Abstract
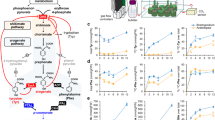
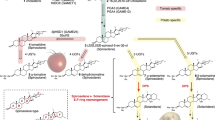
L-Tyrosine (Tyr) is essential for protein synthesis and is a precursor of numerous specialized metabolites crucial for plant and human health. Tyr can be synthesized via two alternative routes by different key regulatory TyrA family enzymes, prephenate dehydrogenase (PDH, also known as TyrAp) or arogenate dehydrogenase (ADH, also known as TyrAa), representing a unique divergence of primary metabolic pathways. The molecular foundation underlying the evolution of these alternative Tyr pathways is currently unknown. Here we characterized recently diverged plant PDH and ADH enzymes, obtained the X-ray crystal structure of soybean PDH, and identified a single amino acid residue that defines TyrA substrate specificity and regulation. Structures of mutated PDHs co-crystallized with Tyr indicate that substitutions of Asn222 confer ADH activity and Tyr sensitivity. Reciprocal mutagenesis of the corresponding residue in divergent plant ADHs further introduced PDH activity and relaxed Tyr sensitivity, highlighting the critical role of this residue in TyrA substrate specificity that underlies the evolution of alternative Tyr biosynthetic pathways in plants.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Moghe, G.D. & Last, R.L. Something old, something new: conserved enzymes and the evolution of novelty in plant specialized metabolism. Plant Physiol. 169, 1512–1523 (2015).
Weng, J.-K., Philippe, R.N. & Noel, J.P. The rise of chemodiversity in plants. Science 336, 1667–1670 (2012).
Gowik, U. & Westhoff, P. The path from C3 to C4 photosynthesis. Plant Physiol. 155, 56–63 (2011).
Torruella, G., Suga, H., Riutort, M., Peretó, J. & Ruiz-Trillo, I. The evolutionary history of lysine biosynthesis pathways within eukaryotes. J. Mol. Evol. 69, 240–248 (2009).
Jensen, R.A. & Pierson, D.L. Evolutionary implications of different types of microbial enzymology for L-tyrosine biosynthesis. Nature 254, 667–671 (1975).
Schenck, C.A., Chen, S., Siehl, D.L. & Maeda, H.A. Non-plastidic, tyrosine-insensitive prephenate dehydrogenases from legumes. Nat. Chem. Biol. 11, 52–57 (2015).
Maeda, H. & Dudareva, N. The shikimate pathway and aromatic amino acid biosynthesis in plants. Annu. Rev. Plant Biol. 63, 73–105 (2012).
Tzin, V. & Galili, G. New insights into the shikimate and aromatic amino acids biosynthesis pathways in plants. Mol. Plant 3, 956–972 (2010).
Fernstrom, J.D. & Fernstrom, M.H. Tyrosine, phenylalanine, and catecholamine synthesis and function in the brain. J. Nutr. 137 (Suppl. 1), 1539S–1547S (2007).
Gleadow, R.M. & Møller, B.L. Cyanogenic glycosides: synthesis, physiology, and phenotypic plasticity. Annu. Rev. Plant Biol. 65, 155–185 (2014).
Hagel, J.M. & Facchini, P.J. Benzylisoquinoline alkaloid metabolism: a century of discovery and a brave new world. Plant Cell Physiol. 54, 647–672 (2013).
Hunter, S.C. & Cahoon, E.B. Enhancing vitamin E in oilseeds: unraveling tocopherol and tocotrienol biosynthesis. Lipids 42, 97–108 (2007).
Millner, P.A. & Barber, J. Plastoquinone as a mobile redox carrier in the photosynthetic membrane. FEBS Lett. 169, 1–6 (1984).
Strack, D., Vogt, T. & Schliemann, W. Recent advances in betalain research. Phytochemistry 62, 247–269 (2003).
Barros, J. et al. Role of bifunctional ammonia-lyase in grass cell wall biosynthesis. Nat. Plants 2, 16050 (2016).
Bonner, C.A. et al. Cohesion group approach for evolutionary analysis of TyrA, a protein family with wide-ranging substrate specificities. Microbiol. Mol. Biol. Rev. 72, 13–53 (2008).
Fischer, R. & Jensen, R. Prephenate dehydrogenase (monofunctional). Methods Enzymol. 142, 503–507 (1987).
Hudson, G.S., Wong, V. & Davidson, B.E. Chorismate mutase/prephenate dehydrogenase from Escherichia coli K12: purification, characterization, and identification of a reactive cysteine. Biochemistry 23, 6240–6249 (1984).
Kuramitsu, S., Inoue, K., Ogawa, T., Ogawa, H. & Kagamiyama, H. Aromatic amino acid aminotransferase of Escherichia coli: nucleotide sequence of the tyrB gene. Biochem. Biophys. Res. Commun. 133, 134–139 (1985).
Dal Cin, V. et al. Identification of genes in the phenylalanine metabolic pathway by ectopic expression of a MYB transcription factor in tomato fruit. Plant Cell 23, 2738–2753 (2011).
Dornfeld, C. et al. Phylobiochemical characterization of class-Ib aspartate/prephenate aminotransferases reveals evolution of the plant arogenate phenylalanine pathway. Plant Cell 26, 3101–3114 (2014).
Graindorge, M. et al. Identification of a plant gene encoding glutamate/aspartate-prephenate aminotransferase: the last homeless enzyme of aromatic amino acids biosynthesis. FEBS Lett. 584, 4357–4360 (2010).
Maeda, H., Yoo, H. & Dudareva, N. Prephenate aminotransferase directs plant phenylalanine biosynthesis via arogenate. Nat. Chem. Biol. 7, 19–21 (2011).
Connelly, J.A. & Conn, E.E. Tyrosine biosynthesis in Sorghum bicolor: isolation and regulatory properties of arogenate dehydrogenase. Z. Naturforsch. C 41, 69–78 (1986).
Gaines, C.G., Byng, G.S., Whitaker, R.J. & Jensen, R.A. L-tyrosine regulation and biosynthesis via arogenate dehydrogenase in suspension-cultured cells of Nicotiana silvestris Speg. et Comes. Planta 156, 233–240 (1982).
Rippert, P. & Matringe, M. Purification and kinetic analysis of the two recombinant arogenate dehydrogenase isoforms of Arabidopsis thaliana. Eur. J. Biochem. 269, 4753–4761 (2002).
Keller, B., Keller, E. & Lingens, F. Arogenate dehydrogenase from Streptomyces phaeochromogenes. Purification and properties. Biol. Chem. Hoppe Seyler 366, 1063–1066 (1985).
Mayer, E., Waldner-Sander, S., Keller, B., Keller, E. & Lingens, F. Purification of arogenate dehydrogenase from Phenylobacterium immobile. FEBS Lett. 179, 208–212 (1985).
Gamborg, O.L. & Keeley, F.W. Aromatic metabolism in plants. I. A study of the prephenate dehydrogenase from bean plants. Biochim. Biophys. Acta 115, 65–72 (1966).
Rubin, J.L. & Jensen, R.A. Enzymology of L-tyrosine biosynthesis in mung bean (Vigna radiata [L.] Wilczek). Plant Physiol. 64, 727–734 (1979).
Legrand, P. et al. Biochemical characterization and crystal structure of Synechocystis arogenate dehydrogenase provide insights into catalytic reaction. Structure 14, 767–776 (2006).
Song, J., Bonner, C.A., Wolinsky, M. & Jensen, R.A. The TyrA family of aromatic-pathway dehydrogenases in phylogenetic context. BMC Biol. 3, 13 (2005).
Christendat, D., Saridakis, V.C. & Turnbull, J.L. Use of site-directed mutagenesis to identify residues specific for each reaction catalyzed by chorismate mutase-prephenate dehydrogenase from Escherichia coli. Biochemistry 37, 15703–15712 (1998).
Sun, W. et al. The crystal structure of Aquifex aeolicus prephenate dehydrogenase reveals the mode of tyrosine inhibition. J. Biol. Chem. 284, 13223–13232 (2009).
Christendat, D. & Turnbull, J.L. Identifying groups involved in the binding of prephenate to prephenate dehydrogenase from Escherichia coli. Biochemistry 38, 4782–4793 (1999).
Sun, W., Singh, S., Zhang, R., Turnbull, J.L. & Christendat, D. Crystal structure of prephenate dehydrogenase from Aquifex aeolicus. Insights into the catalytic mechanism. J. Biol. Chem. 281, 12919–12928 (2006).
Lütke-Eversloh, T. & Stephanopoulos, G. Feedback inhibition of chorismate mutase/prephenate dehydrogenase (TyrA) of Escherichia coli: generation and characterization of tyrosine-insensitive mutants. Appl. Environ. Microbiol. 71, 7224–7228 (2005).
Chiu, H.J. et al. The structure of Haemophilus influenzae prephenate dehydrogenase suggests unique features of bifunctional TyrA enzymes. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 66, 1317–1325 (2010).
Ku, H., Park, S., Yang, I. & Kim, S. Expression and functional characterization of prephenate dehydrogenase from Streptococcus mutans. Process Biochem. 45, 607–612 (2010).
Wierenga, R.K., De Maeyer, M.C.H. & Hol, W.G.J. Interaction of pyrophosphate moieties with alpha-helices in dinucleotide-binding proteins. Biochemistry 24, 1346–1357 (1985).
Rippert, P., Puyaubert, J., Grisollet, D., Derrier, L. & Matringe, M. Tyrosine and phenylalanine are synthesized within the plastids in Arabidopsis. Plant Physiol. 149, 1251–1260 (2009).
Wang, M., Toda, K. & Maeda, H.A. Biochemical properties and subcellular localization of tyrosine aminotransferases in Arabidopsis thaliana. Phytochemistry 132, 16–25 (2016).
Westfall, C.S., Xu, A. & Jez, J.M. Structural evolution of differential amino acid effector regulation in plant chorismate mutases. J. Biol. Chem. 289, 28619–28628 (2014).
Cardoso, D. et al. Revisiting the phylogeny of papilionoid legumes: new insights from comprehensively sampled early-branching lineages. Am. J. Bot. 99, 1991–2013 (2012).
Wojciechowski, M.F., Lavin, M. & Sanderson, M.J. A phylogeny of legumes (Leguminosae) based on analysis of the plastid matK gene resolves many well-supported subclades within the family. Am. J. Bot. 91, 1846–1862 (2004).
Reyes-Prieto, A. & Moustafa, A. Plastid-localized amino acid biosynthetic pathways of Plantae are predominantly composed of non-cyanobacterial enzymes. Sci. Rep. 2, 955–967 (2012).
Bonvin, J. et al. Biochemical characterization of prephenate dehydrogenase from the hyperthermophilic bacterium Aquifex aeolicus. Protein Sci. 15, 1417–1432 (2006).
Goodstein, D.M. et al. Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res. 40, D1178–D1186 (2012).
Matasci, N. et al. Data access for the 1,000 Plants (1KP) project. Gigascience 3, 17 (2014).
Dash, S. et al. Legume information system (LegumeInfo.org): a key component of a set of federated data resources for the legume family. Nucleic Acids Res. 44, D1181–D1188 (2016).
Fernández-Pozo, N. et al. EuroPineDB: a high-coverage web database for maritime pine transcriptome. BMC Genomics 12, 366 (2011).
Whelan, S. & Goldman, N. A general empirical model of protein evolution derived from multiple protein families using a maximum-likelihood approach. Mol. Biol. Evol. 18, 691–699 (2001).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 30, 2725–2729 (2013).
Edgar, R.C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Saitou, N. & Nei, M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425 (1987).
Gorrec, F. The current approach to initial crystallization screening of proteins is under-sampled. J. Appl. Crystallogr. 46, 795–797 (2013).
Minor, W., Cymborowski, M., Otwinowski, Z. & Chruszcz, M. HKL-3000: the integration of data reduction and structure solution–from diffraction images to an initial model in minutes. Acta Crystallogr. D Biol. Crystallogr. 62, 859–866 (2006).
Sheldrick, G.M. A short history of SHELX. Acta Crystallogr. A 64, 112–122 (2008).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Terwilliger, T.C. Maximum-likelihood density modification. Acta Crystallogr. D Biol. Crystallogr. 56, 965–972 (2000).
Morris, R.J., Perrakis, A. & Lamzin, V.S. ARP/wARP and automatic interpretation of protein electron density maps. Methods Enzymol. 374, 229–244 (2003).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Maeda, H. et al. RNAi suppression of Arogenate Dehydratase1 reveals that phenylalanine is synthesized predominantly via the arogenate pathway in petunia petals. Plant Cell 22, 832–849 (2010).
Trott, O. & Olson, A.J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461 (2010).
Acknowledgements
We thank K. Wallin (University of Minnesota) and U. (Rosia) Schmidt for the cloning of MtncADH and SlncADH, respectively, and P.-M. Delaux (Universite´ de Toulouse) for help with a legume phylogenetic analysis. This work was supported by the National Science Foundation (IOS-1354971 to H.A.M. and MCB-1614539 to J.M.J.). C.K.H. was supported by the NSF Graduate Research Fellowship Program (DGE-1143954). Portions of this research were carried out at the Argonne National Laboratory Structural Biology Center of the Advanced Photon Source, a national user facility operated by the University of Chicago for the Department of Energy Office of Biological and Environmental Research (DE-AC02-06CH11357).
Author information
Authors and Affiliations
Contributions
C.A.S., C.K.H., J.M.J., and H.A.M. designed the research, C.A.S. performed phylogenetic analyses, and C.A.S., M.R.S., and Y.M. performed site-directed mutagenesis and biochemical characterization of recombinant enzymes. C.K.H. expressed, purified, and crystallized proteins, collected diffraction data, and determined crystal structures. S.G.L. assisted with SAD phasing, C.A.S., C.K.H., J.M.J., and H.A.M. analyzed data, and C.A.S., C.K.H., J.M.J., and H.A.M. wrote the paper with all authors providing editorial input.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Tables 1–3, Supplementary Figures 1–8 (PDF 11055 kb)
Supplementary Dataset 1
Sequences used in the phylogenetic analyses from Figure 1 and Supplementary Figure 1 (XLSX 49 kb)
Supplementary Dataset 2
Full amino acid alignment made in MUSCLE used to construct the phylogenetic analyses in Supplementary Fig. 1 (TXT 278 kb)
Rights and permissions
About this article
Cite this article
Schenck, C., Holland, C., Schneider, M. et al. Molecular basis of the evolution of alternative tyrosine biosynthetic routes in plants. Nat Chem Biol 13, 1029–1035 (2017). https://doi.org/10.1038/nchembio.2414
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2414
This article is cited by
-
Coordinated regulation of the entry and exit steps of aromatic amino acid biosynthesis supports the dual lignin pathway in grasses
Nature Communications (2023)
-
Intervening Role of Tyrosine in Cadmium Detoxification, Balancing of Mineral Ions Homeostasis, Antioxidants, and Secondary Metabolites in Maize
Journal of Soil Science and Plant Nutrition (2023)
-
Using interdisciplinary, phylogeny-guided approaches to understand the evolution of plant metabolism
Plant Molecular Biology (2022)
-
Orphan crops at the food for future conference
Planta (2019)
-
Aromatic amino acid aminotransferases in plants
Phytochemistry Reviews (2018)