Abstract
The study of schizophrenia genetics has confirmed the importance of genes in etiology, but has not so far identified the relationship between observed genetic risks and specific DNA variants, protein alterations or biological processes. In spite of many limitations, numerous regions of the human genome give consistent, although by no means unanimous, support for linkage, which is unlikely to occur by chance. Two recent shifts have been evident in the field. First, a series of studies combining linkage and association analyses in the same family sets have identified promising candidate genes (DTNBP1, NRG1, G72/G30, TRAR4). Although a consensus definition of replication for genetic association in a complex trait remains difficult to achieve, the evidence for two of these (dystrobrevin binding protein 1 (DTNBP1), NRG1) is strong. Second, a series of studies combining association with functional investigation of changes in the associated gene in schizophrenia have also identified several candidate genes (COMT, RGS4, PPP3CC, ZDHHC8, AKT1). Somewhat surprisingly, the loci implicated by these studies have proven less robust in replication, although the number of replication studies remains small in several cases. Assessment of the combined evidence for the DTNBP1 gene gives some insight into the nature of the problems remaining to be solved.
Similar content being viewed by others
Genetic epidemiology
A large body of data collected from families, twins and adoptees over many years has consistently supported the involvement of a major, complex genetic component in liability to schizophrenia and schizophrenia spectrum disorders.
Family studies: does risk aggregate in relatives?
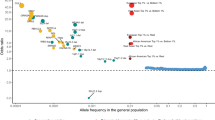
The lifetime morbid risks (MR) from combined results of studies published between 1921 and 19871 are shown in Figure 1. Early family studies were criticized for lack of proper controls, nonsystematic sampling, lack of standardized diagnostic criteria and failure to diagnose family members blind to the status of the index case (or proband). A combined analysis of data from seven later studies that avoided these weaknesses yielded MR for narrowly defined schizophrenia in relatives of patients of 4.8%, 10 times the 0.5% average MR in relatives of controls.2
Lifetime MR for schizophrenia in various classes of relatives of a proband, adapted from Gottesman.1
Twin studies: how large is the genetic component of risk?
Twin studies of schizophrenia, (reviewed in Cardno and Gottesman3) estimate the relative importance of genes and environment in liability and show consistently higher concordance in monozygotic (MZ, ∼50%) than dizygotic (DZ, ∼17%) twins. Both individual twin studies and meta-analyses of twin studies estimate the heritability of liability to schizophrenia to be approximately 80%. Some find evidence that a small proportion of liability to schizophrenia results from shared environmental risk factors. Recent studies have suggested that gene–environment interactions may be components of the overall risk.4 Schizophrenia is thus largely genetically mediated but not genetically determined.
Adoption studies: are genetic factors evident when shared familial environment is controlled?
Across all adoption studies performed, increased risk of schizophrenia was present in the biological relatives of individuals with schizophrenia.5 In 361 families in Finland, 4.9% of adopted away children of schizophrenic mothers have schizophrenia and 9.1% have a schizophrenia spectrum disorder, whereas 1.1% of adopted away offspring of control mothers have schizophrenia.6 In Denmark, schizophrenia was significantly more common in the biological relatives of schizophrenic adoptees than in those of control adoptees.7 The rates of schizophrenia were low and not different in the adoptive families of both affected and control groups.
Segregation analysis: how is the genetic risk transmitted?
Transmission models make specific predictions about risk in various classes of relatives and are tested by assessing how well their predictions match observed patterns of risk. Neither the explicitly monogenic and deterministic generalized single major locus nor the less deterministic and explicitly polygenic multifactorial threshold (MT) models are adequate to explain observed patterns of risk.8 The MT model assumes that the effects of multiple genes combine additively (the total liability from n genes is equal to the sum of the n individual liabilities) and do not interact. A key predictive failure of the MT model is that the observed MZ twin concordance is too high relative to that in siblings and DZ twins. This pattern is more consistent with at least some epistatic interactions between loci, where the total liability is greater than the sum of the n individual liabilities. The inheritance model is unlikely to be fully epistatic, however, because the tetrachoric correlation in MZ relative to DZ twins should be substantially more than observed. It therefore seems that numerous kinds of influences (additive and epistatic genes, environment and G × E interactions) are involved and that the precise model reflecting an individual's or a family's risk may vary depending on the specific risk factors active in them.
Spectrum disorders: how broad is the range of psychiatric illness transmitted and who do we consider affected?
In general, the risk for all psychotic spectrum disorders (including schizoaffective, schizophreniform, delusional, paranoid personality and schizotypal personality disorders) is increased in the relatives of schizophrenics.2 The risk of schizophrenia is also significantly higher in the relatives of individuals with spectrum disorders. Most studies suggest that relatives of schizophrenic patients are not at increased risk of anxiety or alcohol and drug dependence disorders.9 The results for bipolar affective disorder are less clear, but taken together suggest that schizophrenia and bipolar disorder might have both shared and independent genetic risk factors. It is therefore reasonably common to perform several analyses of data using a number of different definitions of illness.
Molecular genetics
Approaches: linkage and association
The issues surrounding the strengths and weaknesses of linkage and association in complex traits have been discussed at length.10 Both are impeded by unclear diagnostic boundaries, multiple loci, epistasis, clinical and etiologic heterogeneity and G × E interactions. Because of their fundamental differences, the two approaches have generally been considered separately. More and more, however, association studies are following linkage evidence. Analytic developments in family-based association testing have furthered association studies in samples originally collected for linkage. The sequential application of these two approaches has produced the most exciting current results, including a small but growing number of specific genes, which we will focus on, for which multiple groups have found support. A second recent and important shift in the field has been towards obtaining functional evidence to support association evidence, and publishing both together. Many of these associations have proven less robust in replication studies that generally do not include replication of the functional component.
A total of 25 genome scans for schizophrenia have now been published, with no evidence for a gene of major effect. To date, eight linkage regions have identified promising candidate genes: 22q12–q13, 8p22–p21, 6p24–p22, 13q14–q32, and 6q21–q22. The interpretation of these results is controversial, particularly as the definition of replication for linkage to a complex trait remains uncertain. For brevity, we omit studies that do not find evidence for linkage in these selected regions, so there is selective bias in the results that follow. Full references for all studies mentioned can be found in Riley and Kendler.10 We do discuss both positive and negative studies of the current candidate genes.
Chromosome 22q linkage studies
In a sample of families from Maryland, three markers spanning ∼23 cM in the 22q13.1 region gave evidence for linkage.11 Two of a number of attempted replications were positive. A total of 11 groups contributed data for the most significant marker from the Maryland sample to the first collaborative schizophrenia linkage study. There was excess sharing of alleles in the affected pairs (P=0.006), particularly in those with data from both parents (P=0.001), but the locus accounts for no more than 2% of total variance in liability.12 Velo-cardio-facial syndrome (VCFS) is caused by deletions nearby at 22q11. Historically, about 10% of VCFS patients were thought to present with a psychotic phenotype, but more recent studies suggest much higher rates of 25–29%.13 The VCFS critical region contains three genes that have been investigated in schizophrenia.
Chromosome 22q candidate genes
COMT
The gene for catechol-O-methyl transferase (COMT), involved in the synthesis and degradation of catecholamines, is functionally polymorphic, with a variable amino acid, Val158Met. Val and Met alleles are of almost identical frequency. Most studies of the COMT gene have tested for association with the low activity (Met) allele (with mixed results). One recent report suggests that the high activity (Val) allele, through increased catabolism of dopamine in the prefrontal cortex, may slightly increase the risk of schizophrenia and may explain some of the observed differences in cognitive performance and prefrontal cortical functioning between cases and controls.14 In a large, homogeneous case–control sample of Ashkenazi Jewish individuals from Israel, the homozygous high activity genotype (Val/Val) and two other SNPs showed significant association with schizophrenia in ∼720 cases and 2000–4000 controls.15
Three recent meta-analyses of COMT association with schizophrenia have provided mixed (though generally negative) results. One found that COMT was associated only in cases of European descent.16 The others did not confirm either association or ethnic differences.17 Finally, the association between COMT and schizophrenia has been re-examined in a recent large-scale European study involving 709 cases, 710 controls and 488 parent–child trios.18 Markers and haplotypes positive in prior studies were tested, but despite substantial power, no evidence of significant association was found.
PRODH2 and ZDHHC8
The proline dehydrogenase (PRODH2) gene gave evidence for association in samples of adult and childhood-onset schizophrenia.19 Marker data from a third independent sample of cases and controls did not reach conventional significance, but analysis of the childhood-onset subset of this case–control sample was significant for association of the same marker alleles with disease. However, most replication studies detect no association between psychotic phenotypes, primarily schizophrenia, and common PRODH2 polymorphisms.20 Association with a third locus in the deletion region, ZDHHC8, was detected by the same group that reported originally on PRODH2.21 Again, follow-up reports have been mixed, but generally have not supported this association.22
Chromosome 8p22–p21 linkage studies
The Maryland family sample also gave the first evidence of linkage to 8p22–p21.23 A multicenter collaborative linkage study supported this putative locus with excess allele sharing at D8S261.24 Data from pedigrees from numerous different ethnic backgrounds all support a locus on 8p, but these results are spread across ∼15 Mb of sequence. One of the key points to note is that although numerous samples support a locus on this chromosome, comparison between individual studies is consistent with the possibility of multiple susceptibility genes in the region.
Chromosome 8p22–p21 candidate genes
NRG1
Following linkage evidence to 8p in Icelandic families, fine mapping with 50 markers across a 30 cM interval identified two risk haplotypes spanning a region of ∼1 Mb within the gene for neuregulin 1 (NRG1).25 Case–control samples from Scotland26 and Ireland27 have provided additional support for this locus and for haplotypes identical or closely related to those identified in the Icelandic cases. Studies in other populations have provided support for association,28 although not with the specific haplotypes seen in the Icelandic, Scottish and Irish samples. A smaller number of studies have found no evidence for association.29 Neuregulin is expressed in CNS synapses and appears to have a role in the expression and activation of neurotransmitter (including glutamate) receptors.
PPP3CC
Parallel strands of evidence implicate the calcineurin A gamma (CNAγ) subunit gene (PPP3CC) in schizophrenia. Calcineurin is a dimeric calcium-dependent serine/threonine phosphatase composed of a regulatory (CNB) and one of three catalytic (CNA) subunits and is highly expressed in the central nervous system. First, case/control association testing of calcineurin-related genes mapping to linkage regions identified association with both individual markers and marker haplotypes in PPP3CC (8p21.3).30 Second, conditional knockout mice lacking CNB expression in the forebrain display traits of potential relevance to schizophrenia, including decreased social interaction, impaired prepulse and latent inhibition and severe working memory deficits.31 Neither of the genes encoding the CNB (2p14) or CNAα (4q24) subunits gave evidence for association. Only one follow-up study of PPP3CC has been published and did not replicate the association.32
Chromosome 6p24–p22 linkage studies
The first evidence for linkage of schizophrenia to the 6p region came from studies of Irish families with a high density of disease.33 In data from 16 markers, evidence for linkage was modest under a narrow diagnostic model, but increased substantially as the diagnostic definition broadened to include spectrum disorders. Evidence for linkage fell when the definition was broadened further to include nonspectrum disorders. To date, nine independent reports of analyses of this region of 6p have been published, six of which are positive.10
Chromosome 6p24–p22 candidate genes
DTNBP1
Follow-up work in the Irish family set demonstrated a positive association in the dystrobrevin binding protein 1 or dysbindin (DTNBP1) gene.34, 35 Replication studies have been generally supportive.36, 37, 38, 39, 40, 41, 42, 43 These results are discussed in detail below. The function of DTNBP1 in the brain is unknown, but expression of DTNBP1 is reduced in certain brain regions of patients with schizophrenia at both RNA44 and protein45 levels. Reduced protein expression is associated with additional changes consistent with a role in glutamatergic neurotransmission, currently a system of great interest for schizophrenia liability. Overexpression of DTNBP1 is associated with increased phosphorylation and activity of AKT1 in neuronal culture, suggesting that DTNBP1 also interacts with the AKT signaling pathway, which mediates cell survival. In this context, it is interesting to note that the AKT1 gene has recently shown association with schizophrenia.46 This association has been confirmed in two of the three published follow-up studies.47, 48, 49
Chromosome 13q14–q32 linkage studies
Data from a mixed sample of UK and Japanese families initially suggested linkage to 13q14.1–q32,50 which is of interest as the region contains the 5HT-2A receptor gene. Data from five studies of six independent samples provided support for the region, although some results were extremely distant from other findings in the region. The results from chromosome 13 are particularly difficult to interpret because of the very large distances between positive markers. Overall, the combined linkage reports are spread over a region of ∼60 Mb, containing ∼120 known or putative genes. On the other hand, although locations are much less certain on 13q than in other linkage regions, this chromosome has produced some of the most significant linkage evidence seen in the studies of schizophrenia.
Chromosome 13q14–q32 candidate genes
G72 and DAAO
An elegant recent study examined markers in the distal 5 Mb of this broad linkage region, site of one of the most significant findings on chromosome 13.51 Nearly 200 SNPs were tested across the region and identified two regions of association. In one of these regions, two genes (G72, now called D-amino acid oxidase activator, DAOA) and G30 were investigated. Of note, the exons of these genes could not be predicted by any computational method tested, suggesting that they are highly novel in their sequence and organization. Both genes show alternative transcripts in the brain and other tissues. Association studies of SNPs within G72 have not yet provided a clear pattern. One of these is nonsynonymous and is significant alone. The nature of the amino-acid change (lysine to arginine) is conservative, but has major functional consequences in some proteins. However, the overall pattern of results is probably most consistent with the existence of further unidentified predisposing variants in this gene.
D-amino acid oxidase (DAO) is activated by the protein product of G72. Four SNPs in the DAO gene on 12q24 were significantly associated. Results of this kind (showing association in two interacting genes in the same sample) are rare, so this study had a unique opportunity to test for an epistatic genetic interaction. Evidence for epistasis was observed for one pair of DAAO and G72 genotypes, supporting a potential interaction between them in risk for schizophrenia. Replication studies have generally provided confirmation of a role for G72.52, 53, 54, 55, 56, 57 Only one negative report has appeared.58
Chromosome 6q21–q22 linkage studies
A sample of 53 US families provided initial evidence for a susceptibility locus on 6q21–q22.3.59 This study is unique in that two additional independent samples of families held by this group supported the results from the first sample. An interval of ∼8 Mb gave the strongest results with an LOD of 3.82 and highly significant excess allele sharing. A collaborative study of this region was highly significant.60
Chromosome 6q21–q22 candidate genes
TRAR4
A recent paper from the original group following up their linkage findings with linkage disequilibrium (LD) studies in the same samples has identified association in the trace amine receptor 4 (TRAR4) gene located at 6q23.2.61 Two replication studies of this locus did not find evidence for association,62, 63 although neither examined samples of similar ethnicity to the original report.
Chromosome 1q32–q42 and 1q21–q22
Interest in chromosome 1 in schizophrenia began with reports of a balanced 1:11 translocation segregating with serious mental illness in a large pedigree from Scotland. The chromosome 1 breakpoint lies at 1q42.1, and two groups reported suggestive linkage findings in this region, in a population isolate from Finland and in the Maryland sample. Genome scans of families from a mixed sample of US/UK families64 and Canada65 provided evidence for a 1q21–q22 locus. The latter provided very strong LOD scores but this region was not replicated by a large collaborative study.66 Unlike the results discussed above, candidate loci on 1q have not been identified through family-based association follow-up of linkage data, but have come instead from studies of chromosomal rearrangements and microarray data.
Chromosome 1q42 candidate genes
DISC1
Ongoing work in the Scottish pedigree has now identified three genes disrupted by the breakpoint, one of which, disrupted in schizophrenia 1 (DISC1), has been intensively studied.67 Studies of this locus in samples with no translocation have been mixed, but positive results68, 69, 70, 71 outnumber negative ones.72, 73, 74 A frameshift mutation has also been identified in a single multiply affected family.75 DISC1 appears to have a role in cytoskeletal regulation, and thus may affect neuronal migration, neurite outgrowth and intracellular transport.76, 77
Chromosome 1q21–q22 candidate genes
RGS4
Microarray studies of post-mortem schizophrenic brain suggested that the regulator of G-protein signalling 4 (RGS4) gene showed altered expression in schizophrenia.78 RGS4 maps to the 1q21–q22 linkage region. A study of US and Indian pedigrees showed association with the same markers in both, although with different specifics.79 One replication study supported the RGS4 association with schizophrenia liability,80 whereas others have not.81, 82, 83, 84
Meta-analyses of genome screen data
Meta-analysis of whole genome screen data represents a first approximation of a very large, multisample genome screen. The strongest method ranks 30 cM bins of the genome from most positive to least positive for each study, sums the ranks for each bin and calculates significance by simulation. As this method uses the actual marker LOD scores from each component study, it can identify regions of the genome showing modest positive results across many samples but which may have been overlooked in individual studies. Results of this approach supported linkage to chromosomes 6p, 8p and 10p.85 However, the strongest evidence for a potential locus was on 2q, a region suggested by only a few studies and not widely followed up, and on 3p, the site of an early linkage finding which could never be replicated. Finally, significant evidence of linkage was also detected for two regions never previously implicated by an individual study, on chromosomes 11q and 14p.
Discussion
Current linkage regions and candidate genes
The best supported linkage regions have been replicated in other samples in addition to the one in which they were first reported but have not been replicated in all studies. Collaborative replication has been at least suggestive of an effect. The collected evidence for current candidate genes displays both similarities and contrasts to the linkage evidence. The reported associations for several of these candidates, DTNBP1, NRG1, G72, RGS4, DISC1 and AKT1 have been replicated in other samples. Critically, positive replications outnumber negative ones for all these loci. Some candidates, such as TRAR4, await the collection of sufficient data to interpret the validity of the original studies, whereas others, such as COMT, have not yielded conclusive results despite intense study, including large sample association and meta-analysis.
One exciting feature of most of the candidates discussed above is that they can be related to potential pathophysiology through dysfunction in glutamatergic neurotransmission, which may be an important systemic element in the etiology of schizophrenia. Detailed discussion of this theory is outside the scope of this review, but recent reviews of the genetic86 and neurobiological87 evidence discuss the positions of the gene products of NRG1, COMT, DAAO, G72 and RGS4, among others, in the biochemical and functional pathways influencing the glutamatergic system. Currently, all of the regions and candidate genes discussed in the sections above are promising, but further assessment of each is still needed, both to clarify patterns of linkage or association and to elucidate their contribution to the neurobiology of schizophrenia.
Limitations of current knowledge
No DNA variant widely associated with schizophrenia has been shown to produce a distinct change in expression, structure or function of any of the candidates. Even in the data from the two genes (DTNBP1 and NRG1) where most replication studies detect evidence for association, important questions remain. No single high-risk haplotype is associated across all samples. It is unclear whether we should expect to observe association with the same specific haplotype, or only with the gene, across samples, as we have no idea of the age or number of liability variants. A single, evolutionarily recent variant would be expected to yield association with the same haplotype across samples, whereas a single ancient variant may have been extensively shuffled by recombination, yielding haplotypes that are differentially represented in, or even specific to, different populations. As we show below (Table 3), there is evidence that such differences in haplotypes exist for some genes even between closely related Northern European groups. Multiple variants of any age would be expected to occur on different haplotypes with or without recombination.
Furthermore, there is no reason to expect that a single variant will be responsible for all risk in a particular gene: multiple mutations are already well documented in the Mendelian cystic fibrosis gene, for example, and multiple variants in a gene may also cause liability to complex traits. If multiple liability variants of any age do exist, population genetic effects and stochastic sampling variation would then determine which variants a particular sample could detect. The complex patterns of association also provide the basis for specific hypotheses about variants that might be shared across multiple backgrounds. We have chosen DTNBP1 as an example, as this locus has the largest number of positive replications, and therefore the most data to assess.
Patterns of association in DTNBP1
The initial report of association between schizophrenia and DTNBP1 in 270 multiply affected Irish families included 12 SNPs within the gene, of which six showed some degree of nominally significant association (0.00004<P<0.0466).34 Subsequent reanalysis of this data set showed that a set of eight of these markers (p1792–p1635) satisfied criteria for a haplotype block. The eight markers cover 30.1 kb from intron 1 to intron 4 and 6 common haplotypes (frequency>1%) define 94.7% of haplotypes observed (Table 1). The six common haplotypes fit a stable clade (Figure 2) with only low-frequency haplotype 6 apparently arising from historical recombination based on alleles at p1325 and p1635.35 Although there is some variation in the haplotype complement observed across different samples, the pattern of specific haplotypes and their relative frequencies are generally maintained.
The clade produced from DTNBP1 haplotypes in Table 1.
Results from the original study and eight published positive replications are shown in Table 2. The replications include 13 independent (two family, three triad, and eight case/control) samples and use different but overlapping subsets of the markers from the original study. In our original haplotype analyses, the association in the Irish family sample is limited to haplotype 2 in Table 1 and Figure 2.35 Two other haplotypes (1 and 5 in Table 1 and Figure 2) show evidence for association in different samples. Significance levels in the following summaries are haplotype-specific unless otherwise noted.
Three (and possibly four) studies observe association with the common haplotype 1. A study of a German, Israeli and Hungarian sample (78 sib-pair families and 125 triads, six shared markers) observed allelic association with five of the markers tested and numerous associated haplotypes (depending on the number of markers included). These were always composed of the common alleles and thus consistent only with haplotype 1 in Table 1 and Figure 2.36 The most significant single haplotype (shown in Table 2) was composed of the common alleles of p1320 (rs760761), p1765 (rs2619528) and p1325 (rs1011313), P=0.00002. This study includes the most shared markers with the original study, and thus is the most useful for assessing the stability of the haplotype picture shown in Table 1 and Figure 2. Comparative haplotype reconstructions from the Irish and German/Israeli/Hungarian sample are shown in Table 3.
Five of the six common Irish haplotypes are present in both samples at comparable frequencies. One low-frequency haplotype from the Irish sample and two from the German/Israeli/Hungarian sample are not observed in the other group at frequencies above our 1% cutoff. Assessment of the allele patterns in Table 1 and Figure 2 shows that all three of these are inconsistent with origins from SNP mutation alone, and thus reflect historical recombination events. As noted in Table 3, haplotype 8 may represent the result of the first of the two events giving rise to haplotype 6. The second of these events and that giving rise to haplotype 9 appear to be sample or population-specific recombinations.
The first study of a Han Chinese sample (233 triads, five shared markers) observed association with a haplotype containing the minor allele at p1583 (rs909706), the common alleles of p1578 (rs1018381) and p1763 (rs2619522) and the G allele (common in the Han sample) at p1655 (rs2619539), P=0.01.37 Considering only p1578 (rs1018381) and p1763 (rs2619522), which appear in the data in Table 1 and Figure 2, this pattern is consistent with any of haplotypes 1, 3, 4 or 6. However, two additional pieces of information suggest strongly that the principal contributor to the observed association is haplotype 1. First, our sequence data show that both major and minor alleles of p1583 (rs909706) and p1655 (rs2619539) are present on multiple haplotypes (including haplotype 1), consistent with recombination since the SNPs arose and our observed block structure which excludes them. Second, the associated haplotype is the most common observed in the Han sample. P1328 (rs742106) was also included in the analysis and the minor allele was present on the associated haplotype. In our data, P1328 (rs742106) shows almost no LD with the more 5′ SNPs, so it does not assist in relating the identity of the associated haplotype to those in Table 1 and Figure 2.
Independent studies of samples of 708/711 UK cases/controls42 and 219/231 Irish case/controls88 observed no evidence for association with any of the markers defining the haplotype associated with the Irish family sample. However, when these samples were analyzed together for a different set of SNPs in DTNBP1, association with the same haplotype was observed in both.42 The associated haplotype is composed of common alleles of SNP A (rs2619538), p1635 (rs3213207) and p1655 (rs2619539), P=0.000056. Both major and minor alleles of SNP A (rs2619538) also occur on the most common background, and this associated haplotype is only consistent with a subgroup (defined by the alleles at SNP A (rs2619538) and p1655 (rs2619539)) of haplotype 1 in Table 1 and Figure 2. This may explain why the UK/Irish sample did not detect association with haplotype 1 using the original marker set.
Some important features of the data from these two studies must be noted. Based on the Irish haplotype structure, these last two studies appear to detect association on completely nonoverlapping subgroups of haplotype 1 in Table 1 and Figure 2 defined by the allele at p1655 (rs2619539). However, the sample populations are very different, and we cannot assume that the haplotype structures or their frequencies are the same in Han Chinese as in Europeans. For example, the associated alleles of p1655 (rs2619539) reported at this marker in the studies of Tang et al37 and Williams et al42 are different, but are the common alleles in both samples. The allele frequencies are very close in European samples (C∼0.52, G∼0.48) but appear to be inverted in the Han sample data (G>0.63). Critically, both studies support the conclusion of the German study that a liability variant may be present on the oldest, highest frequency background haplotype. The implications of this are discussed further below.
A study of a Bulgarian sample (488 trios, four overlapping markers) identifies association with the common alleles of p1757 (rs2005976) and p1635 (rs3213207).40 Numerous haplotypes gave evidence of association, and the authors were unable to identify a specific risk haplotype in this sample. The combination of these two common alleles is consistent with haplotypes 1, 3 or 4 in Table 1 and Figure 2. Although ambiguous, it seems likely that this sample also provides evidence for association of the common haplotype 1.
Two further replication studies identify association with haplotype 5 in Table 1 and Figure 2. These consist of four independent samples (142/272 Swedish, 418/285 German and 294/113 Polish cases/controls, five shared markers38) and 524/573 US cases/controls of various ethnicities (258/467 Caucasian, 215/74 African-American and 51/32 Hispanic cases/controls, six shared markers39). The Swedish, US Caucasian and US Hispanic samples unambiguously detect association with haplotype 5 in Table 1 and Figure 2. The African-American, German and Polish samples show no evidence for association with any DTNBP1 haplotype.
One replication study identifies association with the same risk haplotype observed in the Irish family sample.41 This study of 670/588 Japanese cases/controls gave nominal evidence of association with several single markers tested, particularly the minor allele of p1635 (rs3213207). This allele tags the Irish risk haplotype, and is only observed on that specific background. Other associated alleles and haplotypes are all consistent with the haplotype associated in the Irish family sample. Haplotypes of various lengths from two to six markers were assessed, and many produced significant evidence of association. Case and control frequencies for the associated haplotype vary only slightly depending on the number of SNPs included, with an average value of 0.01 in controls and 0.03 in cases. The maximum observed case–control difference is for the two-marker haplotype of p1635 (rs3213207) and p1325 (rs1011313), P=0.00028.
Finally, one further replication study in two independent samples (638 Han Chinese families and 580/620 Scottish cases/controls, six shared markers) also detects evidence for association.43 The Chinese sample shows individual marker association with p1765 (rs2619528), P=0.002, and p1635 (rs3213207), P=0.02. No individual markers were associated in the Scottish sample. Only haplotypes of two markers were analyzed in this study. In the Chinese sample, the associated haplotype is composed of the common alleles of p1757 (rs2005976) and p1765 (rs2619528), P=0.00005. In the Scottish sample, a haplotype composed of the common allele at p1320 (rs760761) and the rare allele at p1757 (rs2005976) is associated, P=0.0006. The reporting of only two-marker haplotypes and lack of marker overlap with the other study of Han samples makes interpretation ambiguous, but the associated haplotype is again the most frequent, so seems likely to support the previous Han Chinese results, and to represent haplotype 1. The haplotype associated in the Scottish sample is not observed in Table 1 and Figure 2. One additional study observed no evidence for association in any of three separate samples tested.89
In summary, association between schizophrenia and DTNBP1 has been replicated in seven of eight studies (excluding the negative study of the Irish case/control sample) or in 10 of 15 independent samples (the US sample described in Funke et al39 was analyzed separately by Caucasian, Hispanic and African-American ethnicity, but was collected, and is treated here, as a single independent sample).
Two very different hypotheses can be advanced given the data outlined above. First, and perhaps most intuitive, these results may reflect the presence of more than one liability variant in the DTNBP1 gene. As discussed above, there is no biological reason why this may not be the case for at least some genes in complex traits. The evolutionary distance between haplotype 1 and haplotypes 2 and 5 argues in favor of this idea. It remains unclear whether the same or different variants might be responsible for the associations observed on haplotypes 2 and 5; the close evolutionary relationship between them would argue in favor of a variant common to both. However, it would also suggest that these two haplotypes (and markers common to both) should be associated to the same degree in the samples where one or other is associated, and this has not generally been reported. More recent conditional analyses of the Irish families, excluding those transmitting haplotype 2 to affected offspring, suggest that there may be some residual association with haplotype 5 not detected in our original assessment. Interpretation of this is complicated by the fact that, in our sample, both the sequence data and the LD pattern show this haplotype to be the product of ancestral recombination. In contrast, the HAPMAP data in this region show no such recombination. Until further markers are analyzed, it remains unclear whether the reported associations are on the recombinant or nonrecombinant version of this haplotype. Our sequence study of haplotype 2 identified numerous polymorphisms specific to or enriched on our associated haplotype. One coding variant enriched on this background is not associated with schizophrenia. A number of variants occur in evolutionarily conserved regions, and are being followed up currently.
However, a second hypothesis is that there may be variation common to the different associated backgrounds. We observe a number of such variants (with alleles shared by haplotypes 1, 2 and 5) in our sequence data, so their existence is not in question. They are somewhat less intuitive as putative liability factors for disease, however. We tend to think of disease predisposing alleles as changes from a functional ‘common’ norm to a dysfunctional ‘rare’ variant, analogous to Mendelian mutations. The alleles shared across evolutonarily distant haplotypes are almost certainly ancestral rather than variant. This is consistent with the idea (based on reduction in fertility in patients) that predisposing alleles may be advantageous individually, outweighing the higher risk in the smaller number of individuals carrying multiple such alleles. As expected, they tend to have higher minor allele frequencies and evidence for substantial recombination (suggesting greater age); p1655 (rs2619539) is one such marker. A number of these variants also occur in evolutionarily conserved regions, and are also being followed up currently.
Conclusions
Certainly the most important development in the last several years has been the emergence of a number of replicated linkage findings and the identification of associations within genes in these target regions. Replication is critical, but there are many reasons why a ‘true’ finding might not be replicated including genetic variation between populations and differences in statistical power, diagnostic methods and statistical approaches. Given the evidence of linkage replication across several groups for regions on 22q, 8p, 6p, 13q, 5q, 10p, 6q, 1q and 15q, and association replication for DTNBP1, NRG1 and other genes, it seems increasingly unlikely that all of these findings represent false positives. It is difficult to conceive of an inherent bias that would produce spuriously positive results across multiple groups (especially given the wide differences between studies described above) in the same chromosomal region.
We have focused only on the molecular genetic study of schizophrenia in this review. In generalizing to other psychiatric (and other complex) phenotypes, the broad conclusions from the study of schizophrenia seem likely to hold: (1) such phenotypes are genetically influenced but not genetically determined, (2) a number of genes (which may even vary between individual family members) are likely to be involved, (3) the liability variants in these genes are generally expected to be within the range of normal human variation and to have low risk associated with them individually, (4) some of the variants may interact with others or with environmental risk factors.
An emerging number of identified positional candidate genes are, in some cases, now being replicated in independent samples. Given the major focus on this area of multiple research groups throughout the world, it is likely that within several years these or other loci might emerge as widely replicated susceptibility genes for schizophrenia. If this occurs, it will represent a true watershed event in the history of schizophrenia research. Although the step of gene-identification will itself represent a major advance, it will also represent the beginning of several new lines of research including (1) rational drug design based on knowledge of basic pathophysiology, (2) characterization of genotype–phenotype relationships based on knowledge of specific pathogenic mutations, (3) identification of environmental risk factors that interact with specific genes and (4) realistic prevention research given our ability to identify high-risk individuals.
References
Gottesman II : Schizophrenia Genesis. New York: W H Freeman, 1991.
Kendler KS, Diehl SR : The genetics of schizophrenia: a current, genetic-epidemiologic perspective. Schizophr Bull 1993; 19: 261–285.
Cardno AG, Gottesman II : Twin studies of schizophrenia: from bow-and-arrow concordances to Star Wars Mx and functional genomics. Am J Med Genet 2000; 97: 12–17.
Tienari P, Wynne LC, Sorri A et al: Genotype–environment interaction in schizophrenia-spectrum disorder. Long-term follow-up study of Finnish adoptees. Br J Psychiatry 2004; 184: 216–222.
Prescott CA, Gottesman II : Genetically mediated vulnerability to schizophrenia. Psychiatr Clin N Am 1993; 16: 245–267.
Tienari P : Interaction between genetic vulnerability and family environment: the Finnish adoptive family study of schizophrenia. Acta Psychiatr Scand 1991; 84: 460–465.
Kety SS, Wender PH, Jacobsen B et al: Mental illness in the biological and adoptive relatives of schizophrenic adoptees: replication of the Copenhagen study in the rest of Denmark. Arch Gen Psychiatry 1994; 51: 442–455.
McGue M, Gottesman I, Rao DC : Resolving genetic models for the transmission of schizophrenia. Genet Epidemiol 1985; 2: 99–110.
Kendler KS, McGuire M, Gruenberg AM, Spellman M, O'Hare A, Walsh D : The Roscommon family study: II. The risk of nonschizophrenic nonaffective psychoses in relatives. Arch Gen Psychiatry 1993; 50: 645–652.
Riley B, Kendler KS : Genetics of schizophrenia: linkage and association studies; in Kendler KS, Eaves LJ (eds): Psychiatric Genetics. Washington DC: American Psychiatric Publishing Incorporated, 2005, pp 95–140.
Pulver AE, Karayiorgou M, Wolyniec PS et al: Sequential strategy to identify a susceptibility gene for schizophrenia: Report of potential linkage on chromosome 22q12–q13.1: Part 1. Am J Med Genet 1994; 54: 36–43.
Gill M, Vallada H, Collier D et al: A combined analysis of D22S278 marker alleles in affected sib-pairs: support for a susceptibility locus for schizophrenia at chromosome 22q12. Am J Med Genet Neuropsychiatr Genet 1996; 67: 40–45.
Murphy KC, Jones LA, Owen MJ : High rates of schizophrenia in adults with velo-cardio-facial syndrome. Arch Gen Psychiatry 1999; 56: 940–945.
Egan MF, Goldberg TE, Kolachana BS et al: Effect of COMT Val108/158 Met genotype on frontal lobe function and risk for schizophrenia. Proc Natl Acad Sci USA 2001; 98: 6917–6922.
Shifman S, Bronstein M, Sternfeld M et al: A highly significant association between a COMT haplotype and schizophrenia. Am J Hum Genet 2002; 71: 1296–1302.
Glatt SJ, Faraone SV, Tsuang MT : Association between a functional catechol O-methyltransferase gene polymorphism and schizophrenia: meta-analysis of case–control and family-based studies. Am J Psychiatry 2003; 160: 469–476.
Fan JB, Zhang CS, Gu NF et al: Catechol-O-methyltransferase gene Val/Met functional polymorphism and risk of schizophrenia: a large-scale association study plus meta-analysis. Biol Psychiatry 2005; 57: 139–144.
Williams HJ, Glaser B, Williams NM et al: No association between schizophrenia and polymorphisms in COMT in two large samples. Am J Psychiatry 2005; 162: 1736–1738.
Liu H, Heath SC, Sobin C et al: Genetic variation at the 22q11 PRODH2/DGCR6 locus presents an unusual pattern and increases susceptibility to schizophrenia. Proc Natl Acad Sci USA 2002; 99: 3717–3722.
Fan JB, Ma J, Zhang CS et al: A family-based association study of T1945C polymorphism in the proline dehydrogenase gene and schizophrenia in the Chinese population. Neurosci Lett 2003; 338: 252–254.
Mukai J, Liu H, Burt RA et al: Evidence that the gene encoding ZDHHC8 contributes to the risk of schizophrenia. Nat Genet 2004; 36: 725–731.
Chen WY, Shi YY, Zheng YL et al: Case–control study and transmission disequilibrium test provide consistent evidence for association between schizophrenia and genetic variation in the 22q11 gene ZDHHC8. Hum Mol Genet 2004; 13: 2991–2995.
Pulver AE, Lasseter VK, Kasch L et al: Schizophrenia: a genome scan targets chromosomes 3p and 8p as potential sites of susceptibility genes. Am J Med Genet Neuropsychiatr Genet 1995; 60: 252–260.
Levinson DF, Wildenauer DB, Schwab SG et al: Additional support for schizophrenia linkage on chromosomes 6 and 8: a multicenter study. Am J Med Genet Neuropsychiatr Genet 1996; 67: 580–594.
Stefansson H, Sigurdsson E, Steinthorsdottir V et al: Neuregulin 1 and susceptibility to schizophrenia. Am J Hum Genet 2002; 71: 877–892.
Stefansson H, Sarginson J, Kong A et al: Association of neuregulin 1 with schizophrenia confirmed in a Scottish population. Am J Hum Genet 2003; 72: 83–87.
Corvin AP, Morris DW, McGhee K et al: Confirmation and refinement of an ‘at-risk’ haplotype for schizophrenia suggests the EST cluster, Hs.97362, as a potential susceptibility gene at the Neuregulin-1 locus. Mol Psychiatry 2004; 9: 208–213.
Williams NM, Preece A, Spurlock G et al: Support for genetic variation in neuregulin 1 and susceptibility to schizophrenia. Mol Psychiatry 2003; 8: 485–487.
Thiselton DL, Webb BT, Neale BM et al: No evidence for linkage or association of neuregulin-1 (NRG1) with disease in the Irish study of high-density schizophrenia families (ISHDSF). Mol Psychiatry 2004; 9: 777–783.
Gerber DJ, Hall D, Miyakawa T et al: Evidence for association of schizophrenia with genetic variation in the 8p21.3 gene, PPP3CC, encoding the calcineurin gamma subunit. Proc Natl Acad Sci USA 2003; 100: 8993–8998.
Miyakawa T, Leiter LM, Gerber DJ et al: Conditional calcineurin knockout mice exhibit multiple abnormal behaviors related to schizophrenia. Proc Natl Acad Sci USA 2003; 100: 8987–8992.
Kinoshita Y, Suzuki T, Ikeda M et al: No association with the calcineurin A gamma subunit gene (PPP3CC) haplotype to Japanese schizophrenia. J Neural Trans 2005; 112: 1255–1262.
Straub RE, MacLean CJ, O'Neill FA et al: A potential vulnerability locus for schizophrenia on chromosome 6p24–22: evidence for genetic heterogeneity. Nat Genet 1995; 11: 287–293.
Straub RE, Jiang Y, MacLean CJ et al: Genetic variation in the 6p22.3 gene DTNBP1, the human ortholog of mouse dysbindin, is associated with schizophrenia. Am J Hum Genet 2002; 71: 337–348.
van den Oord E, Sullivan PF, Chen X, Kendler KS, Riley B : Identification of a high risk haplotype for the dystrobrevin binding protein 1 (DTNBP1) gene in the Irish study of high density schizophrenia families. Mol Psychiatry 2003; 8: 499–510.
Schwab SG, Knapp M, Mondabon S et al: Support for association of schizophrenia with genetic variation in the 6p22.3 gene, dysbindin, in sib-pair families with linkage and in an additional sample of triad families. Am J Hum Genet 2003; 72: 185–190.
Tang JX, Zhou J, Fan JB et al: Family-based association study of DTNBP1 in 6p22.3 and schizophrenia. Mol Psychiatry 2003; 8: 717–718.
Van Den Bogaert A, Schumacher J, Schulze TG et al: The DTNBP1 (dysbindin) gene contributes to schizophrenia, depending on family history of the disease. Am J Hum Genet 2003; 73: 1438–1443.
Funke B, Finn CT, Plocik AM et al: Association of the DTNBP1 locus with schizophrenia in a U.S. population. Am J Hum Genet 2004; 75: 891–898.
Kirov G, Ivanov D, Williams NM et al: Strong evidence for association between the dystrobrevin binding protein 1 gene (DTNBP1) and schizophrenia in 488 parent–offspring trios from Bulgaria. Biol Psychiatry 2004; 55: 971–975.
Numakawa T, Yagasaki Y, Ishimoto T et al: Evidence of novel neuronal functions of dysbindin, a susceptibility gene for schizophrenia. Hum Mol Genet 2004; 13: 2699–2708.
Williams NM, Preece A, Morris DW et al: Identification in 2 independent samples of a novel schizophrenia risk haplotype of the dystrobrevin binding protein gene (DTNBP1). Arch Gen Psychiatry 2004; 61: 336–344.
Li T, Zhang F, Liu X et al: Identifying potential risk haplotypes for schizophrenia at the DTNBP1 locus in Han Chinese and Scottish populations. Mol Psychiatry 2005; 10: 1037–1044.
Weickert CS, Straub RE, McClintock BW et al: Human dysbindin (DTNBP1) gene expression in normal brain and in schizophrenic prefrontal cortex and midbrain. Arch Gen Psychiatry 2004; 61: 544–555.
Talbot K, Eidem WL, Tinsley CL et al: Dysbindin-1 is reduced in intrinsic, glutamatergic terminals of the hippocampal formation in schizophrenia. J Clin Invest 2004; 113: 1353–1363.
Emamian ES, Hall D, Birnbaum MJ, Karayiorgou M, Gogos JA : Convergent evidence for impaired AKT1-GSK3beta signaling in schizophrenia. Nat Genet 2004; 36: 131–137.
Ikeda M, Iwata N, Suzuki T et al: Association of AKT1 with schizophrenia confirmed in a Japanese population. Biol Psychiatry 2004; 56: 698–700.
Ohtsuki T, Inada T, Arinami T : Failure to confirm association between AKT1 haplotype and schizophrenia in a Japanese case–control population. Mol Psychiatry 2004; 9: 981–983.
Schwab SG, Hoefgen B, Hanses C et al: Further evidence for association of variants in the AKT1 gene with schizophrenia in a sample of European sib-pair families. Biol Psychiatry 2005; 58: 446–450.
Lin MW, Curtis D, Williams N et al: Suggestive evidence for linkage of schizophrenia to markers on chromosome 13q14.1–q32. Psychiatr Genet 1995; 5: 117–126.
Chumakov I, Blumenfeld M, Guerassimenko O et al: Genetic and physiological data implicating the new human gene G72 and the gene for D-amino acid oxidase in schizophrenia. Proc Natl Acad Sci USA 2002; 99: 13675–13680.
Korostishevsky M, Kaganovich M, Cholostoy A et al: Is the G72/G30 locus associated with schizophrenia? Single nucleotide polymorphisms, haplotypes, and gene expression analysis. Biol Psychiatry 2004; 56: 169–176.
Zou F, Li C, Duan S et al: A family-based study of the association between the G72/G30 genes and schizophrenia in the Chinese population. Schizophr Res 2005; 73: 257–261.
Korostishevsky M, Kremer I, Kaganovich M et al: Transmission disequilibrium and haplotype analyses of the G72/G30 locus: suggestive linkage to schizophrenia in Palestinian Arabs living in the North of Israel. Am J Med Genet B Neuropsychiatr Genet 2005, [E-pub ahead of print].
Wang X, He G, Gu N et al: Association of G72/G30 with schizophrenia in the Chinese population. Biochem Biophys Res Commun 2004; 319: 1281–1286.
Addington AM, Gornick M, Sporn AL et al: Polymorphisms in the 13q33.2 gene G72/G30 are associated with childhood-onset schizophrenia and psychosis not otherwise specified. Biol Psychiatry 2004; 55: 976–980.
Schumacher J, Jamra RA, Freudenberg J et al: Examination of G72 and D-amino-acid oxidase as genetic risk factors for schizophrenia and bipolar affective disorder. Mol Psychiatry 2004; 9: 203–207.
Mulle JG, Chowdari KV, Nimgaonkar V, Chakravarti A : No evidence for association to the G72/G30 locus in an independent sample of schizophrenia families. Mol Psychiatry 2005; 10: 431–433.
Cao Q, Martinez M, Zhang J et al: Suggestive evidence for a schizophrenia susceptibility locus on chromosome 6q and a confirmation in an independent series of pedigrees. Genomics 1997; 43: 1–8.
Levinson DF, Holmans P, Straub RE et al: Multicenter linkage study of schizophrenia candidate regions on chromosomes 5q, 6q, 10p, and 13q: schizophrenia linkage collaborative group III. Am J Hum Genet 2000; 67: 652–663.
Duan J, Martinez M, Sanders AR et al: Polymorphisms in the trace amine receptor 4 (TRAR4) gene on chromosome 6q23.2 are associated with susceptibility to schizophrenia. Am J Hum Genet 2004; 75: 624–638.
Duan S, Du J, Xu Y et al: Failure to find association between TRAR4 and schizophrenia in the Chinese Han population. J Neural Transm 2005, [E-pub ahead of print].
Ikeda M, Iwata N, Suzuki T et al: No association of haplotype-tagging SNPs in TRAR4 with schizophrenia in Japanese patients. Schizophr Res 2005; 78: 127–130.
Shaw SH, Kelly M, Smith AB et al: A genome-wide search for schizophrenia susceptibility genes. Am J Med Genet Neuropsychiatr Genet 1998; 81: 364–376.
Brzustowicz LM, Hodgkinson KA, Chow EWC, Honer WG, Bassett AS : Location of a major susceptibility locus for familial schizophrenia on chromosome 1q21–q22. Science 2000; 288: 678–682.
Levinson DF, Holmans PA, Laurent C et al: No major schizophrenia locus detected on chromosome 1q in a large multicenter sample. Science 2002; 296: 739–741.
Millar JK, Wilson-Annan JC, Anderson S et al: Disruption of two novel genes by a translocation co-segregating with schizophrenia. Hum Mol Genet 2000; 9: 1415–1423.
Hennah W, Varilo T, Kestila M et al: Haplotype transmission analysis provides evidence of association for DISC1 to schizophrenia and suggests sex-dependent effects. Hum Mol Genet 2003; 12: 3151–3159.
Hodgkinson CA, Goldman D, Jaeger J et al: Disrupted in schizophrenia 1 (DISC1): association with schizophrenia, schizoaffective disorder, and bipolar disorder. Am J Hum Genet 2004; 75: 862–872.
Thomson PA, Wray NR, Millar JK et al: Association between the TRAX/DISC locus and both bipolar disorder and schizophrenia in the Scottish population. Mol Psychiatry 2005; 10: 657–668, 616.
Callicott JH, Straub RE, Pezawas L et al: Variation in DISC1 affects hippocampal structure and function and increases risk for schizophrenia. Proc Natl Acad Sci USA 2005; 102: 8627–8632.
Devon RS, Anderson S, Teague PW et al: Identification of polymorphisms within Disrupted in Schizophrenia 1 and Disrupted in Schizophrenia 2, and an investigation of their association with schizophrenia and bipolar affective disorder. Psychiatr Genet 2001; 11: 71–78.
Kockelkorn TT, Arai M, Matsumoto H et al: Association study of polymorphisms in the 5′ upstream region of human DISC1 gene with schizophrenia. Neurosci Lett 2004; 368: 41–45.
Zhang X, Tochigi M, Ohashi J et al: Association study of the DISC1/TRAX locus with schizophrenia in a Japanese population. Schizophr Res 2005; 79: 175–180.
Sachs NA, Sawa A, Holmes SE et al: A frameshift mutation in Disrupted in Schizophrenia 1 in an American family with schizophrenia and schizoaffective disorder. Mol Psychiatry 2005; 10: 758–764.
Miyoshi K, Honda A, Baba K et al: Disrupted-In-Schizophrenia 1, a candidate gene for schizophrenia, participates in neurite outgrowth. Mol Psychiatry 2003; 8: 685–694.
Ozeki Y, Tomoda T, Kleiderlein J et al: Disrupted-in-Schizophrenia-1 (DISC-1): mutant truncation prevents binding to NudE-like (NUDEL) and inhibits neurite outgrowth. Proc Natl Acad Sci USA 2003; 100: 289–294.
Mirnics K, Middleton FA, Lewis DA, Levitt P : Analysis of complex brain disorders with gene expression microarrays: schizophrenia as a disease of the synapse. Trends Neurosci 2001; 24: 479–486.
Chowdari KV, Mirnics K, Semwal P et al: Association and linkage analyses of RGS4 polymorphisms in schizophrenia. Hum Mol Genet 2002; 11: 1373–1380.
Chen X, Dunham C, Kendler S et al: Regulator of G-protein signaling 4 (RGS4) gene is associated with schizophrenia in Irish high density families. Am J Med Genet 2004; 129B: 23–26.
Cordeiro Q, Talkowski ME, Chowdari KV, Wood J, Nimgaonkar V, Vallada H : Association and linkage analysis of RGS4 polymorphisms with schizophrenia and bipolar disorder in Brazil. Genes Brain Behav 2005; 4: 45–50.
Sobell JL, Richard C, Wirshing DA, Heston LL : Failure to confirm association between RGS4 haplotypes and schizophrenia in Caucasians. Am J Med Genet B Neuropsychiatr Genet 2005; 139: 23–27.
Morris DW, Rodgers A, McGhee KA et al: Confirming RGS4 as a susceptibility gene for schizophrenia. Am J Med Genet 2004; 125B: 50–53.
Williams NM, Preece A, Spurlock G et al: Support for RGS4 as a susceptibility gene for schizophrenia. Biol Psychiatry 2004; 55: 192–195.
Lewis CM, Levinson DF, Wise LH et al: Genome scan meta-analysis of schizophrenia and bipolar disorder, part II: Schizophrenia. Am J Hum Genet 2003; 73: 34–48.
Harrison PJ, Owen MJ : Genes for schizophrenia? Recent findings and their pathophysiological implications. Lancet 2003; 361: 417–419.
Moghaddam B : Bringing order to the glutamate chaos in schizophrenia. Neuron 2003; 40: 881–884.
Morris DW, McGhee KA, Schwaiger S et al: No evidence for association of the dysbindin gene [DTNBP1] with schizophrenia in an Irish population-based study. Schizophr Res 2003; 60: 167–172.
Hall D, Gogos JA, Karayiorgou M : The contribution of three strong candidate schizophrenia susceptibility genes in demographically distinct populations. Genes Brain Behav 2004; 3: 240–248.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Riley, B., Kendler, K. Molecular genetic studies of schizophrenia. Eur J Hum Genet 14, 669–680 (2006). https://doi.org/10.1038/sj.ejhg.5201571
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201571
Keywords
This article is cited by
-
Effect of VNTR Polymorphism of the AS3MT Gene and Obstetrical Complications on the Severity of Schizophrenia
Bulletin of Experimental Biology and Medicine (2019)
-
Can Psychological, Social and Demographical Factors Predict Clinical Characteristics Symptomatology of Bipolar Affective Disorder and Schizophrenia?
Psychiatric Quarterly (2016)
-
Genome-wide pharmacogenomic analysis of response to treatment with antipsychotics
Molecular Psychiatry (2011)
-
Dysbindin-1, a schizophrenia-related protein, facilitates neurite outgrowth by promoting the transcriptional activity of p53
Molecular Psychiatry (2011)
-
Impaired glycolytic response in peripheral blood mononuclear cells of first-onset antipsychotic-naive schizophrenia patients
Molecular Psychiatry (2011)