Abstract
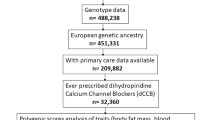
Sulfonylureas, a commonly used class of medication used to treat type 2 diabetes, have been associated with an increased risk of cardiovascular disease. Their effects on QT interval duration and related electrocardiographic phenotypes are potential mechanisms for this adverse effect. In 11 ethnically diverse cohorts that included 71 857 European, African-American and Hispanic/Latino ancestry individuals with repeated measures of medication use and electrocardiogram (ECG) measurements, we conducted a pharmacogenomic genome-wide association study of sulfonylurea use and three ECG phenotypes: QT, JT and QRS intervals. In ancestry-specific meta-analyses, eight novel pharmacogenomic loci met the threshold for genome-wide significance (P<5 × 10−8), and a pharmacokinetic variant in CYP2C9 (rs1057910) that has been associated with sulfonylurea-related treatment effects and other adverse drug reactions in previous studies was replicated. Additional research is needed to replicate the novel findings and to understand their biological basis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 print issues and online access
$259.00 per year
only $43.17 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Nathan DM . Diabetes: advances in diagnosis and treatment. JAMA 2015; 314: 1052–1062.
Hampp C, Borders-Hemphill V, Moeny DG, Wysowski DK . Use of antidiabetic drugs in the u.s., 2003-2012. Diabetes Care 2014; 37: 1367–1374.
Meinert CL, Knatterud GL, Prout TE, Klimt CR . A study of the effects of hypoglycemic agents on vascular complications in patients with adult-onset diabetes. II. Mortality results. Diabetes 1970; 19 (Suppl): 789–830.
Monami M, Genovese S, Mannucci E . Cardiovascular safety of sulfonylureas: a meta-analysis of randomized clinical trials. Diabetes Obes Metab 2013; 15: 938–953.
Simpson SH, Lee J, Choi S, Vandermeer B, Abdelmoneim AS, Featherstone TR . Mortality risk among sulfonylureas: a systematic review and network meta-analysis. Lancet Diabetes Endocrinol 2015; 3: 43–51.
Ikeda T . QT prolongation in type 2 diabetes mellitus treated with glibenclamide. Diabete Metab 1994; 20: 565–567.
Najeed SA, Khan IA, Molnar J, Somberg JC . Differential effect of glyburide (glibenclamide) and metformin on QT dispersion: a potential adenosine triphosphate sensitive K+ channel effect. Am J Cardiol 2002; 90: 1103–1106.
Schwartz PJ, Wolf S . QT interval prolongation as predictor of sudden death in patients with myocardial infarction. Circulation 1978; 57: 1074–1077.
Zhang Y, Post WS, Blasco-Colmenares E, Dalal D, Tomaselli GF, Guallar E . Electrocardiographic QT interval and mortality: a meta-analysis. Epidemiology 2011; 22: 660–670.
Zhang Y, Post WS, Dalal D, Blasco-Colmenares E, Tomaselli GF, Guallar E . QT-interval duration and mortality rate: results from the Third National Health and Nutrition Examination Survey. Arch Intern Med 2011; 171: 1727–1733.
Chow E, Bernjak A, Williams S, Fawdry RA, Hibbert S, Freeman J et al. Risk of cardiac arrhythmias during hypoglycemia in patients with type 2 diabetes and cardiovascular risk. Diabetes 2014; 63: 1738–1747.
Heller S, Darpo B, Mitchell MI, Linnebjerg H, Leishman DJ, Mehrotra N et al. Considerations for assessing the potential effects of antidiabetes drugs on cardiac ventricular repolarization: a report from the Cardiac Safety Research Consortium. Am Heart J 2015; 170: 23–35.
Lasser KE, Allen PD, Woolhandler SJ, Himmelstein DU, Wolfe SM, Bor DH . Timing of new black box warnings and withdrawals for prescription medications. JAMA 2002; 287: 2215–2220.
Qureshi ZP, Seoane-Vazquez E, Rodriguez-Monguio R, Stevenson KB, Szeinbach SL . Market withdrawal of new molecular entities approved in the United States from 1980 to 2009. Pharmacoepidemiol Drug Saf 2011; 20: 772–777.
Shah RR . Drugs, QTc interval prolongation and final ICH E14 guideline : an important milestone with challenges ahead. Drug Saf 2005; 28: 1009–1028.
Hanson B, Tuna N, Bouchard T, Heston L, Eckert E, Lykken D et al. Genetic factors in the electrocardiogram and heart rate of twins reared apart and together. Am J Cardiol 1989; 63: 606–609.
Newton-Cheh C, Larson MG, Corey DC, Benjamin EJ, Herbert AG, Levy D et al. QT interval is a heritable quantitative trait with evidence of linkage to chromosome 3 in a genome-wide linkage analysis: The Framingham Heart Study. Heart Rhythm 2005; 2: 277–284.
Arking DE, Pulit SL, Crotti L, van der Harst P, Munroe PB, Koopmann TT et al. Genetic association study of QT interval highlights role for calcium signaling pathways in myocardial repolarization. Nat Genet 2014; 46: 826–836.
Zhou K, Donnelly L, Burch L, Tavendale R, Doney AS, Leese G et al. Loss-of-function CYP2C9 variants improve therapeutic response to sulfonylureas in type 2 diabetes: a Go-DARTS study. Clin Pharmacol Ther 2010; 87: 52–56.
Holstein A, Plaschke A, Ptak M, Egberts EH, El-Din J, Brockmoller J et al. Association between CYP2C9 slow metabolizer genotypes and severe hypoglycaemia on medication with sulphonylurea hypoglycaemic agents. Br J Clin Pharmacol 2005; 60: 103–106.
Feng Y, Mao G, Ren X, Xing H, Tang G, Li Q et al. Ser1369Ala variant in sulfonylurea receptor gene ABCC8 is associated with antidiabetic efficacy of gliclazide in Chinese type 2 diabetic patients. Diabetes Care 2008; 31: 1939–1944.
Javorsky M, Klimcakova L, Schroner Z, Zidzik J, Babjakova E, Fabianova M et al. KCNJ11 gene E23K variant and therapeutic response to sulfonylureas. Eur J Intern Med 2012; 23: 245–249.
Sesti G, Laratta E, Cardellini M, Andreozzi F, Del Guerra S, Irace C et al. The E23K variant of KCNJ11 encoding the pancreatic beta-cell adenosine 5'-triphosphate-sensitive potassium channel subunit Kir6.2 is associated with an increased risk of secondary failure to sulfonylurea in patients with type 2 diabetes. J Clin Endocrinol Metab 2006; 91: 2334–2339.
Cho HJ, Lee SY, Kim YG, Oh SY, Kim JW, Huh W et al. Effect of genetic polymorphisms on the pharmacokinetics and efficacy of glimepiride in a Korean population. Clin Chim Acta 2011; 412: 1831–1834.
Avery CL, Sitlani CM, Arking DE, Arnett DK, Bis JC, Boerwinkle E et al. Drug-gene interactions and the search for missing heritability: a cross-sectional pharmacogenomics study of the QT interval. Pharmacogenomics J 2014; 14: 6–13.
Sitlani CM, Rice KM, Lumley T, McKnight B, Cupples LA, Avery CL et al. Generalized estimating equations for genome-wide association studies using longitudinal phenotype data. Stat Med 2015; 34: 118–130.
Akylbekova EL, Payne JP, Newton-Cheh C, May WL, Fox ER, Wilson JG et al. Gene-environment interaction between SCN5A-1103Y and hypokalemia influences QT interval prolongation in African Americans: the Jackson Heart Study. Am Heart J 2014; 167: 116–122 e111.
Psaty BM, O'Donnell CJ, Gudnason V, Lunetta KL, Folsom AR, Rotter JI et al. Cohorts for Heart and Aging Research in Genomic Epidemiology (CHARGE) Consortium: design of prospective meta-analyses of genome-wide association studies from 5 cohorts. Circulation Cardiovascular genetics 2009; 2: 73–80.
Skol AD, Scott LJ, Abecasis GR, Boehnke M . Joint analysis is more efficient than replication-based analysis for two-stage genome-wide association studies. Nat Genet 2006; 38: 209–213.
Thomas DC, Casey G, Conti DV, Haile RW, Lewinger JP, Stram DO . Methodological Issues in Multistage Genome-wide Association Studies. Stat Sci 2009; 24: 414–429.
Hanley JA, Negassa A, Edwardes MD, Forrester JE . Statistical analysis of correlated data using generalized estimating equations: an orientation. Am J Epidemiol 2003; 157: 364–375.
International HapMap Consortium. The International HapMap Project. Nature 2003; 426: 789–796.
International HapMap Consortium. A haplotype map of the human genome. Nature 2005; 437: 1299–1320.
International HapMap Consortium International HapMap Consortium Altshuler DM, International HapMap Consortium Gibbs RA, International HapMap Consortium Peltonen L, International HapMap Consortium Altshuler DM, International HapMap Consortium Gibbs RA et al. Integrating common and rare genetic variation in diverse human populations. Nature 2010; 467: 52–58.
The 1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 2010; 467: 1061–1073.
The 1000 Genomes Project Consortium. An integrated map of genetic variation from 1,092 human genomes. Nature 2012; 491: 56–65.
Kent WJ, Sugnet CW, Furey TS, Roskin KM, Pringle TH, Zahler AM et al. The Human Genome Browser at UCSC. Genome Res 2002; 12: 996–1006.
Arizona Center for Education and Research on Therapeutics QTDrugs Lists, available at https://www.crediblemeds.org/ (accessed 17 November 2014).
Satterthwaite FE . An approximate distribution of estimates of variance components. Biometrics 1946; 2: 110–114.
Willer CJ, Li Y, Abecasis GR . METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 2010; 26: 2190–2191.
Ganesh SK, Zakai NA, van Rooij FJ, Soranzo N, Smith AV, Nalls MA et al. Multiple loci influence erythrocyte phenotypes in the CHARGE Consortium. Nat Genet 2009; 41: 1191–1198.
Nalls MA, Couper DJ, Tanaka T, van Rooij FJ, Chen MH, Smith AV et al. Multiple loci are associated with white blood cell phenotypes. PLoS Genet 2011; 7: e1002113.
Kent WJ, Sugnet CW, Furey TS, Roskin KM, Pringle TH, Zahler AM et al. The human genome browser at UCSC. Genome Res 2002; 12: 996–1006.
Welter D, MacArthur J, Morales J, Burdett T, Hall P, Junkins H et al. The NHGRI GWAS Catalog, a curated resource of SNP-trait associations. Nucleic Acids Res 2014; 42 (Database issue): D1001–D1006.
Ward LD, Kellis M . HaploReg: a resource for exploring chromatin states, conservation, and regulatory motif alterations within sets of genetically linked variants. Nucleic Acids Res 2012; 40 (Database issue): D930–D934.
Zhang X, Gierman HJ, Levy D, Plump A, Dobrin R, Goring HH et al. Synthesis of 53 tissue and cell line expression QTL datasets reveals master eQTLs. BMC Genomics 2014; 15: 532.
Ramos E, Doumatey A, Elkahloun AG, Shriner D, Huang H, Chen G et al. Pharmacogenomics, ancestry and clinical decision making for global populations. Pharmacogenomics J 2013; 14: 217–222.
Thomas D . Gene–environment-wide association studies: emerging approaches. Nat Rev Genetics 2010; 11: 259–272.
Morris AP . Transethnic meta-analysis of genomewide association studies. Genet Epidemiol 2011; 35: 809–822.
Becker ML, Aarnoudse AJ, Newton-Cheh C, Hofman A, Witteman JC, Uitterlinden AG et al. Common variation in the NOS1AP gene is associated with reduced glucose-lowering effect and with increased mortality in users of sulfonylurea. Pharmacogenet Genomics 2008; 18: 591–597.
Pearson ER, Donnelly LA, Kimber C, Whitley A, Doney AS, McCarthy MI et al. Variation in TCF7L2 influences therapeutic response to sulfonylureas: a GoDARTs study. Diabetes 2007; 56: 2178–2182.
Holstein A, Hahn M, Korner A, Stumvoll M, Kovacs P . TCF7L2 and therapeutic response to sulfonylureas in patients with type 2 diabetes. BMC Med Genet 2011; 12: 30.
Holstein A, Hahn M, Stumvoll M, Kovacs P . The E23K variant of KCNJ11 and the risk for severe sulfonylurea-induced hypoglycemia in patients with type 2 diabetes. Horm Metab Res 2009; 41: 387–390.
GTEx Consortium. Human genomics. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 2015; 348: 648–660.
Koopmann TT, Adriaens ME, Moerland PD, Marsman RF, Westerveld ML, Lal S et al. Genome-wide identification of expression quantitative trait loci (eQTLs) in human heart. PLoS One 2014; 9: e97380.
Abecasis GR, Auton A, Brooks LD, DePristo MA, Durbin RM, Handsaker RE et al. An integrated map of genetic variation from 1,092 human genomes. Nature 2012; 491: 56–65.
Sotoodehnia N, Isaacs A, de Bakker PI, Dorr M, Newton-Cheh C, Nolte IM et al. Common variants in 22 loci are associated with QRS duration and cardiac ventricular conduction. Nat Genet 2010; 42: 1068–1076.
Kirsten H, Al-Hasani H, Holdt L, Gross A, Beutner F, Krohn K et al. Dissecting the genetics of the human transcriptome identifies novel trait-related trans-eQTLs and corroborates the regulatory relevance of non-protein coding locidagger. Hum Mol Genet 2015; 24: 4746–4763.
Hao K, Bosse Y, Nickle DC, Pare PD, Postma DS, Laviolette M et al. Lung eQTLs to help reveal the molecular underpinnings of asthma. PLoS Genet 2012; 8: e1003029.
Westra HJ, Peters MJ, Esko T, Yaghootkar H, Schurmann C, Kettunen J et al. Systematic identification of trans eQTLs as putative drivers of known disease associations. Nat Genet 2013; 45: 1238–1243.
Larson NB, McDonnell S, French AJ, Fogarty Z, Cheville J, Middha S et al. Comprehensively evaluating cis-regulatory variation in the human prostate transcriptome by using gene-level allele-specific expression. Am J Hum Genet 2015; 96: 869–882.
Olalla L, Gutierrez A, Campos JA, Khan ZU, Alonso FJ, Segura JA et al. Nuclear localization of L-type glutaminase in mammalian brain. J Biol Chem 2002; 277: 38939–38944.
Slavin TP, Feng T, Schnell A, Zhu X, Elston RC . Two-marker association tests yield new disease associations for coronary artery disease and hypertension. Hum Genet 2011; 130: 725–733.
Teumer A, Holtfreter B, Volker U, Petersmann A, Nauck M, Biffar R et al. Genome-wide association study of chronic periodontitis in a general German population. J Clin Periodontol 2013; 40: 977–985.
Kraev A, Quednau BD, Leach S, Li XF, Dong H, Winkfein R et al. Molecular cloning of a third member of the potassium-dependent sodium-calcium exchanger gene family, NCKX3. J Biol Chem 2001; 276: 23161–23172.
Schumacher MA, Rivard AF, Bachinger HP, Adelman JP . Structure of the gating domain of a Ca2+-activated K+ channel complexed with Ca2+/calmodulin. Nature 2001; 410: 1120–1124.
Van Booven D, Marsh S, McLeod H, Carrillo MW, Sangkuhl K, Klein TE et al. Cytochrome P450 2C9-CYP2C9. Pharmacogenet Genomics 2010; 20: 277–281.
Tornio A, Niemi M, Neuvonen PJ, Backman JT . Drug interactions with oral antidiabetic agents: pharmacokinetic mechanisms and clinical implications. Trends Pharmacol Sci 2012; 33: 312–322.
Chung WH, Chang WC, Lee YS, Wu YY, Yang CH, Ho HC et al. Genetic variants associated with phenytoin-related severe cutaneous adverse reactions. JAMA 2014; 312: 525–534.
Jorgensen AL, FitzGerald RJ, Oyee J, Pirmohamed M, Williamson PR . Influence of CYP2C9 and VKORC1 on patient response to warfarin: a systematic review and meta-analysis. PLoS One 2012; 7: e44064.
Yang J, Chen Y, Li X, Wei X, Chen X, Zhang L et al. Influence of CYP2C9 and VKORC1 genotypes on the risk of hemorrhagic complications in warfarin-treated patients: a systematic review and meta-analysis. Int J Cardiol 2013; 168: 4234–4243.
Saldana-Cruz AM, Leon-Moreno LC, Sanchez-Corona J, Marquez-de Santiago DA, Mendoza-Carrera F, Castro-Martinez XH et al. CYP2C9 and CYP2C19 allele and haplotype distributions in four Mestizo populations from Western Mexico: an Interethnic Comparative Study. Genet Test Mol Biomarkers 2016; 20: 702–709.
Claudio-Campos K, Duconge J, Cadilla CL, Ruano G . Pharmacogenetics of drug-metabolizing enzymes in US Hispanics. Drug Metab Pers Ther 2015; 30: 87–105.
Valentin II, Rivera G, Nieves-Plaza M, Cruz I, Renta JY, Cadilla CL et al. Pharmacogenetic association study of warfarin safety endpoints in Puerto Ricans. P R Health Sci J 2014; 33: 97–104.
Klen J, Dolzan V, Janez A . CYP2C9, KCNJ11 and ABCC8 polymorphisms and the response to sulphonylurea treatment in type 2 diabetes patients. Eur J Clin Pharmacol 2014; 70: 421–428.
Gaita F, Giustetto C, Bianchi F, Wolpert C, Schimpf R, Riccardi R et al. Short QT Syndrome: a familial cause of sudden death. Circulation 2003; 108: 965–970.
Wolpert C, Schimpf R, Veltmann C, Giustetto C, Gaita F, Borggrefe M . Clinical characteristics and treatment of short QT syndrome. Expert Rev Cardiovasc Ther 2005; 3: 611–617.
Iribarren C, Round AD, Peng JA, Lu M, Klatsky AL, Zaroff JG et al. Short QT in a cohort of 1.7 million persons: prevalence, correlates, and prognosis. Ann Noninvasive Electrocardiol 2014; 19: 490–500.
Holbrook M, Malik M, Shah RR, Valentin JP . Drug induced shortening of the QT/QTc interval: an emerging safety issue warranting further modelling and evaluation in drug research and development? J Pharmacol Toxicol Methods 2009; 59: 21–28.
Ioannidis JP, Tarone R, McLaughlin JK . The false-positive to false-negative ratio in epidemiologic studies. Epidemiology 2011; 22: 450–456.
Aslibekyan S, Claas SA, Arnett DK . To replicate or not to replicate: the case of pharmacogenetic studies: establishing validity of pharmacogenomic findings: from replication to triangulation. Circ Cardiovasc Genet 2013; 6: 409–412, discussion 412.
Psaty BM, Lee M, Savage PJ, Rutan GH, German PS, Lyles M . Assessing the use of medications in the elderly: methods and initial experience in the Cardiovascular Health Study. The Cardiovascular Health Study Collaborative Research Group. J Clin Epidemiol 1992; 45: 683–692.
Sudlow C, Gallacher J, Allen N, Beral V, Burton P, Danesh J et al. UK biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med 2015; 12: e1001779.
Gaziano JM, Concato J, Brophy M, Fiore L, Pyarajan S, Breeling J et al. Million Veteran Program: a mega-biobank to study genetic influences on health and disease. J Clin Epidemiol 2016; 70: 214–223.
Acknowledgements
Age, Gene/Environment Susceptibility – Reykjavik Study (AGES): This study has been funded by NIH contracts N01-AG-1-2100 and 271201200022C, the NIA Intramural Research Program, Hjartavernd (the Icelandic Heart Association) and the Althingi (the Icelandic Parliament). The study is approved by the Icelandic National Bioethics Committee, VSN: 00-063. The researchers are indebted to the participants for their willingness to participate in the study.
Atherosclerosis Risk in Communities (ARIC): The Atherosclerosis Risk in Communities Study is carried out as a collaborative study supported by National Heart, Lung and Blood Institute Contracts (HHSN268201100005C, HHSN268201100006C, HHSN268201100007C, HHSN268201100008C, HHSN268201100009C, HHSN268201100010C, HHSN268201100011C and HHSN268201100012C), R01HL087641, R01HL59367 and R01HL086694; National Human Genome Research Institute Contract U01HG004402; and National Institutes of Health Contract HHSN268200625226C. We thank the staff and participants of the ARIC study for their important contributions. Infrastructure was partly supported by Grant No. UL1RR025005, a component of the National Institutes of Health and NIH Roadmap for Medical Research.
Cardiovascular Health Study (CHS): This CHS research was supported by NHLBI contracts HHSN268201200036C, HHSN268200800007C, N01HC55222, N01HC85079, N01HC85080, N01HC85081, N01HC85082, N01HC85083 and N01HC85086; and NHLBI Grants U01HL080295, R01HL087652, R01HL105756, R01HL103612, R01HL120393 and R01HL085251 with additional contribution from the National Institute of Neurological Disorders and Stroke (NINDS). Additional support was provided through R01AG023629 from the National Institute on Aging (NIA). A full list of principal CHS investigators and institutions can be found at CHS-NHLBI.org. The provision of genotyping data was supported in part by the National Center for Advancing Translational Sciences, CTSI Grant UL1TR000124 and the National Institute of Diabetes and Digestive and Kidney Disease Diabetes Research Center (DRC) Grant DK063491 to the Southern California Diabetes Endocrinology Research Center. NS was supported by R01HL116747 and RO1HL111089. JSF was supported by K08HL116640.
Health, Aging, and Body Composition (Health ABC): This research was supported by NIA Contracts N01AG62101, N01AG62103 and N01AG62106. The genome-wide association study was funded by NIA Grant 1R01AG032098-01A1 to Wake Forest University Health Sciences and genotyping services were provided by the Center for Inherited Disease Research (CIDR). CIDR is fully funded through a federal contract from the National Institutes of Health to The Johns Hopkins University, Contract No. HHSN268200782096C. This research was supported in part by the Intramural Research Program of the NIH, National Institute on Aging.
Hispanic Community Health Study/Study of Latinos (HCHS/SOL): We thank the participants and staff of the HCHS/SOL study for their contributions to this study. The baseline examination of HCHS/SOL was carried out as a collaborative study supported by contracts from the National Heart, Lung, and Blood Institute (NHLBI) to the University of North Carolina (N01-HC65233), University of Miami (N01-HC65234), Albert Einstein College of Medicine (N01-HC65235), Northwestern University (N01-HC65236) and San Diego State University (N01-HC65237). The following Institutes/Centers/Offices contributed to the first phase of HCHS/SOL through a transfer of funds to the NHLBI: National Institute on Minority Health and Health Disparities, National Institute on Deafness and Other Communication Disorders, National Institute of Dental and Craniofacial Research (NIDCR), National Institute of Diabetes and Digestive and Kidney Diseases, National Institute of Neurological Disorders and Stroke and NIH Institution-Office of Dietary Supplements. The Genetic Analysis Center at University of Washington was supported by NHLBI and NIDCR contracts (HHSN268201300005C AM03 and MOD03). Genotyping efforts were supported by NHLBI HSN 26220/20054C, NCATS CTSI Grant UL1TR000124, and NIDDK Diabetes Research Center (DRC) Grant DK063491.
Jackson Heart Study (JHS): We thank the Jackson Heart Study (JHS) participants and staff for their contributions to this work. The JHS is supported by contracts HHSN268201300046C, HHSN268201300047C, HSN268201300048C, HHSN268201300049C and HHSN268201300050C from the National Heart, Lung, and Blood Institute and the National Institute on Minority Health and Health Disparities.
Multi-Ethnic Study of Atherosclerosis (MESA): MESA and MESA SNP Health Association Resource (SHARe) are conducted and supported by the National Heart, Lung and Blood Institute (NHLBI) in collaboration with MESA investigators. Support is provided by grants and contracts N01 HC-95159, N01-HC-95160, N01-HC-95161, N01-HC-95162, N01-HC-95163, N01-HC-95164, N01-HC-95165, N01-HC-95166, N01-HC-95167, N01-HC-95168, N01-HC-95169 and RR-024156. Additional funding was supported in part by the Clinical Translational Science Institute Grant UL1RR033176 and is now at the National Center for Advancing Translational Sciences, CTSI Grant UL1TR000124. We also thank the other investigators, the staff and the participants of the MESA study for their valuable contributions. A full list of participating MESA investigators and institutions can be found at http://www.mesa-nhlbi.org.
Netherlands Epidemiology of Obesity (NEO): The authors of the NEO study thank all individuals who participated in the Netherlands Epidemiology in Obesity study, all participating general practitioners for inviting eligible participants and all research nurses for collection of the data. We thank the NEO study group, Pat van Beelen, Petra Noordijk and Ingeborg de Jonge for the coordination, lab and data management of the NEO study. The genotyping in the NEO study was supported by the Centre National de Génotypage (Paris, France), headed by Jean-Francois Deleuze. The NEO study is supported by the participating Departments, the Division and the Board of Directors of the Leiden University Medical Center and by the Leiden University, Research Profile Area Vascular and Regenerative Medicine. Dennis Mook-Kanamori is supported by Dutch Science Organization (ZonMW-VENI Grant 916.14.023).
Prospective Study of Pravastatin in the Elderly at Risk (PROSPER): The PROSPER study was supported by an investigator-initiated grant obtained from Bristol-Myers Squibb. Professor Dr J W Jukema is an Established Clinical Investigator of the Netherlands Heart Foundation (Grant No. 2001 D 032). Support for genotyping was provided by the seventh framework program of the European commission (Grant No. 223004) and by the Netherlands Genomics Initiative (Netherlands Consortium for Healthy Aging Grant 050-060-810).
Rotterdam Study (RS): The RS is supported by the Erasmus Medical Center and Erasmus University Rotterdam; The Netherlands Organization for Scientific Research; The Netherlands Organization for Health Research and Development (ZonMw); the Research Institute for Diseases in the Elderly; The Netherlands Heart Foundation; the Ministry of Education, Culture and Science; the Ministry of Health Welfare and Sports; the European Commission; and the Municipality of Rotterdam. Support for genotyping was provided by The Netherlands Organization for Scientific Research (NWO) (175.010.2005.011, 911.03.012) and Research Institute for Diseases in the Elderly (RIDE). This study was supported by The Netherlands Genomics Initiative (NGI)/Netherlands Organization for Scientific Research (NWO) Project No. 050-060-810. This collaborative effort was supported by an award from the National Heart, Lung and Blood Institute (R01-HL-103612, PI BMP).
Women’s Health Initiative Clinical Trial (WHI CT): The Women’s Health Initiative clinical trials were funded by the National Heart, Lung, and Blood Institute, National Institutes of Health, US Department of Health and Human Services through Contracts HHSN268201100046C, HHSN268201100001C, HHSN268201100002C, HHSN268201100003C, HHSN268201100004C and HHSN271201100004C. All contributors to WHI science are listed at https://www.whi.org/researchers/Documents%20%20Write%20a%20Paper/WHI%20Investigator%20Long%20List.pdf. ELB was supported in part by a grant from the National Cancer Institute (5T32CA009001). WHI GARNET: Within the Genomics and Randomized Trials Network, a GWAS of Hormone Treatment and CVD and Metabolic Outcomes in the WHI was funded by the National Human Genome Research Institute, National Institutes of Health, US Department of Health and Human Services through cooperative agreement U01HG005152 (Reiner). All contributors to GARNET science are listed at https://www.genome.gov/27541119/genomics-and-randomized-trials-network-garnet/: The Modification of PM-Mediated Arrhythmogenesis in Populations was funded by the National Institute of Environmental Health Sciences, National Institutes of Health, US Department of Health and Human Services through Grant R01ES017794 (Whitsel). WHI SHARe: The SNP Health Association Resource project was funded by the National Heart, Lung and Blood Institute, National Institutes of Health, US Department of Health and Human Services through contract N02HL64278 (Kooperberg). WHI WHIMS: The Women's Health Initiative Memory Study (WHIMS+) Genome-Wide Association Study was funded by the National Heart, Lung, and Blood Institute, National Institutes of Health, US Department of Health and Human Services through Contract HHSN268201100046C (Anderson).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
BMP serves on the DSMB of a clinical trial of a device funded by the manufacturer (Zoll LifeCor) and on the Steering Committee of the Yale Open Data Access Project funded by Johnson & Johnson. The other authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the The Pharmacogenomics Journal website
Supplementary information
PowerPoint slides
Rights and permissions
About this article
Cite this article
Floyd, J., Sitlani, C., Avery, C. et al. Large-scale pharmacogenomic study of sulfonylureas and the QT, JT and QRS intervals: CHARGE Pharmacogenomics Working Group. Pharmacogenomics J 18, 127–135 (2018). https://doi.org/10.1038/tpj.2016.90
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/tpj.2016.90
This article is cited by
-
Genome-wide association study identifies novel risk variants from RPS6KA1, CADPS, VARS, and DHX58 for fasting plasma glucose in Arab population
Scientific Reports (2020)
-
Objectives, design and main findings until 2020 from the Rotterdam Study
European Journal of Epidemiology (2020)
-
Genomic approaches for the elucidation of genes and gene networks underlying cardiovascular traits
Biophysical Reviews (2018)
-
The Rotterdam Study: 2018 update on objectives, design and main results
European Journal of Epidemiology (2017)