Abstract
Despite investment in toxicogenomics, nonclinical safety studies are still used to predict clinical liabilities for new drug candidates. Network-based approaches for genomic analysis help overcome challenges with whole-genome transcriptional profiling using limited numbers of treatments for phenotypes of interest. Herein, we apply co-expression network analysis to safety assessment using rat liver gene expression data to define 415 modules, exhibiting unique transcriptional control, organized in a visual representation of the transcriptome (the ‘TXG-MAP’). Accounting for the overall transcriptional activity resulting from treatment, we explain mechanisms of toxicity and predict distinct toxicity phenotypes using module associations. We demonstrate that early network responses complement traditional histology-based assessment in predicting outcomes for longer studies and identify a novel mechanism of hepatotoxicity involving endoplasmic reticulum stress and Nrf2 activation. Module-based molecular subtypes of cholestatic injury derived using rat translate to human. Moreover, compared to gene-level analysis alone, combining module and gene-level analysis performed in sequence identifies significantly more phenotype-gene associations, including established and novel biomarkers of liver injury.
Similar content being viewed by others
Introduction
Safety remains a major cause of attrition during clinical trials.1, 2, 3, 4, 5 Prior to clinical testing, all clinical candidates are evaluated in animals to define the spectrum of toxicities that might occur in human subjects and safe doses for clinical testing.6 However, continued occurrences of clinical safety terminations calls into question the value of nonclinical testing in predicting human risk.7, 8 Nonetheless, when confidence in nonclinical safety data is high compounds are more likely to be safe in humans.9 Uncertainty regarding safety predictions occurs at three major transition points in biopharmaceutical testing: (1) the transition inherent in using simple in vitro models to predict in vivo nonclinical (animal) results early in discovery; (2) the transition from nonclinical testing to human clinical trials; and (3) the transition from testing in well-controlled clinical trials to the larger diverse patient population post approval. In other work, we addressed the first transition by associating chemical properties with toxicity early10 and by developing a systems level framework using co-expression networks to evaluate how well mechanisms extrapolate from primary cell cultures to the same organ in vivo.11 Here we address the second transition by investigating the utility of network-based toxicogenomic approaches for predicting mechanisms of drug-induced liver injury and the translation from rodent to human.
Considerable effort has been invested applying transcript profiling to risk assessment using methodologies such as gene signatures,12 pathway-based enrichment analysis,13 co-expression networks,14, 15 and adverse outcome pathways.16 However, toxicogenomic approaches to safety testing remain challenging and have achieved only modest utility in addressing uncertainty in safety predictions, largely as an investigative tool. Nonclinical safety testing remains largely dependent on traditional clinical chemistry and histologic evaluation. Gene signatures are effective as classifiers but their development requires large and costly compendia of transcript profiles and may not translate to other models and mechanisms. Limitations in measurement technologies and the inherent stochastic nature of biological systems pose additional analytical challenges to establishing the relationship between thousands of variables (genes) and toxicity properties using small sets of training compounds. Pathway or Gene Ontology (GO) enrichment analysis can reduce noise but are biased toward known biology captured in existing repositories.13, 17
Unsupervised methods that organize high-dimensional data into networks based on biologically relevant coalescent properties reduce noise and boost signal detection.18, 19 This seems intuitive since organisms demonstrate modularity and conservation of biology across evolution.20, 21 One such approach, weighted gene co-expression network analysis (WGCNA), uses the property of co-expression to organize genes into gene networks or modules.22 Here we develop a co-expression framework called the ‘toxicogenomic module associations with pathogenesis’ (the TXG-MAP) and integrate it with standard pathology evaluation to characterize mechanisms of drug-induced liver injury. We demonstrate the utility of the TXG-MAP for common applications. First, we illustrate how co-expression modules reveal mechanisms of pathogenesis concurrent with or preceding toxicity phenotypes. Second, we illustrate the utility of modules for identifying marker genes in small data sets, while controlling for false discovery. Third, we use case studies to illustrate utility in elucidating specific mechanisms of liver injury. Fourth, we identify transcription factors that couple upstream signals to co-expression changes. Finally, we demonstrate that module-based molecular phenotypes for rodent liver injury translate to human liver disease.
Materials and methods
Microarray data processing from Drug Matrix, TG-GATEs and GEO
The Drug Matrix (DM)23 and open TG-GATEs (TG)24 databases constitute two large publicly available resources describing the effects of drugs and other compounds in rat liver. They contain gene expression data from Affymetrix microarrays, linked to traditional histology and clinical chemistry results for 3528 treatment groups from TG and 654 from DM. A treatment group denotes three or more animals receiving a given dose of drug or vehicle, usually administered daily by oral gavage, and killed following drug exposures lasting from 3 h to 29 days. The treatment groups analyzed in this work are given in Supplementary Table S1. Methods for obtaining, processing and analyzing rat liver microarray data from DM and TG are described in detail elsewhere;11 details for Gene Expression Omnibus (GEO) sets are provided in Supplementary Methods.
Rat liver co-expression networks
Using 654 treatments from DM rat liver and a subset of 9071 liver-expressed genes, 415 co-expression networks were obtained using WGCNA22 as described in reference.11 Co-expression modules represent gene sets that show correlated behavior (that is, co-induced or co-repressed) across the DM rat liver database. A given module summarizes (log) fold change of its constituent genes, weighted by their individual correlation with the first principal component (eigengene) of the module. For each pair of 415 modules, the Pearson correlation of module scores was calculated using 654 DM rat liver experiments. Modules with Pearson R⩾0.8 behave similarly across the DM database (induced or repressed similarly upon drug treatment), and were merged to reduce redundancy among modules; merged modules contain the suffix ‘m’, for example, 46m was merged from 46 and 49. A set of 284 merged modules and unmerged modules with R<0.8 versus all others were labeled as ‘base’ modules. The TXG-MAP was obtained by clustering modules in R applying Ward’s algorithm to a 415 × 415 Pearson distance matrix (1−Pearson R). The resulting dendrogram was imported as a network into Cytoscape and rendered using the yFiles circular layout with small manual adjustments to improve node placement. The Cytoscape session file is available in Supplementary Information.
For each chemical treatment, we obtained a single numeric eigengene (score) per module that aggregates fold-change values for the underlying genes. The scores for each module were normalized across the 654 liver experiments to have a unit variance. As such, a score denotes the degree of induction or repression achieved by a given treatment in the context of a large collection of drug perturbations (a score of 2 is exceeded ~4% of the time, 3 exceeded ~1% of the time and so on).
Defining toxicity phenotypes
Pathology assessment of drug-induced liver injury is typically defined in nonclinical studies by combining morphological assessment (histology) and biochemical measures (clinical chemistry). Occurrence of histology findings or changes in clinical chemistry results can occur in isolation or in combination. In isolation, they may be considered non-adverse, for example, an observation of hypertrophy or an increase in alanine aminotransferase (ALT) <300% with no other observation, or adverse, for example, ALT >100% with a finding of necrosis.
We use the term ‘anchoring phenotype’ to designate treatments resulting in a given histologically defined lesion (for example, single-cell necrosis) or clinical chemistry result (for example, increased total bilirubin). In order to study the association between module scores and different toxicity phenotypes, we identified treatments from TG in Supplementary Table S1 resulting in a severity score above a pre-defined threshold (usually a severity score ⩾0.67) for a given anchoring phenotype and severity scores ⩽0.67 for all other histology findings except infiltration/inflammation (Supplementary Methods). The latter was not considered in defining toxicity phenotypes as it co-occurs with many other histology findings and is generally not considered adverse in isolation. Since certain histology findings frequently co-occur, we defined different levels of selectivity favoring the anchoring phenotype. An example of the impact of stratification by level on the number of treatments can be seen for the anchor ‘single-cell necrosis’ that we treat herein as a morphological correlate of apoptosis. For ‘single-cell necrosis, (4) any other finding’, there were 65 TG treatments in Supplementary Table S1 with score ⩾0.67 for single-cell necrosis and ⩽0.67 for congestion, necrosis, dilation, fibrosis, increased glycogen, hematopoiesis, hyperplasia, hypertrophy, mitosis or vacuolation. Here, ‘any other finding’, unless further clarified in the phenotype label, means ‘any other finding at severity ⩽0.67’ and we use the shorter term throughout. Since hypertrophy and necrosis co-occur most frequently with single-cell necrosis, level 3 (‘single-cell necrosis, (3) allowing hypertrophy or necrosis’) eliminated all treatments causing any of the aforementioned findings except hypertrophy or necrosis at maximal severity of 0.67, leaving 54 treatments. Level 2 (‘single-cell necrosis, (2) allowing hypertrophy’) only allowed low-severity co-occurring hypertrophy (49 treatments) and level 1 (‘single-cell necrosis, (1) no other findings’) left 44 treatments where the only lesion present was single-cell necrosis. For other anchoring phenotypes, such as bile duct hyperplasia (BDH), the trade-off from level 3 to level 1 (33 versus 12 treatments) was more marked, and the latter includes nine 2-AAF treatments (that is, high purity, but few examples and probably not generally representative of BDH-causing compounds given dominance of one compound). In total, we defined 36 toxicity phenotypes each of which was anchored by a primary finding, with 1–4 levels of selectivity.
Association of co-expression modules with toxicity phenotypes
For a given module, the association between module score and the occurrence of toxicity phenotypes using TG data (for example, positive versus negative for BDH) was quantified using Cohen’s d, a measure of effect size. We noted that the average absolute module eigengene (avgAbsEG), a measure of overall transcriptional activity, had a large effect size for many toxicity phenotypes. Logistic regression was performed to determine the contribution of a given module in explaining the residual odds of toxicity after accounting for avgAbsEG, and the significance of the module represented as p-adj (Supplementary Methods).
Comparing one-stage versus two-stage marker gene identification
Molecular markers for toxicity (and other) phenotypes are usually defined via genome-wide analyses with adjustment of P-values to control the false discovery rate. For each toxicity phenotype, we repeated the above logistic regression analysis using each individual genes’ log fold change at 1 day to predict toxicity phenotypes manifesting at 4 days or later (that is, 20 269 regressions, one for each of 20 269 probe sets mapping to one or more of 14 078 genes represented on the array). The resulting p-adj values were converted to q-adj reflecting 20 269 tests.
A two-stage approach was employed for identifying genes associated with toxicity phenotypes: (1) identify modules with strong association to the phenotype, that is, q-adj <0.01, then (2) analyze the association of individual genes that make up all significant modules and perform false discovery rate adjustment only for those genes (rather than the complete genome; Supplementary Methods).
Module similarity analysis using case studies
To identify compounds with similar mechanism and pathogenesis, we searched all module profiles using a treatment with an established mechanism as ‘bait’. We modeled each experiment as a vector of 284 base module scores and compared the bait experiment to all other experiments by calculating the Pearson correlation R of the two vectors. Bait experiments were selected from representative compounds or treatments with clearly defined mechanisms of action and interpretations regarding mechanisms of toxicity. For tunicamycin (ER stress) and butylated hydroxyanisole (BHA; oxidative stress), we selected the dose–time combination with the maximum avgAbsEG as the bait experiment (that is, the dose–time combination giving maximum transcriptional response).
Bile duct ligation study
To study the interplay between cholestatic injury progression and liver gene expression, we performed a rat bile duct ligation study comparing three ligated versus three sham animals at time points ranging from 3 h to 14 days post surgery. Clinical chemistry and histology assessment of liver tissue were performed for each animal, and liver gene expression analyzed using Affymetrix RG230-2 microarrays (Affymetrix, Santa Clara, CA, USA). We also performed ChiP-seq on pooled tissue from animals following 24 h and 14 days ligation, using the following transcription factors: cJun, Sp1, Fxr, Hnf1a. Full details on the study are provided in Supplementary Methods.
The expression and ChIP-seq data have been deposited in NCBI’s Gene Expression Omnibus25 and are accessible through GEO Series accession number GSE87696 (http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE87696) and GSE87730.
Ethics statement
The rat bile duct ligation study was conducted at Covance Laboratories (Greenfield, IN, USA), a facility accredited by the Association for Assessment and Accreditation of Laboratory Animal Care International (AAALAC). Studies were approved by the Eli Lilly Animal Care and Use Committee at Covance Laboratories and conducted in accordance with the Guide for the Care and Use of Laboratory Animals as adopted and promulgated by the US National Institutes of Health, and were approved by the Eli Lilly Animal Care and Use Committee. Rodent and de-identified human gene expression data from Drug Matrix, open TG-GATEs and the Gene Expression Omnibus (GEO) repository are freely available to the public. No institutional review board approval was sought to analyze those data.
Results
Liver co-expression networks and the TXG-MAP
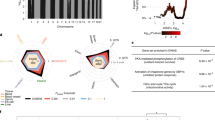
To develop the TXG-MAP, we utilized data from two large publicly available rat liver expression repositories, DM23 and open TG.24 Using WGCNA and 9071 liver-expressed genes, we defined 415 networks consisting of 5–656 co-expressed genes across all 654 DM experiments (Supplementary Table S1). The module correlation patterns based on scores across all treatments were represented in a simplified visual dendrogram, the TXG-MAP, analogous to phylogenetic trees used for conveying sequence similarity (Figure 1). Modules in close proximity on the map (measured by traversal of branches) tend to be induced or repressed coordinately. A labeling system facilitates navigation of the map (Supplementary Table S2 and Figure 1b). Using gene set enrichment, we assigned putative biological significance and predicted transcriptional regulators for each module (Supplementary Table S2). For example, the A-branch is highly enriched in mitochondrial genes with adjacent sub-branches enriched in Gene Ontology (GO) terms pertaining to lipid metabolism. As expected, modules in these branches respond to drugs such as statins and fibrates (Supplementary Figure S1). When we compared gene set analysis to co-expression modules, we noted that some modules yield equivalent results (for example, 46m module scores versus REACTOME cholesterol biosynthesis; Figure 1d), but many unique modules capture biology not represented by canonical pathways (Figure 1e). As noted below, we found that some of these unique modules were highly correlated with specific liver toxicity phenotypes. A reference table of all modules discussed in the text is provided in Supplementary Table S3.
Overview of the toxicogenomic module associations with pathogenesis (TXG-MAP) for analysis of liver gene expression changes. (a) Module 69 consists of co-induced genes; genes are ordered clockwise starting at the hub gene (Mbd2), with decreasing correlation against the module eigengene. Nodes are colored by log10 fold change upon treatment with lipopolysaccharides (LPS) (1.25 mg kg−1, 6 h). (b) 415 Modules are organized on the map using a branch labeling system, with modules in proximity (by branch traversal) reflecting the degree to which they are similarly perturbed across 654 Drug Matrix (DM) rat liver experiments. Each module can be summarized via one numeric score (the eigengene), with induction (red) or repression (blue) denoted in node color after treatment with LPS. Module functions were assigned using GO term enrichments based on gene membership. (c) Comparing the module 69 score for the LPS treatment in panel b treatment against the database of 4182 rat liver treatments from DM and TG-GATEs provides context for the magnitude of the effect. (d) Comparing scoring of 654 DM experiments using module 46m, which contains genes involved in cholesterol biosynthesis, to gene set analysis (GSA) scoring of the REACTOME cholesterol biosynthesis pathway. (e) Graphical representation indicating the number of pathways or GO terms (size) and highest Pearson correlation obtained by comparing scores for each module to GSA scoring of 1840 canonical pathways and GO-biological process terms for 654 DM experiments.
Modules reveal mechanisms of pathogenesis concurrent with liver injury
We analyzed relationships between module scores, reflecting induction or repression of underlying genes, and the presence (positives) or absence (negatives) of liver findings for each of 36 toxicity phenotypes anchored to a histology finding (Supplementary Table S4). Module associations with toxicity phenotypes, measured as effect sizes (Cohen’s d), are either positive (induction) or negative (repression) and considered ‘concurrent’ since liver findings and expression changes occurred at the same time. Effect sizes greater than 0.8 or less than −0.8 are considered large.26
Module signatures for toxicity phenotypes were defined as vectors of 415 effect sizes—one per module. Hierarchical clustering on these signatures revealed limited discrimination of several histologically distinct adverse phenotypes (Figure 2a). We noted that the average absolute module score (avgAbsEG), a measure of the overall transcriptional activity, was strongly associated with many adverse toxicity phenotypes (Supplementary Table S4). This result is intuitive: the occurrence of histologically defined lesions is indicative of liver injury, and injured cells have altered expression for many genes. Therefore, we used logistic regression treating avgAbsEG as a covariate and quantified the P-value for each module in explaining the odds of toxicity. We refer to this P-value as p-adj throughout, that is, after adjusting for avgAbsEG. Ranking module associations with toxicity based on p-adj versus effect size revealed greater differences among toxicity phenotypes having large effect sizes for avgAbsEG (Supplementary Figure S2 and Supplementary Table S5). For example, module 8 (GO-CC, ‘proteasome complex’, Supplementary Table S3) has a modest effect size versus single-cell necrosis, ranking 341 out of 415, but ranks 20th (p-adj of 5.7 × 10−5) when controlling for avgAbsEG.
Pairwise similarity and bi-clustering of toxicity phenotypes. Each toxicity phenotype anchored by a signal histology finding or clinical chemistry result (methods) was defined using 415 values (one per module) reflecting; (a) effect size, and (b) signed log10 p-adj, which accounts for the contribution of avgAbsEG (average module score) in explaining a toxicity phenotype. Both values represent the statistical association between module score and manifestation of the toxicity phenotype. Region (1) in a aggregates histologically distinct toxicity phenotypes, which are resolved into three distinct regions aligning with a unique anchoring histology in b. (c) Module signed log10 p-adj values for representative toxicity phenotypes on the TXG-MAP; positive/negative values indicate that module induction or repression was observed when the lesion was present. The dendrogram was cut at a height corresponding to R=0.7. The numerical value following the toxicity phenotype denotes its level of selectivity, for example, 1=no co-occurring pathology (Supplementary Table S4). Module size corresponds to the number of genes in a module.
We investigated the utility of p-adj for creating module-based signatures of toxicity phenotypes (Figure 2b). This approach substantially enhanced our ability to identify modules, and the underlying biological processes, uniquely associated with a toxicity phenotype or a grouping of related toxicity phenotypes anchored by common histology findings, for example, single-cell necrosis. We identified 98 phenotype pairs having Pearson R⩾0.7 when clustering on signed log10 p-adj, versus 228 using effect sizes (Supplementary Table S6). Phenotypes related to necrosis, single-cell necrosis and BDH that were not differentiated when clustered on effect size emerged as three distinct clusters using p-adj. The use of avgAbsEG as a covariate distinguishes phenotypes that have apparent similarity due to their common association with large transcriptional effects, but little overlap in underlying mechanisms characterizing the injury.
Using one representative phenotype per cluster (see below), we identified the most strongly associated modules for each toxicity phenotype, among those modules having false discovery rate-controlled p-adj lower than 0.01 (Table 1; referred to as q-adj throughout). Some top ranked modules were located on the map in areas with less biological annotation, for example, C.I and C.II (Figure 2c). Mechanism can be inferred from enriched GO terms, for example, hypertrophy and ‘glutathione metabolic process’ for module 42m or single-cell necrosis and ‘MAPK signaling’ for module 29 (Table 1). Modules 70 (enriched with cell cycle arrest genes, including Ddit3) and 29 (enriched for MAPK signaling consistent with apoptosis processes in liver27) were associated with single-cell necrosis, but not with necrosis (non-programmed cell death; single-cell necrosis is synonymous with programmed cell death or apoptosis in histological evaluation28). Thus, clustering on p-adj revealed distinct module associations with distinct forms of cell death not observed when clustering on effect size. The similarity between necrosis and BDH in Figure 2b arises due to the association of both toxicity phenotypes with modules related to inflammation and leukocyte activation (55m, 2m), and several in the E.II branch associated with decreased hepatocellular function.
All analyses of module versus toxicity relationships in this work focused on the TG expression, histology and clinical chemistry results, owing to the larger size of the database and diversity of toxicity phenotypes compared to DM. For BDH, similar results were found when using DM for defining module versus toxicity relationships. For three other phenotypes, we found few significant associations in DM (Supplementary Results).
Modules reveal mechanisms of pathogenesis preceding liver injury
The module–phenotype associations described above provide insight into mechanisms concurrent with injury. Identifying mechanisms that precede injury is also of great interest, that is induction or repression of genes or modules within the first 24 h after a single dose that are prodromal for the occurrence of specific toxicity phenotypes in multi-dose studies of longer duration. We focused on 361 drug–dose combinations with expression data from single-dose studies of 3–24 h duration, and labeled them as positive or negative for the occurrence of a toxicity phenotype between 4 and 29 days. We limited analysis to a subset of 11 toxicity phenotypes, selected as representative of expression clusters in Figure 2b, balancing selectivity of the histology anchor against the number of positive drug–dose pairs (Supplementary Table S4; all further analysis refers exclusively to this subset). After a single dose, we found more significant module–phenotype associations at 24 h prognostic of adversity after 4–29 days (for example, 2, 10, 0 and 19 modules with q-adj <0.01 for BDH at 3, 6, 9 and 24 h, respectively; Supplementary Table S5). The importance of controlling for avgAbsEG was highlighted by the large effect size for certain pathologies, notably BDH (Supplementary Table S4). In general, modules lost significance after accounting for avgAbsEG, however in some cases significance improved markedly. For example, when predicting the occurrence of BDH from univariate logistic regression, module 39 was marginally significant with P=0.08, ranking 298 of 415 modules. However, including avgAbsEG as a covariate improved significance to p-adj=2e−8 and ranked first of 415 modules. Notably, module 39 contains Krt19, a marker for bile duct epithelial cells.29, 30 Thus, for a given avgAbsEG, higher expression of module 39 genes predicts BDH indicating a mechanism involving extracellular matrix remodeling (Table 1). Further, multiple modules can be combined within classifiers for predicting 29-day study outcomes (adverse versus non-adverse) using compound-induced expression changes at 1 or 4 days, improving on the sensitivity observed when using histology-based assessment (Supplementary Results).
Utilizing modules to enable marker gene identification via a two-step approach
Molecular markers for toxicity (and other) phenotypes are usually defined at the gene level. We hypothesized that co-expression modules may have advantages (for example, reduced risk of false positive findings due to lower number of endpoints examined—415 versus ~20 000) but also disadvantages (missing a gene with strong association to a phenotype). Therefore, we repeated the above analysis using each individual genes’ log fold change at 1 day to predict toxicity phenotypes manifesting at 4 days or later (that is, one-step approach). We identified 19 genes having q-adj <0.01 and ⩾3 fold-increased odds of observing BDH upon a one s.d. change in expression; 11 of these genes were from modules meeting the same criteria (Supplementary Table S7). Conversely, by starting with 19 BDH-associated modules and evaluating only their 514 constituent genes, 116 genes were significant predictors using the same criteria (Supplementary Table S8). In summary, we identified approximately six times more predictive genes at the same level of false discovery control (1%), using the two-step approach. Further, 8 of 19 modules have more favorable p-adj for predicting BDH than any of their constituent genes implying that individual genes in the network have synergistic roles in pathogenesis. Thus, by first linking a module to the phenotype of interest and then studying its constituent genes for the same phenotype, we found that module- and gene-level analyses play complementary roles for predicting toxicity and understanding mechanism.
Utilizing co-expression modules to understand stress–response pathways in mechanisms of drug-induced liver injury
Understanding mechanisms for unexpected safety findings is critical in framing the clinical risk based on nonclinical results. The mechanism-based monitoring strategy as well as an understanding of whether injury is adaptive and reversible or progressive will impact the clinical dosing schedule. For example, oxidative stress and antioxidant response play well-established roles in drug-induced liver injury.31 We investigated module responses for several known Nrf2-inducing treatments and identified modules that differentiated the role of Nrf2 activation in adaptive hepatocellular hypertrophy versus non-adaptive progression to hepatocellular necrosis and fibrosis (Supplementary Results). In the next two sections, we focus on novel applications of the TXG-MAP for understanding liver injury.
Case study: endoplasmic reticulum stress as a novel mechanism for ethionamide hepatotoxicity
Tunicamycin causes unfolded protein to accumulate in the endoplasmic reticulum (ER), activating the unfolded protein response (ER stress). ER stress genes, such as Atf4 and Hspa5 (Grp78), help alleviate excess protein load; with extended ER stress activation of genes such as Ddit3 (Chop) contribute to cell death. When describing treatments using a vector of module scores, the drug most similar to tunicamycin in the database was ethionamide, a second line therapy for mycobacterial infection with hepatotoxicity liability32 (Figure 3). We found no literature reports that ethionamide causes ER stress. The only histological finding for tunicamycin was single-cell necrosis, which was also the dominant finding for ethionamide treatment across dose–time combinations (Supplementary Table S1).
Comparison of module scores for tunicamycin and ethionamide in rat liver. Eigengene scores (module expression) for; (a) tunicamycin 300 μg kg−1 at 9 h (single dose), (b) ethionamide, 100 mg kg−1 at 6 h (single dose), (c) correlation of module scores for tunicamycin and ethionamide, (d) primary rat hepatocytes were treated with either ethionamide or tunicamycin for 24 h, and immunoblots were run on the cell lysates as indicated. Functional assignments of labeled modules are discussed in Results and summarized in Supplementary Table S3.
Among modules most induced by tunicamycin and ethionamide, module 70 contains Ddit3 and was top-ranked for single-cell necrosis/apoptosis (Table 1), and module 75 contains canonical ER chaperones Hspa5 and Hsp90b1 (Grp94) (Supplementary Table S3). Notably, module 85 (B.I branch) was also induced by both compounds and contains Atf4 and Atf4-target genes (Supplementary Table S3). Thus, based on the composition of constituent genes, module responses were consistent with ER stress and then overload leading the cell death.33, 34
Module changes in rat hepatocyte treated with tunicamycin and ethionamide were similar including the four ER stress modules in C.I.b (Supplementary Figure S3). Treating rat primary hepatocytes with ethionamide increased Atf4, Hspa5 and Hsp90b1 protein and processing of Atf6 to the transcriptionally active form, evidence of ER stress (Figure 3d). Thus, we identified and validated a putative mechanism for ethionamide hepatotoxicity and predicted the associated pathology (single-cell necrosis). Chemical structure-based approaches, such as QSAR and read-across for toxicity prediction, would not have led to this result as ethionamide is a small molecule and tunicamycin a complex natural product.35
Case study: cellular mechanisms of bile duct hyperplasia
The prior case studies focus on findings that are primarily hepatocellular. Progressive cholestatic injury involves cell types other than hepatocytes.29 BDH (expansion of bile duct epithelia), a common sequela of cholestasis, may occur with or without co-occurring fibrosis (stellate cell activation), oval cell hyperplasia (expansion of hepatic precursor cells), and inflammation due to Kupffer cell activation and/or invading lymphocytes.30, 36, 37 Characterizing BDH as adverse or non-adverse may depend on the severity of hyperplasia, the morphology of the biliary epithelium, occurrence of inflammation, and other factors.38 We conducted a rat bile duct ligation study, a common model for cholestatic liver injury and BDH,39, 40 and monitored injury and expression changes from 3 h through 14 days. Necrosis, BDH, fibrosis and inflammation progressed over time, as expected (Figures 4a and b).
Liver pathogenesis and transcriptional control after bile duct ligation. Liver pathology and modular transcriptomic analysis were compared to time-matched sham surgical animals from 3 h to 14 days after bile duct ligation. (a) Percent change from levels in sham animals for select clinical chemistry endpoints, (b) severity scores for histology findings in ligated animals and (c) scores for selected modules in ligated versus sham animals. Scores for all modules on the TXG-MAP at 1 day (d) and 14 days (e). Livers from animals ligated for 1 and 14 days were further analyzed using ChIP-seq on four transcription factors, and enrichment of genes showing increased or decreased binding in bile duct ligation versus sham calculated for each module (f); modules are ordered by branch and only modules containing 20 or more genes are shown for improved clarity. For ChIP, a signed hypergeometric P-value is calculated for each module–factor pair, reflecting enrichment of differentially bound genes. Module eigengene (EG) and signed log10 hypergeometric TF-binding enrichment are scaled for each color to the maximum absolute module response at 1 or 14 days; 1 unit on the ordinate corresponds to the maximum response for a given color. Maximum response: 1 day=3.0 (module EG) and 8.3 (Jun), 2.3 (Sp1), 7.2 (FXR) and 3.8 (HNF1), 14 day=5.5 (module EG) and 20.1 (Jun), 5.9 (Sp1), 12.8 (FXR) and 12.7 (NHF1). Some module locations for enriched module in f are indicated in e, a complete list of module branch locations can be found in Supplementary Table S2. ALT, alanine aminotransferase.
Thirteen compound treatments were similar to bile duct ligation days 1–14, of which 11 also caused BDH and/or fibrosis (Supplementary Table S9). We identified induced modules associated with BDH, both concurrent and predictive (that is, expression change at 24 h preceding histological changes), as well as modules associated with co-occurring necrosis (Figure 4c and Table 1). Module 55 m was associated with necrosis (rank 11 of 415 modules, concurrent), and BDH (ranks 7 and 4 for predictive and concurrent, respectively). Module 55 m annotation was not extensive but included GO terms ‘cytoskeletal organization’ and ‘membrane raft organization’, due to membership of the hub genes Anxa2 (annexin 2) and its binding partner S100a10, along with other genes involved in membrane and cytoskeletal organization41 (Table 1). Module 88 was strongly associated with BDH (ranks 3 and 7 for predictive and concurrent, respectively) and contains keratins 8 and 18, components of intermediate filaments in hepatocytes and other simple epithelial cells. Release of cytokeratin 18 is a well-known biomarker of liver injury and a target for activated caspases.42 Among the modules associated with BDH, module 39 was particularly interesting, since it was top-ranked at 1 day (Table 1) and uniquely associated when induced but was repressed in association with hypertrophy and other non-adverse phenotypes (Supplementary Table S5). Enrichment for GO terms included ‘extracellular matrix organization’ and ‘collagen fibril organization’. These modules exhibit time-dependent induction for several BDH and fibrosis-inducing compounds (Supplementary Figure S4).
We noted that module 39 contains genes top-ranked for BDH when considered individually, including Tspan8, Krt19, Sox4, Loxl1 in the top 30 (genome-wide) concurrent with BDH, and Lgals1, Col1a2, Fbn1, Tagln, Krt19 and Tpm2 in the top 30 predictive of BDH (Supplementary Table S8). Some of these genes are markers for histological correlates of the ductular reaction, for example, proliferation of duct-like cells (Krt19) or activation of hepatic stellate cells (HSCs) and deposition of collagen (Col1a1, Col1a2, Fbn1).29, 43 Two interesting novel genes had strong association (concurrent) with BDH and lower induction by CCl4 (which causes fibrosis without BDH) relative to BDH-inducing treatments (Supplementary Figure S5): Tspan8 (rank 1 genome-wide, p-adj=2e−35, effect size=4.6) and Sox4 (rank 10, p-adj=1e−27; effect size=3.0; Supplementary Table S8). Tspan8 is overexpressed and prognostic of survival in several cancers,44, 45, 46 regulates cell adhesion and promotes migration.47 It has measurable protein expression in normal human biliary epithelial cells,48 and was identified in GWAS studies for type 2 diabetes.49, 50 Sox4 is a member of the SRY-related HMG-box family of transcription factors expressed in human hepatocytes48 and is required for biliary tract development in mice.51 It is overexpressed in and prognostic of several cancers, including cholangiocarcinoma and hepatocellular carcinoma.52, 53 These genes are worthy of further investigation given the relative paucity of the literature implicating them in BDH in comparison to other well-studied genes in the module, such as Krt19.
Upstream signals couple co-expression to toxicity phenotypes
To gain insight into how upstream signals couple hepatocellular injury to activation or repression of co-expression modules, we identified putative transcription factors (TFs) regulating module induction or repression using the bile duct ligation model and literature annotation of TF versus target gene pairs (Supplementary Table S10). Four factors were enriched among a plurality of induced (cJun and Sp1) and repressed modules (Hnf1a and Fxr). Jun, associated with immediate early response module 29, and Jun kinase (JNK) play important roles in liver pathogenesis.27, 54 Fxr and Hnf1a are implicated broadly in cholestatic liver injury.55, 56, 57, 58 Reports that Sp1 plays a role in liver injury are more anecdotal.59, 60, 61
To confirm the role of these four factors, we performed ChIP-seq on liver samples 1 and 14 days after ligation. Increased Jun binding occurred at both time points. Binding for other factors increased at 1 day and decreased at 14 days (Table 2). Of 33 module-TF associations predicted for these four factors, we validated 27 by ChIP-seq. In addition, we identified 111 additional module–TF associations. In total, 76 modules were enriched in target genes for any of the four factors, with 45 enriched for two or more factors (that is, possible multi-factor associations; Supplementary Table S12); 49 of the modules had ranks ⩽20 in association with BDH or necrosis.
We also examined the extent to which progression of injury and changes in module expression aligned with changes in TF binding. Modules in branches of the TXG-MAP showing strong association with BDH, for example, C.II.a (55m and 18m) and C.II.e (69 and 88), were characterized by strong induction (Figures 4d and e) and increased concomitant binding upon ligation by three or four of the factors at 1 day (Figure 4f). At 14 days, expression and cJun/HNF1 binding for modules 2/2m (immune systems process), 55m (membrane raft assembly) and 18m (leukocyte adhesion) overlapped strongly. Conversely, repression in E.II.b region (modules relevant to liver-specific function, Supplementary Table S3) overlapped with diminished binding by all but cJun, suggesting coordinated regulation of expression. Regions characterized by strong expression but devoid of binding for the four factors, for example, C.I at 1 day, consisted of modules related to cell cycle, ribosomal biogenesis and related functions, where other factors are predicted to regulate expression (Supplementary Table S2).
Module-based molecular phenotypes translate to human liver disease
BDH is both a common findings and an important histological determinant of liver pathogenesis in rodent and human.29, 38 To identify molecular phenotypes for BDH, we first performed hierarchical clustering using module scores for 57 rat liver treatments that caused BDH, and identified four distinct clusters (Figure 5). Bile duct ligation samples segregated across three clusters, based on time after ligation, and were labeled accordingly. One cluster (sub-acute BDH) had lower module perturbation. Notably, two of three treatments in the sub-acute cluster exhibited hyperplasia at 4–8 days that resolved at the later time points (2 mg kg−1 lomustine, 30 mg kg−1 4-MDA, Supplementary Table S1). The fourth cluster contained only BDH-inducing carcinogens such as 2-acetylaminofluorene and methapyrilene. Recursive partitioning identified the modules most responsible for grouping experiments into clusters. Notably, modules 8 (proteasome), 15m (mitochondrial biogenesis) and 37m (cell cycle) were induced by BDH-causing carcinogens; module 39 (extracellular matrix) and 2m (inflammation) correlated with progression to fibrosis and differentiated acute from chronic BDH while module 55m distinguished sub-acute BDH from other clusters. Modules 13m (complement cascade) and 23 (secreted proteins, for example, Alb, Apoe, Apoh, Fgg and mitochondrial coded genes; Supplementary Table S3) were more strongly repressed in chronic BDH and carcinogen clusters. Thus, module associations identified molecular subtyping of BDH linked to specific biological mechanisms useful for differentiating disease etiology and progression.
Molecular subtypes of BDH. Hierarchical clustering of 57 treatments from TG causing BDH in rats, using 415 co-expression modules, with treatment, dose in mg kg−1 and days of dosing indicated on rows; four distinct subtypes were identified from the dendrogram. The heatmap on the left shows a subset of modules selected with recursive partitioning that recapitulate the (all-module) clustering and severity of liver histology findings. Abbreviations are bile duct ligation (BDL), 4,4′ methylenedianiline (MDA), 1-naphthyl isothiocyanate (ANIT), 2-acetylaminofluorene (2AAF), N-nitrosodiethylamine (DEN). The heat map on the right shows the histopathology scores for the same treatments.
Compared to gene-level analysis, modules improved the correlation for expression changes across species.11 Therefore, it was of particular interest to determine if module-based molecular phenotypes identified in rat translate to human liver. Although transcriptomic data are not available for drug-induced liver injury in human, data are available for diseases that share etiology with drug-induced liver injury in rodents, such as cholestasis, lipid accumulation (steatosis) and inflammation/fibrosis. The latter includes a spectrum of diseases that may or may not progress to liver failure, including alcoholic liver disease (ALD), non-alcoholic steatohepatitis (NASH), and hepatitis B and C infection. Therefore, we evaluated liver expression data from the GEO for normal and disease samples including: type 2 diabetes, non-alcoholic fatty liver (NAFL also termed steatosis) and NASH, ALD, biliary atresia (BA; a congenital cholestatic condition in infants), hepatitis B infection-mediated acute liver failure, hepatitis C infection-mediated cirrhosis, hepatocellular carcinoma (HCC) and hepatoblastoma.
We assessed the similarity of module scores for human liver diseases compared to rat liver toxicity phenotypes (Supplementary Table S13). Module responses in liver tissue from patients with non-alcoholic fatty liver (NAFL) or type 2 diabetes had low similarity to all rat liver treatments. However, module responses in BA, ALD, NASH, acute liver failure and cirrhotic liver all tended to resemble rat liver treatments causing bile duct hyperplasia accompanied by co-occurring fibrosis and/or necrosis. Among all treatments, bile duct ligation most closely mimicked biliary atresia. Bile duct ligation combined with methionine-choline deficient diet and lipopolysaccharide treatment more closely mimicked NASH (Supplementary Table S14). Of interest, other common rodent NASH models, such as the CCl4-liver injury model and diet-induced models (for example, high cholesterol, high fructose and high fat), show lower concordance with human disease (Supplementary Table S14).
Since the molecular profile of human BA, ALD and NASH generally resembled rat treatments resulting in BDH, we evaluated the modules that defined the rat BDH subtypes in human liver disease, both in terms of their magnitude of change compared to normal human liver (Figure 6), and relative rank across the 415 rodent modules (Supplementary Table S15). In particular, module 37m was strongly induced and ranked in ⩾97th percentile (100=most induced) across HCC etiologies and hepatoblastoma; strongest induction of 37m in rat was for BDH carcinogens and ranked 4th in association with the increased mitosis across all liver treatments (Supplementary Table S3). Module 39 was most induced in NASH, ALD, BA, acute liver failure and cirrhosis. Module 2m was induced in all diseases except HCC, consistent with the immunological suppression to evade cell death in cancer.62 Modules 13m (complement activation) and 23 (very-low-density lipoprotein) were downregulated in all states, most substantially in cirrhosis, acute liver failure and HCC. In summary, modules most significantly perturbed in rat BDH show similar effects in human liver disease, supporting translational utility. Further, when applying the two-stage approach for disease gene identification (analyzing module versus phenotype relations first, then examining their constituent genes), several genes implicated in liver fibrosis were identified (Figure 7 and Supplementary Results).
Comparison of BDH molecular subtypes in rats versus human liver disease. Module scores are averaged across treatments in each BDH subtype (rats) from Figure 5 (top heatmap) and human samples (bottom heatmap) available in each Gene Expression Omnibus (GEO) series, identified via their accession number.
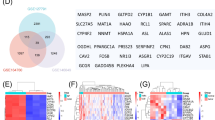
Analysis of modules and genes in human NASH. Distinguishing liver samples from patients with non-alcoholic steatohepatitis (NASH) versus those with normal liver or non-alcoholic fatty liver (NAFL). (a) Waterfall plot showing the significance of individual genes in distinguishing NASH from non-NASH samples in two GEO series, indicating whether genes are pre-selected by membership of key rat bile duct hyperplasia (BDH)-associated modules. The nine rat modules used for preselection contain 15 or more genes, have abs(coef) >1.09 (odds ratio of 3) from logistic regression and q-adj <0.01 for both prediction of rat BDH at 1 day and concurrent with BDH. The waterfall plot includes 560 genes with P-value (non-adj) ⩽0.05 for GSE48452 (NASH versus pooled NAFL/obese) and GSE49541 (NASH versus NAFL). Illustrating application of module 39 (b) and 88 (c) in distinguishing human liver samples, with number of samples indicated in parentheses. Comparing the performance of the best individual genes within module 88 selected in GSE48452 (d; KRT8) and GSE49541 (e; TPM1), showing that the individual genes perform notably worse than the module in the data set not used for its selection. All P-values shown in b–e are from t-tests assuming unequal variance.
Discussion
Use of toxicogenomics in drug safety assessment has expanded in the last decade, driven in part by improvements and standardization of measurement technology.63, 64, 65 However, inclusion of genomic studies in formal drug safety assessment remains limited,66 with relatively few reports describing significant insights from expression profiling that complement traditional nonclinical studies.67, 68 This may be due in part to a focus on gene signatures as chemical classifiers and a poor understanding of how selected genes from case-by-case analyses fit into the larger context of organ injury. We reasoned that co-expression is a ‘coalescent property’19 of liver and might order gene expression data into biologically relevant networks of interacting elements extensible to other species. To test this hypothesis, we developed the TXG-MAP approach, an integrated analysis and visualization framework, using a large corpus of transcriptomic and pathology data. Central to our approach was the use of co-expression network analysis to identify genes behaving similarly across hundreds of drug perturbations to reduce the dimensionality of transcriptomic data and to facilitate visual interpretation of results. We illustrated the utility of the approach to identify mechanisms of pathogenesis and demonstrated translation from nonclinical models to human, two central questions in drug safety assessment.
Summarizing expression changes using co-expression modules addresses a significant challenge in toxicogenomics: the challenge of ‘large p, small n’ that arises when analyzing phenotypic outcomes for a few hundred treatments (typically with few positive class examples) with thousands of genes.18 We illustrated the utility of a two-stage approach, whereby modules are first linked to a phenotype followed by gene-level analysis performed within significant modules to identify constituent genes that underpin the modular behavior and association with the phenotype. Compared to gene-level analysis alone, this approach identified ~6-fold more genes predictive of the later occurrence of bile duct hyperplasia using expression at 1 day. Several genes implicated in BDH by the literature that fail to reach genome-wide significance (q-value >0.05) were successfully identified using this approach (for example, Epcam, Pdgfrb).29
Consistent with the observations by Foster et al.,69 we highlighted the strong association between avgAbsEG, a measure of overall transcriptional effect,11 and adverse phenotypes, for example, necrosis, single-cell necrosis (apoptosis) and BDH. Treating avgAbsEG as a covariate allowed us to identify groups of module–phenotype relationships on the TXG-MAP that uniquely associate with distinct histology findings, both concurrent (induced concurrent with the lesion) and as predictors of toxicity (for example, induced prior to the lesion). This distinction is important, as models that ‘predict’ injury when the lesion is visible, without providing mechanistic information, are of little practical utility if histology assessment is conducted in parallel. Notably, we identified modules most predictive of toxicity phenotypes in areas of the TXG-MAP having little overlap with canonical pathways. Moreover, pathways and gene ontology did not perform as well as modules when expression data from studies of 1- or 4-day duration were used to predict for adverse outcomes in 29-day studies, study durations required for entering phase I studies in humans.5 Notably, adding expression-derived features (using modules or genes), to 1-day studies was sufficient to predict adverse outcomes in 29-day studies, suggesting it may be feasible to incorporate safety endpoints into standard 1-day toxicokinetic studies.
The pathogenesis of cholestatic liver disease and biliary tract injury involves complex interactions among cholangiocytes, hepatocytes, stellate cells and other non-parenchymal cells and lymphocytes. Expression changes observed in whole liver tissue reflect changes in the baseline state of all underlying cells responding to injury (for example, upregulation of detoxifying enzymes in hepatocytes; activation of stellate cells), cell proliferation and trans-differentiation across cell types (for example, the ductular reaction70). We demonstrated similar histology and module response for bile duct ligation and compound-induced BDH or fibrosis involving these cell types and identified known as well as potential marker genes useful in the study of these toxicity phenotypes.
We also identified upstream transcription factors (TF) associated with modules changes by validating literature-based prediction of TF-target gene pairs with ChiP-seq analysis for four factors, cJun, Sp1, FXR and HNF1 following 1 and 14 days of bile duct ligation. cJun binding was enriched in the C.II branch, an area rich in modules associated with toxicity phenotypes. Notably, cJun and its upstream kinase Jnk are implicated in liver disease pathogenesis.54 Approximately 50% of C.II modules significantly associated with adverse pathology have constituent genes with increased cJun binding at one or both time points. Although GO and pathway enrichment was lower across C.II, enriched terms were related to actin cytoskeletal and cell–cell contact, suggesting perturbation of these components may be important early predictors of hepatocellular injury.
The development of biomarkers of altered liver function and progression of liver disease is critical.71 However, establishing the relevance of insights obtained from transcriptomic analysis in preclinical species is challenging due to the paucity of human liver data. Analyses based on conservation of biological properties (coalescent properties) across species, including co-expression modules, seemed a tractable approach that avoids the impracticality of sampling human subjects before and after drug administration. Key modules associated with drug-induced BDH (and fibrosis) in rats were highly perturbed for several human diseases with similar etiology. Further, using rat-derived co-expression networks, we identified known and putative gene markers diagnostic for human liver disease that would be missed in false discovery rate-controlled genome-wide analysis.
In summary, we developed the TXG-MAP, a network-based visualization framework for analyzing transcriptomic data. This approach allows new experiments to be interpreted in a physiologically relevant context and exemplify translation to human disease. We suggest that this approach offers significant advantages in human risk assessment for drug candidates.
References
Arrowsmith J . Trial watch: phase II failures: 2008-2010. Nat Rev Drug Discov 2011; 10: 328–329.
Arrowsmith J . Trial watch: phase III and submission failures: 2007-2010. Nat Rev Drug Discov 2011; 10: 87.
Waring MJ, Arrowsmith J, Leach AR, Leeson PD, Mandrell S, Owen RM et al. An analysis of the attrition of drug candidates from four major pharmaceutical companies. Nat Rev Drug Discov 2015; 14: 475–486.
Bai JPF, Abernethy DR . Systems pharmacology to predict drug toxicity: integration across levels of biological organization. Annu Rev Pharmacol Toxicol 2013; 53: 451–473.
Stevens JL . Future of toxicology—mechanisms of toxicity and drug safety: where do we go from here? Chem Res Toxicol 2006; 19: 1393–1401.
Greaves P, Williams A, Eve M . First dose of potential new medicines to humans: how animals help. Nat Rev Drug Discov 2004; 3: 226–236.
Peers IS, South MC, Ceuppens PR, Bright JD, Pilling E . Can you trust your animal study data? Nat Rev Drug Discov 2014; 13: 560.
Hartung T . Toxicology for the twenty-first century. Nature 2009; 460: 208–212.
Cook D, Brown D, Alexander R, March R, Morgan P, Satterthwaite G et al. Lessons learned from the fate of AstraZeneca’s drug pipeline: a five-dimensional framework. Nat Rev Drug Discov 2014; 13: 419–431.
Sutherland JJ, Raymond JW, Stevens JL, Baker TK, Watson DE . Relating molecular properties and in vitro assay results to in vivo drug disposition and toxicity outcomes. J Med Chem 2012; 55: 6455–6466.
Sutherland JJ, Jolly RA, Goldstein KM, Stevens JL . Assessing concordance of drug-induced transcriptional response in rodent liver and cultured hepatocytes. PLoS Comput Biol 2016; 12: e1004847.
Afshari CA, Hamadeh HK, Bushel PR . The evolution of bioinformatics in toxicology: advancing toxicogenomics. Toxicol Sci Off J Soc Toxicol 2011; 120 (Suppl 1): S225–S237.
Smalley JL, Gant TW, Zhang S-D . Application of connectivity mapping in predictive toxicology based on gene-expression similarity. Toxicology 2010; 268: 143–146.
Tawa GJ, AbdulHameed MDM, Yu X, Kumar K, Ippolito DL, Lewis JA et al. Characterization of chemically induced liver injuries using gene co-expression modules. PLoS ONE 2014; 9: e107230.
Te JA, AbdulHameed MDM, Wallqvist A . Systems toxicology of chemically induced liver and kidney injuries: histopathology-associated gene co-expression modules. J Appl Toxicol 2016; 36: 1137–1149.
Bell SM, Angrish MM, Wood CE, Edwards SW . Integrating publicly available data to generate computationally predicted adverse outcome pathways for fatty liver. Toxicol Sci Off J Soc Toxicol 2016; 150: 510–520.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 2005; 102: 15545–15550.
Ideker T, Dutkowski J, Hood L . Boosting signal-to-noise in complex biology: prior knowledge is power. Cell 2011; 144: 860–863.
Califano A, Butte AJ, Friend S, Ideker T, Schadt E . Leveraging models of cell regulation and GWAS data in integrative network-based association studies. Nat Genet 2012; 44: 841–847.
Wagner GP, Pavlicev M, Cheverud JM . The road to modularity. Nat Rev Genet 2007; 8: 921–931.
Barabási A-L, Gulbahce N, Loscalzo J . Network medicine: a network-based approach to human disease. Nat Rev Genet 2011; 12: 56–68.
Zhang B, Horvath S . A general framework for weighted gene co-expression network analysis. Stat Appl Genet Mol Biol 2005; 4, Article17.
Ganter B, Tugendreich S, Pearson CI, Ayanoglu E, Baumhueter S, Bostian KA et al. Development of a large-scale chemogenomics database to improve drug candidate selection and to understand mechanisms of chemical toxicity and action. J Biotechnol 2005; 119: 219–244.
Igarashi Y, Nakatsu N, Yamashita T, Ono A, Ohno Y, Urushidani T et al. Open TG-GATEs: a large-scale toxicogenomics database. Nucleic Acids Res 2015; 43: D921–D927.
Edgar R, Domrachev M, Lash AE . Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res 2002; 30: 207–210.
Cohen D. Statistical Power Analysis for the Behavioral Sciences, 2nd edn. Lawrence Erlbaum Associates: Hillsdale, NJ, 1988.
Brenner C, Galluzzi L, Kepp O, Kroemer G . Decoding cell death signals in liver inflammation. J Hepatol 2013; 59: 583–594.
Elmore SA, Dixon D, Hailey JR, Harada T, Herbert RA, Maronpot RR et al. Recommendations from the INHAND Apoptosis/Necrosis Working Group. Toxicol Pathol 2016; 44: 173–188.
Williams MJ, Clouston AD, Forbes SJ . Links between hepatic fibrosis, ductular reaction, and progenitor cell expansion. Gastroenterology 2014; 146: 349–356.
Boulter L, Lu W-Y, Forbes SJ . Differentiation of progenitors in the liver: a matter of local choice. J Clin Invest 2013; 123: 1867–1873.
Tang W, Jiang Y-F, Ponnusamy M, Diallo M . Role of Nrf2 in chronic liver disease. World J Gastroenterol 2014; 20: 13079–13087.
Livertox database. Available athttp://livertox.nlm.nih.gov/ Accessed 13 Jun2016.
Walter P, Ron. D . The unfolded protein response: from stress pathway to homeostatic regulation. Science 2011; 334: 1081–1086.
Wang M, Kaufman RJ . Protein misfolding in the endoplasmic reticulum as a conduit to human disease. Nature 2016; 529: 326–335.
Ellison CM, Enoch SJ, Cronin MTD . A review of the use of in silico methods to predict the chemistry of molecular initiating events related to drug toxicity. Expert Opin Drug Metab Toxicol 2011; 7: 1481–1495.
Wallace MC, Friedman SL, Mann DA . Emerging and disease-specific mechanisms of hepatic stellate cell activation. Semin Liver Dis 2015; 35: 107–118.
Bataller R, Brenner DA . Liver fibrosis. J Clin Invest 2005; 115: 209–218.
Hailey JR, Nold JB, Brown RH, Cullen JM, Holder JC, Jordan HL et al. Biliary proliferative lesions in the Sprague-Dawley rat: adverse/non-adverse. Toxicol Pathol 2013; 42: 844–854.
Rodríguez-Garay EA . Cholestasis: human disease and experimental animalmodels. Ann Hepatol 2003; 2: 150–158.
Woolbright BL, Jaeschke H . Novel insight into mechanisms of cholestatic liver injury. World J Gastroenterol 2012; 18: 4985–4993.
Hitchcock JK, Katz AA, Schäfer G . Dynamic reciprocity: the role of annexin A2 in tissue integrity. J Cell Commun Signal 2014; 8: 125–133.
Eguchi A, Wree A, Feldstein AE . Biomarkers of liver cell death. J Hepatol 2014; 60: 1063–1074.
Hernandez-Gea V, Friedman SL . Pathogenesis of liver fibrosis. Annu Rev Pathol 2011; 6: 425–456.
Zhu H, Wu Y, Zheng W, Lu S . CO-029 is overexpressed in gastric cancer and mediates the effects of EGF on gastric cancer cell proliferation and invasion. Int J Mol Med 2015; 35: 798–802.
Pan S-J, Wu Y-B, Cai S, Pan Y-X, Liu W, Bian L-G et al. Over-expression of tetraspanin 8 in malignant glioma regulates tumor cell progression. Biochem Biophys Res Commun 2015; 458: 476–482.
Park CS, Kim T-K, Kim HG, Kim Y-J, Jeoung MH, Lee WR et al. Therapeutic targeting of tetraspanin8 in epithelial ovarian cancer invasion and metastasis. Oncogene 2016; 35: 4540–4548.
Guo Q, Xia B, Zhang F, Richardson MM, Li M, Zhang JS et al. Tetraspanin CO-029 inhibits colorectal cancer cell movement by deregulating cell-matrix and cell-cell adhesions. PLoS ONE 2012; 7: e38464.
Human Protein Atlas. Available athttp://www.proteinatlas.org/. Accessed 16 Jun 2016.
Prabhanjan M, Suresh RV, Murthy MN, Ramachandra NB . Type 2 diabetes mellitus disease risk genes identified by genome wide copy number variation scan in normal populations. Diabetes Res Clin Pract 2016; 113: 160–170.
Ho MM, Yoganathan P, Chu KY, Karunakaran S, Johnson JD, Clee SM . Diabetes genes identified by genome-wide association studies are regulated in mice by nutritional factors in metabolically relevant tissues and by glucose concentrations in islets. BMC Genet 2013; 14: 10.
Poncy A, Antoniou A, Cordi S, Pierreux CE, Jacquemin P, Lemaigre FP . Transcription factors SOX4 and SOX9 cooperatively control development of bile ducts. Dev Biol 2015; 404: 136–148.
Wang W, Zhang J, Zhan X, Lin T, Yang M, Hu J et al. SOX4 is associated with poor prognosis in cholangiocarcinoma. Biochem Biophys Res Commun 2014; 452: 614–621.
Hur W, Rhim H, Jung CK, Kim JD, Bae SH, Jang JW et al. SOX4 overexpression regulates the p53-mediated apoptosis in hepatocellular carcinoma: clinical implication and functional analysis in vitro. Carcinogenesis 2010; 31: 1298–1307.
Seki E, Brenner DA, Karin M . A liver full of JNK: signaling in regulation of cell function and disease pathogenesis, and clinical approaches. Gastroenterology 2012; 143: 307–320.
Beuers U, Trauner M, Jansen P, Poupon R . New paradigms in the treatment of hepatic cholestasis: from UDCA to FXR, PXR and beyond. J Hepatol 2015; 62: S25–S37.
Geier A, Zollner G, Dietrich CG, Wagner M, Fickert P, Denk H et al. Cytokine-independent repression of rodent Ntcp in obstructive cholestasis. Hepatology 2005; 41: 470–477.
Pontoglio M, Barra J, Hadchouel M, Doyen A, Kress C, Bach JP et al. Hepatocyte nuclear factor 1 inactivation results in hepatic dysfunction, phenylketonuria, and renal Fanconi syndrome. Cell 1996; 84: 575–585.
Qian H, Deng X, Huang Z-W, Wei J, Ding C-H, Feng R-X et al. An HNF1α-regulated feedback circuit modulates hepatic fibrogenesis via the crosstalk between hepatocytes and hepatic stellate cells. Cell Res 2015; 25: 930–945.
Iqbal J, McRae S, Banaudha K, Mai T, Waris G . Mechanism of hepatitis C virus (HCV)-induced osteopontin and its role in epithelial to mesenchymal transition of hepatocytes. J Biol Chem 2013; 288: 36994–37009.
Bao W-D, Fan Y, Deng Y-Z, Long L-Y, Wang J-J, Guan D-X et al. Iron overload in hereditary tyrosinemia type 1 induces liver injury through the Sp1/Tfr2/hepcidin axis. J Hepatol 2016; 65: 137–145.
Hart ML, Gorzolla IC, Schittenhelm J, Robson SC, Eltzschig HK . SP1-dependent induction of CD39 facilitates hepatic ischemic preconditioning. J Immunol 2010; 184: 4017–4024.
Makarova-Rusher OV, Medina-Echeverz J, Duffy AG, Greten TF . The yin and yang of evasion and immune activation in HCC. J Hepatol 2015; 62: 1420–1429.
Shi L, Reid LH, Jones WD, Shippy R, Warrington JA, Baker SC et al. The MicroArray Quality Control (MAQC) project shows inter- and intraplatform reproducibility of gene expression measurements. Nat Biotechnol 2006; 24: 1151–1161.
Fan X, Lobenhofer EK, Chen M, Shi W, Huang J, Luo J et al. Consistency of predictive signature genes and classifiers generated using different microarray platforms. Pharmacogenomics J 2010; 10: 247–257.
Su Z, Labaj PP, Thierry-Mieg J, Thierry-Mieg D, Shi W et al. A comprehensive assessment of RNA-seq accuracy, reproducibility and information content by the Sequencing Quality Control Consortium. Nat Biotechnol 2014; 32: 903–914.
Chen M, Zhang M, Borlak J, Tong W . A decade of toxicogenomic research and its contribution to toxicological science. Toxicol Sci 2012; 130: 217–228.
Adler M, Leich E, Ellinger-Ziegelbauer H, Hewitt P, Dekant W, Rosenwald A et al. Application of RNA interference to improve mechanistic understanding of omics responses to a hepatotoxic drug in primary rat hepatocytes. Toxicology 2014; 326: 86–95.
Wang T, Papoutsi M, Wiesmann M, DeCristofaro M, Keselica MC, Skuba E et al. Investigation of correlation among safety biomarkers in serum, histopathological examination, and toxicogenomics. Int J Toxicol 2011; 30: 300–312.
Foster WR, Chen S-J, He A, Truong A, Bhaskaran V, Nelson DM et al. A retrospective analysis of toxicogenomics in the safety assessment of drug candidates. Toxicol Pathol 2007; 35: 621–635.
Tarlow BD, Pelz C, Naugler WE, Wakefield L, Wilson EM, Finegold MJ et al. Bipotential adult liver progenitors are derived from chronically injured mature hepatocytes. Cell Stem Cell 2014; 15: 605–618.
Lomonaco R, Sunny NE, Bril F, Cusi K . Nonalcoholic fatty liver disease: current issues and novel treatment approaches. Drugs 2013; 73: 1–14.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on The Pharmacogenomics Journal website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Sutherland, J., Webster, Y., Willy, J. et al. Toxicogenomic module associations with pathogenesis: a network-based approach to understanding drug toxicity. Pharmacogenomics J 18, 377–390 (2018). https://doi.org/10.1038/tpj.2017.17
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/tpj.2017.17
This article is cited by
-
Progress in toxicogenomics to protect human health
Nature Reviews Genetics (2024)
-
The evolving role of investigative toxicology in the pharmaceutical industry
Nature Reviews Drug Discovery (2023)
-
Mapping the cellular response to electron transport chain inhibitors reveals selective signaling networks triggered by mitochondrial perturbation
Archives of Toxicology (2022)
-
The human hepatocyte TXG-MAPr: gene co-expression network modules to support mechanism-based risk assessment
Archives of Toxicology (2021)
-
De novo generation of hit-like molecules from gene expression signatures using artificial intelligence
Nature Communications (2020)