complex pathways, such as sequential metal trafficking and assembly to copper sites via NosDFY41. The exact cause is presently unknown, but the aforementioned factors may define the upper limit of N2O reduction activity in bradyrhizobia.
Previous studies have shown the single or two transcription start sites of nosR in B. diazoefficiens USDA110T under free-living (aerobic), NO3--respiring (anaerobic), and symbiotic conditions24,33,42. In the current study, we present the first analysis of transcription start point under N2O-respiring condition. In B. diazoefficiens USDA110T, single transcription start point was observed regardless of different electron acceptors (N2O or NO3-). In contrast, B. ottawaense has a variable transcription start point depending on the electron acceptors: two transcription start points were detected under N2O respiration conditions in both SG09 and OO99T. Changes in the transcription start site depending on two different electron acceptor have been reported in studies on Geobactor43. Also, genome-wide analysis of transcription start sites in Clostridium identified several metabolism-related genes with multiple transcription start sites that change depending on the substrate44. Although the importance of having multiple transcription start sites has not been fully elucidated, it is considered an important regulatory mechanism of gene expression because it largely influences transcription efficiency, translation initiation, and protein abundance45. Changes in the transcription start sites of B. ottawaense nosR depend on the type of electron acceptor, which may be part of the nos genes expression regulatory mechanism in the denitrification system. Mutations in the transcription start sites will clarify their importance in the regulation of nos gene expression.
Genome sequence comparisons of high and low N2O-reducing activity strains revealed a novel determinant of activity. Incidentally, 56 bp deletion in the upstream region including translational start codon (ATG) of nosR was detected specifically in the low N2O-reducing activity strains, SF12 and SF19 (Fig. 4 and Supplementary Fig. 7), and introducing the deletion in the high N2O-reducing activity strains (SG09 and OO99T) reduced nosZ gene expression and N2O-reducing activity (Table 3). These results are consistent with previous studies where decreased nosZ expression was observed in artificially generated nosR-deleted Pseudomonas aeruginosa strains in which the nos genes were encoded in a single operon similar to that of Bradyrhizobium46. Since NosR functions as an electron donor for NosZ and regulates the transcription of nos genes47, the 56 bp deletion including the translational start codon may impair the function of NosR protein, causing a decrease in nosZ gene expression and N2O-reducing activity. As partial gene deletion is among the driving forces for environmental adaptation or functional evolution in bacteria48,49,50, the strains with natural deletion isolated in the present study may have evolved to adapt to environments with limited denitrification substrates. Accordingly, examining the distribution and abundance of high and low N2O-reducing activity strains in various environments may reveal the importance of N2O-reducing activity in environmental adaptation.
In summary, we demonstrated that the N2O-reducing activity of B. ottawaense is significantly higher than that of conventional strains, and this activity is achieved via high nosZ expression. Since N2O is a GHG mainly generated in agricultural lands, developing strategies for reducing N2O emissions from agricultural lands is an urgent task. The B. ottawaense we reported here has great potential for GHG mitigation in the rhizosphere owing to its high N2O-reducing activity. In addition, the regulatory mechanism of nos gene expression we elucidated in this study will be useful for developing and identifying bacteria with higher GHG-reducing ability. Further studies on the ecology of B. ottawaense including its compatibility with legume crops and competitiveness with other indigenous rhizobacteria are needed to improve its utility on actual agricultural land.
Methods
Bacterial strains, isolation, and genome analysis
The type strain B. ottawaense OO99T was purchased from the Microbial Domain Biological Resource Centre HAMBI (Helsinki, Finland). The type strain B. diazoefficiens USDA110T was provided by Dr. Michael J. Sadowsky at University of Minnesota. We used the culture stock of the nosZ deleted mutant B. diazoefficiens USDA110ΔnosZ generated by Hirayama et al. 201151. The B. ottawaense strains used in this study are listed in Supplementary Table 1. Eight strains (SG09, SG10, SG20, SG23, TM102, TM233, and TM239) have been reported by Wasai-Hara et al.17, and the other strains were isolated by the same procedures. For whole genome sequencing, genomic DNA was extracted using a Bacteria GenomicPrep Mini Spin Kit (Cytiva, Tokyo, Japan). DNA libraries were prepared using a Nextera Sample Preparation Kit (Illumina, San Diego, CA, USA), and the 300-bp paired-end libraries were sequenced using Illumina Miseq (Illumina). Subsequently, 20 bp of the 5′ and 3′ ends were trimmed, and the genomes were assembled using CLC Genomics Workbench ver. 8.5.1 (Illumina). Genome annotation was performed using DFAST52.
N2O-reducing activity
N2O-reducing activity was determined by culturing the bacteria under anaerobic conditions with 1% N2O supplemented as the sole electron acceptor. The N2O concentration was measured using a gas chromatograph (GC2014; Shimadzu, Kyoto, Japan) equipped with a thermal conductivity detector and Porapak Q column (GL Sciences, Tokyo, Japan). Bacterial strains were aerobically cultured for over 6 h in a 75-mL test tube with an air-permeable plug containing 10 mL HM liquid medium53 supplemented with 0.1% (w/v) arabinose and 0.025% (w/v) yeast extract at 28 °C with shaking at 200 rpm. Thereafter, the appropriate volume of bacterial culture was added to new tubes containing 10 mL HM medium to reach an optical density (OD) at 660 nm (OD660) of 0.05. The OD was measured using a test tube 25 mm diameter (TEST25NP; AGC Techno Glass Co., Ltd., Shizuoka, Japan). The test tube was closed with a butyl rubber cap, and the gas phase was replaced with 4.98% N2O + 95.02% N2 gas following overnight (12–14 h) culture to induce N2O reduction metabolism. Subsequently, the gas phase was again replaced with 100% N2 gas, after which 100% N2O was supplemented to adjust to a final concentration of 1%. Finally, the test tube was incubated at 28 ºC with shaking at 200 rpm, and 100 µL of gas phase was withdrawn every 1–3 h and subjected to the gas chromatography. To calculate the N2O-reducing activity per total protein, calibration curves for protein content at OD660 were developed for both B. ottawaense and B. diazoefficiens. The total protein content was calculated from the measured OD660. Protein content in the supernatants was measured using a protein assay kit (Bio-Rad Laboratories, Hercules, CA, USA).
N2O flux experiment
N2O flux in the soybean rhizosphere was measured using a previously described method with modifications11 (see Supplementary Fig. 4). Bacterial strains (B. japonicum USDA6T, B. diazoefficiens USDA110T and B. ottawaense SG09) were aerobically cultured in HM liquid medium at 30 °C for 1 week, after which the prepared bacterial suspension was adjusted to 1 × 108 cells mL−1 using sterilized water. Soybean seeds (Glycine max, cv. Enrei (GmJMC025) seeds acquired from Genebank Project NARO, Japan) were sterilized using 0.5% sodium hypochlorite were sown in Leonardo Jar pots—at three seeds per pot—containing sterilized vermiculite and were inoculated with 1 mL of bacterial suspension. Four pots were prepared per each inoculated strain. The seeds were cultivated in a growth chamber at 25 °C for 16 h in light and 8 h in the dark. Thinning was performed on the third day after sowing, leaving an individual plant that was in the best germination state, and cultivation continued for another 33 days. A nitrogen-free hydroponic solution was periodically added to the pot during cultivation. After cultivation, plant shoots system were decapitated, and the root system was gently immersed in water to remove excess vermiculite. The root system was transferred to a 100-mL glass vial containing 30 mL soil obtained from the Kashimadai experimental field (38°27′36.0″N 141°05′24.0″E, at the permission of Tohoku University, Japan). The soil was previously sieved through a 2mm mesh sieve to remove large soil aggregates and stones. In addition, 5 ml of sterile distilled water was added to the glass vials. Thereafter, the vials with the roots, soil, and water were incubated aerobically at 25 °C for 20 days to induce nodule degradation: the vials were covered with soft wiping cloth to maintain aeration during the incubation period. After the incubation period, for determining the N2O flux, the vials were sealed with butyl-rubber caps and kept for 240–360 min in the following treatments: (1) atmospheric condition and (2) with the addition of 50 ppm of N2O, in four replicates. N2O concentration in the gas phase of vials was determined a gas chromatograph (GC2014; Shimadzu) equipped with a 63Ni electron capture detector and tandem Porapak Q columns (GL Sciences; 80/100 mesh; 3.0 mm × 1.0 m and 3.0 mm × 2.0 m).
Expression analysis
nosZ gene expression levels were measured under N2O- and NO3--respiring conditions. For N2O-respiring conditions, cells were prepared the same as described above. 3 h after exposure to 1% N2O conditions, a 1 mL phenol solution (10% phenol in ethanol) was added to the 1-mL culture to stop metabolism. After centrifugation, the pellets were stored at − 80 °C until further processing. For NO3- respiring conditions, cells were anaerobically grown in 20 mL HM medium supplemented with 10 mM KNO3 in a 75-mL test tube. The OD660 was initially adjusted to 0.05 and monitored to induce the exponential growth phase of cells. When OD660 reached 0.1, the cells were collected as described above. Subsequently, total RNA was isolated using the hot-phenol method as described previously54, followed by DNase I treatment (RQ1; Promega, Madison, WI, USA) and further purification using RNA Clean & Concentrator-5 (Zymo Research, Irvine, CA, USA). First-strand cDNA was synthesized using 500 ng RNA as a template and SuperScript IV Reverse Transcriptase (Invitrogen, Waltham, MA, USA) according to the manufacturer’s instructions. RT-qPCR was performed using a LightCycler Nano Instrument (Roche, Basel, Switzerland), LightCycler® FastStart DNA MasterPLUS SYBR® Green I (Roche), and specific primers for sigA (sigAf/sigAr) and nosZ (nosZ_qPCR_F/ nosZ_qPCR_R) (see Supplementary Table 4) at an annealing temperature of 60 ºC for 50 cycles. Relative expression calculated using the 2−ΔΔCt method55 was normalized to sigA expression.
NosZ activity staining
B. ottawaense SG09 and B. diazoefficiens USDA110T were cultured overnight under N2O-respiring conditions. After centrifugation of the cultured cells, total protein was extracted using lysis buffer (CelLytic B; Sigma-Aldrich, St. Louis, MO, USA) and sonication (BIORUPTOR; BM Equipment Co., Ltd., Tokyo, Japan). Protein content in the supernatants was measured using a DC-protein assay (Bio-Rad). For DOC-PAGE, approximately 9.6 µg of total protein was used as the non-diluted sample (1×, Fig. 3). After electrophoresis, each gel was immersed in the buffer containing 25 mM Tris, 192 mM glycine, and 1 mM methyl viologen (pH 8.3). Subsequently, Ti(III)-citrate was used to reduce methyl viologen, and N2O-saturated H2O was added for the in-gel N2O-reducing enzymatic reaction. Band signal intensity was determined using an Image J macro, Band/Peak Quantification56. All experiments except for protein quantification were performed under an N2 atmosphere. CBB staining was performed after N2O-reducing activity staining to confirm protein content.
nasS, nosZ, and 56-bp deletion mutants
nasS deletion mutants were generated using the in-frame markerless method. The deletion region was determined as in the B. diazoefficiens USDA110T nasS mutant strain (5M09) reported by Sánchez et al.25. Briefly, pK18mobsacB-Ω was created by replacing the kanamycin resistance gene coding region of the suicide vector pK18mobsacB57 with a streptomycin-spectinomycin resistance gene (aadA). pK18mobsacB was digested with NcoI and BglII to obtain 5.1 kb of linear DNA with 0.6 kb of the kanamycin resistance gene partially deleted. The aadA fragment to be introduced was amplified by PCR using primers aadA_F_IF and aadA_R_IF and Prime STAR® Max DNA Polymerase (Takara Bio Inc., Shiga, Japan) (details on the primers are shown in Supplementary Table 4) with pHP45Ω58 as a template. Thereafter, 1.1 kb of amplified DNA was extracted using Wizard® SV Gel and a PCR Clean-Up System (Promega). The resulting linear pK18mobsacB and aadA were combined using an In-fusion HD cloning kit (Takara Bio Inc.) and transformed E. coli DH5α according to the manufacturer’s instructions.
The up- and downstream regions of the nasS gene were amplified by PCR using primers Bo_nasSdel_F1/R1 and Bo_nasSdel_F2/R2 (see Supplementary Table 4 for details on primers) and Prime STAR® Max DNA Polymerase (Takara Bio Inc.). Amplified fragments were combined by overlap extension PCR and inserted into the SmaI site of pK18mobsacB-Ω using an In-fusion HD Cloning Kit (Takara Bio Inc.). The sequence of the introduced fragment was confirmed by sequencing, and the resultant plasmid was designated pMS187. Transmission of pMS187 to B. ottawaense strains and homologous recombination of the nasS region were performed by triparental mating with a mobilizing E. coli HB101 strain harboring the pRK201359 helper plasmid. Next, B. ottawaense SG09 or OO99T, E. coli DH5α harboring pMS187, and E. coli HB101 harboring pRK2013 were mixed and cultured for mating. Transconjugants were selected by resistance to streptomycin (Sp, 100 µg/mL), spectinomycin (Sm, 100 µg/mL), and polymyxin (Px, 50 µg/mL) and sensitivity to sucrose (10%). The single crossover strains were further cultured in HM medium without antibiotics, and deletion mutants that showed Sp/Sm sensitivity and sucrose resistance—SG09ΔnasS and OO99TΔnasS—were obtained.
ΔnosZ and 56 bp deletion mutants were generated using the same methods as for the nasS mutants with the pK18mobsacB-Ω vector. Briefly, the up- and downstream regions of the nosZ gene were amplified by PCR using primers SG09_nos-1F/1R and SG09_nos-2F/2R (see Supplementary Table 4 for details on the primers) and Prime STAR® max DNA Polymerase (Takara Bio Inc.). The amplified fragments were combined by overlap extension PCR using primers SG09_nos-1F/1R and SG09_nos-2F/2R (see Supplementary Table 4) and Prime STAR® max DNA Polymerase (Takara Bio Inc.). The PCR fragments and pK18mobsacB-Ω were digested with EcoRI and HindIII and then ligated using a DNA Ligation Kit (< Mighty Mix > ; Takara Bio Inc.). Thereafter, triparental mating was performed using the sequence-introduced vector as described above.
For 56-bp deletion mutants, the up- and downstream regions of the 56-bp region were amplified by PCR using primers 56del_F1/R1 and 56del_F2/R2 (see Supplementary Table 4) and then combined and inserted into the SmaI site of pK18mobsacB-Ω using an In-fusion HD Cloning Kit (Takara Bio Inc.). Thereafter, triparental mating was performed using the sequence-introduced vector as described above. The generated 56 bp deletion mutants were designated SG09Δ56 and OO99TΔ56.
5′ RACE
5′ RACE experiments were performed using a 5′/3′ RACE kit, 2nd Generation (Roche). Briefly, the total RNA of B. ottawaense and B. diazoefficiens strains were isolated from cells grown under N2O- and NO3-- respiring conditions using the hot-phenol method as described above. cDNA synthesis and amplification of the 5′- region of nosR were conducted according to the manufacturer’s instructions using the primers listed in Supplementary Table 4 (Bw_SP1, SP2, and SP3 for B. ottawaense strains, R_SP1, SP2, and SP3 for B. diazoefficiens strains). The amplified fragments were sequenced to determine the transcription start site.
Statistical analysis
Differences in N2O reducing activities between all strains tested were evaluated using Tukey’s test after ANOVA analysis. Differences in N2O flux and nosZ gene expression between the two strains were evaluated using Student’s t tests at a significance level of 0.05.
Data availability
Genome data are available in NCBI (https://www.ncbi.nlm.nih.gov/), and accession numbers are detailed in the Supplementary Information files.
References
Prather, M. J. et al. Measuring and modeling the lifetime of nitrous oxide including its variability. J. Geophys Res. 120, 5693–5705 (2015).
IPCC 2021, A. R. Climate change 2021: The physical science basis. In Working Group I Contribution to the IPCC Sixth Assessment Report (2021).
Tian, H. et al. A comprehensive quantification of global nitrous oxide sources and sinks. Nature 586, 248–256 (2020).
McKenney, D. J., Wang, S. W., Drury, C. F. & Findlay, W. I. Denitrification and mineralization in soil amended with legume, grass and corn residues. Soil. Sci. Soc. Am. J. 57, 1013–1020 (1993).
Uchida, Y. & Akiyama, H. Mitigation of postharvest nitrous oxide emissions from soybean ecosystems: A review. Soil Sci. Plant Nutr. 59, 477–487 (2013).
Thomson, A. J., Giannopoulos, G., Pretty, J., Baggs, E. M. & Richardson, D. J. Biological sources and sinks of nitrous oxide and strategies to mitigate emissions. Philos. Trans. R. Soc. B: Biol. Sci. 367, 1157–1168 (2012).
Kuypers, M. M. M., Marchant, H. K. & Kartal, B. The microbial nitrogen-cycling network. Nat. Rev. Microbiol. 16, 263–276 (2018).
Hallin, S., Philippot, L., Loffler, F. E., Sanford, R. A. & Jones, C. M. Genomics and ecology of novel N2O-reducing microorganisms. Trends Microbiol. 1485, 43–55 (2017).
Della, C. T. et al. Higher than expected N2O emissions from soybean crops in the Pampas Region of Argentina: Estimates from DayCent simulations and field measurements. Sci. Total Environ. 835, 155408 (2022).
Sameshima-Saito, R. et al. Symbiotic Bradyrhizobium japonicum reduces N2O surrounding the soybean root system via nitrous oxide reductase. Appl. Environ. Microbiol. 72, 2526–2532 (2006).
Itakura, M. et al. Mitigation of nitrous oxide emissions from soils by Bradyrhizobium japonicum inoculation. Nature Clim. Change 3, 208–212 (2013).
Akiyama, H. et al. Mitigation of soil N2O emission by inoculation with a mixed culture of indigenous Bradyrhizobium diazoefficiens. Sci. Rep. 6, 32869 (2016).
Hénault, C., Barbier, E., Hartmann, A. & Revellin, C. New insights into the use of rhizobia to mitigate soil N2O emissions. Agriculture 12, 271 (2022).
Melissa, O. et al. Evaluation of nitrous oxide emission by soybean inoculated with Bradyrhizobium strains commonly used as inoculants in South America. Plant Soil 472, 311–328 (2022).
Bueno, E. et al. Anoxic growth of Ensifer meliloti 1021 by N2O-reduction, a potential mitigation strategy. Front. Microbiol. 6, 537 (2015).
Shiina, Y. et al. Relationship between soil type and N2O reductase genotype (nosZ) of indigenous soybean bradyrhizobia: nosZ-minus populations are dominant in Andosols. Microbes Environ. 29, 420–426 (2014).
Wasai-Hara, S. et al. Diversity of Bradyrhizobium in non-leguminous sorghum plants: B. ottawaense isolates unique in genes for N2O reductase and lack of the type VI secretion system. Microbes Environ. 35, 19102 (2020).
Sánchez, C. & Minamisawa, K. Nitrogen cycling in soybean rhizosphere: Sources and sinks of nitrous oxide (N2O). Front. Microbiol. 10, 1943 (2019).
Yang, L. F. & Cai, Z. C. The effect of growing soybean (Glycine max L.) on N2O emission from soil. Soil Biol. Biochem. 37, 1205–1209 (2005).
Inaba, S. et al. Nitrous oxide emis- sion and microbial community in the rhizosphere of nodulated soybeans during the late growth period. Microbes Environ. 24, 64–67 (2009).
Inaba, S. et al. N2O Emission from degraded soybean nodules depends on denitrification by Bradyrhizobium japonicum and other microbes in the rhizosphere. Microbes Environ. 27, 470–476 (2012).
Sameshima-Saito, R., Chiba, K. & Minamisawa, K. New method of denitrification analysis of bradyrhizobium field isolates by gas chromatographic determination of 15N-labeled N2. Appl. Environ. Microbiol. 70, 2886–2891 (2004).
Bonnet, M. et al. The structure of Bradyrhizobium japonicum transcription factor FixK2 unveils sites of DNA binding and oxidation. J. Biol. Chem. 288, 14238–14246 (2013).
Torres, M. J. et al. FixK2 Is the main transcriptional activator of Bradyrhizobium diazoefficiens nosRZDYFLX genes in response to low oxygen. Front. Microbiol. 8, 1621 (2017).
Sánchez, C. et al. The nitrate-sensing NasST system regulates nitrous oxide reductase and periplasmic nitrate reductase in Bradyrhizobium japonicum. Environ. Microbiol. 16, 3263–3274 (2014).
Torres, M. J. et al. The global response regulator RegR controls expression of denitrification genes in Bradyrhizobium japonicum. PLoS ONE 9, e99011 (2014).
Zumft, W. G. & Kroneck, P. M. Respiratory transformation of nitrous oxide (N2O) to dinitrogen by bacteria and archaea. Adv. Microb. Physiol. 52, 107–227 (2007).
Velasco, L., Mesa, S., Xu, C., Delgado, M. J. & Bedmar, E. J. Molecular characterization of nosRZDFYLX genes coding for denitrifying nitrous oxide reductase of Bradyrhizobium japonicum. Anton. Van Leeuwenh. 85, 229–235 (2004).
Yu, X., Cloutier, S., Tambong, J. T. & Bromfield, E. S. P. Bradyrhizobium ottawaense sp. nov., a symbiotic nitrogen fixing bacterium from root nodules of soybeans in Canada. Int. J. Syst. Evol. Microbiol. 64, 3202–3207 (2014).
Win, et al. Synergistic N2-fixation and salt stress mitigation in soybean through dual inoculation of ACC deaminase-producing Pseudomonas and Bradyrhizobium. Sci. Rep. 13, 17050 (2023).
Nguyen, H. D. T., Cloutier, S. & Bromfield, E. S. P. Complete genome sequence of Bradyrhizobium ottawaense OO99T, an efficient nitrogen-fixing symbiont of soybean. Microbiol. Resour. Announc. 7, e01477-e1518 (2018).
Wu, M. & Eisen, J. A. A simple, fast, and accurate method of phylogenomic inference. Genome Biol. 9, 151 (2008).
Sánchez, C., Mitsui, H. & Minamisawa, K. Regulation of nitrous oxide reductase genes by NasT-mediated transcription antitermination in Bradyrhizobium diazoefficiens. Environ. Microbiol. Rep. 9, 389–396 (2017).
Itakura, M. et al. Generation of Bradyrhizobium japonicum mutants with increased N2O reductase activity by selection after introduction of a mutated dnaQ gene. Appl. Environ. Microbiol. 74, 7258–7264 (2008).
Tang, J., Bromfield, E. S., Rodrigue, N., Cloutier, S. & Tambong, J. T. Microevolution of symbiotic Bradyrhizobium populations associated with soybeans in east North America. Ecol. Evol. 2, 2943–2961 (2012).
Yan, J. et al. Genetic diversity of indigenous soybean-nodulating rhizobia in response to locally-based long term fertilization in a Mollisol of Northeast China. World J. Microbiol. Biotechnol. 33, 6 (2017).
Minakata, C., Wasai-Hara, S., Fujioka, S., Sano, S. & Matsumura, A. Unique rhizobial communities dominated by Bradyrhizobium liaoningense and Bradyrhizobium ottawaense were found in vegetable soybean nodules in Osaka Prefecture, Japan. Microbes Environ. 38, ME22081 (2023).
Mania, D., Woliy, K., Degefu, T. & Frostegård, Å. A common mechanism for efficient N2O reduction in diverse isolates of nodule-forming bradyrhizobia. Environ. Microbiol. 22, 17–31 (2020).
Shao, S., Chen, M., Liu, W., Hu, X. & Li, Y. Long-term monoculture reduces the symbiotic rhizobial biodiversity of peanut. Syst. Appl. Microbiol. 43, 126101 (2020).
Gao, Y. et al. Competition for electrons favours N2O reduction in denitrifying Bradyrhizobium isolates. Environ. Microbiol. 23, 2244–2259 (2021).
Müller, C. et al. Molecular interplay of an assembly machinery for nitrous oxide reductase. Nature 608, 626–631 (2022).
Čuklina, J. et al. Genome-wide transcription start site mapping of Bradyrhizobium japonicum grown free-living or in symbiosis—a rich resource to identify new transcripts, proteins and to study gene regulation. BMC Genom. 17, 302 (2016).
González, G. et al. Global transcriptional start site mapping in Geobacter sulfurreducens during growth with two different electron acceptors. FEMS Microbiol. Lett. 363, fnw175 (2016).
Boutard, M. et al. Global repositioning of transcription start sites in a plant-fermenting bacterium. Nat. Commun. 7, 13783 (2016).
Arribere, J. A. & Gilbert, W. V. Roles for transcript leaders in translation and mRNA decay revealed by transcript leader sequencing. Genome Res. 23, 977–987 (2013).
Arai, H., Mizutani, M. & Igarashi, Y. Transcriptional regulation of the nos genes for nitrous oxide reductase in Pseudomonas aeruginosa. Microbiology 149, 29–36 (2003).
Honisch, U. & Zumft, W. G. Operon structure and regulation of the nos gene region of Pseudomonas stutzeri, encoding an ABC-Type ATPase for maturation of nitrous oxide reductase. J. Bacteriol. 185, 1895–1902 (2003).
D’Souza, G. et al. Less is more: Selective advantages can explain the prevalent loss of biosynthetic genes in bacteria. Evolution 68, 2559–2570 (2014).
Godfroid, M. et al. Insertion and deletion evolution reflects antibiotics selection pressure in a Mycobacterium tuberculosis outbreak. PLoS Pathog. 30, e1008357 (2020).
Bottai, D. et al. TbD1 deletion as a driver of the evolutionary success of modern epidemic Mycobacterium tuberculosis lineages. Nat. Commun. 11, 684 (2020).
Hirayama, J., Eda, S., Mitsui, H. & Minamisawa, K. Nitrate-dependent N2O emission from intact soybean nodules via denitrification by Bradyrhizobium japonicum bacteroids. Appl. Environ. Microbiol. 77, 8787–8790 (2011).
Tanizawa, Y., Fujisawa, T., Kaminuma, E., Nakamura, Y. & Arita, M. DFAST and DAGA: Web-based integrated genome annotation tools and resources. Biosci. Microbiota Food Health 35, 173–184 (2016).
Cole, M. A. & Elkan, G. H. Transmissible resistance to penicillin G, neomycin, and chloramphenicol in Rhizobium japonicum. Antimicrob. Agents Chemother. 4, 248–253 (1973).
Hauser, F. et al. Dissection of the Bradyrhizobium japonicum NifA+σ54 regulon, and identification of a ferredoxin gene (fdxN) for symbiotic nitrogen fixation. Mol. Genet. Genom. 278, 255–271 (2007).
Schmittgen, T. D. & Livak, K. J. Analyzing real-time PCR data by the comparative CT method. Nat. Protoc. 3, 1101–1108 (2008).
Ohgane, K. & Yoshioka H. Quantification of gel bands by an image J Macro, Band/Peak Quantification Tool. Protocols.io. https://doi.org/10.17504/protocols.io.7vghn3w (2019).
Schäfer, A. et al. Small mobilizable multi-purpose cloning vectors derived from the Escherichia coli plasmids pK18 and pK19: Selection of defined deletions in the chromosome of Corynebacterium glutamicum. Gene 145, 69–73 (1994).
Prentki, P. & Krisch, H. M. In vitro insertional mutagenesis with a selectable DNA fragment. Gene 29, 303–313 (1984).
Figurski, D. H. & Helinski, D. R. Replication of an origin-containing derivative of plasmid RK2 dependent on a plasmid function provided in trans. Proc. Natl. Acad. Sci. U. S. A. 76, 1648–1652 (1979).
Acknowledgements
This work was supported by a Grant-in-Aid for JSPS Fellows [Grant Numbers: 20J12228, 22J01397] and project JPNP18016, commissioned by the New Energy and Industrial Technology Development Organization (NEDO). We would like to thank Cristina Sánchez Gomes for valuable discussion, and Yukiko Fujisawa, Kaori Kakizaki, Kanako Tago, and Yoshiyuki Ohtsubo for their technical support.
Author information
Authors and Affiliations
Contributions
S.W.-H. carried out experiments, data analysis, and genome analysis and drafted the original manuscript. M.I., A.F.S., and D.T. designed and performed experiments and contributed to manuscript writing. M.S. contributed to the generation of gene deletion mutants and interpretation of data. N.I. and T.Y. supervised protein experiments and contributed to the interpretation of data. H.M., S.S., H.I-A, Y.S., and K.M. supervised the conduct of this study. Y.S. and K.M. contributed to manuscript finalization and revision. All authors reviewed the results and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wasai-Hara, S., Itakura, M., Fernandes Siqueira, A. et al. Bradyrhizobium ottawaense efficiently reduces nitrous oxide through high nosZ gene expression. Sci Rep 13, 18862 (2023). https://doi.org/10.1038/s41598-023-46019-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-46019-w
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.