Abstract
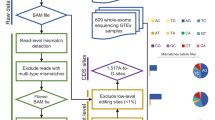
The accurate and thorough genome-wide detection of adenosine-to-inosine editing, a biologically indispensable process, has proven challenging. Here, we present a discovery pipeline in adult Drosophila, with 3,581 high-confidence editing sites identified with an estimated accuracy of 87%. The target genes and specific sites highlight global biological properties and functions of RNA editing, including hitherto-unknown editing in well-characterized classes of noncoding RNAs and 645 sites that cause amino acid substitutions, usually at conserved positions. The spectrum of functions that these gene targets encompass suggests that editing participates in a diverse set of cellular processes. Editing sites in Drosophila exhibit sequence-motif preferences and tend to be concentrated within a small subset of total RNAs. Finally, editing regulates expression levels of target mRNAs and strongly correlates with alternative splicing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Mallela, A. & Nishikura, K. A-to-I editing of protein coding and noncoding RNAs. Crit. Rev. Biochem. Mol. Biol. 47, 493–501 (2012).
Wulff, B.E., Sakurai, M. & Nishikura, K. Elucidating the inosinome: global approaches to adenosine-to-inosine RNA editing. Nat. Rev. Genet. 12, 81–85 (2011).
Danecek, P. et al. High levels of RNA-editing site conservation amongst 15 laboratory mouse strains. Genome Biol. 13, 26 (2012).
Peng, Z. et al. Comprehensive analysis of RNA-Seq data reveals extensive RNA editing in a human transcriptome. Nat. Biotechnol. 30, 253–260 (2012).
Ramaswami, G. et al. Identifying RNA editing sites using RNA sequencing data alone. Nat. Methods 10, 128–132 (2013).
Sam, L.T. et al. A comparison of single molecule and amplification based sequencing of cancer transcriptomes. PLoS ONE 6, e17305 (2011).
Kapranov, P. et al. The majority of total nuclear-encoded non-ribosomal RNA in a human cell is 'dark matter' un-annotated RNA. BMC Biol. 8, 149 (2010).
Raz, T. et al. Protocol dependence of sequencing-based gene expression measurements. PLoS ONE 6, e19287 (2011).
Wu, D., Lamm, A.T. & Fire, A.Z. Competition between ADAR and RNAi pathways for an extensive class of RNA targets. Nat. Struct. Mol. Biol. 18, 1094–1101 (2011).
Hoopengardner, B., Bhalla, T., Staber, C. & Reenan, R. Nervous system targets of RNA editing identified by comparative genomics. Science 301, 832–836 (2003).
Breiman, L. Random forests. Mach. Learn. 45, 5–32 (2001).
Ho, T. The random subspace method for constructing decision forests. IEEE Trans. Pattern Anal. Mach. Intell. 20, 832–844 (1998).
Palladino, M.J., Keegan, L.P., O'Connell, M.A. & Reenan, R.A. A-to-I pre-mRNA editing in Drosophila is primarily involved in adult nervous system function and integrity. Cell 102, 437–449 (2000).
Stark, A. et al. Discovery of functional elements in 12 Drosophila genomes using evolutionary signatures. Nature 450, 219–232 (2007).
Laurencikiene, J., Kallman, A.M., Fong, N., Bentley, D.L. & Ohman, M. RNA editing and alternative splicing: the importance of co-transcriptional coordination. EMBO Rep. 7, 303–307 (2006).
Rodriguez, J., Menet, J.S. & Rosbash, M. Nascent-seq indicates widespread cotranscriptional RNA editing in Drosophila. Mol. Cell 47, 27–37 (2012).
Ryman, K., Fong, N., Bratt, E., Bentley, D.L. & Ohman, M. The C-terminal domain of RNA Pol II helps ensure that editing precedes splicing of the GluR-B transcript. RNA 13, 1071–1078 (2007).
Reenan, R.A. Molecular determinants and guided evolution of species-specific RNA editing. Nature 434, 409–413 (2005).
Bass, B.L. RNA editing by adenosine deaminases that act on RNA. Annu. Rev. Biochem. 71, 817–846 (2002).
Lehmann, K.A. & Bass, B.L. Double-stranded RNA adenosine deaminases ADAR1 and ADAR2 have overlapping specificities. Biochemistry 39, 12875–12884 (2000).
Li, J.B. et al. Genome-wide identification of human RNA editing sites by parallel DNA capturing and sequencing. Science 324, 1210–1213 (2009).
Polson, A.G. & Bass, B.L. Preferential selection of adenosines for modification by double-stranded RNA adenosine deaminase. EMBO J. 13, 5701–5711 (1994).
Riedmann, E.M., Schopoff, S., Hartner, J.C. & Jantsch, M.F. Specificity of ADAR-mediated RNA editing in newly identified targets. RNA 14, 1110–1118 (2008).
Eggington, J.M., Greene, T. & Bass, B.L. Predicting sites of ADAR editing in double-stranded RNA. Nat. Commun. 2, 319 (2011).
Stefl, R. et al. The solution structure of the ADAR2 dsRBM-RNA complex reveals a sequence-specific readout of the minor groove. Cell 143, 225–237 (2010).
Graveley, B.R. et al. The developmental transcriptome of Drosophila melanogaster. Nature 471, 473–479 (2011).
Larschan, E. et al. X chromosome dosage compensation via enhanced transcriptional elongation in Drosophila. Nature 471, 115–118 (2011).
Park, Y. et al. Sequence-specific targeting of Drosophila roX genes by the MSL dosage compensation complex. Mol. Cell 11, 977–986 (2003).
Reichow, S.L., Hamma, T., Ferre-D'Amare, A.R. & Varani, G. The structure and function of small nucleolar ribonucleoproteins. Nucleic Acids Res. 35, 1452–1464 (2007).
St Laurent, G. III, Savva, Y.A. & Reenan, R. Enhancing non-coding RNA information content with ADAR editing. Neurosci. Lett. 466, 89–98 (2009).
Gardner, E.J., Nizami, Z.F., Talbot, C.C. Jr. & Gall, J.G. Stable intronic sequence RNA (sisRNA), a new class of noncoding RNA from the oocyte nucleus of Xenopus tropicalis. Genes Dev. 26, 2550–2559 (2012).
St Laurent, G. III et al. Intronic RNAs constitute the major fraction of the non-coding RNA in mammalian cells. BMC Genomics 13, 504 (2012).
Penalva, L.O. & Sanchez, L. RNA binding protein sex-lethal (Sxl) and control of Drosophila sex determination and dosage compensation. Microbiol Mol Biol Rev 67, 343–359 (2003).
Scadden, A.D. The RISC subunit Tudor-SN binds to hyper-edited double-stranded RNA and promotes its cleavage. Nat. Struct. Mol. Biol. 12, 489–496 (2005).
Huang, W., Sherman, B.T. & Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Witten, J.T. & Ule, J. Understanding splicing regulation through RNA splicing maps. Trends Genet. 27, 89–97 (2011).
Savva, Y.A. et al. Auto-regulatory RNA editing fine-tunes mRNA re-coding and complex behaviour in Drosophila. Nat. Commun. 3, 790 (2012).
Jepson, J.E. et al. Engineered alterations in RNA editing modulate complex behavior in Drosophila: regulatory diversity of adenosine deaminase acting on RNA (ADAR) targets. J. Biol. Chem. 286, 8325–8337 (2011).
Bhogal, B. et al. Modulation of dADAR-dependent RNA editing by the Drosophila fragile X mental retardation protein. Nat. Neurosci. 14, 1517–1524 (2011).
You, F.M. et al. BatchPrimer3: a high throughput web application for PCR and sequencing primer design. BMC Bioinformatics 9, 253 (2008).
Chun, H. & Keles, S. Sparse partial least squares regression for simultaneous dimension reduction and variable selection. J. R. Stat. Soc. Series B Stat. Methodol. 72, 3–25 (2010).
Liaw, A. & Weiner, M. Classification and regression by random forest. R News 2, 18–22 (2002).
Díaz-Uriarte, R. & Alvarez de Andres, S. Gene selection and classification of microarray data using random forest. BMC Bioinformatics 7, 3 (2006).
Sing, T., Sander, O., Beerenwinkel, N. & Lengauer, T. ROCR: visualizing classifier performance in R. Bioinformatics 21, 3940–3941 (2005).
R Development Core Team. R: A language and environment for statistical computing. 409 (2009).
Kuhn, M. Building predictive models in R using the caret package. J. Stat. Softw. 28, 1–26 (2008).
Crooks, G.E., Hon, G., Chandonia, J.M. & Brenner, S.E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Schneider, T.D. & Stephens, R.M. Sequence logos: a new way to display consensus sequences. Nucleic Acids Res. 18, 6097–6100 (1990).
Acknowledgements
We wish to thank Y. Vyatkin for help with the bioinformatics analysis, D. Jones for help with SMS, M. Mazaitis for help with figure preparation, R. Bell for help with validations and J. Finch for expert technical assistance. We also would like to thank L. Sugden and members of the Reenan laboratory for helpful discussions. R.A.R. was supported as an Ellison Medical Research Foundation Senior Investigator.
Author information
Authors and Affiliations
Contributions
G.S.L. and R.A.R. designed the experiments. M.R.T., Y.A.S. and G.S.L. conducted the experiments. S.N., D.S., D.A., G.S.L. and P.K. did the bioinformatics analyses. G.S.L., R.A.R., C.E.L., D.S., M.R.T., P.K. and D.A. evaluated the results. R.M. generated mutants by homologous recombination and performed RNA editing analysis. M.R.T., S.N., P.K., G.S.L. and D.A. produced the figures. G.S.L., R.A.R., Y.A.S., C.E.L. and P.K. wrote the manuscript, with contributions from all the authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Notes 1–6 (PDF 3709 kb)
Supplementary Tables
Supplementary Tables 1–29 (XLSX 4627 kb)
Rights and permissions
About this article
Cite this article
St Laurent, G., Tackett, M., Nechkin, S. et al. Genome-wide analysis of A-to-I RNA editing by single-molecule sequencing in Drosophila. Nat Struct Mol Biol 20, 1333–1339 (2013). https://doi.org/10.1038/nsmb.2675
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2675
This article is cited by
-
Deep transcriptome profiling reveals limited conservation of A-to-I RNA editing in Xenopus
BMC Biology (2023)
-
Lactobacillus for ribosome peptide editing cancer
Clinical and Translational Oncology (2023)
-
Membrane and synaptic defects leading to neurodegeneration in Adar mutant Drosophila are rescued by increased autophagy
BMC Biology (2020)
-
Adar RNA editing-dependent and -independent effects are required for brain and innate immune functions in Drosophila
Nature Communications (2020)
-
Evolutionarily significant A-to-I RNA editing events originated through G-to-A mutations in primates
Genome Biology (2019)