Abstract
Trace amine-associated receptor 1 (TAAR1), the founding member of a nine-member family of trace amine receptors, is responsible for recognizing a range of biogenic amines in the brain, including the endogenous β-phenylethylamine (β-PEA)1 as well as methamphetamine2, an abused substance that has posed a severe threat to human health and society3. Given its unique physiological role in the brain, TAAR1 is also an emerging target for a range of neurological disorders including schizophrenia, depression and drug addiction2,4,5. Here we report structures of human TAAR1–G-protein complexes bound to methamphetamine and β-PEA as well as complexes bound to RO5256390, a TAAR1-selective agonist, and SEP-363856, a clinical-stage dual agonist for TAAR1 and serotonin receptor 5-HT1AR (refs. 6,7). Together with systematic mutagenesis and functional studies, the structures reveal the molecular basis of methamphetamine recognition and underlying mechanisms of ligand selectivity and polypharmacology between TAAR1 and other monoamine receptors. We identify a lid-like extracellular loop 2 helix/loop structure and a hydrogen-bonding network in the ligand-binding pockets, which may contribute to the ligand recognition in TAAR1. These findings shed light on the ligand recognition mode and activation mechanism for TAAR1 and should guide the development of next-generation therapeutics for drug addiction and various neurological disorders.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Density maps and structure coordinates have been deposited in the Electron Microscopy Data Bank (EMDB) and the Protein Data Bank (PDB) with accession codes EMD-37347 and 8W87 for METH–TAAR1–Gs complex; EMD-37348 and 8W88 for SEP-363856-bound TAAR1–Gs complex; and EMD-37349 and 8W89 for β-PEA-bound TAAR1–Gs complex; EMD-37350 and 8W8A for RO5256390–TAAR1–Gs complex; and EMD-37351 and 8W8B for SEP-363856 bounded 5-HT1AR–Gi protein complex. Source data are provided with this paper.
References
Gainetdinov, R. R., Hoener, M. C. & Berry, M. D. Trace amines and their receptors. Pharmacol. Rev. 70, 549–620 (2018).
Liu, J., Wu, R. & Li, J.-X. TAAR1 and psychostimulant addiction. Cell. Mol. Neurobiol. 40, 229–238 (2020).
Most overdose deaths involve illicitly manufactured fentanyls. CDC www.cdc.gov/drugoverdose/featured-topics/overdose-deaths-data.html (2020).
Berry, M. D., Gainetdinov, R. R., Hoener, M. C. & Shahid, M. Pharmacology of human trace amine-associated receptors: therapeutic opportunities and challenges. Pharmacol. Ther. 180, 161–180 (2017).
Dodd, S. et al. Trace amine-associated receptor 1 (TAAR1): a new drug target for psychiatry? Neurosci. Biobehavi. Rev. 120, 537–541 (2021).
Heffernan, M. L. R. et al. Ulotaront: A TAAR1 agonist for the treatment of schizophrenia. ACS Med. Chem. Lett. 13, 92–98 (2022).
Nair, P. C. et al. Binding of SEP-363856 within TAAR1 and the 5HT1A receptor: implications for the design of novel antipsychotic drugs. Mol. Psychiatry 27, 88–94 (2022).
Schep, L. J., Slaughter, R. J. & Beasley, D. M. The clinical toxicology of metamfetamine. Clin. Toxicol. 48, 675–694 (2010).
Jayanthi, S., Daiwile, A. P. & Cadet, J. L. Neurotoxicity of methamphetamine: main effects and mechanisms. Exp. Neurol. 344, 113795 (2021).
Chan, B. et al. Pharmacotherapy for methamphetamine/amphetamine use disorder—a systematic review and meta-analysis. Addiction 114, 2122–2136 (2019).
Bisagno, V. & Cadet, J. L. in Handbook of Neurotoxicity (ed. Kostrzewa, R.M.) 563–585 (Springer, 2022).
Grandy, D. K. in Neuropathology of Drug Addictions and Substance Misuse (ed. Preedy, V. R.) 108–116 (Academic Press, 2016).
Miller, G. M. The emerging role of trace amine-associated receptor 1 in the functional regulation of monoamine transporters and dopaminergic activity. J. Neurochem. 116, 164–176 (2011).
Berry, M. D. Mammalian central nervous system trace amines. Pharmacologic amphetamines, physiologic neuromodulators. J. Neurochem. 90, 257–271 (2004).
Tonelli, M. & Cichero, E. Trace amine associated receptor 1 (TAAR1) modulators: a patent review (2010–present). Expert Opin.Ther. Pat. 30, 137–145 (2020).
Xu, Z. & Li, Q. TAAR agonists. Cell. Mol. Neurobiol. 40, 257–272 (2020).
Achat-Mendes, C., Lynch, L. J., Sullivan, K. A., Vallender, E. J. & Miller, G. M. Augmentation of methamphetamine-induced behaviors in transgenic mice lacking the trace amine-associated receptor 1. Pharmacol. Biochem. Behav. 101, 201–207 (2012).
Cotter, R. et al. The trace amine-associated receptor 1 modulates methamphetamine’s neurochemical and behavioral effects. Front. Neurosci. 9, 39 (2015).
Underhill, S. M. et al. Amphetamines signal through intracellular TAAR1 receptors coupled to Gα13 and GαS in discrete subcellular domains. Mol. Psychiatry 26, 1208–1223 (2021).
Eyun, S.-i, Moriyama, H., Hoffmann, F. G. & Moriyama, E. N. Molecular evolution and functional divergence of trace amine–associated receptors. PLoS ONE 11, e0151023 (2016).
Accorroni, A. & Zucchi, R. in Trace Amines and Neurological Disorders (eds Farooqui, T. & Farooqui, A. A.) 151–164 (Academic Press, 2016).
Liberles, S. D. Trace amine-associated receptors: ligands, neural circuits, and behaviors. Curr. Opin. Neurobiol. 34, 1–7 (2015).
Alnefeesi, Y. et al. Trace amine-associated receptor 1 (TAAR1): potential application in mood disorders: a systematic review. Neurosci. Biobehav. Rev. 131, 192–210 (2021).
Lothar, L. et al. Trace amine-associated receptor 1 modulates dopaminergic activity. J. Pharmacol. Exp. Ther. 324, 948 (2008).
Revel, F. G. et al. A new perspective for schizophrenia: TAAR1 agonists reveal antipsychotic- and antidepressant-like activity, improve cognition and control body weight. Mol. Psychiatry 18, 543–556 (2013).
Wu, R. & Li, J.-X. Potential of ligands for trace amine-associated receptor 1 (TAAR1) in the management of substance use disorders. CNS Drugs 35, 1239–1248 (2021).
Galley, G., Stalder, H., Goergler, A., Hoener, M. C. & Norcross, R. D. Optimisation of imidazole compounds as selective TAAR1 agonists: discovery of RO5073012. Bioorg. Med. Chem. Lett. 22, 5244–5248 (2012).
Galley, G. et al. Discovery and characterization of 2-aminooxazolines as highly potent, selective, and orally active TAAR1 agonists. ACS Med. Chem. Lett. 7, 192–197 (2016).
Raony, Í., Domith, I., Lourenco, M. V., Paes-de-Carvalho, R. & Pandolfo, P. Trace amine-associated receptor 1 modulates motor hyperactivity, cognition, and anxiety-like behavior in an animal model of ADHD. Prog. Neuropsychopharmacol. Biol. Psychiatry 117, 110555 (2022).
Pei, Y. et al. Activation of the trace amine-associated receptor 1 prevents relapse to cocaine seeking. Neuropsychopharmacology 39, 2299–2308 (2014).
Barak, L. S. et al. Pharmacological characterization of membrane-expressed human trace amine-associated receptor 1 (TAAR1) by a bioluminescence resonance energy transfer cAMP biosensor. Mol. Pharmacol. 74, 585–594 (2008).
Saarinen, M. et al. TAAR1 dependent and independent actions of the potential antipsychotic and dual TAAR1/5-HT(1A) receptor agonist SEP-383856. Neuropsychopharmacology 47, 2319–2329 (2022).
Xu, P. et al. Structural insights into the lipid and ligand regulation of serotonin receptors. Nature 592, 469–473 (2021).
Reese, E. A., Bunzow, J. R., Arttamangkul, S., Sonders, M. S. & Grandy, D. K. Trace amine-associated receptor 1 displays species-dependent stereoselectivity for isomers of methamphetamine, amphetamine, and para-hydroxyamphetamine. J. Pharmacol. Exp. Ther. 321, 178–186 (2007).
Chun, E. et al. Fusion partner toolchest for the stabilization and crystallization of G protein-coupled receptors. Structure 20, 967–976 (2012).
Duan, J. et al. Cryo-EM structure of an activated VIP1 receptor-G protein complex revealed by a NanoBiT tethering strategy. Nat. Commun. 11, 4121 (2020).
Liu, P. et al. The structural basis of the dominant negative phenotype of the Gαi1β1γ2 G203A/A326S heterotrimer. Acta Pharmacol. Sin. 37, 1259–1272 (2016).
Maeda, S. et al. Development of an antibody fragment that stabilizes GPCR/G-protein complexes. Nat. Commun. 9, 3712 (2018).
Isberg, V. et al. Generic GPCR residue numbers - aligning topology maps while minding the gaps. Trends Pharmacol. Sci. 36, 22–31 (2015).
Zhuang, Y. et al. Structural insights into the human D1 and D2 dopamine receptor signaling complexes. Cell 184, 931–942.e918 (2021).
Tan, E. S. et al. The molecular basis of species-specific ligand activation of trace amine-associated receptor 1 (TAAR(1)). ACS Chem. Biol. 4, 209–220 (2009).
Cichero, E., Espinoza, S., Gainetdinov, R. R., Brasili, L. & Fossa, P. Insights into the structure and pharmacology of the human trace amine-associated receptor 1 (hTAAR1): homology modelling and docking studies. Chem. Biol. Drug Des. 81, 509–516 (2013).
Liao, S., Pino, M. J. Jr., Deleon, C., Lindner-Jackson, M. & Wu, C. Interaction analyses of hTAAR1 and mTAAR1 with antagonist EPPTB. Life Sci. 300, 120553 (2022).
Reese, E. A. et al. Exploring the determinants of trace amine-associated receptor 1’s functional selectivity for the stereoisomers of amphetamine and methamphetamine. J. Med. Chem. 57, 378–390 (2014).
Huang, S. et al. GPCRs steer G(i) and G(s) selectivity via TM5-TM6 switches as revealed by structures of serotonin receptors. Mol. Cell 82, 2681–2695.e2686 (2022).
Su, M. et al. Structures of β1-adrenergic receptor in complex with Gs and ligands of different efficacies. Nat. Commun. 13, 4095 (2022).
Kaplan, A. L. et al. Bespoke library docking for 5-HT2A receptor agonists with antidepressant activity. Nature 610, 582–591 (2022).
Tan, Y. et al. Structural insights into the ligand binding and Gi coupling of serotonin receptor 5-HT5A. Cell Discov. 8, 50 (2022).
Borowsky, B. et al. Trace amines: Identification of a family of mammalian G protein-coupled receptors. Proc. Natl Acad. Sci. USA 98, 8966–8971 (2001).
Zhang, Y. et al. Single-particle cryo-EM structural studies of the β2AR–Gs complex bound with a full agonist formoterol. Cell Discov. 6, 45 (2020).
Imai, S. et al. Structural equilibrium underlying ligand-dependent activation of β2-adrenoreceptor. Nat. Chem. Biol. 16, 430–439 (2020).
Staus, D. P. et al. Allosteric nanobodies reveal the dynamic range and diverse mechanisms of G-protein-coupled receptor activation. Nature 535, 448–452 (2016).
Lam, V. M. et al. Behavioral effects of a potential novel TAAR1 antagonist. Front. Pharmacol. 9, 953 (2018).
Bradaia, A. et al. The selective antagonist EPPTB reveals TAAR1-mediated regulatory mechanisms in dopaminergic neurons of the mesolimbic system. Proc. Natl Acad. Sci. USA 106, 20081–20086 (2009).
Guo, L. et al. Structural basis of amine odorant perception by a mammal olfactory receptor. Nature 618, 193–200 (2023).
Carpenter, B., Nehmé, R., Warne, T., Leslie, A. G. & Tate, C. G. Structure of the adenosine A(2A) receptor bound to an engineered G protein. Nature 536, 104–107 (2016).
García-Nafría, J., Lee, Y., Bai, X., Carpenter, B. & Tate, C. G. Cryo-EM structure of the adenosine A(2A) receptor coupled to an engineered heterotrimeric G protein. eLife 7, e35946 (2018).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Pettersen, E. F. et al. UCSF Chimera–a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Croll, T. I. ISOLDE: a physically realistic environment for model building into low-resolution electron-density maps. Acta Crystallogr. D 74, 519–530 (2018).
Adams, P. D. et al. Recent developments in the PHENIX software for automated crystallographic structure determination. J. Synchrotron Radiat. 11, 53–55 (2004).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Wang, F. I., Ding, G., Ng, G. S., Dixon, S. J. & Chidiac, P. Luciferase-based GloSensor™ cAMP assay: temperature optimization and application to cell-based kinetic studies. Methods 203, 249–258 (2022).
Revel, F. G. et al. Brain-specific overexpression of trace amine-associated receptor 1 alters monoaminergic neurotransmission and decreases sensitivity to amphetamine. Neuropsychopharmacology 37, 2580–2592 (2012).
Olsen, R. H. J. et al. TRUPATH, an open-source biosensor platform for interrogating the GPCR transducerome. Nat. Chem. Biol. 16, 841–849 (2020).
Webb, B. & Sali, A. Comparative Protein Structure Modeling Using MODELLER. Curr. Protoc. Bioinformatics 54, 5.6.1–5.6.37 (2016).
Heo, L. & Feig, M. Multi-state modeling of G-protein coupled receptors at experimental accuracy. Proteins 90, 1873–1885 (2022).
Olsson, M. H., Søndergaard, C. R., Rostkowski, M. & Jensen, J. H. PROPKA3: consistent treatment of internal and surface residues in empirical pKa predictions. J. Chem. Theory Comput. 7, 525–537 (2011).
Vanommeslaeghe, K. et al. CHARMM general force field: a force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J. Comput. Chem. 31, 671–690 (2010).
Wu, E. L. et al. CHARMM-GUI Membrane Builder toward realistic biological membrane simulations. J. Comput. Chem. 35, 1997–2004 (2014).
Huang, J. et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2017).
Salomon-Ferrer, R., Götz, A. W., Poole, D., Le Grand, S. & Walker, R. C. Routine microsecond molecular dynamics simulations with AMBER on GPUs. 2. Explicit solvent particle mesh Ewald. J. Chem. Theory Comput. 9, 3878–3888 (2013).
Miller, B. R. 3rd et al. MMPBSA.py: an efficient program for end-state free energy calculations. J. Chem. Theory Comput. 8, 3314–3321 (2012).
Acknowledgements
This work was partially supported by the Ministry of Science and Technology of China (2018YFA0507002 to H.E.X., 2020YFA0509102 to S.W.); the National Natural Science Foundation of China (32071194 to F.X., 32130022 and 82121005 to H.E.X., 52272087 to W.L., 82225025 and 32071197 to S.W.); Shanghai Municipal Science and Technology Major Project (2019SHZDZX02 to H.E.X.); CAS Strategic Priority Research Programme (XDB37030103 to H.E.X.); the Lingang Laboratory, grant no. LG-GG-202204-01 (H.E.X.); and the National Key R&D Program of China (2022YFC2703105 to H.E.X.). The cryo-EM data were collected at the Shanghai Advanced Electron Microscope Center, Shanghai Institute of Material Medica, as well as at the Bio-EM platform of ShanghaiTech University. We thank Q. Yuan, K. Wu, W. Hu and S. Zhang for providing technical support and assistance during data collection at the Shanghai Advanced Electron Microscope Center, Shanghai Institute of Material Medica; we thank Q. Sun, Z. Zhang, L. Wang, D. Liu and Y. Liu at the Bio-EM facility at ShanghaiTech University; and Q. Tan, Q. Shi, J. Liu, N. Chen, S. Hu and W. Xiao from ShanghaiTech University for protein cloning, expression, assay and data collection support. We thank S. Zhao from ShanghaiTech University, J. Wang from Institute of Biophysics, CAS, and Q. Li from Shanghai Jiaotong University to provide initial sets of TAAR1 tool compounds and suggestions for this work. We also thank the support from Shanghai Frontiers Science Center for Biomacromolecules and Precision Medicine at ShanghaiTech University.
Author information
Authors and Affiliations
Contributions
F.X. and H.E.X. initiated the project. H.L. designed and screened the expression constructs of TAAR1 and prepared protein samples of TAAR1–METH–Gs, TAAR1–SEP-363856–Gs, 5-HT1AR–SEP-363856–Gi complexes for cryo-EM data collection, performed cryo-EM grids preparation, data acquisition, and structure determination, participated in GloSensor cAMP assay, and prepared the draft of the manuscript and figures. Y.Z. designed and screened the expression constructs of TAAR1 and prepared protein samples of TAAR1–β-PEA-Gs, TAAR1–RO5256390-Gs, complexes for cryo-EM data collection, performed cryo-EM grids preparation, data acquisition and structure determination, and participated in manuscript editing and figure preparation. Y.W. performed GloSensor cAMP assay, G-protein recruitment assay and participated in figure preparation. Y.W. performed the radioligand-binding assay and Gαiβγ dissociation assay. X.H. performed MD simulation analysis. P.X. and S.H. participated in protein sample preparation and structure determination. Q.Y. participated in data acquisition. X.Z. participated in G-protein recruitment assay. L.W. participated in GloSensor cAMP assay. K.J. assisted in some protein preparation work. H.C. assisted in compound preparation and synthesis. Z.L. helped construct the mutations for function assays. W.L. supervised compound preparation and synthesis. S.W. supervised ligand binding and functional assay. F.X. and H.E.X. conceived and supervised the project and participated in manuscript editing. F.X. and H.E.X. wrote the manuscript with input from H.L., Y.Z. and Y.W.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Thomas Keck, Tao Che and Harald Sitte for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Functional assessment of TAAR1 by four agonists and structural feature of aminergic receptors.
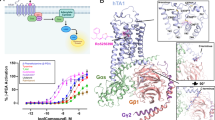
(a-b) Concentration–response curves of the four agonists activating TAAR1 evaluated by miniGs recruitment assay (a) and radio-ligand-binding assay (b). Data are mean ± S.E.M. from 3 independent experiments (n = 3). (c) Structural comparison of ECL2s from TAAR1 and other aminergic receptors, ECL2 that forms a short helix in the structures is highlighted with red dotted circle.
Extended Data Fig. 2 Purification and structure determination of METH and β-PEA-bound TAAR1-Gs complexes.
(a) Cartoon models of the two TAAR1 constructs used in this study. (b) Representative size exclusion chromatography (SEC) profiles and SDS-PAGE analysis of TAAR1-Gs complex activated by METH. Experiment was repeated at least three times with similar results. (c-d) Representative cryo-EM image from 6,686 movies (c) and 2D classification averages (d) of METH–TAAR1–Gs. (e) Cryo-EM data processing flowcharts of METH-TAAR1-Gs by cyroSPARC 3.2. (f) The Fourier shell correlation (FSC) curves of METH-TAAR1-Gs. The global resolution of the final processed density map estimated at the FSC = 0.143 is 2.8 Å. (g) The global density map of METH-TAAR1-Gs colored by local resolutions. (h) The density maps of helices TM1–TM7 of transmembrane domain, extracellular loop ECL2 of TAAR1, α5 helix of Gαs, and METH in METH-TAAR1-Gs complex. (i) Representative size exclusion chromatography (SEC) profiles and SDS-PAGE analysis of TAAR1–Gs complex activated by β-PEA. Experiment was repeated at least three times with similar results. (j-k) Representative cryo-EM image from 4,371 movies (j) and 2D classification averages (k) of β-PEA-TAAR1-Gs. (l) Cryo-EM data processing flowcharts of β-PEA-TAAR1-Gs by cyroSPARC 3.2. (m) The Fourier shell correlation (FSC) curves of β-PEA-TAAR1-Gs. The global resolution of the final processed density map estimated at the FSC = 0.143 is 3.0 Å. (n) The global density map of β-PEA-TAAR1-Gs colored by local resolutions. (o) The density maps of helices TM1–TM7 of transmembrane domain, extracellular loop ECL2 of TAAR1, α5 helix of Gαs, and β-PEA in β-PEA-TAAR1-Gs complex.
Extended Data Fig. 3 Mutational effects of agonists and MD simulation analysis of TAAR1.
(a) Concentration–response curves of wild-type/mutants of TAAR1 induced by the four agonists. Every response of mutant was normalized to corresponding wild-type as 100%. Data are mean ± S.E.M. from 3 independent experiments (n = 3). (b-c) RMSD plot of METH (b) or β-PEA (c) in TAAR1 pocket during simulations. (d) The free energy contribution of key residues in TAAR1 pocket for METH or β-PEA binding. Data are mean ± S.E.M. from 6 independent experiments (n = 6).
Extended Data Fig. 4 Purification and structure determination of SEP-363856 bound TAAR1-Gs and 5HT1AR-Gi complexes.
(a) Representative size exclusion chromatography (SEC) profiles and SDS-PAGE analysis of TAAR1-Gs complex activated by SEP-363856. Experiment was repeated at least three times with similar results. (b-c) Representative cryo-EM image from 7,262 movies (b) and 2D classification averages (c) of SEP-363856-TAAR1-Gs. (d) Cryo-EM data processing flowcharts of SEP-363856-TAAR1-Gs by cyroSPARC 3.2. (e) The Fourier shell correlation (FSC) curves of SEP-363856-TAAR1-Gs. The global resolution of the final processed density map estimated at the FSC = 0.143 is 2.6 Å. (f) The global density map of SEP-363856-TAAR1-Gs colored by local resolutions. (g) The density maps of helices TM1–TM7 of transmembrane domain, extracellular loop ECL2 of TAAR1, α5 helix of Gαs, and SEP-363856 in SEP-363856-TAAR1-Gs complex. (h) Representative size exclusion chromatography (SEC) profiles and SDS-PAGE analysis of 5HT1AR -Gi complex activated by SEP-363856. Experiment was repeated at least three times with similar results. (i-j) Representative cryo-EM image from 6,222 movies (i) and 2D classification averages (j) of SEP-363856-5HT1AR -Gi. (k) Cryo-EM data processing flowcharts of SEP-363856-5HT1AR-Gi complex by cyroSPARC 3.2. (l) The Fourier shell correlation (FSC) curves of SEP-363856-5HT1AR -Gi complex. The global resolution of the final processed density map estimated at the FSC = 0.143 is 3.0 Å. (m) The global density map of SEP-363856-5HT1AR-Gi colored by local resolutions. (n) The density maps of helices TM1–TM7 of transmembrane domain, α5 helix of Gαi, and SEP-363856 in 5HT1AR -Gi complex.
Extended Data Fig. 5 Comparison of the SEP-363856 bound pockets of TAAR1 and 5HT1AR, and purification, structure determination of RO5256390 bound TAAR1-Gs complex.
(a) Binding pockets of SEP-363856, aripiprazole (PDB ID: 7e2z) and serotonin (PDB ID: 7e2y) in 5HT1AR. (b) Binding pocket of SEP-363856 in TAAR1. (c) Alignment of key residues constituting the ligand-binding pockets in SEP-383656 bound 5HT1AR or TAAR1 structures, and serotonin and aripiprazole bound 5HT1AR structures. The green background indicated 100% conserved residues and orange background indicated residues that are conserved in 5HT1AR but not in TAAR1. (d) Representative size exclusion chromatography (SEC) profiles and SDS-PAGE analysis of TAAR1-Gs complex activated by RO5256390. Experiment was repeated at least three times with similar results. (e-f) Representative cryo-EM image from 5,135 movies (e) and 2D classification averages (f) of RO5256390-TAAR1-Gs. (g) Cryo-EM data processing flowcharts of RO5256390-TAAR1-Gs complex by cyroSPARC 3.2. (h) The Fourier shell correlation (FSC) curves of RO5256390-TAAR1-Gs. The global resolution of the final processed density map estimated at the FSC = 0.143 is 2.8 Å. (i) The global density map of RO5256390-TAAR1-Gs colored by local resolutions. (j) The density maps of helices TM1–TM7 of transmembrane domain, extracellular loop ECL2 of TAAR1, α5 helix of Gαs, and RO5256390 in RO5256390-TAAR1-Gs complex.
Extended Data Fig. 6 Molecular mechanism of the high potency and selectivity of RO5256390.
(a) Interactions of RO5256390 and key residues in TAAR1 pocket. (b) RO5256390 inserted deep into the cavity formed by D1033x32, S1073x36, W2646x48, Y2947x43. (c-d) Concentration–response curves for mutations of key residues in TAAR1 induced by RO5256390 using Glo-sensor assay. Data are mean ± S.E.M. from 3 independent experiments (n = 3). (e) and (g) Evaluation of RO5256390 and SEP-363856 activating 5HT1AR using Gi protein dissociation assay (e) or Glo-sensor assay (g), 5HT was used as reference. Data are mean ± S.E.M. from 3 independent experiments (n = 3). (f) and (h) Evaluation of RO5256390 activating 5HT1AR with C120 substituted with alanine or serine, using Gi protein dissociation assay (f) or Glo-sensor assay (h), 5HT was used as reference. Data are mean ± S.E.M. from 3 independent experiments (n = 3).
Extended Data Fig. 7 Conservation of key residues for TAAR1 activating and role of ECL2 in aminergic receptors.
(a) Sequence alignment of key residues in ECL2 from representative aminergic receptors. (b) Display of residues in ECL2s from TAAR1 and other aminergic receptors that may take part in ligand recognition and binding pocket formation.
Extended Data Fig. 8 Functional assessment of four agonists activating human TAAR family subtypes.
Human TAAR family subtypes (hTAAR1/2/5/6/8/9) were cloned into pcDNA3.1 vector and the activity was measured by Glo-Sensor assay. The negative control was pcDNA3.1 vector and DMSO solvent. The results showed that only TAAR1 can be activated by the four agonist. Data are mean ± S.E.M. from 3 independent experiments (n = 3).
Extended Data Fig. 9 Sequence alignment of human TAAR family, TAAR1 activation mechanism, and analysis of METH binding in human and rodent TAAR1.
(a-b) Sequence alignment of residues important for TAAR1-ligand binding (a) or activating (b) from human TAAR family and mTAAR9. (c-d) Superposition of activated TAAR1 (dark cyan) with active mTAAR9 (light green; PDB ID: 8iw7), β2AR (orange; PDB ID: 6kr8) and inactive β2AR (gray; PDB ID: 5jqh). Notable conformational changes occur at extracellular end of TM1 and intracellular ends of TM6, TM7 and H8 upon receptor activation, side view (c) and bottom view (d). (e) The “toggle switch”, W2646.48, of TAAR1 display relative rotameric change when sensing agonist. (f–h) The key P-I-F6.44 (f), D-R3.50-Y (g), and N-P7.50-xx-Y7.53 (h) motifs displayed conformational rearrangement in activated TAAR1 structure. (i–k) RMSD plot of METH binding in rat, mouse, or human TAAR1 pocket during simulations. (l) Bar graph showing RMSD differences for METH binding in rat, mouse, or human TAAR1 pocket during simulations. Data are mean ± S.E.M. of RMSD from 3 independent experiments (n = 3) for rat and mouse TAAR1 and 6 independent experiments (n = 6) for human TAAR1. (m) Sequence alignment for residues that are important for ligand binding to TAAR1 from rat, mouse or human.
Supplementary information
Supplementary Information
Supplementary Tables 1–6 and Supplementary Fig. 1.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, H., Zheng, Y., Wang, Y. et al. Recognition of methamphetamine and other amines by trace amine receptor TAAR1. Nature 624, 663–671 (2023). https://doi.org/10.1038/s41586-023-06775-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-023-06775-1
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.