Abstract
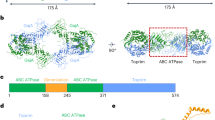
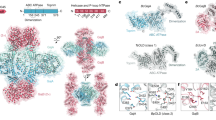
Prokaryotes have evolved intricate innate immune systems against phage infection1,2,3,4,5,6,7. Gabija is a highly widespread prokaryotic defence system that consists of two components, GajA and GajB8. GajA functions as a DNA endonuclease that is inactive in the presence of ATP9. Here, to explore how the Gabija system is activated for anti-phage defence, we report its cryo-electron microscopy structures in five states, including apo GajA, GajA in complex with DNA, GajA bound by ATP, apo GajA–GajB, and GajA–GajB in complex with ATP and Mg2+. GajA is a rhombus-shaped tetramer with its ATPase domain clustered at the centre and the topoisomerase–primase (Toprim) domain located peripherally. ATP binding at the ATPase domain stabilizes the insertion region within the ATPase domain, keeping the Toprim domain in a closed state. Upon ATP depletion by phages, the Toprim domain opens to bind and cleave the DNA substrate. GajB, which docks on GajA, is activated by the cleaved DNA, ultimately leading to prokaryotic cell death. Our study presents a mechanistic landscape of Gabija activation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Cryo-EM maps and the atomic coordinates of GajA have been deposited in the Electron Microscopy Data Bank (EMDB) and Protein Data Bank (PDB) under the accession codes EMD-36541 and 8JQ9. Cryo-EM maps and the atomic coordinates of ATP-bound GajA have been deposited in the EMDB and PDB under the accession codes EMD-37915 and 8WY4. Cryo-EM maps and the atomic coordinates of GajA–dsDNA have been deposited in the EMDB and PDB under the accession codes EMD-37916 and 8WY5. Cryo-EM maps and the atomic coordinates of dsDNA-bound GajA dimer (focused refinement) have been deposited in the EMDB and PDB under the accession codes EMD-38058 and 8X51. Cryo-EM maps and the atomic coordinates of the 4:1 GajA–GajB complex have been deposited in the EMDB and PDB under the accession codes EMD-36569 and 8JQC. Cryo-EM maps and the atomic coordinates of the 4:4 GajA–GajB complex have been deposited in the EMDB and PDB under the accession codes EMD-36563 and 8JQB. Cryo-EM maps and the atomic coordinates of ATP and Mg2+-bound GajA–GajB have been deposited in the EMDB and PDB under the accession codes EMD-38071 and 8X5N. Cryo-EM maps and the atomic coordinates of ATP and Mg2+-bound GajA (focused refinement) have been deposited in the EMDB and PDB under the accession codes EMD-38070 and 8X5I. Source data are provided with this paper.
References
Stern, A. & Sorek, R. The phage-host arms race: shaping the evolution of microbes. BioEssays 33, 43–51 (2011).
Hampton, H. G., Watson, B. N. J. & Fineran, P. C. The arms race between bacteria and their phage foes. Nature 577, 327–336 (2020).
Dy, R. L., Richter, C., Salmond, G. P. C. & Fineran, P. C. Remarkable mechanisms in microbes to resist phage infections. Annu. Rev. Virol. 1, 307–331 (2014).
Labrie, S. J., Samson, J. E. & Moineau, S. Bacteriophage resistance mechanisms. Nat. Rev. Microbiol. 8, 317–327 (2010).
Millman, A. et al. An expanded arsenal of immune systems that protect bacteria from phages. Cell Host Microbe 30, 1556–1569.e1555 (2022).
Rousset, F. et al. Phages and their satellites encode hotspots of antiviral systems. Cell Host Microbe 30, 740–753.e745 (2022).
Vassallo, C. N., Doering, C. R., Littlehale, M. L., Teodoro, G. I. C. & Laub, M. T. A functional selection reveals previously undetected anti-phage defence systems in the E. coli pangenome. Nat. Microbiol. 7, 1568–1579 (2022).
Doron, S. et al. Systematic discovery of antiphage defense systems in the microbial pangenome. Science 359, eaar4120 (2018).
Cheng, R. et al. A nucleotide-sensing endonuclease from the Gabija bacterial defense system. Nucleic Acids Res. 49, 5216–5229 (2021).
Tesson, F. et al. Systematic and quantitative view of the antiviral arsenal of prokaryotes. Nat. Commun. 13, 2561 (2022).
Payne, L. J. et al. PADLOC: a web server for the identification of antiviral defence systems in microbial genomes. Nucleic Acids Res. 50, W541–W550 (2022).
Aravind, L., Leipe, D. D. & Koonin, E. V. Toprim—a conserved catalytic domain in type IA and II topoisomerases, DnaG-type primases, OLD family nucleases and RecR proteins. Nucleic Acids Res. 26, 4205–4213 (1998).
Koonin, E. V. & Gorbalenya, A. E. The superfamily of UvrA-related ATPases includes three more subunits of putative ATP-dependent nucleases. Protein Seq. Data Anal. 5, 43–45 (1992).
Berger, J. M., Gamblin, S. J., Harrison, S. C. & Wang, J. C. Structure and mechanism of DNA topoisomerase II. Nature 379, 225–232 (1996).
Allemand, F., Mathy, N., Brechemier-Baey, D. & Condon, C. The 5S rRNA maturase, ribonuclease M5, is a Toprim domain family member. Nucleic Acids Res. 33, 4368–4376 (2005).
Keck, J. L., Roche, D. D., Lynch, A. S. & Berger, J. M. Structure of the RNA polymerase domain of E. coli primase. Science 287, 2482–2486 (2000).
Podobnik, M., McInerney, P., O’Donnell, M. & Kuriyan, J. A TOPRIM domain in the crystal structure of the catalytic core of Escherichia coli primase confirms a structural link to DNA topoisomerases. J. Mol. Biol. 300, 353–362 (2000).
Corbett, K. D. & Berger, J. M. Structure, molecular mechanisms, and evolutionary relationships in DNA topoisomerases. Annu. Rev. Biophys. Biomol. Struct. 33, 95–118 (2004).
Schiltz, C. J., Lee, A., Partlow, E. A., Hosford, C. J. & Chappie, J. S. Structural characterization of class 2 OLD family nucleases supports a two-metal catalysis mechanism for cleavage. Nucleic Acids Res. 47, 9448–9463 (2019).
Schiltz, C. J., Adams, M. C. & Chappie, J. S. The full-length structure of Thermus scotoductus OLD defines the ATP hydrolysis properties and catalytic mechanism of class 1 OLD family nucleases. Nucleic Acids Res. 48, 2762–2776 (2020).
Cheng, R. et al. Prokaryotic Gabija complex senses and executes nucleotide depletion and DNA cleavage for antiviral defense. Cell Host Microbe 31, 1331–1344 e1335 (2023).
Oh, H. et al. Structural and functional investigation of GajB protein in Gabija anti-phage defense. Nucleic Acids Res. 51, 11941–11951 (2023).
Champoux, J. J. DNA topoisomerases: structure, function, and mechanism. Annu. Rev. Biochem. 70, 369–413 (2001).
Laponogov, I. et al. Trapping of the transport-segment DNA by the ATPase domains of a type II topoisomerase. Nat. Commun. 9, 2579 (2018).
Dong, K. C. & Berger, J. M. Structural basis for gate-DNA recognition and bending by type IIA topoisomerases. Nature 450, 1201–1205 (2007).
Lee, J. Y. & Yang, W. UvrD helicase unwinds DNA one base pair at a time by a two-part power stroke. Cell 127, 1349–1360 (2006).
Lopatina, A., Tal, N. & Sorek, R. Abortive infection: bacterial suicide as an antiviral immune strategy. Annu. Rev. Virol. 7, 371–384 (2020).
Sather, L. M. et al. A broadly distributed predicted helicase/nuclease confers phage resistance via abortive infection. Cell Host Microbe 31, 343–355.e345 (2023).
Millman, A., Melamed, S., Amitai, G. & Sorek, R. Diversity and classification of cyclic-oligonucleotide-based anti-phage signalling systems. Nat. Microbiol. 5, 1608–1615 (2020).
Cohen, D. et al. Cyclic GMP-AMP signalling protects bacteria against viral infection. Nature 574, 691–695 (2019).
Gao, L. et al. Diverse enzymatic activities mediate antiviral immunity in prokaryotes. Science 369, 1077–1084 (2020).
Gao, L. A. et al. Prokaryotic innate immunity through pattern recognition of conserved viral proteins. Science 377, eabm4096 (2022).
Wu, H. Higher-order assemblies in a new paradigm of signal transduction. Cell 153, 287–292 (2013).
Burroughs, A. M. & Aravind, L. Identification of uncharacterized components of prokaryotic immune systems and their diverse eukaryotic reformulations. J. Bacteriol. 202, e00365–20 (2020).
Ye, Q. Z. et al. HORMA domain proteins and a Trip13-like ATPase regulate bacterial cGAS-like enzymes to mediate bacteriophage immunity. Mol. Cell 77, 709–722.e7 (2020).
Bernheim, A. et al. Prokaryotic viperins produce diverse antiviral molecules. Nature 589, 120–124 (2021).
Hogrel, G. et al. Cyclic nucleotide-induced helical structure activates a TIR immune effector. Nature 608, 808–812 (2022).
Johnson, A. G. et al. Bacterial gasdermins reveal an ancient mechanism of cell death. Science 375, 221–225 (2022).
Kibby, E. M. et al. Bacterial NLR-related proteins protect against phage. Cell 186, 2410–2424.e18 (2023).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Sanchez-Garcia, R. et al. DeepEMhancer: a deep learning solution for cryo-EM volume post-processing. Commun. Biol. 4, 874 (2021).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Mirdita, M. et al. ColabFold: making protein folding accessible to all. Nat. Methods 19, 679–682 (2022).
Drozdetskiy, A., Cole, C., Procter, J. & Barton, G. J. JPred4: a protein secondary structure prediction server. Nucleic Acids Res. 43, W389–W394 (2015).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D 75, 861–877 (2019).
Williams, C. J. et al. MolProbity: More and better reference data for improved all-atom structure validation. Protein Sci. 27, 293–315 (2018).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Mazzocco, A., Waddell, T. E., Lingohr, E. & Johnson, R. P. Enumeration of bacteriophages using the small drop plaque assay system. Methods Mol. Biol. 501, 81–85 (2009).
Kropinski, A. M., Mazzocco, A., Waddell, T. E., Lingohr, E. & Johnson, R. P. Enumeration of bacteriophages by double agar overlay plaque assay. Methods Mol. Biol. 501, 69–76 (2009).
Acknowledgements
The authors thank H. Wu and C. Dong for critical feedback; and C. Zhao for lending us bench space. Cryo-EM data were collected with the assistance of D. Li, X. Li and Y. Zeng at the Core Facility of Wuhan University. This work was supported by National Key R&D Program of China (2022YFA0912200 and 2022YFA0912202), a startup fund from Wuhan University to Longfei Wang, Large-scale Instrument And Equipment Sharing Foundation of Wuhan University, National Natural Science Foundation of China (grant 32150009 to B.Z., 32100025 to R.C.), and fund from Science, Technology and Innovation Commission of Shenzhen Municipality (grant JCYJ20210324115811032 to B.Z.).
Author information
Authors and Affiliations
Contributions
Longfei Wang and B.Z conceived the project. J.L., R.C., Z.W., W.Y., J.X. and X.Z. purified proteins. J.L. prepared the cryo-EM samples and performed data collection. Longfei Wang and J.L. solved the cryo-EM structures. X.D. and Longfei Wang carried out structure predictions. Z.W. and Longfei Wang performed model building. J.L. and W.Y. performed molecular cloning for mutants. Longfei Wang and Z.W. analysed the structures and designed experiments. R.C. performed phage, ATPase, gel shift and DNA cleavage assays. Longfei Wang, J.L. and Z.W. wrote the manuscript with input from R.C., W.Y., S.X., Lianrong Wang and B.Z.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Jack Bravo, Ryan Jackson and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 DNA cleavage activity and 3D reconstruction of GajA.
a, Full-length GajA was expressed in E.coli BL21 (DE3) cells and was purified using Ni-NTA agarose. SDS-PAGE showing the fractions collected at each step of the purification with concentrations of imidazole labeled in eluate. Images are representative of three biological replicates. b, The eluate was subjected to size-exclusion chromatography (10/300 Superose 6). The molecular weight of GajA tetramer was about 276 kDa. The retention volume of rabbit aldolase (158 kDa) is indicated by the dotted line. c, The protein purity was visualized by SDS-PAGE followed by Coomassie-blue staining. Images are representative of three biological replicates. d, The recognize sequence and cleavage site of GajA. e, Cleavage of pUC19-955 DNA by GajA in the presence of 1 mM Mg2+. The assays were repeated three times with similar results. f, Flow chart of cryo-EM data processing and 3D reconstruction of GajA. g, The gold standard threshold FSC curve for the map of GajA and the map to model FSC curve for model refined against the overall map of GajA. h, The orientation distribution plot of the 3D reconstruction of GajA. i, The final model of GajA tetramer fitted in the density. j, Cryo-EM density validation and close-up view of Toprim/ATPase dimer interface. The Cryo-EM densities are displayed as gray mesh. k, Cryo-EM density validation and close-up view of the ATPase dimer interface. l, Residues in the GajA Toprim active site. m, Metal coordinating residues in the BpOLD active site.
Extended Data Fig. 2 The effect on phage resistance and DNA cleavage activity of GajA mutations.
a, Structure-guided alignment of GajA proteins from indicated bacteria colored according to amino acid conservation. The determined Bacillus cereus GajA secondary structure is displayed, and active-site and interface residues are annotated according to the key below. b, Plaque assay with 10-fold serial dilutions using T4 and T7 phages to infect E. coli B expressing the WT Gabija system and indicated mutants of GajA. Images are representative of three biological replicates. c, The purified GajA and GajA E117K/D135R, E397K/E399K and E388K/D392R/E529K mutants were visualized by SDS-PAGE followed by Coomassie-blue staining. d, The molecular weight of GajA and indicated mutants were shown on native PAGE. e, GajA-E399K was subjected to size-exclusion chromatography (10/300 Superose 6) (Top). The line-dashed line indicated GajA tetramer was about 276 kDa. The retention volume of rabbit aldolase (158 kDa) is indicated by the dotted line. SDS-PAGE gel at the bottom shows the fractions of GajA-E399K collected in size-exclusion chromatography. For c–e, images are representative of three biological replicates. f, g, Cleavage of pUC19-955 DNA by GajA or indicated mutants in the presence of 1 mM Mg2+. The assays were repeated three times with similar results.
Extended Data Fig. 3 3D reconstruction of the GajA/DNA complex.
a, The sequences of designed dsDNA substrates in different lengths. b, The recognition sequence and cleavage sites of dsDNA substrate for GajA. c, Cleavage of the designed dsDNA substrates by GajA in the presence of 1 mM Mg2+. The assays were repeated three times with similar results. d, Flow chart of cryo-EM data processing and 3D reconstruction of the GajA/DNA complex. e, The gold standard threshold FSC curve for the map of the GajA/DNA complex and the map to model FSC curve for model refined against the overall map of the GajA/DNA complex. f, The orientation distribution plot of the 3D reconstruction of the GajA/DNA complex. g, The final model of GajA/DNA complex fitted in the density. h, Local refinement of the GajA/DNA complex. i, The gold standard threshold FSC curve for the map of the GajA/DNA complex after local refinement and the map to model FSC curve for model refined against the overall map of the GajA/DNA complex after local refinement. j, The orientation distribution plot of the 3D reconstruction of the GajA/DNA complex after local refinement. k, The final model of GajA/DNA complex after local refinement fitted in the density. l, Cryo-EM density validation and close-up view of interface between KxxK motif (cyan) and bound-DNA. m, Cryo-EM density validation and close-up view of catalytic residues at the Toprim active site (cyan) that can interact with bound-DNA via Ca2+ (gray).
Extended Data Fig. 4 Interaction between the GajA tetramer and dsDNA.
a, A schematic of the intermolecular interactions between GajA and the 21-bp dsDNA in the GajA/DNA complex. b–e, Cleavage of pUC19-955 DNA by GajA or indicated mutants in the presence of 1 mM Mg2+. f, Cleavage of pUC19 plasmid by GajA or indicated mutants in the presence of 1 mM Mg2+. g, h, Binding of pUC19-955 DNA by GajA or indicated mutants in the presence of 5 mM Ca2+. For b–h, the assays were repeated three times with similar results. i, Plaque assay with 10-fold serial dilutions using T4 and T7 phages to infect E. coli B expressing the WT Gabija system and indicated mutants of GajA. Images are representative of three biological replicates.
Extended Data Fig. 5 Structure-guided alignment and active site of GajB.
a, Structure-guided alignment of GajB proteins from indicated bacteria colored according to amino acid conservation. The determined Bacillus cereus GajB secondary structure is displayed, and active-site and GajB-GajA interface residues are annotated according to the key below. b, Residues in the GajB active site. c, Residues in the EcUvrD active site.
Extended Data Fig. 6 3D reconstruction of the 4:1 GajA-GajB complex and 4:4 GajA-GajB complex.
a, Flow chart of cryo-EM data processing and 3D reconstruction of the 4:1 GajA-GajB complex. b, The gold standard threshold FSC curve for the map of the 4:1 GajA-GajB complex and the map to model FSC curve for model refined against the overall map of the 4:1 GajA-GajB complex. c, The orientation distribution plot of the 3D reconstruction of the 4:1 GajA-GajB complex. d, The final model of the 4:1 GajA-GajB complex fitted in the density. e, Flow chart of cryo-EM data processing and 3D reconstruction of the 4:4 GajA-GajB complex. f, The gold standard threshold FSC curve for the map of the 4:4 GajA-GajB complex and the map to model FSC curve for model refined against the overall map of the 4:4 GajA-GajB complex. g, The orientation distribution plot of the 3D reconstruction of the 4:4 GajA-GajB complex. h, The final model of the 4:4 GajA-GajB complex fitted in the density. i, j, Cryo-EM density validation and close-up view of GajA-GajB interfaces at 1B (i) and 2B (j) domains of GajB.
Extended Data Fig. 7 The function of GajB and GajB-Q155A/Y347A.
a, SDS-PAGE showing the fractions collected at each step of the purification of GajB. b, The eluate of GajB was subjected to size-exclusion chromatography (16/600 Superdex 200). The molecular weight of GajB was about 59.9 kDa. c, The protein purity of GajB was visualized by SDS-PAGE. d, Overlaid structures of 4:4 GajA-GajB complex and EcUvrD with DNA. e, f, Superimposed structures of GajB (colored) and EcUvrD in apo (gray) (e) or EcUvrD with DNA (gray) (f). g–i, Structures of GajB (g), EcUvrD in apo (h), and EcUvrD with DNA (i). j, The diagram of various Gabija constructs. k, The protein expression levels of GajA and GajB in different constructs are indicated in the SDS-PAGE gel. l, SDS-PAGE gel showing purified GajA-GajB (His-tagged GajA and co-purified non-tagged GajB) and GajA+B (His-tagged GajA and co-purified non-tagged GajB). m, Plaque assay with 10-fold serial dilutions using T4 and T7 phages to infect E. coli B expressing the WT Gabija system and indicated mutants. Images are representative of three biological replicates. n, Superimposed cryo-EM structures of the 4:1 GajA-GajB complex (brown) and 4:4 GajA-GajB complex (gray). o, SDS-PAGE showing the fractions collected at each step of the purification of GajB-Q155A/Y347A. p, The eluate of GajB-Q155A/Y347A was subjected to size-exclusion chromatography (16/600 Superdex 200). The molecular weight of GajB-Q155A/Y347A was about 59.7 kDa. q, The protein purity of GajB-Q155A/Y347A was visualized by SDS-PAGE. r, GajA with GajB or GajB-Q155A/Y347A at different molar ratio was incubated and shown on native PAGE. s, t, Co-purification of His-tagged GajA and non-tagged GajB (s) or GajB-Q155A/Y347A (t) from bacteria expressing the WT GajA-GajB gene cassette under T7 bacterial promoter (schematic is shown above the gel). SDS-PAGE showing the fractions collected at each step during the purification, different concentrations of imidazole in eluate has been labeled. For a, c, k–l, o, q–t, images are representative of three biological replicates.
Extended Data Fig. 8 3D reconstruction of the ATP-bound GajA complex and ATP-bound GajA-GajB complex.
a, Flow chart of cryo-EM data processing and 3D reconstruction of the ATP-bound GajA-GajB complex. b, The gold standard threshold FSC curve for the map of the ATP-bound GajA-GajB complex and the map to model FSC curve for model refined against the overall map of the ATP-bound GajA-GajB complex. c, The orientation distribution plot of the 3D reconstruction of the ATP-bound GajA-GajB complex. d, Local refinement of the ATP-bound GajA-GajB complex. e, The gold standard threshold FSC curve for the map of the ATP-bound GajA-GajB complex after local refinemen and the map to model FSC curve for model refined against the overall map of the ATP-bound GajA-GajB complex after local refinement. f, The orientation distribution plot of the 3D reconstruction of the ATP-bound GajA-GajB complex after local refinement. g, The final model of the ATP-bound GajA-GajB complex after local refinement fitted in the density. h, Flow chart of cryo-EM data processing and 3D reconstruction of the ATP-bound GajA complex. i, The gold standard threshold FSC curve for the map of the ATP-bound GajA complex and the map to model FSC curve for model refined against the overall map of the ATP-bound GajA complex. j, The orientation distribution plot of the 3D reconstruction of the ATP-bound GajA complex. k, The final model of the ATP-bound GajA complex fitted in the density.
Extended Data Fig. 9 Interaction between the GajA tetramer and ATP.
a, Titration of ATP in various concentrations on GajA endonuclease activity in the presence of 5 mM Mg2+. b, c, Cryo-EM density validation of GajA tetramer and close-up view of insertion region. d, e, Cryo-EM density validation and close-up view of the ATP binding pocket at the ABC ATPase domain. f–h, Cleavage of pUC19-955 DNA by GajA or indicated mutants in the presence of 1 mM Mg2+ and with or without 0.5 mM ATP. i, Cryo-EM density validation and close-up view of Toprim domain of ATP-bound GajA. j, Overview of the structure of GajA in complex with ATP. k, l, Enlarged view of the ATP-binding site at the Toprim domain. Two GajA protomers are colored in cyan and gray. The mesh indicates the EM density of ATP. m–r, Cleavage of pUC19-955 DNA by GajA or indicated mutants in the presence of 1 mM Mg2+ and with or without 0.5 mM ATP. s, Binding of pUC19-955 DNA by GajA or GajA-H535R with or without 0.75 mM ATP. t, ATP hydrolysis assays to compare the activation of GajB or GajB-D167A/E168A by intact T7 genomic DNA or T7 genomic DNA treated by GajA or GajA-E379A. GajB-D167A/E168A is an active site mutant of GajB and GajA-E379A is an Toprim active site mutant of GajA. Data are the average of three biological replicates, with individual data points shown. For a, f–h, and m–s, the assays were repeated three times with similar results.
Supplementary information
Supplementary Figure 1
All original source images.
Supplementary Table 1
Comparison of cleavage or binding of pUC19-955 DNA by GajA and GajA mutants on potential DNA binding sites.
Supplementary Table 2
Comparison of cleavage of pUC19-955 DNA by GajA and GajA mutants on potential ATP binding sites.
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, J., Cheng, R., Wang, Z. et al. Structures and activation mechanism of the Gabija anti-phage system. Nature 629, 467–473 (2024). https://doi.org/10.1038/s41586-024-07270-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-024-07270-x
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.