Abstract
Ribosomes are highly sophisticated translation machines that have been demonstrated to be heterogeneous in the regulation of protein synthesis1,2. Male germ cell development involves complex translational regulation during sperm formation3. However, it remains unclear whether translation during sperm formation is performed by a specific ribosome. Here we report a ribosome with a specialized nascent polypeptide exit tunnel, RibosomeST, that is assembled with the male germ-cell-specific protein RPL39L, the paralogue of core ribosome (RibosomeCore) protein RPL39. Deletion of RibosomeST in mice causes defective sperm formation, resulting in substantially reduced fertility. Our comparison of single-particle cryo-electron microscopy structures of ribosomes from mouse kidneys and testes indicates that RibosomeST features a ribosomal polypeptide exit tunnel of distinct size and charge states compared with RibosomeCore. RibosomeST predominantly cotranslationally regulates the folding of a subset of male germ-cell-specific proteins that are essential for the formation of sperm. Moreover, we found that specialized functions of RibosomeST were not replaceable by RibosomeCore. Taken together, identification of this sperm-specific ribosome should greatly expand our understanding of ribosome function and tissue-specific regulation of protein expression pattern in mammals.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The MS proteomics data have been deposited at the ProteomeXchange Consortium via the PRIDE under project accessions: PXD020874, PXD020864, PXD020922 and PXD037257. Sequencing data were deposited at the National Center for Biotechnology Information (NCBI) Sequence Read Archive under BioProject PRJNA657531. The cryo-EM density maps reported in this paper are available at the Electron Microscopy Data Bank under accession codes EMD-30432 (80S ribosome from mouse kidney) and EMD-30433 (80S ribosome from mouse testis); atomic coordinates are reported at the PDB under accession codes 7CPU and 7CPV, respectively. The human 40S and 60S ribosome structures used in this study were obtained from PDB 6EK0. Source data are provided with this paper.
References
Genuth, N. R. & Barna, M. The discovery of ribosome heterogeneity and its implications for gene regulation and organismal life. Mol. Cell 71, 364–374 (2018).
Ghulam, M. M., Catala, M. & Abou Elela, S. Differential expression of duplicated ribosomal protein genes modifies ribosome composition in response to stress. Nucleic Acids Res. 48, 1954–1968 (2020).
Wang, M. et al. Single-cell RNA sequencing analysis reveals sequential cell fate transition during human spermatogenesis. Cell Stem Cell 23, 599–614 (2018).
Shi, Z. et al. Heterogeneous ribosomes preferentially translate distinct subpools of mRNAs genome-wide. Mol. Cell 67, 71–83 (2017).
Gupta, V. & Warner, J. R. Ribosome-omics of the human ribosome. RNA 20, 1004–1013 (2014).
Chaillou, T., Zhang, X. & McCarthy, J. J. Expression of muscle-specific ribosomal protein L3-like impairs myotube growth. J. Cell. Physiol. 231, 1894–902 (2016).
Kao, B. R. et al. Knockdown of muscle-specific ribosomal protein L3-like enhances muscle function in healthy and dystrophic mice. Nucleic Acid Ther. 31, 457–464 (2021).
O’Leary, M. N. et al. The ribosomal protein Rpl22 controls ribosome composition by directly repressing expression of its own paralog, Rpl22l1. PLoS Genet. 9, e1003708 (2013).
Rao, S. et al. RPL22L1 induction in colorectal cancer is associated with poor prognosis and 5-FU resistance. PLoS ONE 14, e0222392 (2019).
Kondrashov, N. et al. Ribosome-mediated specificity in Hox mRNA translation and vertebrate tissue patterning. Cell 145, 383–397 (2011).
Simsek, D. et al. The mammalian ribo-interactome reveals ribosome functional diversity and heterogeneity. Cell 169, 1051–1065 (2017).
Slavov, N., Semrau, S., Airoldi, E., Budnik, B. & van Oudenaarden, A. Differential stoichiometry among core ribosomal proteins. Cell Rep. 13, 865–873 (2015).
Jiang, L. et al. RPL10L is required for male meiotic division by compensating for RPL10 during meiotic sex chromosome inactivation in mice. Curr. Biol. 27, 1498–1505 (2017).
Gamalinda, M. & Woolford Jr, J. L. Deletion of L4 domains reveals insights into the importance of ribosomal protein extensions in eukaryotic ribosome assembly. RNA 20, 1725–1731 (2014).
Anger, A. M. et al. Structures of the human and Drosophila 80S ribosome. Nature 497, 80–85 (2013).
Young, S. A. et al. CABYR is essential for fibrous sheath integrity and progressive motility in mouse spermatozoa. J. Cell Sci. 129, 4379–4387 (2016).
Yang, K., Adham, I. M., Meinhardt, A. & Hoyer-Fender, S. Ultra-structure of the sperm head-to-tail linkage complex in the absence of the spermatid-specific LINC component SPAG4. Histochem. Cell Biol. 150, 49–59 (2018).
Shang, Y. et al. Essential role for SUN5 in anchoring sperm head to the tail. eLife 6, e28199 (2017).
Yuan, S. et al. Spata6 is required for normal assembly of the sperm connecting piece and tight head-tail conjunction. Proc. Natl Acad. Sci. USA 112, E430–E439 (2015).
Wang, Y. Y. et al. Deficiency of the Tbc1d21 gene causes male infertility with morphological abnormalities of the sperm mitochondria and flagellum in mice. PLoS Genet. 16, e1009020 (2020).
Farias-Rico, J. A., Goetz, S. K., Marino, J. & von Heijne, G. Mutational analysis of protein folding inside the ribosome exit tunnel. FEBS Lett. 591, 155–163 (2017).
Kramer, G., Boehringer, D., Ban, N. & Bukau, B. The ribosome as a platform for co-translational processing, folding and targeting of newly synthesized proteins. Nat. Struct. Mol. Biol. 16, 589–597 (2009).
Cassaignau, A. M. E., Cabrita, L. D. & Christodoulou, J. How does the ribosome fold the proteome. Annu. Rev. Biochem. 89, 389–415 (2020).
Ott, M., Amunts, A. & Brown, A. Organization and regulation of mitochondrial protein synthesis. Annu. Rev. Biochem. 85, 77–101 (2016).
Bhushan, S. et al. α-Helical nascent polypeptide chains visualized within distinct regions of the ribosomal exit tunnel. Nat. Struct. Mol. Biol. 17, 313–317 (2010).
Balchin, D., Hayer-Hartl, M. & Hartl, F. U. In vivo aspects of protein folding and quality control. Science 353, aac4354 (2016).
Li, B. et al. A comprehensive mouse transcriptomic BodyMap across 17 tissues by RNA-seq. Sci. Rep. 7, 4200 (2017).
Wang, D. et al. LYPD4, mouse homolog of a human acrosome protein, is essential for sperm fertilizing ability and male fertility. Biol. Reprod. 102, 1033–1044 (2020).
Mordret, E. et al. Systematic detection of amino acid substitutions in proteomes reveals mechanistic basis of ribosome errors and selection for translation fidelity. Mol. Cell 75, 427–441 (2019).
Zou, Q. et al. Proteostasis regulated by testis-specific ribosomal protein RPL39L maintains mouse spermatogenesis. iScience 24, 103396 (2021).
Petrone, P. M., Snow, C. D., Lucent, D. & Pande, V. S. et al. Side-chain recognition and gating in the ribosome exit tunnel. Proc. Natl Acad. Sci. USA 105, 16549–16554 (2008).
Lu, J. & Deutsch, C. Electrostatics in the ribosomal tunnel modulate chain elongation rates. J. Mol. Biol. 384, 73–86 (2008).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Varadi, M. et al. AlphaFold protein structure database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res. 50, D439–D444 (2022).
Buhr, F. et al. Synonymous codons direct cotranslational folding toward different protein conformations. Mol. Cell 61, 341–351 (2016).
Simpson, A. J., Caballero, O. L., Jungbluth, A., Chen, Y. T. & Old, L. J. Cancer/testis antigens, gametogenesis and cancer. Nat. Rev. Cancer 5, 615–625 (2005).
Chen, S. et al. Trimethylamine N-oxide binds and activates PERK to promote metabolic dysfunction. Cell Metab. 30, 1141–1151 (2019).
Walter, P. & Ron, D. The unfolded protein response: from stress pathway to homeostatic regulation. Science 334, 1081–1086 (2011).
Pallares, I. & Ventura, S. Understanding and predicting protein misfolding and aggregation: insights from proteomics. Proteomics 16, 2570–2581 (2016).
Piatek, R., Bruzdziak, P., Wojciechowski, M., Zalewska-Piatek, B. & Kur, J. The noncanonical disulfide bond as the important stabilizing element of the immunoglobulin fold of the Dr Fimbrial DraE subunit. Biochemistry 49, 1460–1468 (2010).
Tokuhiro, K. et al. OAZ-t/OAZ3 is essential for rigid connection of sperm tails to heads in mouse. PLoS Genet. 5, e1000712 (2009).
Koonin, E. V. Orthologs, paralogs, and evolutionary genomics. Annu. Rev. Genet. 39, 309–338 (2005).
Komili, S., Farny, N. G., Roth, F. P. & Silver, P. A. Functional specificity among ribosomal proteins regulates gene expression. Cell 131, 557–571 (2007).
Gerst, J. E. Pimp my ribosome: ribosomal protein paralogs specify translational control. Trends Genet. 34, 832–845 (2018).
Amirbeigiarab, S. et al. Invariable stoichiometry of ribosomal proteins in mouse brain tissues with aging. Proc. Natl Acad. Sci. USA 116, 22567–22572 (2019).
Rinaldi, V. D. et al. An atlas of cell types in the mouse epididymis and vas deferens. eLife 9, e55474 (2020).
Gur, Y. & Breitbart, H. Mammalian sperm translate nuclear-encoded proteins by mitochondrial-type ribosomes. Genes Dev. 20, 411–416 (2006).
Bryant, J. M., Meyer-Ficca, M. L., Dang, V. M., Berger, S. L. & Meyer, R. G. Separation of spermatogenic cell types using STA-PUT velocity sedimentation. J. Vis. Exp. https://doi.org/10.3791/50648 (2013).
Wang, J. P., Page, D. C. & McCarrey, J. R. Differential expression of sex-linked and autosomal germ-cell-specific genes during spermatogenesis in the mouse. Hum. Mol. Genet. 14, 2911–2918 (2005).
Guo, X. et al. Proteomic analysis of male 4C germ cell proteins involved in mouse meiosis. Proteomics 11, 298–308 (2011).
Zheng, W. et al. DDB1 regulates sertoli cell proliferation and testis cord remodeling by TGFβ pathway. Genes 10, 974 (2019).
Hua, R. et al. FBXO47 regulates telomere-inner nuclear envelope integration by stabilizing TRF2 during meiosis. Nucleic Acids Res. 47, 11755–11770 (2019).
Mahadevaiah, S. K., Costa, Y. & Turner, J. M. Using RNA FISH to study gene expression during mammalian meiosis. Methods Mol. Biol. 558, 433–444 (2009).
Wang, J. et al. Quantitative phosphoproteomics analysis reveals a key role of insulin growth factor 1 receptor (IGF1R) tyrosine kinase in human sperm capacitation. Mol. Cell Proteom. 14, 1104–1112 (2015).
Fan, Y. et al. Phosphoproteomic analysis of neonatal regenerative myocardium revealed important roles of checkpoint kinase 1 via activating mammalian target of rapamycin C1/ribosomal protein S6 kinase b-1 pathway. Circulation 141, 1554–1569 (2020).
Hao, P., Ren, Y., Dutta, B. & Sze, S. K. Comparative evaluation of electrostatic repulsion-hydrophilic interaction chromatography (ERLIC) and high-pH reversed phase (Hp-RP) chromatography in profiling of rat kidney proteome. J. Proteom. 82, 254–262 (2013).
Tyanova, S., Temu, T. & Cox, J. The MaxQuant computational platform for mass spectrometry-based shotgun proteomics. Nat. Protoc. 11, 2301–2319 (2016).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Peterson, A. C., Russell, J. D., Bailey, D. J., Westphall, M. S. & Coon, J. J. Parallel reaction monitoring for high resolution and high mass accuracy quantitative, targeted proteomics. Mol. Cell. Proteom. 11, 1475–1488 (2012).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & N G, CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Fernandez-Leiro, R. & Scheres, S. H. W. A pipeline approach to single-particle processing in RELION. Acta Crystallogr. D 73, 496–502 (2017).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Natchiar, S. K., Myasnikov, A. G., Kratzat, H., Hazemann, I. & Klaholz, B. P. Visualization of chemical modifications in the human 80S ribosome structure. Nature 551, 472–477 (2017).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D 75, 861–877 (2019).
Kopylova, E., Noe, L. & Touzet, H. SortMeRNA: fast and accurate filtering of ribosomal RNAs in metatranscriptomic data. Bioinformatics 28, 3211–3217 (2012).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Tokumoto, S. et al. Development of a Tet-on inducible expression system for the anhydrobiotic cell line, Pv11. Insects 11, 781 (2020).
Hao Zeng, W. X. Ctr9, a key subunit of PAFc, affects global estrogen signaling and drives ERα-positive breast tumorigenesis. Genes Dev. 29, 2153–2167 (2015).
Mauthe, M. et al. Chloroquine inhibits autophagic flux by decreasing autophagosome-lysosome fusion. Autophagy 14, 1435–1455 (2018).
Go, Y. M. & Jones, D. P. Thioredoxin redox western analysis. Curr. Protoc. Toxicol. 41, 17.12.1–17.12.12 (2009).
Manning, M. & Colón, W. Structural basis of protein kinetic stability: resistance to sodium dodecyl sulfate suggests a central role for rigidity and a bias toward beta-sheet structure. Biochemistry 43, 11248–11254 (2004).
Piatek, R. et al. Molecular aspects of biogenesis of Escherichia coli Dr Fimbriae: characterization of DraB-DraE complexes. Infect. Immun. 73, 135–145 (2005).
Hu, M. L. Measurement of protein thiol groups and glutathione in plasma. Methods Enzymol. 233, 380–385 (1994).
Acknowledgements
We thank D. Li (Center for Excellence in Molecular Cell Science, Chinese Academy of Sciences (CAS)) and S. Qian (University of Cornell) for the assistance during the experimental design; B. Zhu, X. Huang and G. Ji (Institute of Biophysics (IBP), CAS) for cryo-EM data collection at the Center for Biological Imaging (CBI, http://cbi.ibp.ac.cn), IBP, CAS; D. Fan, L. Zhang and B. Huangfu (IBP, CAS) for assistance during cryo-EM sample screening; Q. Dong (IBP, CAS) for artwork preparation; W. Li (Institute of Zoology, CAS) for assistance in sperm fluorescence in situ hybridization analyses; and S. Eckardt for help with paper preparation. This work was supported by the National Key R&D Program (2018YFC1003500, 2018YFA0106900 and 2021YFC2700200) and the National Natural Science Foundation of China (31890784, 31830043).
Author information
Authors and Affiliations
Contributions
H.L., Y.H., X.H., L.Y., H.Z., Y. Cui and H.X. conducted majority of experiments and analysed the data. H.L., X.H., Y. Cui, H.X., W.X., S.Z., H.T. and Y.Z. performed functional studies in mouse and cell lines. H.L., Y.H., X.H., D.Z., X.C. and T.J. performed ribosomal and structural analyses. L.Y., Y.W., Y. Cheng and Y.G. performed proteomics analysis. H.Z. performed bioinformatics analysis. J.S., Y.Q. and X.G. initiated the project, designed the experiments, supervised the whole project and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Jon Oatley, Petra Van Damme and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Cell type specific ribosome: Ribosomal proteome profiles and RP identification from 80S monosomes across mouse tissues and different stages of postnatal development testis.
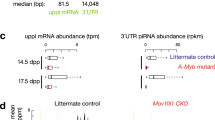
a, Separation of cytoplasmic ribosomes through sucrose gradient fractionation using different adult mouse tissues. b, Venn diagram of the identified RPs in a comparative analysis of this and earlier studies. c, Distribution of RP paralogues in different mouse tissues identified in this study. d, Correlation matrix of cytoplasmic ribosomes from different mouse tissues. e, Density plot of maximal SPM of RPs of nine mouse tissues. f, The relationship between retrogene Rpl39l and its X-linked parental gene Rpl39. g, mRNA levels of Rpl39 and Rpl39l in 15 mouse tissues. n = 3. h, Relative quantification of RPL3L in nine mouse tissues using a PRM method. n = 3. i, Separation of cytoplasmic ribosomes through sucrose gradient fractionation using mouse testicular tissue at different postnatal developmental stages. j, Correlation matrix for cytoplasmic ribosomes from mouse testicular tissue at different developmental stages. k, Relative protein quantification of RPL4 and RPL39L in non-ribosome, 40S, 60S and ribosome fractions by PRM. n = 3. l, Morphology and specific staining of mouse testicular cells: testicular interstitial cells (IC), Sertoli cells (SC), spermatogonia (SG), pachytene spermatocytes (PS), round spermatids (RS), and elongated spermatids (ES). Scale bar, 20 μm. m, Purities of SG (82.60 %), PS (82.69 %), RS (87.76 %), ES (81.9 %), IC (92.63 %) and SC (96.35 %) cells. n = 3. n, Quantification of Rpl39 and Rpl39l mRNA levels in testicular cells by qPCR. n = 3. o, Absolute protein quantification of RPL10, RPL10L, and RPL22, RPL22L1 in testicular cells by PRM. n = 3. For g, h, k, m, n and o, data are mean ± s.e.m. n values represent the number of samples.
Extended Data Fig. 2 The CRISPR-Cas9 knockout strategy and phenotype of RibosomeST-/- mice.
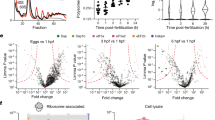
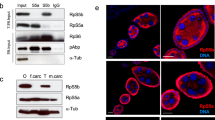
a, Schematic of CRISPR-Cas9 and sgRNAs used. b, Sequences of the three mutant strains generated by CRISPR-Cas9. c,d, Relative protein expression levels of RPL39L (pachytene spermatocytes: P = 5.63 × 10−6, spermatids: P = 1.47 × 10−6) (c) and RPL39 (pachytene spermatocytes: P = 1.04 × 10−4, spermatids: P = 1.09 × 10−4) (d) in RibosomeST-/- and wild-type mouse pachytene spermatocytes and spermatids, as evaluated by PRM. n = 3. e, RibosomeST-/- and wild-type testes with the 40S (P = 0.664), 60S (P = 0.664), 80S (P = 1.25 × 10−6), 2-6 (P = 0.712) and 7+ polysome (P = 0.489) peaks annotated and statistically compared. n = 5. f, Testis morphology and size of both RibosomeST-/- and wild-type mice. n = 3. Scale bar, 3 mm. g, Sperm morphology and prevalence of head (P = 7.86 × 10−4) and tail (P = 9.02 × 10−6) defects in RibosomeST-/- and wild-type sperm. n = 4. Scale bar, 10 μm. h, Levels of testosterone (P = 0.662), luteinizing hormone (LH) (P = 0.491) and follicle-stimulating hormone (FSH) (P = 0.437) in RibosomeST-/- and wild-type mice. n = 3. i, Average frequency of vaginal plugs per female mouse for RibosomeST-/- or wild-type male mice during fertility tests (P = 0.902, n = 3). j, Histology of wild-type and RibosomeST-/- testis showing seminiferous tubules at stage VII-VIII, circled by dashed lines, and amplified in the bottom panels. n = 3. PL: preleptotene spermatocyte, black arrowhead; PS: pachytene spermatocyte, red arrowhead; RS: round spermatid, green arrowhead; ES: elongated spermatid, blue arrowhead; SC: Sertoli cell. Top panels scale bar, 50 μm; Bottom panels scale bar, 10 μm. k, Histology of wild-type and RibosomeST-/- testis showing complete testicular section and stage IX sections together with statistics of percentages of stage IX (P = 0.179) and average number of unreleased sperms per stage IX tubule (P = 5.87 × 10−4). The red dotted circle indicates the seminiferous tubules of stage IX, and the black dotted line indicates delay-released sperms. Left panels scale bar, 500 μm; Right panels scale bar, 20 μm. n = 3. l, Histology of wild-type and RibosomeST-/- testis showing unreleased sperms at stage IX. Areas marked by dashed lines are shown at high magnification to the bottom. n = 3. L: leptotene spermatocyte, black arrowhead; PS: pachytene spermatocyte, red arrowhead; ES: elongated spermatid, blue arrowhead; SC: Sertoli cell; RB: residual body. Top panels scale bar, 50 μm; Bottom panels scale bar, 10 μm. m, The images of testes and testis/body weight ratio (P = 5.63 × 10−4, n = 6) of the three RibosomeST-/- strains and wild-type mice. Scale bar, 3 mm. n, Epididymal sperm count (Strain 2: P = 0.048, Strain 3: P = 0.045), motility (Strain 2: P = 4.03 × 10−4, Strain 3: P = 4.14 × 10−4) and progressive motility (Strain 2: P = 3.77 × 10−4, Strain 3: P = 4.05 × 10−4) in two more RibosomeST-/- strains and wild-type mice. n = 3. o, HE staining of the testis and epidydimal sperms from the three RibosomeST-/- strains. n = 3. Scale bar, 10 μm. p, Fertility of female RibosomeST-/- mice when mated with wild-type male mice (P = 0.606, n = 3). For c, d, e, g, h, i, k, m, n and p, data are mean ± s.e.m. P values were determined using two-tailed Student’s t-tests. n values represent the number of biologically independent animals.
Extended Data Fig. 3 The phenotypic analysis of RibosomeST-/- mice.
a, Immunofluorescence detection of LIN28 (red) with nuclei stained by Hoechst 33342 (blue), and the quantification of positively stained cells in three RibosomeST-/- strains and wild-type mice (Strain 1: P = 0.144, n = 4) (Strain 2: P = 0.759, Strain 3: P = 0.320, n = 3). Left panels scale bar, 500 μm; Right panels scale bar, 20 μm. b, Immunofluorescence detection of SYCP3 (green) and γH2AX (red) in leptotene, zygotene, pachytene and diplotene spermatocytes in RibosomeST-/- and wild-type testis and the quantification of each stage in RibosomeST-/- and wild-type mice. Scale bar, 10 μm. c, Immunofluorescence detection of SOX9 (red) with nuclei stained by Hoechst 33342 (blue), and the quantification of positively stained cells in RibosomeST-/- and wild-type mice (P = 0.161, n = 3). Left panels scale bar, 50 μm; Right panels scale bar, 20 μm. d, Fluorescence in situ hybridization of RibosomeST-/- and wild-type sperms using probes specific to X and Y chromosomes, and their corresponding ratio bar plot. Scale bar, 2 μm. e,f, Transmission electron microscopy analysis and schematic of abnormalities of RibosomeST-/- and wild-type sperms in epididymis. Red arrowhead: defects in doublet microtubules 4-7 of axoneme, blue arrowhead: other defects of axoneme. Scale bar, 1 μm. (e); The percentage of doublet microtubules 4-7 defects and other defects in axoneme of RibosomeST-/- sperm tail (P = 3.74 × 10−4, n = 3) (f). g, TUNEL staining (red) of RibosomeST-/- and wild-type testes with Hoechst 33342 stained nuclei (blue), and the quantification statistics of TUNEL positive cells per tubule (P = 0.009) and percentage of TUNEL positive tubules (P = 0.003, n = 3). Left panels scale bar, 50 μm; Right panels scale bar, 20 μm. For a, c, f and g, data are mean ± s.e.m. P values were determined using two-tailed Student’s t-tests. n values represent the number of biologically independent animals.
Extended Data Fig. 4 Proteomics and bioinformatics analysis of differential proteins in RibosomeST-/- spermatocytes and ESs.
a, Morphology and Hoechst 33342 (blue) staining of spermatocytes and ESs purified by Fluorescence-Activated Cell Sorting or STA-PUT from RibosomeST-/- and wild-type mice. n = 3. Scale bar, 20 μm. b, The protein expression changes in spermatocytes showing log2 (Fold change) (RibosomeST-/- vs. wild-type) and –log10 (FDR-adjusted q). c, Enriched GO terms for proteins downregulated in RibosomeST-/- ESs. d, Enriched GO terms for proteins upregulated in RibosomeST-/- ESs. e, Validation of PRM2 (P = 4.06 × 10−5), SMCP (P = 7.70 × 10−6), ATP1A4 (P = 1.23 × 10−5), H1FNT (P = 2.46 × 10−4), SPAG4 (P = 1.41 × 10−5), PGK2 (P = 1,84 × 10−5), ROPN1 (P = 6.09 × 10−7), ODF1 (P = 1.00 × 10−5), SPATA6 (P = 8.43 × 10−6), TSSK2 (P = 8.22 × 10−5), CST13 (P = 1.80 × 10−6), CRISP2 (P = 2.62 × 10−7), CCIN (P = 5.45 × 10−5), DBIL5 (P = 4.53 × 10−7), TCP11 (P = 6.34 × 10−9), UBE2J1 (P = 5.30 × 10−7), DCUN1D1 (P = 7.57 × 10−4), and DYNC1LI2 (P = 0.118) in RibosomeST-/- spermatids by relative targeted quantification using PRM (n = 3). f, Western blot analysis of PRM2 (P = 2.47 × 10−7), PGK2 (P = 0.017), SPATA6 (P = 2.06 × 10−4), TSSK2 (P = 6.99 × 10−4), CRISP2 (P = 0.047), TCP11 (P = 5.92 × 10−4), UBE2J1 (P = 0.007), DCUN1D1 (P = 2.38 × 10−4) and DYNC1LI2 (P = 1.00) in RibosomeST-/- spermatids. n = 3. g, Heat map of clustered mRNA and protein expression patterns of downregulated proteins with testis-specific expression at different stages of germ cell development. In clusters 1 and 2, the mRNA levels peaked in either spermatocytes or round spermatids, while the protein levels peaked at ESs. For clusters 3 and 4, all mRNA expression levels peaked in ESs, which agrees with the observed protein expression patterns. h, Distribution of the testis-specific proteins in downregulated and upregulated differentially expressed proteins in RibosomeST-/- ESs. i, Enriched GO terms for downregulated testis-specific proteins in RibosomeST-/- ESs. j, Overlap in differentially expressed proteins between RibosomeST-/- and control ESs, and differential genes in 80S, 2-6 and 7+ polysomes between RibosomeST-/- and wild-type testes by RNC-seq. k, Heat map of mRNA levels in the 80S, 2-6 or 7+ polysomes between RibosomeST-/- and wild-type testes by RNC-seq. l, Western blotting of nascent peptides using anti-Puromycin in spermatocytes (left) and spermatids (right) following O-propargyl-puromycin assays, β-Tubulin was used as a control. n = 3. m, Analysis of Ccin, Crisp2, Cst13, Dbil5, Dcun1d1, H1fnt, Odf1, Pgk2, Prm2, Ropn1, Spata6, Tcp11, Tssk2, Ube2j1, Atp1a4, Smcp, Spag4 and Dync1li2 mRNA levels in all polysomal fractions. n = 3. n, Translation ratio of differential proteomics in the 80S, 2-6 or 7+ polysomes between RibosomeST-/- and wild-type testes. o, Matrix of substitution identifications. Each entry in the matrix represents the number of independent substitutions detected for the corresponding (original codon, destination amino acid) pair in the MOPS-complete dataset. The logarithmic colour bar highlights the dynamic range of detection. Grey squares indicate substitutions from a codon to its cognate amino acid, substitutions from the stop codon, or substitutions undetectable via our method because they are indistinguishable from one of the PTMs or artifacts in the UniMod (http://www.unimod.org/) database. For e, f and m, data are mean ± s.e.m. P values were determined using two-tailed Student’s t-tests. n values represent the number of samples. Gel source data are provided in Supplementary Fig. 1.
Extended Data Fig. 5 Cryo-EM data processing and resolution evaluation of kidney and testis ribosomes.
a, Multiple sequence alignment of RPL39 and RPL39L from human, mouse, rat, Drosophila, C.elegans, and yeast. b, Representative micrograph of the kidney ribosome. c, Flow chart of the kidney ribosome cryo-EM data processing. d, Local resolution variation of kidney ribosome final map estimated with ResMap. e, Gold standard Fourier shell correlation (FSC) curves of the kidney ribosome final map showing that the resolution is 2.82 Å at FSC = 0.143. f, Representative micrograph of the testis ribosome. g, Flow chart of the testis ribosome cryo-EM data processing. h, Local resolution variation of testis ribosome final map estimated with ResMap. i, Gold standard Fourier shell correlation (FSC) curves of the testis ribosome final map showing that the resolution is 3.03 Å at FSC = 0.143.
Extended Data Fig. 6 Structure comparison of kidney RibosomeCore, testis RibosomeCore and testis RibosomeST.
a, Structure superposition of 60S subunits of kidney RibosomeCore (pink), testis RibosomeCore (grey), and testis RibosomeST (brick red). 40S ribosomes were omitted for clarity. c, Close view of RPL39 from kidney RibosomeCore (pink), RPL39 from testis RibosomeCore (grey), and RPL39L from testis RibosomeST (brick red). b,d, Density of RPL39 from kidney ribosome and density of RPL39 and RPL39L from testis ribosome (level 1.4). f, Close view of sidechains of R28 and R36 of RPL39 from kidney RibosomeCore (pink), R28 and R36 of RPL39 from testis RibosomeCore (grey),and Q28 and M36 of RPL39L from testis RibosomeST (brick red), indicating orientation of the sidechains. e,g, Close view of the density of R28 and R36 of RPL39 from kidney RibosomeCore (e); the density of R28 and R36 of RPL39 from testis RibosomeCore, and Q28 and M36 of RPL39L from testis RibosomeST (g). h,Close view of R28 and R36 of RPL39 from kidney RibosomeCore (pink), with density map omitted for clarity. i,j, Close view of R28 and R36 of RPL39 from testis RibosomeCore (grey; i), and Q28 and M36 of RPL39L from testis RibosomeST (brick red; j), with density map omitted for clarity.
Extended Data Fig. 7 Structural characteristic analysis of proteins downregulated in RibosomeST-/- ESs.
a, Downregulated proteins exhibited higher percentage of helix at amino acid positions of 10-11, 14-20, and 23-27 compared with unchanged proteins. Upregulated proteins exhibited a lower percentage of helix at amino acid positions from 23 to 38 compared with unchanged proteins. b, 3D structure of CCIN based on the AlphaFold Protein Structure Database (https://alphafold.ebi.ac.uk) with N-terminal 50 amino acids shown in blue, and the C-terminal half sequence shown in grey. c, Downregulated proteins formed more disulfide bonds in C-terminal half of the sequence than non-significance and upregulated proteins. d, 3D structure of CST13 based on the AlphaFold Protein Structure Database (https://alphafold.ebi.ac.uk) with N-terminal 50 amino acids coloured in blue, the C-terminal half sequence coloured in grey, and disulfide bonds are emphasized as red sticks. e, Amino acid enrichment analysis of upregulated proteins in RibosomeST-/- ESs. f, Levels of total thiols (P = 0.547) and glutathione (GSH) (P = 2.27 × 10−4) in RibosomeST-/- and wild-type N2a cell lines. n = 3. For a and f, data are mean ± s.e.m. P values were determined using two-sided Fisher’s exact tests for a; two-tailed Student’s t-tests for f. n values represent the number of independent experiments.
Extended Data Fig. 8 The construction of RibosomeST knockout N2a cell line.
a, Schematics of CRISPR-Cas9 and sgRNA against Rpl39l used in N2a cells. b, Sequences of the RibosomeST-/- N2a cell line generated by CRISPR-Cas9. c, Absorbance of ribosome profiles in RibosomeST-/- and wild-type N2a cell lines. d, Validation of TCP11 (P = 0.028), DCUN1D1 (P = 0.006) and DYNC1LI2 (P = 0.916) in RibosomeST-/- N2a cells by western blotting. n = 3. e, Absolute quantification of RPL39 and RPL39L protein expression levels in GC-1 ribosomes, and in RibosomeST-/- and wild-type N2a ribosomes by PRM (P = 0.004, n = 3). f, Quantification of Rpl39 mRNA in RibosomeST-/- and wild-type N2a cells (P = 0.034, n = 3). g, Western blotting of nascent peptides using anti-Puromycin in N2a cell lines following O-propargyl-puromycin assays, β-Tubulin was used as a control. n = 3. h, Quantification of Prm2, H1fnt, Ccin, and Dync1li2 in cell lines and testis. n = 3. For d, e, f and h, data are mean ± s.e.m. P values were determined using two-tailed Student’s t-tests. n values represent the number of independent experiments. Gel source data are provided in Supplementary Fig. 1.
Extended Data Fig. 9 Functional validation of RibosomeST in spermatids and cell lines.
a, Stability of PRM2 (P = 3.39 × 10−4) and DYNC1LI2 (P = 0.863) in RibosomeST-/- and wild-type spermatids by cycloheximide chase analysis. Protein level of each protein was normalized to β-Actin and then to time point = 0 h. n = 3. b, Stability of H1FNT-HA in RibosomeST-/- and wild-type N2a cells by cycloheximide chase analysis with the treatment of MG-132. Protein level was firstly normalized to eGFP and then to time point = 0 h (P = 0.049, n = 3). c, Western blotting showing the relative LC3II/LC3I ratio in RibosomeST-/- and wild-type N2a cells without (left, P = 0.100, n = 3) or with (right, P = 0.002, n = 5) H1FNT transfection. Stability of H1FNT-HA in RibosomeST-/- and wild-type N2a cells by cycloheximide chase analysis with the treatment of chloroquine (CQ). Protein level was normalized to eGFP and then to time point = 0 h (P = 0.818, n = 3). d, Stability of PRM2-HA (R28Q: P = 0.021, RPL39L: P = 0.004), H1FNT-HA (R28Q: P = 0.006, RPL39L: P = 0.001), CCIN-HA (R28Q: P = 0.044, RPL39L: P = 0.021), and DYNC1LI2-HA (R28Q: P = 0.208, RPL39L: P = 0.362) in RibosomeST-/- N2a cell lines overexpressing RPL39, RPL39(R28Q) and RPL39L by cycloheximide chase analysis, respectively. Protein level of each protein was normalized to eGFP and then to time point = 0 h. n = 3. e, Unfolded protein response signalling pathway analysis by western blotting of PERK (P = 0.804), eIF2α (P = 0.619), phosphorylated eIF2α (P = 0.608) and ATF6α (P = 2.07 × 10−4) with β-Tubulin as loading control, qPCR of Xbp1 (Xbp1s: P = 0.790, Xbp1u: P = 0.437, n = 4) splicing with 18S as an internal standard, and relative protein expression levels of HSP90B1 (FDR- adjusted q = 0.002, fold change < 1.5, Student’s t-test with BH FDR correction) and HSPA5 (FDR- adjusted q = 0.019, fold change < 1.5, Student’s t-test with BH FDR correction) according to TMT-based quantification by LC-MS/MS (Supplementary Table 2) in wild-type and RibosomeST-/-spermatids. n = 3. f, Distribution of endogenous proteins MOV10, GAPDH and β-Tubulin in the soluble (S) and pellet (P) fractions by western blotting in RibosomeST-/- and wild-type N2a cells. n = 3. g, Distribution of H1FNT-HA (R28Q: P = 0.020, RPL39L: P = 0.013) and DYNC1LI2-HA (R28Q: P = 0.393, RPL39L: P = 0.465) in the S and P fractions by western blotting and the protein expression levels shown as S/P ratio in RibosomeST-/- N2a cell lines overexpressing RPL39, RPL39(R28Q) and RPL39L, respectively. n = 3. h, SDS-resistance assay for protein OAZ3 synthesized in RibosomeST-/- and wild-type N2a cells. n = 3. i, Proportion of different wild-type and mutated alleles in wild-type N2a cells, 1st round of Cas9-edited N2a cells, monoclonal heterozygous Rpl39+/- N2a cell line derived from 1st round of edit, Rpl39+/- N2a cells subjected to 2nd round of Cas9-edit and selection for 4 days by blasticidin and puromycin, with and without additional 7 days of culture without selection. Those without additional 7 days of culture were also subjected to single cell sorting to establish monoclonal cell lines. n = 3. j, Morphology of RPL39L-IRES-eGFP and doxycycline (Dox)-inducible RPL39-overexpressing RibosomeCore-/Y GC-1 cells with or without Dox treatment for different ranges of time. The number of cells was corrected according the split ratio during passage. Scale bar, 20 μm. k, Absolute quantification of RPL39 and RPL39L levels in ribosomes of GC-1 wild-type cell, and RPL39L-IRES-eGFP-overexpressing RibosomeCore-/Y GC-1 cells with or without Dox-induced expression of RPL39 by PRM. n = 3. For a, b, c, d, e, g, i and k, data are mean ± s.e.m. P values were determined using two-tailed Student’s t-tests. n values represent the number of independent experiments. Gel source data are provided in Supplementary Fig. 1.
Extended Data Fig. 10 Linearity between the observed and expected signal ratios of the light and heavy peptides by PRM in serial dilution experiments.
The observed and expected signal ratios of the light and heavy peptides by PRM in serial dilution experiments showed good linearity.
Supplementary information
Supplementary Information
Supplementary Fig. 1 and full descriptions for Supplementary Tables 1–10.
Supplementary Table 1
Quantification of RPs in the 80S monosomes.
Supplementary Table 2
Differential proteins observed in different RibosomeST-knockout mouse germ cells.
Supplementary Table 3
GO analysis in this study.
Supplementary Table 4
mRNA and protein expression levels of testis-specific genes in different stages of germ cell development.
Supplementary Table 5
RNC-mRNA expression levels in mouse RibosomeST-knockout testis.
Supplementary Table 6
Cryo-EM data collection and refinement statistics.
Supplementary Table 7
Subunit lists modelled in kidney and testis ribosomes.
Supplementary Table 8
Structural feature analysis of downregulated proteins in mouse RibosomeST-knockout spermatids.
Supplementary Table 9
Primers and the peptide sequences used in this study.
Supplementary Table 10
Extracted ion chromatograms from protein quantification measurements by PRM.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, H., Huo, Y., He, X. et al. A male germ-cell-specific ribosome controls male fertility. Nature 612, 725–731 (2022). https://doi.org/10.1038/s41586-022-05508-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-05508-0
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.