Abstract
Affinity-matured plasma cells (PCs) of varying lifespans are generated through a germinal center (GC) response. The developmental dynamics and genomic programs of antigen-specific PC precursors remain to be elucidated. Here, using a model antigen in mice, we demonstrate biphasic generation of PC precursors, with those generating long-lived bone marrow PCs preferentially produced in the late phase of GC response. Clonal tracing using single-cell RNA sequencing and B cell antigen receptor sequencing in spleen and bone marrow compartments, coupled with adoptive transfer experiments, reveals a new PC transition state that gives rise to functionally competent PC precursors. The latter undergo clonal expansion, dependent on inducible expression of TIGIT. We propose a model for the proliferation and programming of precursors of long-lived PCs, based on extended antigen encounters in the GC.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All FASTQ files of scRNA-seq datasets presented and used for analysis in this study are publicly available on the Gene Expression Omnibus (GSE240813). Data collection and analysis were not performed blind to the conditions of the experiments. Source data are provided with this paper.
Code availability
The R code defining the BCR clones is provided at https://github.com/kairaveet/bcr-clones/. Other software and tools used for generation of bioinformatic analysis and plots in this study include R (3.6.3; 4.2.1), R Studio (1.4), rstatix (0.7.0), 10x Cell Ranger (3.1.0), Seurat (3.1.0; 4.1.1), ICGS2 and cellHarmony in AltAnalyze, R package CountClust (1.12.0), Monocle (2.12.0), FlowJo (0.7.1), Python (3.8.8), Plotly (4.14.3), Pandas (1.2.3; 1.2.4), Matplotlib (3.5.2), Pycircos (0.3.0), Velocyto (0.17.15), scVelo (0.2.5) and Scanpy (1.9).
References
Benner, R., van Oudenaren, A. & de Ruiter, H. Antibody formation in mouse bone marrow: IX. Peripheral lymphoid organs are involved in the initiation of bone marrow antibody formation. Cell. Immunol. 34, 125–137 (1977).
Benner, R., Hijmans, W. & Haaijman, J. J. The bone marrow: the major source of serum immunoglobulins, but still a neglected site of antibody formation. Clin. Exp. Immunol. 46, 1–8 (1981).
Slifka, M. K., Matloubian, M. & Ahmed, R. Bone marrow is a major site of long-term antibody production after acute viral infection. J. Virol. 69, 1895–1902 (1995).
Smith, K. G. C., Hewitson, T. D., Nossal, G. J. V. & Tarlinton, D. M. The phenotype and fate of the antibody-forming cells of the splenic foci. Eur. J. Immunol. 26, 444–448 (1996).
Manz, R. A., Thiel, A. & Radbruch, A. Lifetime of plasma cells in the bone marrow. Nature 388, 133–134 (1997).
Nutt, S. L., Hodgkin, P. D., Tarlinton, D. M. & Corcoran, L. M. The generation of antibody-secreting plasma cells. Nat. Rev. Immunol. 15, 160–171 (2015).
Shlomchik, M. J. & Weisel, F. Germinal center selection and the development of memory B and plasma cells. Immunol. Rev. 247, 52–63 (2012).
Amanna, I. J., Carlson, N. E. & Slifka, M. K. Duration of humoral immunity to common viral and vaccine antigens. N. Engl. J. Med. 357, 1903–1915 (2007).
Halliley, J. L. et al. Long-lived plasma cells are contained within the CD19−CD38hiCD138+ subset in human bone marrow. Immunity 43, 132–145 (2015).
Manakkat Vijay, G. K. & Singh, H. Cell fate dynamics and genomic programming of plasma cell precursors. Immunol. Rev. 303, 62–71 (2021).
Nguyen, D. C. et al. COVID-19 and plasma cells: is there long-lived protection?*. Immunol. Rev. 309, 40–63 (2022).
Weisel, F. J., Zuccarino-Catania, G. V., Chikina, M. & Shlomchik, M. J. A temporal switch in the germinal center determines differential output of memory B and plasma cells. Immunity 44, 116–130 (2016).
O’Connor, B. P., Cascalho, M. & Noelle, R. J. Short-lived and long-lived bone marrow plasma cells are derived from a novel precursor population. J. Exp. Med. 195, 737–745 (2002).
Shi, W. et al. Transcriptional profiling of mouse B cell terminal differentiation defines a signature for antibody-secreting plasma cells. Nat. Immunol. 16, 663–673 (2015).
Njau, M. N. et al. CD28–B7 interaction modulates short- and long-lived plasma cell function. J. Immunol. 189, 2758–2767 (2012).
Ochiai, K. et al. Transcriptional regulation of germinal center B and plasma cell fates by dynamical control of IRF4. Immunity 38, 918–929 (2013).
Shapiro-Shelef, M. et al. Blimp-1 is required for the formation of immunoglobulin secreting plasma cells and pre-plasma memory B cells. Immunity 19, 607–620 (2003).
Shaffer, A. L. et al. Blimp-1 orchestrates plasma cell differentiation by extinguishing the mature B cell gene expression program. Immunity 17, 51–62 (2002).
Shaffer, A. L. et al. XBP1, downstream of Blimp-1, expands the secretory apparatus and other organelles, and increases protein synthesis in plasma cell differentiation. Immunity 21, 81–93 (2004).
Lee, K. et al. IRE1-mediated unconventional mRNA splicing and S2P-mediated ATF6 cleavage merge to regulate XBP1 in signaling the unfolded protein response. Genes Dev. 16, 452–466 (2002).
Chen, D. et al. Coupled analysis of transcriptome and BCR mutations reveals role of OXPHOS in affinity maturation. Nat. Immunol. 22, 904–913 (2021).
Xu, H. et al. Regulation of bifurcating B cell trajectories by mutual antagonism between transcription factors IRF4 and IRF8. Nat. Immunol. 16, 1274–1281 (2015).
Ise, W. et al. T follicular helper cell-germinal center B cell interaction strength regulates entry into plasma cell or recycling germinal center cell fate. Immunity 48, 702–715 (2018).
Zotos, D. et al. IL-21 regulates germinal center B cell differentiation and proliferation through a B cell–intrinsic mechanism. J. Exp. Med. 207, 365–378 (2010).
Luo, W. et al. IL-21R signal reprogramming cooperates with CD40 and BCR signals to select and differentiate germinal center B cells. Sci. Immunol. 8, eadd1823 (2023).
Reiman, D. et al. Pseudocell tracer—a method for inferring dynamic trajectories using scRNAseq and its application to B cells undergoing immunoglobulin class switch recombination. PLoS Comput. Biol. 17, e1008094 (2021).
Kitamura, D., Roes, J., Kühn, R. & Rajewsky, K. A B cell-deficient mouse by targeted disruption of the membrane exon of the immunoglobulin mu chain gene. Nature 350, 423–426 (1991).
Jacob, J. & Kelsoe, G. In situ studies of the primary immune response to (4-hydroxy-3-nitrophenyl)acetyl. II. A common clonal origin for periarteriolar lymphoid sheath-associated foci and germinal centers. J. Exp. Med. 176, 679–687 (1992).
Venkatasubramanian, M., Chetal, K., Schnell, D. J., Atluri, G. & Salomonis, N. Resolving single-cell heterogeneity from hundreds of thousands of cells through sequential hybrid clustering and NMF. Bioinformatics 36, 3773–3780 (2020).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Bergen, V., Lange, M., Peidli, S., Wolf, F. A. & Theis, F. J. Generalizing RNA velocity to transient cell states through dynamical modeling. Nat. Biotechnol. 38, 1408–1414 (2020).
Blier, P. R. & Bothwell, A. L. The immune response to the hapten NP in C57BL/6 mice: insights into the structure of the B-cell repertoire. Immunol. Rev. 105, 27–45 (1988).
DePasquale, E. A. K. et al. cellHarmony: cell-level matching and holistic comparison of single-cell transcriptomes. Nucleic Acids Res. 47, e138 (2019).
Robinson, M. J. et al. Long-lived plasma cells accumulate in the bone marrow at a constant rate from early in an immune response. Sci. Immunol. 7, eabm8389 (2022).
Liu, X., Yao, J., Zhao, Y., Wang, J. & Qi, H. Heterogeneous plasma cells and long-lived subsets in response to immunization, autoantigen and microbiota. Nat. Immunol. 23, 1564–1576 (2022).
Luo, W., Weisel, F. & Shlomchik, M. J. B cell receptor and CD40 signaling are rewired for synergistic induction of the c-Myc transcription factor in germinal center B cells. Immunity 48, 313–326 (2018).
Chen, S. T., Oliveira, T. Y., Gazumyan, A., Cipolla, M. & Nussenzweig, M. C. B cell receptor signaling in germinal centers prolongs survival and primes B cells for selection. Immunity 56, 547–561 (2023).
Caven, T. H., Sturgill, J. L. & Conrad, D. H. BCR ligation antagonizes the IL-21 enhancement of anti-CD40/IL-4 plasma cell differentiation and IgE production found in low density human B cell cultures. Cell. Immunol. 247, 49–58 (2007).
Rasheed, M. A. U. et al. Interleukin-21 is a critical cytokine for the generation of virus-specific long-lived plasma cells. J. Virol. 87, 7737–7746 (2013).
Anderson, A. C., Joller, N. & Kuchroo, V. K. Lag-3, Tim-3 and TIGIT: co-inhibitory receptors with specialized functions in immune regulation. Immunity 44, 989–1004 (2016).
Xiao, S. et al. Checkpoint receptor TIGIT expressed on Tim-1+ B cells regulates tissue inflammation. Cell Rep. 32, 107892 (2020).
Hasan, M. M. et al. Implication of TIGIT+ human memory B cells in immune regulation. Nat. Commun. 12, 1534 (2021).
Joller, N. et al. Cutting edge: TIGIT has T cell-intrinsic inhibitory functions. J. Immunol. 186, 1338–1342 (2011).
Yu, X. et al. The surface protein TIGIT suppresses T cell activation by promoting the generation of mature immunoregulatory dendritic cells. Nat. Immunol. 10, 48–57 (2009).
de Hoog, C. L., Foster, L. J. & Mann, M. RNA and RNA binding proteins participate in early stages of cell spreading through spreading initiation centers. Cell 117, 649–662 (2004).
Diz-Muñoz, A. et al. Control of directed cell migration in vivo by membrane-to-cortex attachment. PLoS Biol. 8, e1000544 (2010).
Bergeman, J., Caillier, A., Houle, F., Gagné, L. M. & Huot, M. -É. Localized translation regulates cell adhesion and transendothelial migration. J. Cell Sci. 129, 4105–4117 (2016).
Zuccarino-Catania, G. V. et al. CD80 and PD-L2 define functionally distinct memory B cell subsets that are independent of antibody isotype. Nat. Immunol. 15, 631–637 (2014).
Callahan, D. et al. Memory B cell subsets have divergent developmental origins that are coupled to distinct imprinted epigenetic states. Nat. Immunol. 15, 562–575 (2024).
Olsson, A. et al. Single-cell analysis of mixed-lineage states leading to a binary cell fate choice. Nature 537, 698–702 (2016).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Acknowledgements
We thank the expert personnel within the Division of Laboratory Animal Resources, the Flow Cytometry Core, the Single Cell Core and UPMC Genome Center at the University of Pittsburgh and Westlake University for their invaluable assistance. We particularly acknowledge T. Tabib for helping with the scRNA-seq and BCR-seq library preparations and S. Hay for setting up the viewer for scRNA-seq datasets. This research was supported by grants from the UPMC ITTC fund and National Institutes of Health (U01AI141990 and RO1 AI145064 to H.S.), the National Natural Science Foundation of China (82325023 and U20A20346 to H.X.), ‘Pioneer’ and ‘Leading Goose’ R&D Program of Zhejiang (2024SSYS0031 to H.X.), HRHI program of Westlake Laboratory of Life Sciences and Biomedicine (202309001 to H.X.) and the Education Foundation of Westlake University (to H.X.).
Author information
Authors and Affiliations
Contributions
H.S. and H.X. conceived and supervised the study. G.K.M.V. and M.Z. performed all the experiments. A.R., A.S.C. and A.J. assisted with some of the immunization and adoptive transfer experiments. G.K.M.V. and K.T. performed the computational analysis. L.C.-W.L., P.H.G., D.C. and K.C. helped with the computational analysis. P.C., J.F. and J.D. helped with some of the data representation. A.J. and L.B. provided critical input during the study. N.S. supervised the computational analysis. G.K.M.V., M.Z., H.X. and H.S. wrote the paper with input from the other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks Wendy Béguelin, Rudolf Grosschedl and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: L. A. Dempsey, in collaboration with the Nature Immunology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Temporal dynamics of BMPC precursors generated during a GC response.
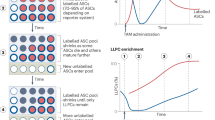
a, A representative flow plot showing the frequency of CD138 expressing cells in the enriched CD138+ splenocytes (positive selection) that were used in adoptive transfer experiments. b,c, Titers of NP-specific IgG1 antibodies in µMT recipients following adoptive transfer of CD138+ splenocytes at 28 d.p.i. (n = 6) (b) and 42 d.p.i. (n = 7) (c) at indicated d.p.t. Data are pooled from three independent experiments and shown as the mean + S.E.M (b,c).
Extended Data Fig. 2 scRNA-seq analysis of PC genomic states and trajectories.
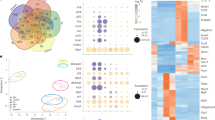
a-b, Violin plots displaying the UMI counts (a) and genes expressed per cell (b) in the scRNA-seq datasets associated with the indicated experiments and cell samples. c, UMAP plot (top left) of scRNA-seq dataset of CD138+ splenocytes (35 d.p.i.) delineating cells using the ICGS2 cluster labels as in Fig. 2. All other plots display expression levels of indicated genes within and across ICGS2 cell clusters on the same UMAP. d, Comb plots displaying the incidence and amplitude of indicated GC B cell, DZ GC and cell cycle genes in designated cluster as in Fig. 2a. e, Plots displaying the incidence and amplitude of indicated PC genes in each cluster as above. f, Plots displaying the incidence and amplitude of indicated marker genes distinguishing the three PC clusters as above. g, Bar plots displaying key gene ontology pathways associated with the top 100 marker genes in PC progenitors, PC-Tigit* and PC-Lag3* clusters with their associated p values. h, Individual plots displaying the indicated GC B cell and PC clusters in the developmental trajectory constructed by Monocle 2 as in Fig. 2c. Statistical significance was tested by Fisher exact test and Benjamini-Hochberg method for correction of multiple hypotheses testing (g).
Extended Data Fig. 3 Delineation of antigen-specific PC genomic states and clonal tracking.
a, Experimental design for isolation and analysis of NP-specific B220+ cells from C57BL6/J mice (14 d.p.i.), analyzed by scRNA-seq and BCR-seq. b, Plot displaying the rank order of top 25 IGHV genes expressed in NP-specific B220+ cells (14 d.p.i.). c, d, Plots displaying the rank order of top 50 IGHV genes expressed in CD138+ splenocytes on 35 d.p.i. (c) and 21 d.p.i. (d). Red bars are NP-specific IGHV genes expressed on 14, 21 and 35 d.p.i., blue bars are presumptive KLH-specific IGHV genes expressed on 21 and 35 d.p.i. e, Mutation frequencies of IGHV1-72*01 gene in splenic PC progenitors (left) and PC clusters (right) on 21 and 35 d.p.i. f, Pie-charts displaying the proportion of IGHV1-72*01 gene segments harboring high affinity W33L mutations in splenic PCs (21 d.p.i. vs. 35 d.p.i.). g, Combined heatmap of genomic states in the enriched CD138+ splenocytes from both 21 and 35 d.p.i., generated using cellHarmony, for which CD138+ splenocytes (35 d.p.i.) were used as a reference dataset. Columns in heatmap represent cells from 21 and 35 d.p.i. (n = 14,068); rows represent MarkerFinder genes (n = 413). h, Heatmap of differentially expressed genes (DEGs) in NP- and KLH-specific PC progenitors and indicated PC genomic states (35 vs. 21 d.p.i.) generated using cellHarmony, for which 35 d.p.i. were used as a reference dataset. Rows represent the expression of differentially expressed genes on 35 d.p.i. Statistical significance was tested by two-tailed Mann-Whitney test (e).
Extended Data Fig. 4 Functional and genomic analysis of BMPC precursors within splenic CD138+ subsets.
a, Representative t-SNE plots based on flow cytometry data showing the indicated markers in the enriched CD138+ splenocytes obtained using positive selection at 35 d.p.i. b, Representative flow plot showing the gating strategy for sorting CD138+ splenic subsets (35 d.p.i.) used in adoptive transfer experiments. Purity of the indicated PC subsets are displayed.
Extended Data Fig. 5 Clonal tracking of antigen-specific PC precursors that migrate from spleen to bone marrow.
a, Experimental design enabling clonal tracking of PC precursors that migrate from spleen to bone marrow of NP-KLH immunized mice (35 d.p.i.). Coupled scRNA-seq and BCR-seq was performed on indicated cells (including PC progenitors), within each compartment, isolated by flow cytometry. b, Circos plot displaying clones and their genomic states in spleen and bone marrow. Colored bars denote distinctive ICGS2 delineated genomic states in spleen and bone marrow. Colored lines represent clones that contain cells with identical V(D)J rearrangements that span two or more genomic states. c, Heatmap displaying the frequencies of clones spanning indicated genomic states in the spleen and the bone marrow.
Extended Data Fig. 6 Inducible TIGIT expression demarcates splenic PC precursors.
a, Representative flow plots showing TIGIT expression in splenic GC B cells and PC subsets from NP-KLH immunized C57Bl6/J mice (35 d.p.i.) in the absence of LPS. b, Representative flow plots showing TIGIT and LAG3 expression in splenic CD138+ cells with isotype controls from C57Bl6/J mice at steady state. c, Representative flow plot showing the gating strategy for sorting TIGIT+ and TIGIT- splenic PC subsets from immunized C57Bl6/J mice (35 d.p.i.) used in adoptive transfer experiments. d, Representative flow plots showing the frequencies of KI-67+ cells in the B220int plasma cell subsets based on TIGIT expression in immunized C57Bl6/J mice (35 d.p.i.). e, Representative flow plot showing the frequency of TIGIT+ cells in NP-specific splenic PCs from B1-8i+/- Balbc/J mice at indicated timepoints. The frequencies displayed in the plots were calculated from the CD44+CD11a+ subset gated from the B220intB220-CD138+NPsurface-NPcytoplasmic+ splenocytes. Gates for each plot were based on fluroscence minus one (FMO).
Extended Data Fig. 7 Tigit deficiency impairs generation of PCs.
a, Experimental design for the bone marrow chimeric model. b, Representative flow plots showing the proportions of CD45.1+ or CD45.2+ cells in each cell compartment of the chimeras. c, d, Quantification of the normalized proportions of CD45.2+ cells in the indicated cell compartments in the spleen (c) and bone marrow (d) of the chimeras. The proportions of CD45.2+ cells in individual compartments were normalized to the proportion of CD45.2+ cells within naïve B cell subset in the spleen of each mouse. e,f, ELISpot analysis of NP-specific IgG+, IgA+ and IgM+ASCs (Tigit+/+ or Tigit−/−) in spleen (e) and bone marrow compartment (f) of chimeric mice after NP-KLH immunization (35 d.p.i.). Splenic and bone marrow cells were sorted based on CD45.1 or CD45.2 expression and plated for ELISpot analyses. g,h, Quantification of the mean fluorescence intensity of TIGIT+ cells (g) and frequency of EdU+ cells (h) in the indicated CD138+ subsets. Each symbol represents an individual mouse (c-h). Data are pooled from two independent experiments and shown as the mean (e-h) or three independent experiments and shown as mean + S.E.M (c,d). Statistical significance was tested by two-tailed Unpaired t-test (c-f) and Ordinary one-way ANOVA with Tukey’s multiple comparison test (g,h).
Extended Data Fig. 8 Dynamics of PC precursors emanating from a germinal center.
The schematic depicts the interplay between CD40, BCR, IL-21 and TIGIT signaling in the generation of PC precursors that emanate from a GC and migrate to the bone marrow. During the peak of the GC response, BCR and CD40 signaling favor cycling of B cells between LZ and DZ and the process of affinity maturation. At later times in the immune response as antigen becomes limiting and BCR signaling begins to diminish, CD40 and IL-21 favor the exit of B cells from the GC and their fate specification as PC progenitors which subsequently give rise to PC precursors. Only the latter are competent to leave the spleen and home to the bone marrow. Concomitantly, extended antigen encounters by B cells in the GC induce the expression of TIGIT which is proposed to antagonize BCR signaling thereby promoting GC exit, in conjunction with CD40 and IL-21. TIGIT signaling additionally promotes the proliferation of PC precursors. Created with BioRender.com.
Supplementary information
Supplementary Tables 1–7
Supplementary Table 1 Information of tissue source, cells used, 10x genomics chemistry and QC metrics for all scRNA-seq and BCR-seq samples (Extended Data Fig. 2a,b). Supplementary Table 2 Top 100 marker genes in the ICGS2-based cell clusters of CD138+ splenocytes isolated on 35 d.p.i. and analyzed using scRNA-seq (Fig. 2a). Supplementary Table 3a GO pathways from the Enricher output associated with the top 100 marker genes in PC progenitors on 35 d.p.i. (Extended Data Fig. 2g). Supplementary Table 3b GO pathways from the Enricher output associated with the top 100 marker genes in PC-Tigit* on 35 d.p.i. (Extended Data Fig. 2g). Supplementary Table 3c GO pathways from the Enricher output associated with the top 100 marker genes in PC-Slpi* on 35 d.p.i. Supplementary Table 3d GO pathways from the Enricher output associated with the top 100 marker genes in PC-Lag3* on 35 d.p.i. (Extended Data Fig. 2g). Supplementary Table 4 Top 25 IGHV genes (14 d.p.i.) and top 50 IGHV genes (21 and 35 d.p.i.) analyzed by BCR-seq using NP-specific B cells (14 d.p.i.) and CD138+ splenocytes (21 and 35 d.p.i.), respectively (Extended data Fig. 3b–d). Supplementary Table 5a NP-specific and KLH-specific clones within the CD138+ splenocytes (35 d.p.i.) (Fig. 3c,d). Supplementary Table 5b NP-specific and KLH-specific clones within the CD138+ splenocytes (21 d.p.i.). Supplementary Table 6a DEGs in antigen-specific PC progenitors within the CD138+ splenocytes (35 versus 21 d.p.i.) (Extended Data Fig. 3h). Supplementary Table 6b DEGs in antigen-specific PC-Tigit* within the CD138+ splenocytes (35 versus 21 d.p.i.) (Extended Data Fig. 3h). Supplementary Table 6c GO pathways from the Enricher output associated with the upregulated DEGs in NP-specific and KLH-specific PC progenitors on 35 d.p.i. (Fig. 3e). Supplementary Table 6d GO pathways from the Enricher output associated with the output of upregulated DEGs in NP-specific and KLH-specific PC-Tigit* on 35 d.p.i. (Fig. 3f). Supplementary Table 7a Traceable NP-specific and KLH-specific clones from spleen to bone marrow within the B220intCD138+CD44+CD11a+ and B220−CD138+CD44+CD11a+ subsets (35 d.p.i.) (Fig. 5b). Supplementary Table 7b Traceable NP-specific and KLH-specific clones from spleen to bone marrow within the B220+CD138+CD44+CD11a+, B220intCD138+CD44+CD11a+ and B220−CD138+CD44+CD11a+ subsets (35 d.p.i.) (Extended Data Fig. 5b).
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Next-generation sequencing source data.
Source Data Fig. 3
Next-generation sequencing source data.
Source Data Fig. 4
Next-generation sequencing and statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 3
Next-generation sequencing source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Manakkat Vijay, G.K., Zhou, M., Thakkar, K. et al. Temporal dynamics and genomic programming of plasma cell fates. Nat Immunol (2024). https://doi.org/10.1038/s41590-024-01831-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41590-024-01831-y