Abstract
PTEN hamartoma tumour syndrome is characterised by mutations in the human PTEN gene. We performed transcriptomic and proteomic analyses of neural tissues and primary cultures from heterozygous and homozygous Pten-knockout mice. The somatosensory cortex of heterozygous Pten-knockout mice was enriched in immune response and oligodendrocyte development Gene Ontology (GO) terms. Parallel proteomic analysis revealed differentially expressed proteins (DEPs) related to dendritic spine development, keratinisation and hamartoma signatures. However, primary astrocytes (ASTs) from heterozygous Pten-knockout mice were enriched in the extracellular matrix GO term, while primary cortical neurons (PCNs) were enriched in immediate-early genes. In ASTs from homozygous Pten-knockout mice, cilium-related activity was enriched, while PCNs exhibited downregulation of forebrain neuron generation and differentiation, implying an altered excitatory/inhibitory balance. By integrating DEPs with pre-filtered differentially expressed genes, we identified the enrichment of traits of intelligence, cognitive function and schizophrenia, while DEPs in ASTs were significantly associated with intelligence and depression.
Similar content being viewed by others
Introduction
PTEN hamartoma tumour syndrome (PHTS) is a spectrum of disorders caused by heterozygous germline mutations of the phosphatase and tensin homologue deleted on chromosome 10 (PTEN) tumour suppressor gene, which is a gatekeeper of the phosphoinositide 3-kinase (PI3K) pathway1. PHTS is characterised by a hamartomatous growth that develops in various parts of the body2,3 and neurodevelopmental disorders, such as macrocephaly, learning disabilities, developmental delays, and autism spectrum disorder (ASD)4. The prevalence of ASD in PHTS is estimated to be ~ 22%5,6 and the frequency of PTEN mutation in ASD patients with macrocephaly is ~ 10%2,7. The results of a recent clinical study suggest the disruption of frontal lobe systems, including full-scale IQ, attention, motor coordination, cognitive function and sensory deficits, in patients with PHTS6. Indeed, heterozygous Pten-knockout (Pten+/−) mice exhibit broad brain overgrowth with excess neurons at birth and cortical glia in adulthood, deficits in social behaviour, acquired repetitive behaviours8,9,10 and enhanced axonal branching and connectivity in projections of the medial prefrontal cortex to the basolateral amygdala axons11. Moreover, CaMK2-Cre+/−; Pten+/loxP mice display reduced intrinsic excitability of pyramidal neurons due to increased expression levels of small-conductance calcium-activated potassium channels12. At the cellular level, the expression profile of neural precursor cells prepared from the subventricular zone (SVZ) of adult Pten+/− mice shows increases in vascular endothelial growth factor and doublecortin (DCX) expression levels and a shift in DCX-positive cells from the SVZ to the olfactory bulb13.
Most of the other functions of PTEN in the brain have been discovered using conditional Pten-knockout mice due to embryonic lethality associated with homozygous germline Pten deletion14. One of the earliest mouse models using GFAP-Cre-mediated deletion of Pten in astrocytes (ASTs) and neurons in the whole brain resulted in macrocephaly, increased dendritic spine density, abnormal enlargement of synaptic structures, layering defects, seizures and ataxia15,16,17,18,19. To examine neuronal-specific effects, Pten was knocked out in dopaminergic neurons using dopamine transporter (DAT)-Cre20,21, in excitatory neurons in the forebrain using calcium/calmodulin-dependent protein kinase II alpha (CamKIIα)-Cre22, in Purkinje neurons using Purkinje cell protein 2 (L7)-Cre23 and in subsets of postmitotic neurons using neuron-specific enolase (NSE)-Cre24,25,26. These knockout strategies resulted in hypertrophic neurons with aberrant dendritic arborisation and increased axonal density and length. Electrophysiologically, Pten knockout driven by CamKIIα-Cre and NSE-Cre induces impairment of hippocampal long-term potentiation and metabotropic glutamate receptor-dependent long-term depression, respectively. Hence, specific behavioural phenotypes, such as impaired spatial memory, abnormal social interaction and abnormal response to sensory stimuli, have been assigned to specific subsets of neurons. In addition, mice with postnatally generated Pten knockout in hippocampal dentate granule cells exhibit hyperexcitability due to an increased excitatory synaptic drive27,28,29,30. Interestingly, Pten deletion in cortical γ-aminobutyric acid (GABA)ergic interneurons using NK2 homeobox 1 (Nkx2.1)-Cre, which targets medial and caudal ganglionic eminences and preoptic area progenitors, resulted in the preferential loss of somatostatin (SST)-positive interneurons, which increased the ratio of parvalbumin/SST interneurons and induced inhibitory actions on glutamatergic cortical neurons, leading to deficits in social behaviour31. The above evidence supports the hypothesis of disrupted excitatory/inhibitory (E/I) balance as a potential mechanism of autism32.
The loss of PTEN leads to the upregulation of mechanistic target of rapamycin (mTOR)-driven transcriptional and translational machinery33,34. Increased PI3-K/mTOR signalling has been implicated in the autistic phenotypes of PHTS patients, as treatment with the pharmacological mTOR inhibitor rapamycin normalises some of the behavioural defects in Pten-knockout mice35. However, the PTEN-regulated genes responsible for the neuropathological or ASD phenotypes in PHTS patients remain largely unknown. Furthermore, homozygous PTEN mutations have not been found in PHTS patients and findings in conditional Pten-knockout models have not been reproduced in heterozygous Pten-knockout models, which imply that the heterozygous loss of PTEN may have very subtle pathological consequences that could not be detected using conventional methods.
In this report, we present a comprehensive analysis of the proteomes and transcriptomes of the somatosensory cortex, which is a region responsible for receiving and processing sensory stimuli, and has been implicated in ASD36. This region develops actively during the early postnatal period, as determined by previous studies of the E/I balance regulatory networks37,38 in heterozygous Pten-knockout mice. Moreover, we dissected the cell-type-specific contribution from the transcriptomes of primary neural progenitor cells (NPCs), cortical ASTs, and primary cortical neurons (PCNs) derived from heterozygous and homozygous Pten-knockout mice. Our findings may provide the underlying molecular mechanism responsible for ASD phenotypes observed in PHTS patients.
Methods
Mice
The permission to conduct animal experimentation was obtained from The Department of Health of the Government of the Hong Kong Special Administrative Region, under the following two animal licences (20-71) in DH/HT&A/8/2/1 Pt.4 and (20-72) in DH/HT&A/8/2/1 Pt.4. All animal protocols were approved by the Animal Experimentation and Ethics Committee of the Chinese University of Hong Kong under the approval number, 14/148/MIS. All methods are reported in accordance with ARRIVE guidelines and were carried out in accordance with relevant guidelines and regulations. All the mice were maintained under selective pathogen-free conditions. All mice had a C57BL/6J background (see Supplementary Table ST4 for a summary). B6.129-Ptentm1Rps/Nci heterozygous (Pten+/−) mice were obtained from the National Cancer Institute’s Mouse Models of Human Cancers Consortium (Frederick, MD, USA)14. The Pten+/− mice were crossed with wild-type (WT) C57BL/6J mice to generate either WT or Pten+/− progeny. B6.129S4-Ptentm1Hwu/J (Ptenfl/fl) and B6.Cg-Tg(Nes-Cre)1Kln/J (Nes-Cre) mice were purchased from the Jackson Laboratory (Bar Harbor, ME, USA)39,40. Ptenfl/fl mice were crossed with Nestin promoter-driven Cre recombinase (Nes-Cre) mice to generate compound heterozygous mice (Nes-Cre:Ptenfl/+). Nes-Cre:Ptenfl/+ mice were then mated with Ptenfl/fl mice to generate the following four genotypes: Ptenfl/+ (Ctrl), Ptenfl/fl (Ctrl), Nes-Cre:Ptenfl/+ (Nes-HET) and Nes-Cre:Ptenfl/fl (Nes-KO). All mice were euthanized with CO2, except for P0 and P2 pups which were euthanized with isoflurane.
Primary cultures
Primary NPCs, PCAs and PCNs were established based on previously published protocols41. The detailed procedures are provided in the Supplementary Information.
Brain microdissection
Mice were perfused with 30 mL of ice-cold phosphate-buffered saline. The SSC (1.82 mm to 1.94 mm anteroposterior from the bregma) was dissected under a stereomicroscope (SMZ660; Nikon, Tokyo, Japan)42. The right hemisphere SSC was snap-frozen in liquid nitrogen, while the left hemisphere SSC was triturated with 1 mL of TRIzol™ Reagent (Thermo Fisher Scientific, Waltham, MA) before snap freezing.
RNA-seq
Total RNA was extracted using TRIzol™ Reagent. Three mice per group were used as biological replicates, except for P30M mice, of which there were only two in the Pten+/− group. The RNA samples were subjected to quantitative RNA-seq by BGI (Hong Kong SAR) or GROKEN Bioscience (Hong Kong SAR). Briefly, polyA-enriched mRNA was reverse-transcribed into cDNA and purified using magnetic beads. Adaptors were ligated to the DNA ends, size fractionated and enriched by polymerase chain reaction (PCR) amplification. The quality of the DNA fragments was determined using an Agilent 2100 Bioanalyzer (Agilent, Santa Clara, CA, USA) and sequenced on a HiSeq2500, HiSeq4000 or X Ten platform (Illumina, San Diego, CA, USA). Further information on data analysis is provided in the Supplementary Information section. Raw data and normalised counts are available on the Gene Expression Omnibus database under series GSE190879.
Proteomic analysis
Snap-frozen SSCs were processed for proteomics analysis based on a previously published protocol43. Approximately 100 µg of soluble protein was subjected to iTRAQ analysis (BGI). Additional experimental procedures and raw tandem mass spectrometry data are provided in the Supplementary Information section. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via PRIDE, under the identifier PXD03057344.
Quantitative reverse transcription PCR
First-strand cDNA was generated from 0.5 μg of DNase I (Thermo Fisher)-treated total RNA using the High-Capacity cDNA Reverse Transcription Kit (Thermo Fisher Scientific) based on the manufacturer’s instructions. Quantitative PCR was performed using the Power SYBR Green PCR Master Mix (Thermo Fisher Scientific) and the ABI QuantStudio 7 Flex Real-Time PCR System (384-well plates, Thermo Fisher Scientific). Primers were designed using the online tool GETprimer (https://gecftools.epfl.ch/getprime). Only intron-spanning primers that also targeted protein-coding transcripts were selected. The specific primers used in this analysis are listed in the supplementary information section. Cycle threshold (CT) values were first normalised to the expression of β-actin, and relative gene expression levels were calculated using the 2−∆∆CT method.
Integration with ASD co-expression modules
Gene lists of ASD co-expression modules were obtained from Parikshak et al.45. The enrichment of the identified DEGs in the SSC within the ASD co-expression modules was calculated using a cumulative hypergeometric test, and the FDR was adjusted using the Benjamini–Hochberg correction for multiple comparisons.
Integration with neurological disease GWAS data and neurobehavioural traits
GWAS catalogue version 1.0.2 was used for analysis (https://www.ebi.ac.uk/gwas/home). Risk genes for the following neuropathologies and neurobehavioural traits were extracted from the raw file: Alzheimer’s disease, Parkinson’s disease, amyotrophic lateral sclerosis, multiple sclerosis, autism, schizophrenia, bipolar disorder, attention deficit hyperactivity disorder, depression, anxiety, language disorder, intelligence and cognitive function. Duplicated genes, non-protein coding genes, GWAS loci in intergenic regions and genes with a p-value > 9 × 10–6 were removed. Genes reported in ‘MAPPED_GENE’ were used only if the ‘REPORTED GENE(S)’ field was empty. Statistical significance between the GWAS gene lists and our DEP and DEG lists against the total number of protein-coding genes (GENCODE Release version 38) as background was calculated using a cumulative hypergeometric test, and the FDR was adjusted using Benjamini–Hochberg correction for multiple comparisons.
Statistical analysis
For western blotting and reverse transcription-quantitative PCR analyses, statistically significant differences between two groups were identified using an unpaired Student’s t-test. Comparisons of age, sex and genotype effects in the SSC were made using a three-way analysis of variance, followed by a Bonferroni post hoc multiple comparisons test. Statistical significance was defined as a p-value < 0.05. The fold change (FC) directionality of DEGs and DEPs in the SSC and the log2FC of DEGs in primary cultures of PCNs and ASTs from Pten+/− and Nes-KO mice (FDR < 0.05) were calculated using Spearman’s correlation test.
Ethics approval and consent to participate
All of the animal procedures were approved by the Animal Experimentation and Ethics Committee of the Chinese University of Hong Kong and maintained under selective pathogen-free conditions.
Results
Transcriptomic analysis reveals increased immune response and oligodendrocyte development in somatosensory cortex
To investigate the effect of the heterozygous loss of Pten on the somatosensory cortex (SSC) at different ages, the activation state of the PI3K/mTOR signalling pathway was first characterised. Postnatal day 30 and 42 (P30 and P42) were selected for analysis, as higher spine density was observed in P29-30 Tsc2-null mice46, and autistic phenotypes were reported in multiple PTEN knockout mouse models from 6 week (P42) onward9,47. However, this analysis failed to reveal significant changes except for the elevated levels of p-S6. (Supplementary Fig. 1A,B).
It is possible that global transcriptomic and proteomic changes could be responsible for the ASD phenotypes in PHTS. We established a workflow to delineate these molecular changes in Pten-deficient mice (Fig. 1A). To delineate the molecular changes in the SSC of Pten+/− mice, RNA-sequencing (RNA-seq) data were analysed using the DESeq2 software package. Irrespective of age and sex, there were 351 differentially expressed genes (DEGs), of which 301 were upregulated and 50 were downregulated in Pten+/− mice compared with their littermate controls (Fig. 1B). Of the 10 most significant DEGs, eight were upregulated (Fig. 1B). ADGRG1, LGI3 and NEFM are known to play roles in neural development and are associated with neurological diseases, such as epilepsy (Supplementary Fig. 1C). Additional upregulated genes, such as C1QC and CSF1R, are known immune-related genes48,49. RPRM, which is involved in the p53-dependent G2 arrest of the cell cycle, was downregulated50. Furthermore, relatively more DEGs were observed in the following two groups—female mice, irrespectively of age, and mice at P42, irrespective of sex (Fig. 1B; Supplementary Fig. 1D–H).
Transcriptomic analysis reveals increased immune response and oligodendrocyte development in somatosensory cortex. (A) Workflow of current study summarizing the genotypes, stages, cell types, and molecular techniques used. (B) Number of protein coding genes that were differentially expressed (false discovery rate < 0.05) in the somatosensory cortex of male (M) and female (F) Pten+/− mice at P30 ( male: n=3 for wild-type, and n=2 for Pten+/−; female: n=3 for wild-type and Pten+/−) and P42 (n = 3 for each group). Differentially expressed genes (DEGs) are stratified by their percentage fold change (Pten+/− vs. wild-type). Age and sex-independent analysis after pooling P30M, P30F, P42M and P42F groups; M (P30 + P42), male-specific analysis after pooling P30M and P42M groups; F (P30 + P42), female-specific analysis; P30 (F + M), P30-specific analysis after pooling P30F and P30M groups; P42 (F + M), P42-specific analysis after pooling P42F and P42M groups. (C) Top five upregulated Gene Ontology (GO) terms in each group. (D) Top five downregulated GO terms in each group.
Gene Ontology (GO) and canonical pathway enrichment analyses revealed that the immune response and vasculature development were amongst the top upregulated pathways in P30 male (P30M) mice (Fig. 1C; Supplementary Fig. 1I,J). Similarly enriched terms, such as artery morphogenesis, cytokine production and leukocyte migration, were upregulated in P30 female (P30F) mice (Fig. 1C). Immune-related terms, such as dendritic cell differentiation, were also enriched in P42M mice. Ensheathment of neurons was an enriched term shared between P42M and P42F mice (Fig. 1C). Downregulated terms and pathways related to ribosomes, protein translation and protein targeting were only found in P30M mice (Fig. 1D; Supplementary Fig. 1K). In a comparison of enriched GO terms between groups, more common GO terms were found to be shared between P30M and P30F mice than between other groups (Supplementary Fig. 1I). Most of these terms were related to the inflammatory/immune response and the regulation of PI3K signalling, while terms related to oligodendrocytes were shared between all groups except P42F mice (Supplementary Fig. 1I). Taken together, the immune response appeared to be commonly enriched in P30 mice from different groups, whereas oligodendrocyte differentiation was a common transcriptomic change in all groups.
As PTEN has been extensively implicated in ASD, we compared the DEGs of the mice in this study with the weighted gene co-expression network of human brain development45. We found that modules 8, 13, 15 and 18, which correspond to the negative regulation of neuron differentiation, synaptic transmission, the response to viruses and the defense response, respectively, were significantly enriched (Supplementary Fig. 1L,M)45. The following nine genes were shared by the neuron/synaptic transmission and virus/defense response modules: LGALS3BP, SLC1A3, CYBRD1, TLN1, LPAR1, ATP1A2, SLCO2B1, CLDN11 and B2M (Supplementary Table ST1). Only module 17 was enriched with downregulated genes. This module was also related to synaptic transmission, with the following 12 DEGs shared between groups: TBRG1, GPRASP1, BEX4, SSTR1, YPEL4, ARPP21, SNCA, CDH9, OPRK1, NCALD, ZNF365 and ABI3BP. Of note, similar transcriptional alterations have been observed in the Ptenm3m4 mutant mouse model51.
Proteomic analysis of the somatosensory cortex reveals perturbations of dendritic spine development, keratinisation and hamartoma signatures
Proteomics analysis was performed using the SSCs of the same mice used for RNA-seq analysis. The P30M group was chosen, because a significant increase in spine density has been observed in Tsc2-null mice with mTOR hyperactivation46. Of the proteins with human orthologs, 545 proteins were found with Benjamini–Hochberg-corrected false discovery rates (FDRs) < 0.05, consisting of 248 upregulated and 297 downregulated proteins (Fig. 2A). By integrating the differentially expressed proteins (DEPs) and DEGs of all pooled groups, we found that only 15 were upregulated and 4 were downregulated consistently (Fig. 2B,C; Supplementary Fig. 2A,B; and Supplementary Table ST2a, ST2b). Intriguingly, by querying the Brain RNA-Seq database52,53, we observed three groups of upregulated proteins. One group, including HAVCR2, CRYBB1, C1QC, MPEG1 and TLN1, was mainly expressed in microglia and macrophages. The second group, including GSN, CERS2 and S100A6, was mainly expressed in oligodendrocytes. The third group was widely expressed in different neural cells and included GRB14 and PLIN3, which are expressed in oligodendrocytes and endothelial cells; CD82, which is expressed in oligodendrocytes, microglia, macrophages and endothelial cells; and BCAN, which is expressed in oligodendrocytes and astrocytes (Supplementary Fig. 2A,B). In contrast, the downregulated protein FXYD6 was mainly expressed in neurons and oligodendrocyte precursor cells (OPCs), and the downregulated proteins ATP2B4 and GPM6A were expressed in ASTs, neurons and OPCs (Supplementary Fig. 2A,B). These results suggest that different neural cells may be affected differently by the heterozygous loss of Pten.
Proteomic analysis of somatosensory cortex of Pten+/− mice. (A) Number of differentially expressed proteins (DEPs; false discovery rate < 0.05) stratified by the percentage fold-change in the somatosensory cortex of male Pten+/− mice (n = 3) and littermate controls (WT) (n = 3) at P30. (B & C) Upset plots show the number of overlapping differentially expressed genes (DEGs) and DEPs in somatosensory cortices, where DEPup indicates upregulated proteins and DEPdown indicates downregulated proteins. Other groups are DEGs. (D) Gene Ontology terms and pathways and (E) Human Phenotype Ontology terms significantly enriched with three or more proteins. Number in the rightmost blanket on the left indicates the total number of genes in the term/pathway. Number in the circle indicates the number of proteins involved in the enrichment.
Next, we observed that the GO and canonical pathways were only enriched with downregulated DEPs, and postsynaptic specialisation organisation and dendritic spine development were among the top neural-related pathways in the Pten+/− SSC proteome (Fig. 2D). Interestingly, keratinisation, which is one of the clinical diagnostic criteria for Cowden syndrome, was also perturbed, but its function in brain development is unknown. To understand the relationship between the DEPs and enriched pathways, we filtered out proteins with an FDR > 0.05, and then plotted the DEPs against their associated pathways (Supplementary Fig. 2C). We observed that PTEN, RELN, PSEN1, TIAM1 and PPFIA2 shared associations with postsynaptic and dendritic spine development, whereas DSC1, KRT2, KRT15, KRT1 and KRT77 were associated with keratinisation and skin development (Supplementary Table ST2c). NRIP1 inactivation has been shown to be involved in cognitive impairment in mice54 and de novo mutations in KAT555 and WASF156 are associated with cerebral malformations, seizures and developmental delay. GJA1 is predominantly located at astrocytic gap junctions. It has been shown to modulate synaptic plasticity and is predicted to be a driver of Alzheimer’s disease57. Taken together, these results indicate that downregulated proteins in the Pten+/− SSC may affect cognitive function by changing synaptic development. However, their effect on keratinisation requires further investigation.
To understand how these DEPs are associated with known phenotypes, we performed enrichment analysis using the Human Phenotype Ontology database (Fig. 2E, Supplementary Fig. 2D; Supplementary Table ST2d). Benign tumour-like growth is a diagnostic feature of PHTS. Consistent with this phenotype, the DEPs we identified were enriched with gene sets related to aberrant hypertrophic growth. Hamartoma and neoplasm of the eye were associated with KRT1, VHL and SLC25A11. Various skin abnormalities, such as hyperkeratosis, abnormality of the plantar skin of the foot, palmoplantar keratoderma, subcutaneous nodules and erythema, were associated with GJA1, KRT1, KRT2, PSEN1 and INSR. Hyperostosis, abnormalities of odontoid tissue and carious teeth are problems related to bone or hard tissues, and these were associated with common proteins, such as GJA1 and SLC24A4. Collectively, proteins downregulated in the cortex of the Pten+/− mouse model were associated with PHTS pathologies in multiple tissues.
Transcriptomic analysis of Pten-haploinsufficient primary neural cells reveals a major perturbation of immediate early genes
To understand the effect of Pten haploinsufficiency on three important neural cell types, we first investigated the PI3K pathway in primary cultures of NPCs, ASTs and PCNs. NPCs at E12-13 were chosen because they are neural lineage cells that have multiple possible cell fate determination pathways. Moreover, the level of PTEN in Pten+/− NPCs was reduced by > 70%, but the levels of p-AKT and p-S6 were not significantly elevated. (Supplementary Fig. 3A,B). These results suggest that the remaining wild-type allele in Pten+/− NPCs is sufficient to suppress aberrant PI3K hyperactivation. The level of p-AKT, but not p-S6, was significantly increased in primary ASTs from the frontal cortices of Pten+/− mice, concurrent with a greater than 70% reduction in PTEN protein levels (Supplementary Fig. 3C,D). These results suggest that Akt activity may be more sensitive to Pten depletion in primary astrocytes. Finally, for PCNs from the frontal cortices of E15.5 to E16.5 mouse embryos, Pten heterozygosity did not alter p-AKT or p-S6 levels, but PTEN levels were reduced by half as expected (Supplementary Fig. 3E,F). These findings suggest that PI3K signalling is not affected by Pten depletion in PCNs. Collectively, only very subtle or cell-type-specific effects on PI3K signalling were observed in Pten+/− neural cells.
Next, we investigated the transcriptomic changes in neural cells from Pten+/− mice in comparison with those from their littermate controls. Overall, NPCs were the least affected cells, with no DEGs identified. ASTs had 59 upregulated and 38 downregulated genes. Interestingly, PCN had only 5 upregulated genes, but 57 downregulated genes (Fig. 3A; Supplementary Fig. 3G–I). Notably, most of the downregulated DEGs in PCNs from Pten+/− mice were immediate-early response genes (IEGs), namely, Btg2, Dusp1, Erg1, Fos, Fosb, Ier2, Jun, Junb, Maff, Nr4a1, Nr4a2, Npas4, Ptp4a1 and Trib1 (GSE190879). IEGs are known to be involved in synaptic plasticity and memory formation and have been implicated in psychiatric disorders58,59,60,61. It is paradoxical that these IEGs were not detected as DEGs in the SSC from Pten+/− mice (data not shown).
Transcriptomic analysis of Pten+/− primary neural cells reveals major perturbation of immediate early genes in neurons. (A) The differentially expressed DEGs) are stratified by the percentage fold-change in primary cultures of neural progenitor cells (NPCs), astrocytes (ASTs) and primary cortical neurons (PCNs) from Pten+/− (n = 3) and wild-type littermate control (WT) (n = 3) mice. (B) The five most significantly upregulated Gene Ontology (GO) terms (false discovery rate [FDR] < 0.05) enriched in Pten+/− primary neural cells. (C) Downregulated GO terms (FDR < 0.05) enriched in Pten+/− primary neural cells. No enriched GO terms in NPC and AST. (D) The 10 most significantly upregulated canonical pathways (FDR < 0.05) enriched in Pten+/− primary neural cells. (E) Downregulated canonical pathways (FDR < 0.05) enriched in Pten+/− primary neural cells.
Enrichment analysis of GO gene sets and canonical pathways showed that NPCs had the least number of enriched terms. Only a few cellular components and pathways related to the extracellular matrix (ECM), platelet granule and cell adhesion were upregulated, and the circadian clock was the only pathway that was suppressed (Fig. 3B,D,E). More enriched GO terms and pathways were observed in ASTs (Fig. 3B,D,E; Supplementary Fig. 3H). Some of these were similar to those observed in NPCs, such as upregulated ECM-related and cell adhesion pathways. Pathways uniquely enriched in ASTs included blood circulation, muscle contraction, protein kinase and metalloendopeptidase activity, interleukin 2 family signalling and heparin sulphate/heparin metabolism, which were upregulated, and pathways related to meiosis regulation, which were downregulated. Finally, PCNs had many uniquely enriched gene sets (Fig. 3B–E; Supplementary Fig. 3I). The upregulated gene sets in PCNs were mostly related to mitochondrial function and energy production, such as ATP synthesis-coupled electron transport, the respirasome, sugar metabolism and the tricarboxylic acid cycle. Downregulated gene sets included functions such as pre-microRNA transcription by RNA polymerase II, DNA-binding transcription activator activity, long-term synaptic depression and long-term memory. Taken together, these data suggest that the heterozygous loss of Pten induces undetectable to very low gene expression and very mild functional perturbations in different neural cells.
Transcriptomic analysis of Pten-knockout primary neural cells uncovers neural-cell-specific signatures
The heterozygous loss of Pten resulted in subtle transcriptional changes. We hypothesised that the complete knockout of Pten would unambiguously reveal disease-associated DEGs. Nestin (Nes)-Cre transgenic mice were crossed with Ptenfl/fl mice to generate complete Pten knockout (Nes-KO) mice. The brains of Nes-KO embryos were visually distinguishable from littermate controls at E16.5 and were more strikingly different at P0 (Supplementary Fig. 4). Thus, Pten is required for correct brain development at the commencement of neurogenesis62. NPCs, ASTs and PCNs were cultured as aforementioned (Fig. 4A,D,G). Western blotting analysis showed that p-AKT was upregulated in all neural cells relative to control (Fig. 4B,C,E,F,H,I). Interestingly, other PTEN-related signalling molecules, such as p-S6, p-mTOR, p-GSK3β, p-FOXO1 and p-ERK, were not significantly altered in NPCs (Fig. 4B,C). In contrast, levels of p-S6 and p-GSK3β were elevated in ASTs and PCNs, which was consistent with the results of previous studies63,64. These data suggest that different neural cell types have distinct responses to Pten knockout.
Aberrant PI3-K signalling in Pten-knockout neural cells. Representative bright-field images of primary cultures of (A) neural progenitor cells (NPCs), (D) astrocytes (ASTs) and (G) neurons at DIV14. Scale bar = 0.1 mm. Red arrowheads, hypertrophic neurites. Immunoblotting of the indicated proteins in (B) NPCs, (E) ASTs and (H) neurons of Pten-knockout (Nes-KO) and littermate control (Ctrl) mice. Cortical lysate from a wild-type mouse (Ctx) was used as a control. The specific phosphorylation of p-AKT and p-GSK3-β relative to control (Ctrl) are shown in (C) NPCs, (F) ASTs and (I) PCN. An unpaired Student’s t-test was used for statistical analysis with the indicated p-values: * < 0.05, ** < 0.01, *** < 0.001 and **** < 0.0001.
Furthermore, the levels of the early neuronal marker NeuN and the astrocyte marker GFAP remained low in Nes-KO NPCs (Fig. 4B). This indicates that the loss of Pten did not affect the neural differentiation of NPCs obtained at E12-13. In ASTs, we observed elevated expression levels of GFAP in Nes-KO cells (Fig. 4E). Increased GFAP levels may suggest reactive astrogliosis, which is commonly observed in hypertrophic astrocytes17,65. Therefore, the complete knockout of Pten in ASTs may lead to the hyperactivation of the PI3K pathway and the promotion of astrogliosis. In PCNs, we noted that the levels of NF68, a major constituent of the axonal cytoskeleton66,67, were markedly increased in the Nes-KO group when the cultures were matured from 5 Days In Vitro (DIV5) to DIV14 (Fig. 4H). Moreover, hypertrophic neurons with thick neurites were observed at DIV14 (Fig. 4G). Thus, the complete depletion of Pten in neurons leads to PI3K activation and is associated with hypertrophic axonal growth.
RNA-seq analysis showed that Nes-KO NPCs had the fewest DEGs, with 1,070 upregulated and 662 downregulated protein-coding genes, but only 170 upregulated and 9 downregulated genes, based on a log2 fold-change (FC) > 1 (Fig. 5A; Supplementary Fig. 5A–E). Nes-KO ASTs had 3359 upregulated and 2615 downregulated genes, of which 1926 upregulated and 1118 downregulated genes had an absolute log2FC of >1. Strikingly, the maximal expression differences ranged from log2FC − 7.8 to + 12.7. Nes-KO PCNs had 2541 upregulated and 2462 downregulated genes, with 694 and 439 of those, respectively, showing an absolute log2FC > 1. Collectively, the transcriptomes of Nes-KO ASTs showed the greatest perturbations compared with other neural cells after the complete loss of Pten.
Transcriptomic analysis of Pten-knockout primary neural cells uncovers neural cell-specific signatures. (A) Differentially expressed genes (DEGs) in primary cultures of neural progenitor cells (NPCs)(n = 3), astrocytes (ASTs)(n = 3) and primary cortical neurons (PCNs)(n = 3) from Pten-knockout (Nes-KO) mice and littermate controls (Ctrl) (n = 3) are stratified by log2 fold-change. (B) The five most significantly upregulated Gene Ontology (GO) terms (false discovery rate [FDR] < 0.05) enriched in Nes-KO primary neural cells. (C) The five most significantly downregulated GO terms (FDR < 0.05) enriched in Nes-KO primary neural cells.
Gene set enrichment analysis showed that Nes-KO NPCs had upregulated genes related to energy metabolic processes, such as hexose catabolic, NAD and NADH metabolic processes; carbohydrate binding and gluconeogenesis (Fig. 5B; Supplementary Fig. 5A,C), and downregulated ribosome-related processes, such as ribosome and ribonucleoprotein complex biogenesis and processes involving the ribosomal subunit, preribosome, ribosome and spliceosomal complex, along with related molecular functions, such as DNA replication and rRNA binding (Fig. 5C; Supplementary Fig. 5B,C). These results indicate that upregulated energy metabolism and downregulated mRNA translation and protein expression are signatures of Nes-KO NPCs.
In Nes-KO ASTs, components of cilia and ribosomes and mitosis-related pathways, were among the most upregulated pathways (Fig. 5B; Supplementary Fig. 5A,D), whereas relatively few downregulated pathways, such as the insulin-like growth factor-1 receptor signalling pathway, endothelial cell migration, growth factor activity and the JAK-STAT signalling pathway, were identified (Fig. 5C; Supplementary Fig. 5B,D). These results point to upregulated cilial activities, mRNA translation and mitosis as the main signatures of Nes-KO ASTs.
In Nes-KO PCNs, major changes included upregulated ECM components and cell adhesion, sterol metabolic process, leukocyte transendothelial migration and MET activation of PTK2 signalling (Fig. 5B; Supplementary Fig. 5A,E), and downregulated forebrain neuron generation and differentiation (Fig. 5C; Supplementary Fig. 5B,E). Of note, most of these downregulated genes, such as Ascl168, Lhx6, Arx, Dlx1, Dlx2, Dlx569, Erbb470, Prox171 and Foxg1, are confined to GABAergic neurons or act as determinants of differentiation into inhibitory neurons72. At the same time, the levels of some secretory proteins specific to inhibitory neurons, such as RELN, ERBB4, SST and NPY, were decreased (Supplementary Fig. 6). These findings suggest that upregulated genes were involved in multiple functions, whereas downregulated genes implicated in E/I balance were the most notable signatures of Nes-KO PCNs.
Linking Pten haploinsufficiency to intelligence, cognitive function and schizophrenia
Next, we explored the potential neurological disorders and traits associated with Pten haploinsufficiency. First, the gene expression correlations between heterozygous and homozygous Pten-knockout PCNs and ASTs were compared by Spearman’s correlation analysis (Fig. 6A,B). The results showed that the DEGs between PCNs from Pten+/− and Nes-KO mice were significantly correlated (ρ = 6.1e−06, R = 0.68; Fig. 6A), but this was not observed in ASTs (Fig. 6B). Next, we curated gene lists by integrating DEGs with DEPs from the SSC of P30M Pten+/− mice. Of the different neural cells, ASTs from Nes-KO mice showed the greatest number of overlapping DEGs and DEPs, followed by PCNs from Nes-KO mice (Fig. 6C,D; Supplementary Table ST3a, ST3b). Overall, there was a lack of concordance between PCN DEGs of Pten+/− mice and SSC DEPs. This may be due to the differences between neuronal cells in vitro and in vivo. Alternatively, the complete depletion of Pten may have highly distorted the gene expression profiles, compared with those in Pten+/− mice, such that DEPs may not be detected in the SSC of Pten+/− mice.
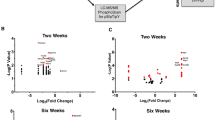
Linking Pten haploinsufficiency to intelligence, cognitive function and schizophrenia. Gene expression fold-change dimensionality comparison of common differentially expressed genes (DEGs) between Pten+/− and Nes-KO (A) primary cultures of neurons and (B) astrocytes (ASTs). R: Spearman’s correlation coefficient. Overlapping (C) upregulated and (D) downregulated differentially expressed proteins (DEPs) in the somatosensory cortex of Pten+/− mice with DEGs from Pten+/− (HET) and Nes-KO (KO) neural cells. Statistical analysis of enrichment was performed using SuperExactTest. (E) Enrichment of DEPs and DEGs in the somatosensory cortex of Pten+/− mice and translated DEGs in neural cells converging with DEPs and all translated DEGs (adding up translated DEGs from all neural cells) within genes identified in GWAS of neurological diseases and traits. Dots enclosed by a black circle represent enrichment with FDR < 0.05. Dots enclosed by a red circle represent enrichment with FDR < 0.1. Statistically significant enrichment was calculated using an FDR-corrected cumulative hypergeometric test.
Finally, to uncover the potential neurological disorders and traits associated with Pten haploinsufficiency, we curated gene lists by integrating DEPs from the SSC of Pten+/− mice and DEGs from neural cells of Pten+/− and Nes-KO mice by filtering out genes encoding non-DEPs from the DEGs of neural cells as NPC(DEP), AST(DEP), PCN(DEP) and curating translated DEGs comprising DEGs from all neural cells that overlapped with DEPs (Supplementary Table ST3c). Next, we performed enrichment analysis between the gene lists and the genetic variants associated with neurological disorders and traits obtained from genome-wide association studies (Fig. 6E; Supplementary Table ST3d)73. Overall, we found significant enrichment (FDR < 0.05) of both DEPs and translated DEGs in the traits of intelligence and cognitive function, and proteins involved in schizophrenia were significantly enriched in the DEPs (Fig. 6E; Supplementary Table ST3e). Further investigation at the cell-type level showed that AST(DEP) was significantly associated with intelligence and depression.
Discussion
In the present study, we made the following key observations (see supplementary Table ST4 for a summary). First, the levels of PTEN in Pten+/− neural cells fluctuated widely, ranging from 10 to 50%. Second, immediate-early genes were enriched as DEGs in Pten+/− PCNs, which implies an important function of PTEN in the regulation of neuronal activity. Unexpectedly, none of these IEGs were found as DEPs in the SSC of Pten+/− mice. One explanation for this finding is that the expression levels of IEGs readily change upon neuronal stimulation or depolarisation74,75. Indeed, cultured PCNs represent an unstimulated, naïve state. In contrast, the somatosensory region of mice is constantly being stimulated, for example, by the whiskers37. Another reason for this finding is that the dynamic fluctuation in PTEN levels may make downstream gene expression and protein functions too transient to be captured. To address this issue, we characterised the effect of homozygous Pten knockout on these neural cells. As expected, the number of DEGs and the number of DEGs that overlapped with DEPs were markedly increased in all neural cells. The drawback of knockout models is the generation of clinically irrelevant effects due to non-physiological transcriptomic changes and hyperactivation of the PI3K pathway. Filtering out these irrelevant genes based on DEPs in brain tissues partially addressed this issue.
A previous transcriptomic study explored the whole brain and cortical tissues from a Ptenm3m4 mutant knock-in mouse line76. These mutant mice have reduced nuclear PTEN localisation and exhibit behavioural deficits similar to those seen in high-functioning ASD. Similar to our findings, the brains of homozygous Ptenm3m4/m3m4 mutant mice contain more DEGs than those of heterozygous Ptenm3m4/+ mutant mice. In addition, we observed that Nes-KO transcriptomes contained many more DEGs that integrated with the proteomes of the SSC of Pten+/− mice. Therefore, analysing neural cells with complete Pten knockout enabled us to identify more PHTS-relevant genes based on the proteomic profile of Pten+/− mice.
Many IEGs are transcription factors and their expression is regulated by neuronal activity, such that neuronal circuits can be adjusted to meet functional needs. PTEN has been found localised to different cellular compartments. More recently, postsynaptic density-localised PTEN has been shown to interact with the synaptic scaffolding molecule, PSD-95, through a PDZ-binding motif, which is required for N-methyl-D-aspartate (NMDA) receptor-dependent long-term depression77. The excitatory neurotransmitter glutamate binds to the NMDA receptor and induces calcium influx and triggers calcium-dependent signalling, leading to changes in the expression of transcription factors, such as FOS and NPAS478. Interestingly, glutamate binds to glutamate receptors to induce protein-synthesis-dependent long-lasting synaptic plasticity by activating mTOR79. Therefore, we speculate that PTEN is involved in both IEG expression and mTOR-dependent protein synthesis.
We uncovered pathways in both ASTs and neurons that may explain the neurological disorders observed in PHTS. In Nes-KO ASTs, cilium-related activity and components and mitosis-related pathways showed the most enriched perturbations. ASTs possess a single, non-motile primary cilium80 that regulates various functions, such as cell division and signal transduction pathways, and plays important roles in sensory functions81. In fact, PTEN regulates cilial turnover by controlling dishevelled phosphorylation82. The dysregulation of ciliogenesis causes behavioural and cognitive defects83 and brain malformations84. Furthermore, Nes-KO neurons exhibited decreased forebrain neuron generation and differentiation, which may lead to disruption of the E/I balance in the brain. This included the downregulation of genes that are required for differentiation into GABAergic neurons. Similarly, conditional Pten knockout driven by Nkx2.1-Cre results in the preferential loss of SST-positive interneurons31. However, whether changes in cilia and interneurons are present in PHTS patients remains to be confirmed.
Through our DEP and translated DEGs lists, we uncovered associations between PHTS and schizophrenia, intelligence and cognitive function. In fact, a pathogenic PTEN mutation (Gln219*) has been reported in a patient with Cowden syndrome complicated by schizophrenia85. There were more than 50 proteins in our list of DEPs that matched with schizophrenia genome-wide association study (GWAS) data, and the genes encoding 18 of these were identified as DEGs in the transcriptomic analysis. One of these proteins was RELN, which is preferentially expressed in GABAergic interneurons of prefrontal cortices, temporal cortex and hippocampus86. The protein level of RELN was downregulated by 17%, but the transcript level was downregulated by 67% (GSE190879). Other PHTS cognitive manifestations, such as intellectual abilities87, were also enriched in our DEP/DEGs list. Furthermore, it is worth noting that AST(DEP) was significantly associated with depression and intelligence, which may provide a new direction for research on the function of PTEN in astrocytes.
Our study has several limitations. While the Pten+/− mouse model recapitulates most of the pathological features of PHTS, potential species-specific differences may have confounded the results. Moreover, we only analysed the SSC region. Other neuroanatomical regions may have molecular changes associated with the pathological features of PHTS. Another caveat is the possibility that neural cell types other than NPC, AST and PCN may play critical roles in PHTS. For instance, microglia have been implicated as key regulators of neuroinflammation and they play a role in neurodevelopmental and neurodegenerative disorders88. Future studies should explore the use of single-cell RNA-seq analyses of fresh brain tissues from Pten+/− mice.
In summary, we identified multiple pathways and genes that are perturbed in Pten-deficient cells and tissues. Further, by integrating transcriptomic and proteomic profiles, we identified several neurological diseases and traits that may be relevant to the neuropathogenesis of PHTS.
Data availability
Transcriptomic data: GEO accession GSE190879. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via PRIDE, under the identifier PXD030573.
Abbreviations
- AST:
-
Astrocytes
- ASD:
-
Autism spectrum disorders
- DEGs:
-
Differential expressed genes
- DEPs:
-
Differentially expressed proteins
- DCX:
-
Doublecortin
- E/I:
-
Excitatory/inhibitory
- ECM:
-
Extracellular matrix
- FDR:
-
False discovery rate
- GO:
-
Gene Ontology
- HPO:
-
Human phenotype ontology
- IEGs:
-
Immediate-early response genes
- mTOR:
-
Target of rapamycin
- NMDA:
-
N-methyl-d-aspartate
- NSE:
-
Neuron-specific enolase
- NPC:
-
Neuroprogenitor cell
- OPC:
-
Oligodendrocyte precursor cell
- PI3K:
-
Phosphoinositide 3-kinase
- PCN:
-
Primary cortical neurons
- PHTS:
-
PTEN hamartoma tumour syndrome
- SSC:
-
Somatosensory cortex
- SST:
-
Somatostatin
- SVZ:
-
Subventricular zone
References
Parsons, R. Human cancer, PTEN and the PI-3 kinase pathway. Semin. Cell Dev. Biol. 15(2), 171–176 (2004).
Butler, M. G. et al. Subset of individuals with autism spectrum disorders and extreme macrocephaly associated with germline PTEN tumour suppressor gene mutations. J. Med. Genet. 42(4), 318–321 (2005).
Goffin, A., Hoefsloot, L. H., Bosgoed, E., Swillen, A. & Fryns, J. P. PTEN mutation in a family with Cowden syndrome and autism. Am. J. Med. Genet. 105(6), 521–524 (2001).
Tilot, A. K., Frazier, T. W. 2nd. & Eng, C. Balancing proliferation and connectivity in PTEN-associated autism spectrum disorder. Neurotherapeutics 12(3), 609–619 (2015).
Tan, M. H. et al. A clinical scoring system for selection of patients for PTEN mutation testing is proposed on the basis of a prospective study of 3042 probands. Am. J. Hum. Genet. 88(1), 42–56 (2011).
Busch, R. M. et al. Neurobehavioral phenotype of autism spectrum disorder associated with germline heterozygous mutations in PTEN. Transl. Psychiatry. 9(1), 253 (2019).
Hansen-Kiss, E. et al. A retrospective chart review of the features of PTEN hamartoma tumour syndrome in children. J. Med. Genet. 54(7), 471–478 (2017).
Clipperton-Allen, A. E. & Page, D. T. Pten haploinsufficient mice show broad brain overgrowth but selective impairments in autism-relevant behavioral tests. Hum. Mol. Genet. 23(13), 3490–3505 (2014).
Clipperton-Allen, A. E. & Page, D. T. Decreased aggression and increased repetitive behavior in Pten haploinsufficient mice. Genes Brain Behav. 14(2), 145–157 (2015).
Chen, Y., Huang, W. C., Sejourne, J., Clipperton-Allen, A. E. & Page, D. T. Pten mutations alter brain growth trajectory and allocation of cell types through elevated beta-catenin signaling. J. Neurosci. 35(28), 10252–10267 (2015).
Huang, W. C., Chen, Y. & Page, D. T. Hyperconnectivity of prefrontal cortex to amygdala projections in a mouse model of macrocephaly/autism syndrome. Nat. Commun. 7, 13421 (2016).
Garcia-Junco-Clemente, P. et al. Overexpression of calcium-activated potassium channels underlies cortical dysfunction in a model of PTEN-associated autism. Proc. Natl. Acad. Sci. USA. 110(45), 18297–18302 (2013).
Li, L., He, F., Litofsky, N. S., Recht, L. D. & Ross, A. H. Profiling of genes expressed by PTEN haploinsufficient neural precursor cells. Mol. Cell Neurosci. 24(4), 1051–1061 (2003).
Podsypanina, K. et al. Mutation of Pten/Mmac1 in mice causes neoplasia in multiple organ systems. Proc. Natl. Acad. Sci. USA. 96(4), 1563–1568 (1999).
Backman, S. A. et al. Deletion of Pten in mouse brain causes seizures, ataxia and defects in soma size resembling Lhermitte-Duclos disease. Nat. Genet. 29(4), 396–403 (2001).
Fraser, M. M., Bayazitov, I. T., Zakharenko, S. S. & Baker, S. J. Phosphatase and tensin homolog, deleted on chromosome 10 deficiency in brain causes defects in synaptic structure, transmission and plasticity, and myelination abnormalities. Neuroscience 151(2), 476–488 (2008).
Fraser, M. M. et al. Pten loss causes hypertrophy and increased proliferation of astrocytes in vivo. Cancer Res. 64(21), 7773–7779 (2004).
Kwon, C. H. et al. Pten regulates neuronal soma size: A mouse model of Lhermitte-Duclos disease. Nat Genet. 29(4), 404–411 (2001).
Yue, Q. et al. PTEN deletion in Bergmann glia leads to premature differentiation and affects laminar organization. Development 132(14), 3281–3291 (2005).
Diaz-Ruiz, O. et al. Selective deletion of PTEN in dopamine neurons leads to trophic effects and adaptation of striatal medium spiny projecting neurons. PLoS ONE 4(9), e7027 (2009).
Domanskyi, A. et al. Pten ablation in adult dopaminergic neurons is neuroprotective in Parkinson’s disease models. FASEB J. 25(9), 2898–2910 (2011).
Sperow, M. et al. Phosphatase and tensin homologue (PTEN) regulates synaptic plasticity independently of its effect on neuronal morphology and migration. J. Physiol. 590(4), 777–792 (2012).
Marino, S. et al. PTEN is essential for cell migration but not for fate determination and tumourigenesis in the cerebellum. Development 129(14), 3513–3522 (2002).
Kwon, C. H. et al. Pten regulates neuronal arborization and social interaction in mice. Neuron 50(3), 377–388 (2006).
Napoli, E. et al. Mitochondrial dysfunction in Pten haplo-insufficient mice with social deficits and repetitive behavior: Interplay between Pten and p53. PLoS ONE 7(8), e42504 (2012).
Takeuchi, K. et al. Dysregulation of synaptic plasticity precedes appearance of morphological defects in a Pten conditional knockout mouse model of autism. Proc. Natl. Acad. Sci. USA. 110(12), 4738–4743 (2013).
Luikart, B. W. et al. Pten knockdown in vivo increases excitatory drive onto dentate granule cells. J. Neurosci. 31(11), 4345–4354 (2011).
Santos, V. R. et al. PTEN deletion increases hippocampal granule cell excitability in male and female mice. Neurobiol. Dis. 108, 339–351 (2017).
Williams, M. R., DeSpenza, T. Jr., Li, M., Gulledge, A. T. & Luikart, B. W. Hyperactivity of newborn Pten knock-out neurons results from increased excitatory synaptic drive. J. Neurosci. 35(3), 943–959 (2015).
Xiong, Q., Oviedo, H. V., Trotman, L. C. & Zador, A. M. PTEN regulation of local and long-range connections in mouse auditory cortex. J. Neurosci. 32(5), 1643–1652 (2012).
Vogt, D., Cho, K. K. A., Lee, A. T., Sohal, V. S. & Rubenstein, J. L. R. The parvalbumin/somatostatin ratio is increased in Pten mutant mice and by human PTEN ASD alleles. Cell Rep. 11(6), 944–956 (2015).
Nelson, S. B. & Valakh, V. Excitatory/inhibitory balance and circuit homeostasis in autism spectrum disorders. Neuron 87(4), 684–698 (2015).
Lee, Y. R., Chen, M. & Pandolfi, P. P. The functions and regulation of the PTEN tumour suppressor: New modes and prospects. Nat. Rev. Mol. Cell Biol. 19(9), 547–562 (2018).
Song, M. S., Salmena, L. & Pandolfi, P. P. The functions and regulation of the PTEN tumour suppressor. Nat. Rev. Mol. Cell Biol. 13(5), 283–296 (2012).
Zhou, J. et al. Pharmacological inhibition of mTORC1 suppresses anatomical, cellular, and behavioral abnormalities in neural-specific Pten knock-out mice. J. Neurosci. 29(6), 1773–1783 (2009).
Iannone, A. F. & De Marco Garcia, N. V. The emergence of network activity patterns in the somatosensory cortex—An early window to autism spectrum disorders. Neuroscience 466, 298–309 (2021).
Chittajallu, R. & Isaac, J. T. Emergence of cortical inhibition by coordinated sensory-driven plasticity at distinct synaptic loci. Nat. Neurosci. 13(10), 1240–1248 (2010).
Jiao, Y. et al. A key mechanism underlying sensory experience-dependent maturation of neocortical GABAergic circuits in vivo. Proc. Natl. Acad. Sci. USA. 108(29), 12131–12136 (2011).
Lesche, R. et al. Cre/loxP-mediated inactivation of the murine Pten tumor suppressor gene. Genesis 32(2), 148–149 (2002).
Groszer, M. et al. Negative regulation of neural stem/progenitor cell proliferation by the Pten tumor suppressor gene in vivo. Science 294(5549), 2186–2189 (2001).
Banker, G. G. Kimberly Culturing Nerve Cells (1998).
Paxinos, G., Franklin, K. B. J. & Franklin, K. B. J. The Mouse Brain in Stereotaxic Coordinates 2nd edn. (Academic Press, 2001).
Tian, L. et al. iTRAQ-based quantitative proteomic analysis provides insight for molecular mechanism of neuroticism. Clin. Proteomics. 16, 38 (2019).
Perez-Riverol, Y. et al. The PRIDE database and related tools and resources in 2019: Improving support for quantification data. Nucleic Acids Res. 47(D1), D442–D450 (2019).
Parikshak, N. N. et al. Integrative functional genomic analyses implicate specific molecular pathways and circuits in autism. Cell 155(5), 1008–1021 (2013).
Tang, G. et al. Loss of mTOR-dependent macroautophagy causes autistic-like synaptic pruning deficits. Neuron 83(5), 1131–1143 (2014).
Lugo, J. N. et al. Deletion of PTEN produces autism-like behavioral deficits and alterations in synaptic proteins. Front. Mol. Neurosci. 7, 27 (2014).
Sontheimer, R. D., Racila, E. & Racila, D. M. C1q: Its functions within the innate and adaptive immune responses and its role in lupus autoimmunity. J. Investig. Dermatol. 125(1), 14–23 (2005).
Olmos-Alonso, A. et al. Pharmacological targeting of CSF1R inhibits microglial proliferation and prevents the progression of Alzheimer’s-like pathology. Brain 139(Pt 3), 891–907 (2016).
Ohki, R. et al. Reprimo, a new candidate mediator of the p53-mediated cell cycle arrest at the G2 phase. J. Biol. Chem. 275(30), 22627–22630 (2000).
Tilot, A. K. et al. Neural transcriptome of constitutional Pten dysfunction in mice and its relevance to human idiopathic autism spectrum disorder. Mol. Psychiatry. 21(1), 118–125 (2016).
Zhang, Y. et al. An RNA-sequencing transcriptome and splicing database of glia, neurons, and vascular cells of the cerebral cortex. J. Neurosci. 34(36), 11929–11947 (2014).
Zhang, Y. et al. Purification and characterization of progenitor and mature human astrocytes reveals transcriptional and functional differences with mouse. Neuron 89(1), 37–53 (2016).
Duclot, F. et al. Cognitive impairments in adult mice with constitutive inactivation of RIP140 gene expression. Genes Brain Behav. 11(1), 69–78 (2012).
Humbert, J. et al. De novo KAT5 variants cause a syndrome with recognizable facial dysmorphisms, cerebellar atrophy, sleep disturbance, and epilepsy. Am. J. Hum. Genet. 107(3), 564–574 (2020).
Ito, Y. et al. De novo truncating mutations in WASF1 cause intellectual disability with seizures. Am. J. Hum. Genet. 103(1), 144–153 (2018).
Kajiwara, Y. et al. GJA1 (connexin43) is a key regulator of Alzheimer’s disease pathogenesis. Acta Neuropathol. Commun. 6(1), 144 (2018).
Minatohara, K., Akiyoshi, M. & Okuno, H. Role of immediate-early genes in synaptic plasticity and neuronal ensembles underlying the memory trace. Front. Mol. Neurosci. 8, 78 (2015).
Gallo, F. T., Katche, C., Morici, J. F., Medina, J. H. & Weisstaub, N. V. Immediate early genes, memory and psychiatric disorders: Focus on c-Fos, Egr1 and Arc. Front Behav Neurosci. 12, 79 (2018).
Marballi, K. K. & Gallitano, A. L. Immediate early genes anchor a biological pathway of proteins required for memory formation, long-term depression and risk for schizophrenia. Front. Behav. Neurosci. 12, 23 (2018).
Kandel, E. R., Dudai, Y. & Mayford, M. R. The molecular and systems biology of memory. Cell 157(1), 163–186 (2014).
Lein, E. S. et al. Genome-wide atlas of gene expression in the adult mouse brain. Nature 445(7124), 168–176 (2007).
de la Iglesia, N. et al. Identification of a PTEN-regulated STAT3 brain tumor suppressor pathway. Genes Dev. 22(4), 449–462 (2008).
Bhattacharya, K., Maiti, S. & Mandal, C. PTEN negatively regulates mTORC2 formation and signaling in grade IV glioma via Rictor hyperphosphorylation at Thr1135 and direct the mode of action of an mTORC1/2 inhibitor. Oncogenesis. 5(5), e227 (2016).
Silver, J. & Miller, J. H. Regeneration beyond the glial scar. Nat. Rev. Neurosci. 5(2), 146–156 (2004).
Hoffman, P. N. Distinct roles of neurofilament and tubulin gene expression in axonal growth. Ciba Found Symp. 138, 192–204 (1988).
Hoffman, P. N. et al. Neurofilament gene expression: A major determinant of axonal caliber. Proc. Natl. Acad. Sci. USA. 84(10), 3472–3476 (1987).
Aydin, B. et al. Proneural factors Ascl1 and Neurog2 contribute to neuronal subtype identities by establishing distinct chromatin landscapes. Nat. Neurosci. 22(6), 897–908 (2019).
Cobos, I., Long, J. E., Thwin, M. T. & Rubenstein, J. L. Cellular patterns of transcription factor expression in developing cortical interneurons. Cereb. Cortex. 16(Suppl 1), i82–i88 (2006).
Yau, H. J., Wang, H. F., Lai, C. & Liu, F. C. Neural development of the neuregulin receptor ErbB4 in the cerebral cortex and the hippocampus: Preferential expression by interneurons tangentially migrating from the ganglionic eminences. Cereb. Cortex. 13(3), 252–264 (2003).
Miyoshi, G. et al. Prox1 regulates the subtype-specific development of caudal ganglionic eminence-derived gabaergic cortical interneurons. J. Neurosci. 35(37), 12869–12889 (2015).
Mariani, J. et al. FOXG1-dependent dysregulation of GABA/glutamate neuron differentiation in autism spectrum disorders. Cell 162(2), 375–390 (2015).
Buniello, A. et al. The NHGRI-EBI GWAS catalog of published genome-wide association studies, targeted arrays and summary statistics 2019. Nucleic Acids Res. 47(D1), D1005–D1012 (2019).
Sheng, M. & Greenberg, M. E. The regulation and function of c-fos and other immediate early genes in the nervous system. Neuron 4(4), 477–485 (1990).
Ginty, D. D. et al. Regulation of CREB phosphorylation in the suprachiasmatic nucleus by light and a circadian clock. Science 260(5105), 238–241 (1993).
Tilot, A. K. et al. Germline disruption of Pten localization causes enhanced sex-dependent social motivation and increased glial production. Hum. Mol. Genet. 23(12), 3212–3227 (2014).
Jurado, S. et al. PTEN is recruited to the postsynaptic terminal for NMDA receptor-dependent long-term depression. EMBO J. 29(16), 2827–2840 (2010).
Ebert, D. H. & Greenberg, M. E. Activity-dependent neuronal signalling and autism spectrum disorder. Nature 493(7432), 327–337 (2013).
Luscher, C. & Huber, K. M. Group 1 mGluR-dependent synaptic long-term depression: Mechanisms and implications for circuitry and disease. Neuron 65(4), 445–459 (2010).
Guemez-Gamboa, A., Coufal, N. G. & Gleeson, J. G. Primary cilia in the developing and mature brain. Neuron 82(3), 511–521 (2014).
Marshall, W. F. & Nonaka, S. Cilia: Tuning in to the cell’s antenna. Curr. Biol. 16(15), R604–R614 (2006).
Shnitsar, I. et al. PTEN regulates cilia through dishevelled. Nat. Commun. 6, 8388 (2015).
Sharma, N., Berbari, N. F. & Yoder, B. K. Ciliary dysfunction in developmental abnormalities and diseases. Curr. Top. Dev. Biol. 85, 371–427 (2008).
Valente, E. M., Rosti, R. O., Gibbs, E. & Gleeson, J. G. Primary cilia in neurodevelopmental disorders. Nat. Rev. Neurol. 10(1), 27–36 (2014).
Kobayashi, Y. et al. Cowden syndrome complicated by schizophrenia: A first clinical report. Eur. J. Med. Genet. 63(8), 103959 (2020).
Pohlkamp, T. et al. Characterization and distribution of Reelin-positive interneuron subtypes in the rat barrel cortex. Cereb. Cortex. 24(11), 3046–3058 (2014).
Busch, R. M. et al. Cognitive characteristics of PTEN hamartoma tumor syndromes. Genet. Med. 15(7), 548–553 (2013).
Salter, M. W. & Stevens, B. Microglia emerge as central players in brain disease. Nat. Med. 23(9), 1018–1027 (2017).
Acknowledgements
We thank the core facilities of the School of Biomedical Sciences at the Chinese University of Hong Kong for technical supports.
Funding
This work was supported by Lo Kwee-Seong Biomedical Research Seed Fund (6903806), Hong Kong Research Grants Council, Collaborative Research Fund (C4014-14G), and Brain and Mind Institute of The Chinese University of Hong Kong Pilot Project Fund (4930742).
Author information
Authors and Affiliations
Contributions
Conceptualization: B.F., K.W.C., R.C.C.C., J.P.H.B., A.S.L.C. and A.M.C.; Methodology and investigation: S.K.K.C., J.K., P.M.Y.O., C.W.W. and A.M.C.; Formal Analysis: S.K.K.C. and A.M.C.; Writing: S.K.K.C. and A.M.C.; and supervision and funding acquisition: A.M.C.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cheung, S.K.K., Kwok, J., Or, P.M.Y. et al. Neuropathological signatures revealed by transcriptomic and proteomic analysis in Pten-deficient mouse models. Sci Rep 13, 6763 (2023). https://doi.org/10.1038/s41598-023-33869-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-33869-7
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.