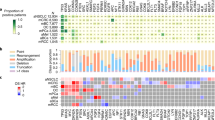
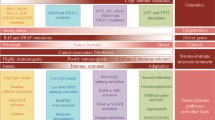
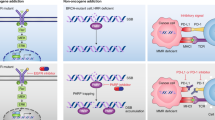
Abstract
The goal of cancer pharmacogenomics is to obtain benefit from personalized approaches of cancer treatment and prevention. Recent advances in genomic research have shed light on the crucial role of genetic variants, mainly involving genes encoding drug-metabolizing enzymes, drug transporters and targets, in driving different treatment responses among individuals, in terms of therapeutic efficacy and safety. Although a considerable amount of new targeted agents have been designed based on a finely understanding of molecular alterations in cancer, a wide gap between pharmacogenomic knowledge and clinical application still persists. This review focuses on the relevance of mutational analyses in predicting individual response to antitumor therapy, in order to improve the translational impact of genetic information on clinical practice.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 print issues and online access
$259.00 per year
only $43.17 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Meyer UA . Pharmacogenetics 5 decades of therapeutic lessons from genetic diversity. Nat Rev Genet 2004; 6: 669–676.
Evans WE, Relling MV . Pharmacogenomics: translating functional genomics into rational therapeutics. Science 1999; 286: 487–491.
McDermott U, Downing JT, Stratton M . Genomics and the continuum of cancer care. N Engl J Med 2011; 36: 340–350.
Stratton MR, Campbell PJ, Futreal PA . The cancer genome. Nature 2009; 458: 719–724.
Pleasance ED, Cheetham RK, Stephens PJ, McBride DJ, Humphray SJ, Greenman CD et al. A comprehensive catalogue of somatic mutations from a human cancer genome. Nature 2010; 463: 191–196.
Huang YT, Heist RS, Chirieac LR, Lin X, Skaug V, Zienolddiny S et al. Genome-wide analysis of survival in early-stage non-small- cell lung cancer. J Clin Oncol 2009; 27: 2660–2667.
Zhang Y, Martens JW, Yu JX, Jiang J, Sieuwerts AM, Smid M et al. Copy number alterations that predict metastatic capability of human breast cancer. Cancer Res 2009; 69: 3795–3801.
Normanno N, De Luca A, Bianco C, Strizzi L, Mancino M, Maiello MR et al. Epidermal growth factor receptor (EGFR) signaling in cancer. Gene 2006; 366: 2–16.
Raymond E, Faivre S, Armand JP . Epidermal growth factor receptor tyrosine kinase as a target for anticancer therapy. Drugs 2000; 60 (suppl 1): 15–23.
Jorissen RN, Walker F, Pouliot N, Garrett TP, Ward CW, Burgess AW . Epidermal growth factor receptor: mechanisms of activation and signalling. Exp Cell Res 2003; 284: 31–53.
Krause DS, Van Etten RA . Tyrosine kinases as targets for cancer therapy. N Engl J Med 2005; 353: 172–187.
De Witt Hamer PC . Small molecule kinase inhibitors in glioblastoma: a systematic review of clinical studies. Neuro Oncol 2010; 12: 304–316.
Campone M, Berton-Rigaud D, Bourbouloux E, Sophie S, Zanetti A, Frenel JS . Her2 positive breast cancer: practices. Bull Cancer 2011; 98: 154–163.
De Vita F, Giuliani F, Silvestris N, Catalano G, Ciardiello F, Orditura M . Human epidermal growth factor receptor 2 (HER2) in gastric cancer: a new therapeutic target. Cancer Treat Rev 2010; 36 (Suppl 3): 11–15.
Bass A . Impact of KRAS and BRAF gene mutations on targeted therapies in colorectal cancer. J Clin Oncol 2011; 29: 2728–2729.
Sigalotti L, Fratta E, Parisi G, Coral S, Maio M . Stability of BRAF V600E mutation in metastatic melanoma: new insights for therapeutic success? Br J Cancer 2011; 105: 327–328.
Cleary JM, Shapiro GI . Development of phosphoinositide-3 kinase pathway inhibitors for advanced cancer. Curr Oncol Rep 2010; 12: 87–94.
Klümpen HJ, Beijnen JH, Gurney H, Schellens JH . Inhibitors of mTOR. Oncologist 2010; 15: 1262–1269.
Dahabreh IJ, Linardou H, Siannis F, Kosmidis P, Bafaloukos D, Murray S . Somatic EGFR mutation and gene copy gain as predictive biomarkers for response to tyrosine kinase inhibitors in non-small cell lung cancer. Clin Cancer Res 2010; 16: 291–303.
Pillai RN, Ramalingam SS . The biology and clinical features of non-small cell lung cancers with EML4-ALK translocation. Curr Oncol Rep 2012; 14: 105–110.
Wardelmann E, Büttner R, Merkelbach-Bruse S, Schildhaus HU . Mutational analysis of gastrointestinal stromal tumors: increasing significance for risk assessment and effective targeted therapy. Virchows Arch 2007; 451: 743–749.
Plesac TP, Hunt JL . KRAS mutation testing in colorectal cancer. Adv Anat Pathol 2009; 16: 196–203.
Allegra CJ, Jessup JM, Somerfield MR, Hamilton SR, Hammond EH, Hayes DF et al. American Society of Clinical Oncology provisional clinical opinion: testing for KRAS gene mutations in patients with metastatic colorectal carcinoma to predict response to anti-epidermal growth factor receptor monoclonal antibody therapy. J Clin Oncol 2009; 27: 2091–2096.
Vultur A, Villanueva J, Herlyn M . Targeting BRAF in advanced melanoma: a first step toward manageable disease. Clin Cancer Res 2011; 17: 1658–1663.
Sim SC, Ingelman-Sundberg M . Pharmacogenomic biomarkers: new tools in current and future drug therapy. Trends Pharmacol Sci 2011; 32: 72–81.
Relling MV, Altman RB, Goetz MP, Evans WE . Clinical implementation of pharmacogenomics: overcoming genetic exceptionalism. Lancet Oncol 2010; 11: 507–509.
Goldstraw P, Ball D, Jett JR, Le Chevalier T, Lim E, Nicholson AG et al. Non-small-cell lung cancer. Lancet 2011; 378: 1727–1740.
Sangha R, Price J, Buttes CA . Adjuvant therapy in non-small cell lung cancer: current and future directions. Oncologist 2010; 15: 862–872.
Ohe Y, Ohashi Y, Kubota K, Tamura T, Nakagawa K, Negoro S et al. Randomized phase III study of cisplatin plus irinotecan versus carboplatin plus paclitaxel, cisplatin plus gemcitabine, and cisplatin plus vinorelbine for ad- vanced non-small-cell lung cancer: Four- Arm Cooperative Study in Japan. Ann Oncol 2007; 18: 317–323.
Cataldo VD, Gibbons DL, Pérez-Soler R, Quintás-Cardama A . Treatment of non-small-cell lung cancer with erlotinib or gefitinib. N Engl J Med 2011; 364: 947–955.
Shepherd FA, Rodrigues Pereira J, Ciuleanu T, Tan EH, Hirsh V, Thongprasert S et al. Erlotinib in previously treated non–small-cell lung cancer. N Engl J Med 2005; 353: 123–132.
Kim ES, Hirsh V, Mok T, Socinski MA, Gervais R, Wu YL et al. Gefitinib versus docetaxel in previously treated non-small-cell lung cancer (INTEREST): a randomised phase III trial. Lancet 2008; 372: 1809–1818.
Oxnard GR, Miller VA . Use of erlotinib or gefitinib as initial therapy in advanced NSCLC. Oncology 2010; 24: 392–399.
Rosell R, Gervais R, Vergnenegre A, Massuti B, Felip E, Cardenal F et al. Erlotinib versus chemotherapy (CT) in advanced non-small cell lung cancer (NSCLC) patients (p) with epidermal growth factor receptor (EGFR) mutations: Interim results of the European Erlotinib Versus Chemotherapy (EURTAC) phase III randomized trial. J Clin Oncol 2011; 29, abstr 7503. Presented at: 47th Annual Meeting of the American Society of Clinical Oncology. Chicago, IL, USA. 3–7 June 2011.
Pao W, Miller V, Zakowski M, Doherty J, Politi K, Sarkaria I et al. EGF receptor gene mutations are common in lung cancers from ‘never smokers’ and are associated with sensitivity of tumors to gefitinib and erlotinib. Proc Natl Acad Sci USA 2004; 101: 13306–13311.
Paez JG, Jänne PA, Lee JC, Tracy S, Greulich H, Gabriel S et al. EGFR mutations in lung cancer: correlation with clinical response to gefitinib therapy. Science 2004; 304: 1497–1500.
Lynch TJ, Bell DW, Sordella R, Gurubhagavatula S, Okimoto RA, Brannigan BW et al. Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N Engl J Med 2004; 350: 2129–2139.
Riely GJ, Politi KA, Miller VA, Pao W . Update on EGFR mutations in non-small cell lung cancer. Clin Cancer Res 2006; 12: 7232–7241.
Toschi L, Cappuzzo F . Understanding the new genetics of responsiveness to epidermal growth factor receptor tyrosine kinase inhibitors. Oncologist 2007; 12: 211–220.
Linardou H, Dahabreh IJ, Bafaloukos D, Kosmidis P, Murray S . Somatic EGFR mutations and efficacy of tyrosine kinase inhibitors in NSCLC. Nat Rev Clin Oncol 2009; 6: 352–366.
Thatcher N, Chang A, Parikh P, Rodrigues Pereira J, Ciuleanu T, von Pawel J et al. Gefitinib plus best supportive care in previously treated patients with refractory advanced non-small-cell lung cancer: results from a randomised, placebo-controlled, multicentre study (Iressa Survival Evaluation in Lung Cancer). Lancet 2005; 366: 1527–1537.
Bell DW, Brannigan BW, Matsuo K, Finkelstein DM, Sordella R, Settleman J et al. Increased prevalence of EGFR-mutant lung cancer in women and in East Asian populations: analysis of estrogen-related polymorphisms. Clin Cancer Res 2008; 14: 4079–4084.
Pao W, Miller VA, Politi KA, Riely GJ, Somwar R, Zakowski MF et al. Acquired resistance of lung adenocarcinomas to gefitinib or erlotinib is associated with a second mutation in the EGFR kinase domain. PLoS Med 2005; 2: e73.
Yun CH, Mengwasser KE, Toms AV, Woo MS, Greulich H, Wong KK et al. The T790M mutation in EGFR kinase causes drug resistance by increasing the affinity for ATP. Proc Natl Acad Sci USA 2008; 105: 2071.
Bell DW, Gore I, Okimoto RA, Godin-Heymann N, Sordella R, Mulloy R et al. Inherited susceptibility to lung cancer may be associated with the T790M drug resistance mutation in EGFR. Nat Genet 2005; 37: 1315–1316.
Maheswaran S, Sequist LV, Nagrath S, Ulkus L, Brannigan B, Collura CV et al. Detection of mutations in EGFR in circulating lung-cancer cells. N Engl J Med 2008; 359: 366–377.
Marchetti A, Normanno N, Pinto C, Taddei GL, Adamo V, Ardizzoni A et al. Recommendations for mutational analysis of EGFR in lung carcinoma. Pathologica 2010; 102: 119–126.
Bean J, Brennan C, Shih JY, Riely G, Viale A, Wang L et al. MET amplification occurs with or without T790M mutations in EGFR mutant lung tumors with acquired resistance to gefitinib or erlotinib. Proc Natl Acad Sci USA 2007; 104: 20932–20937.
Giovannetti E, Zucali PA, Peters GJ, Cortesi F, D’Incecco A, Smit EF et al. Association of polymorphisms in AKT1 and EGFR with clinical outcome and toxicity in non-small cell lung cancer patients treated with gefitinib. Mol Cancer Ther 2010; 9: 581–593.
Morris SW, Kirstein MN, Valentine MB, Dittmer KG, Shapiro DN, Saltman DL et al. Fusion of a kinase gene, ALK, to a nucleolar protein gene, NPM, in non-Hodgkin's lymphoma. Science 1994; 263: 1281–1284.
Palmer RH, Vernersson E, Grabbe C, Hallberg B . Anaplastic lymphoma kinase: signalling in development and disease. Biochem J 2009; 420: 345–361.
Soda M, Choi YL, Enomoto M, Takada S, Yamashita Y, Ishikawa S et al. Identification of the transforming EML4-ALK fusion gene in nonsmall-cell lung cancer. Nature 2007; 448: 561–566.
Inamura K, Takeuchi K, Togashi Y, Nomura K, Ninomiya H, Okui M et al. EML4-ALK fusion is linked to histological characteristics in a subset of lung cancers. J Thorac Oncol 2008; 3: 13–17.
Inamura K, Takeuchi K, Togashi Y, Hatano S, Ninomiya H, Motoi N et al. EML4-ALK lung cancers are characterized by rare other mutations, a TTF-1 cell lineage, an acinar histology, and young onset. Mod Pathol 2009; 22: 508–515.
Koivunen JP, Mermel C, Zejnullahu K, Murphy C, Lifshits E, Holmes AJ et al. EML4-ALK fusion gene and efficacy of an ALK kinase inhibitor in lung cancer. Clin Cancer Res 2008; 14: 4275–4283.
Shinmura K, Kageyama S, Tao H, Bunai T, Suzuki M, Kamo T et al. EML4-ALK fusion transcripts, but no NPM-, TPM3-, CLTC-,ATIC-, or TFG-ALK fusion transcripts, in non-small cell lung carcinomas. Lung Cancer 2008; 61: 163–169.
Martelli MP, Sozzi G, Hernandez L, Pettirossi V, Navarro A, Conte D et al. EML4-ALK rearrangement in non-small cell lung cancer and non-tumor lung tissues. Am J Pathol 2009; 174: 661–670.
Shaw AT, Yeap BY, Mino-Kenudson M, Digumarthy SR, Costa DB, Heist RS et al. Clinical features and outcome of patients with nonsmall-cell lung cancer who harbor EML4-ALK. J Clin Oncol 2009; 27: 4247–4253.
Wong DW, Leung EL, So KK, Tam IY, Sihoe AD, Cheng LC et al. The EML4-ALK fusion gene is involved in various histologic types of lung cancers from nonsmokers with wild-type EGFR and KRAS. Cancer 2009; 115: 1723–1733.
Mano H . Non-solid oncogenes in solid tumors: EML4-ALK fusion genes in lung cancer. Cancer Sci 2008; 99: 2349–2355.
Takeuchi K, Choi YL, Togashi Y, Soda M, Hatano S, Inamura K et al. KIF5B-ALK, a novel fusion oncokinase identified by an immunohistochemistry-based diagnostic system for ALK-positive lung cancer. Clin Cancer Res 2009; 15: 3143–3149.
Soda M, Takada S, Takeuchi K, Choi YL, Enomoto M, Ueno T et al. A mouse model for EML4-ALK-positive lung cancer. Proc Natl Acad Sci USA 2008; 105: 19893–19897.
Kwak EL, Bang YJ, Camidge DR, Shaw AT, Solomon B, Maki RG et al. Anaplastic lymphoma kinase inhibition in non-small-cell lung cancer. N Engl J Med 2010; 363: 1693–1703.
Shaw AT, Yeap BY, Solomon BJ, Riely GJ, Gainor J, Engelman JA et al. Effect of crizotinib on overall survival in patients with advanced non-small-cell lung cancer harbouring ALK gene rearrangement: a retrospective analysis. Lancet Oncol 2011; 12: 1004–1012.
Choi YL, Soda M, Yamashita Y, Ueno T, Takashima J, Nakajima T et al. ALK Lung Cancer Study Group EML4-ALK mutations in lung cancer that confer resistance to ALK inhibitors. N Engl J Med 2010; 363: 1734–1739.
Sasaki T, Okuda K, Zheng W, Butrynski J, Capelletti M, Wang L et al. The neuroblastoma associated F1174L ALK mutation causes resistance to an ALK kinase inhibitor in ALK translocated cancers. Cancer Res 2010; 70: 10038–10043.
Choi YL, Takeuchi K, Soda M, Inamura K, Togashi Y, Hatano S et al. Identification of novel isoforms of the EML4-ALK transforming gene in non-small cell lung cancer. Cancer Res 2008; 68: 4971–4976.
Zhang X, Zhang S, Yang X, Yang J, Zhou Q, Yin L et al. Fusion of EML4 and ALK is associated with development of lung adenocarcinomas lacking EGFR and KRAS mutations and is correlated with ALK expression. Mol Cancer 2010; 9: 188.
Goldstein NS, Armin M . Epidermal growth factor receptor immunohistochemical reactivity in patients with American Joint Committee on Cancer Stage IV colon adenocarcinoma: implications for a standardized scoring system. Cancer 2001; 92: 1331–1346.
Graham J, Muhsin M, Kirkpatrick P . Cetuximab. Nat Rev Drug Discov 2004; 3: 549–550.
Cunningham D, Humblet Y, Siena S, Khayat D, Bleiberg H, Santoro A et al. Cetuximab monotherapy and cetuximab plus irinotecan in irinotecan-refractory metastatic colorectal cancer. N Engl J Med 2004; 351: 337–345.
Saltz L, Easley C, Kirkpatrick P . Panitumumab. Nat Rev Drug Discov 2006; 5: 987–988.
Mendelsohn J, Baselga J . Status of epidermal growth factor receptor anatgonists in the biology and treatment of cancer. J Clin Oncol 2003; 21: 2787–2799.
Camp ER, Summy J, Bauer TW, Liu W, Gallick GE, Ellis LM . Molecular mechanisms of resistance to therapies targeting the epidermal growth factor receptor. Clin Cancer Res 2005; 11: 397–405.
Moroni M, Veronese S, Benvenuti S, Marrapese G, Sartore-Bianchi A, Di Nicolantonio F et al. Gene copy number for epidermal growth factor receptor (EGFR) and clinical response to antiEGFR treatment in colorectal cancer: a cohort study. Lancet Oncol 2005; 6: 279–286.
Amado RG, Wolf M, Peeters M, Van Cutsem E, Siena S, Freeman DJ et al. Wild-type KRAS is required for panitumumab efficacy in patients with metastatic colorectal cancer. J Clin Oncol 2008; 26: 1626–1634.
Lièvre A, Bachet JB, Boige V, Cayre A, Le Corre D, Buc E et al. KRAS mutations as an independent prognostic factor in patients with advanced colorectal cancer treated with cetuximab. J Clin Oncol 2008; 26: 374–379.
Malumbres M, Barbacid M . RAS oncogenes: the first 30 years. Nat Rev Cancer 2003; 3: 459–465.
Martinez-Garza SG, Núñez-Salazar A, Calderon-Garcidueñas AL, Bosques-Padilla FJ, Niderhauser-García A, Barrera-Saldaña HA . Frequency and clinicopathology associations of K-ras mutations in colorectal cancer in a Northeast Mexican population. Digest Dis 1999; 17: 225–229.
Urosević N, Krtolica K, Skaro-Milić A, Knezević-Usaj S, Dujić A . Prevalence of G-to-T transversions among K-ras oncogene mutations in human colorectal tumors in Yugoslavia. Int J Cancer 1993; 54: 249–254.
Palmirotta R, Savonarola A, Formica V, Ludovici G, Del Monte G, Roselli M et al. A novel K-ras mutation in colorectal cancer. A case report and literature review. Anticancer Res 2009; 29: 3369–3374.
Palmirotta R, Savonarola A, Ludovici G, Marchis ML, Covello R, Ettorre GM et al. Concurrent mutation in exons 1 and 2 of K-ras oncogene in colorectal cancer. Folia Histochemica Et Cytobiologica 2011; 49: 729–733.
Edkins S, O’Meara S, Parker A, Stevens C, Reis M, Jones S et al. Recurrent KRAS codon 146 mutations in human colorectal cancer. Cancer Biol Ther 2006; 5: 928–932.
Normanno N, Tejpar S, Morgillo F, De Luca A, Van Cutsem E, Ciardiello F . Implications for KRAS status and EGFR-targeted therapies in metastatic CRC. Nat Rev Clin Oncol 2009; 6: 519–527.
Andreyev HJ, Norman AR, Cunningham D, Oates J, Dix BR, Iacopetta BJ et al. Kirsten ras mutations in patients with colorectal cancer: the ‘RASCAL II’ study. Br J Cancer 2001; 85: 692–696.
Neumann J, Zeindl-Eberhart E, Kirchner T, Jung A . Frequency and type of KRAS mutations in routine diagnostic analysis of metastatic colorectal cancer. Pathol Res Pract 2009; 205: 858–862.
Forbes S, Clements J, Dawson E, Bamford S, Webb T, Dogan A et al. Cosmic 2005. Br J Cancer 2006; 94: 318–322.
Van Cutsem E, Köhne CH, Láng I, Folprecht G, Nowacki MP, Cascinu S et al. Cetuximab plus irinotecan, fluorouracil, and leucovorin as first-line treatment for metastatic colorectal cancer: updated analysis of overall survival according to tumor KRAS and BRAF mutation status. J Clin Oncol 2011; 29: 2011–2019.
Bokemeyer C, Bondarenko I, Hartmann JT, de Braud F, Schuch G, Zubel A et al. Efficacy according to biomarker status of cetuximab plus FOLFOX-4 as first-line treatment for metastatic colorectal cancer: the OPUS study. Ann Oncol 2011; 22: 1535–1546.
Douillard JY, Siena S, Cassidy J, Tabernero J, Burkes R, Barugel M et al. Randomized, phase III trial of panitumumab with infusional fluorouracil, leucovorin, and oxaliplatin (FOLFOX4) versus FOLFOX4 alone as first-line treatment in patients with previously untreated metastatic colorectal cancer: the PRIME study. J Clin Oncol 2010; 28: 4697–4705.
Di Nicolantonio F, Martini M, Molinari F, Sartore-Bianchi A, Arena S, Saletti P et al. Wild-type BRAF is required for response to panitumumab or cetuximab in metastatic colorectal cancer. J Clin Oncol 2008; 26: 5705–5712.
Sartore-Bianchi A, Martini M, Molinari F, Veronese S, Nichelatti M, Artale S et al. PIK3CA mutations in colorectal cancer are associated with clinical resistance to EGFR-targeted monoclonal antibodies. Cancer Res 2009; 69: 1851–1857.
Loupakis F, Pollina L, Stasi I, Ruzzo A, Scartozzi M, Santini D et al. PTEN expression and KRAS mutations on primary tumors and metastases in the prediction of benefit from cetuximab plus irinotecan for patients with metastatic colorectal cancer. J Clin Oncol 2009; 27: 2622–2629.
Jacobs B, De Roock W, Piessevaux H, Van Oirbeek R, Biesmans B, De Schutter J et al. Amphiregulin and epiregulin mRNA expression in primary tumors predicts outcome in metastatic colorectal cancer treated with cetuximab. J Clin Oncol 2009; 27: 5068–5074.
Rubin BP, Heinrich MC, Corless CL . Gastrointestinal stromal tumor. Lancet 2007; 369: 1731–1741.
Miettinen M, Lasota J . Gastrointestinal stromal tumors: definition, clinical, histological, immunohistochemical, and molecular genetic features and differential diagnosis. Virchows Arch 2001; 438: 1–12.
Rubin BP, Singer S, Tsao C, Duensing A, Lux ML, Ruiz R et al. KIT activation is a ubiquitous feature of gastrointestinal stromal tumors. Cancer Res 2001; 61: 8118–8121.
Heinrich MC, Corless CL, Duensing A, McGreevey L, Chen CJ, Joseph N et al. PDGFRA activating mutations in gastrointestinal stromal tumors. Science 2003; 299: 708–710.
Antonescu C, Sommer G, Sarran L, Tschernyavsky S, Riedel E, Woodruff J et al. Association of KIT exon 9 mutations with nongastric primary site and aggressive behavior: KIT mutation analysis and clinical correlates of 120 gastrointes- tinal stromal tumors. Clin Cancer Res 2003; 9: 3329–3337.
Kinoshita K, Isozaki K, Hirota S, Nishida T, Chen H, Nakahara M et al. c-Kit gene mutation at exon 17 or 13 is very rare in sporadic gastrointestinal stromal tumors. J Gastroenterol Hepatol 2003; 18: 147.
Gafter-Gvili A, Leader A, Gurion R, Vidal L, Ram R, Shacham-Abulafia A et al. High-dose Imatinib for newly diagnosed chronic phase chronic myeloid leukemia patients-systematic review and meta-analysis. Am J Hematol 2011; 86: 657–662.
Joensuu H, Roberts PJ, Sarlomo-Rikala M, Andersson LC, Tervahartiala P, Tuveson D et al. Effect of the tyrosine kinase inhibitor STI571 in a patient with a metastatic gastrointestinal stromal tumor. N Engl J Med 2001; 344: 1052–1056.
Dematteo RP, Ballman KV, Antonescu CR, Maki RG, Pisters PW, Demetri GD et al. Adjuvant imatinib mesylate after resection of localised, primary gastrointestinal stromal tumor: a randomised, double-blind, placebo-controlled trial. Lancet 2009; 373: 1097–1104.
Eisenberg BL, Trent JC . Adjuvant and neoadjuvant imatinib therapy: current role in the management of GIST. Int J Cancer 2011; 129: 2533–2542.
Debiec-Rychter M, Sciot R, Le Cesne A, Schlemmer M, Hohenberger P, van Oosterom AT et al. KIT mutations and dose selection for imatinib in patients with advanced gastrointestinal stromal tumors. Eur J Cancer 2006; 42: 1093–1103.
Gastrointestinal Stromal Tumor Meta-Analysis Group (Meta-GIST). Comparison of two doses of imatinib for the treatment of unresectable or metastatic gastrointestinal stromal tumors: a meta-analysis of 1,640 patients. J Clin Oncol 2010; 28: 1247–1253.
Demetri GD, van Oosterom AT, Garrett CR, Blackstein ME, Shah MH, Verweij J et al. Efficacy and safety of sunitinib in patients with advanced gastrointestinal stromal tumour after failure of imatinib: a randomised controlled trial. Lancet 2006; 368: 1329–1338.
Corless C, Schroeder A, Griffith D, Town A, McGreevey L, Harrell P et al. PDGFRA mutations in gastrointestinal stromal tumors: frequency, spectrum and in vitro sensitivity to imatinib. J Clin Oncol 2005; 23: 5357–5364.
Wang WL, Conley A, Reynoso D, Nolden L, Lazar AJ, George S et al. Mechanisms of resistance to imatinib and sunitinib in gastrointestinal stromal tumor. Cancer Chemother Pharmacol 2011; 67 (1s): S15–S24.
Martín-Broto J, Rubio L, Alemany R, López-Guerrero JA . Clinical implications of KIT and PDGFRA genotyping in GIST. Clin Transl Oncol 2010; 12: 670–676.
Mouawad R, Sebert M, Michels J, Bloch J, Spano JP, Khayat D . Treatment for metastatic malignant melanoma: old drugs and new strategies. Crit Rev Oncol/Hematol 2010; 74: 27–39.
Kumar R, Angelini S, Snellman E, Hemminki K . BRAF mutations are common somatic events in melanocytic nevi. J Invest Dermatol 2004; 122: 342–348.
Davies H, Bignell GR, Cox C, Stephens P, Edkins S, Clegg S et al. Mutations of the BRAF gene in human cancer. Nature 2002; 417: 949–954.
Dong J, Phelps RG, Qiao R, Yao S, Benard O, Ronai Z et al. BRAF oncogenic mutations correlate with progression rather than initiation of human melanoma. Cancer Res 2003; 63: 3883–3885.
Patton EE, Widlund HR, Kutok JL, Kopani KR, Amatruda JF, Murphey RD et al. BRAF mutations are sufficient to promote nevi formation and cooperate with p53 in the genesis of melanoma. Curr Biol 2005; 15: 249–254.
Dhomen N, Reis-Filho JS, da Rocha Dias S, Hayward R, Savage K, Delmas V et al. Oncogenic braf induces melanocyte senescence and melanoma in mice. Cancer Cell 2009; 15: 294–303.
Wan PT, Garnett MJ, Roe SM, Lee S, Niculescu-Duvaz D, Good VM et al. Mechanism of activation of the RAF-ERK signaling pathway by oncogenic mutations of B-RAF. Cell 2004; 116: 855.
Wellbrock C, Ogilvie L, Hedley D, Karasarides M, Martin J, Niculescu-Duvaz D et al. V599EBRAF is an oncogene in melanocytes. Cancer Res 2004; 64: 2338–2342.
Chapman PB, Hauschild A, Robert C, Haanen JB, Ascierto P, Larkin J et al. Improved survival with vemurafenib in melanoma with BRAF V600E mutation. N Engl J Med 2011; 364: 2507–2516.
Flaherty KT, Puzanov I, Kim KB, Ribas A, McArthur GA, Sosman JA et al. Inhibition of mutated, activated BRAF in metastatic melanoma. N Engl J Med 2010; 363: 809–819.
Kefford R, Arkenau H, Brown MP, Millward M, Infante JR, Long GV et al. Phase I/II study of GSK2118436, a selective inhibitor of oncogenic mutant BRAF kinase, in patients with metastatic melanoma and other solid tumors. J Clin Oncol 2010; 28, abstract 8503. Presented at 2010 ASCO Annual Meeting: Chicago, IL.
Ribas A, Kim KB, Schucter LM, Gonzalez R, Pavlick AC, Weber JS et al. BRIM-2: an openlabel, multicenter phase II study of vemurafenib in previously treated patients with BRAF V600E mutation-positive metastatic melanoma. J Clin Oncol 2011; 29, abstract 8509. Presented at 2011 ASCO Annual Meeting: Chicago, IL.
Joseph EW, Pratilas CA, Poulikakos PI, Tadi M, Wang W, Taylor BS et al. The RAF inhibitor PLX4032 inhibits ERK signaling and tumor cell proliferation in a V600E BRAF-selective manner. Proc Natl Acad Sci USA 2010; 107: 14903–14908.
Johannessen CM, Boehm JS, Kim SY, Thomas SR, Wardwell L, Johnson LA et al. COT drives resistance to RAF inhibition through MAP kinase pathway reactivation. Nature 2010; 468: 968–972.
Nazarian R, Shi H, Wang Q, Kong X, Koya RC, Lee H et al. Melanomas acquire resistance to B-RAF(V600E) inhibition by RTK or N-RAS upregulation. Nature 2010; 468: 973–977.
Wagle N, Emery C, Berger MF, Davis MJ, Sawyer A, Pochanard P et al. Dissecting therapeutic resistance to RAF inhibition in melanoma by tumor genomic profiling. J Clin Oncol 2011; 29: 3085–3096.
Reya T, Morrison SJ, Clarke MF, Weissman IL . Stem cells, cancer, and cancer stem cells. Nature 2001; 414: 105–111.
Sánchez-García I, Vicente-Dueñas C, Cobaleda C . The theoretical basis of cancer-stem-cell-based therapeutics of cancer: can it be put into practice? Bioessays 2007; 29: 1269–1280.
Crea F, Duhagon MA, Farrar WL, Danesi R . Pharmacogenomics and cancer stem cells: a changing landscape? Trends Pharmacol Sci 2011; 32: 487–494.
Tzvetkov M, von Ahsen N . Pharmacogenetic screening for drug therapy: from single gene markers to decision making in the next generation sequencing era. Pathology 2012; 44: 166–180.
Reis-Filho JS . Next-generation sequencing. Breast Cancer Res 2009; 11 (Suppl 3): S12.
Wagle N, Berger MF, Davis MJ, Blumenstiel B, DeFelice M, Pochanard P et al. High-throughput detection of actionable genomic alterations in clinical tumor samples by targeted, massively parallel sequencing. Cancer Discovery 2011; 2: 82–93.
Holbrook JD, Parker JS, Gallagher KT, Halsey WS, Hughes AM, Weigman VJ et al. Deep sequencing of gastric carcinoma reveals somatic mutations relevant to personalized medicine. J Transl Med 2011; 25: 119.
Schweiger MR, Kerick M, Timmermann B, Isau M . The power of NGS technologies to delineate the genome organization in cancer: from mutations to structural variations and epigenetic alterations. Cancer Metastasis Rev 2011; 30: 199–210.
Borràs E, Jurado I, Hernan I, Gamundi MJ, Dias M, Martí I et al. Clinical pharmacogenomic testing of KRAS, BRAF and EGFR mutations by high resolution melting analysis and ultra-deep pyrosequencing. BMC Cancer 2011; 24: 406.
Inukai M, Toyooka S, Ito S, Asano H, Ichihara S, Soh J et al. Presence of epidermal growth factor receptor gene T790M mutation as a minor clone in non-small cell lung cancer. Cancer Res 2006; 66: 7854–7858.
Heinrich MC, Maki RG, Corless CL, Antonescu CR, Harlow A, Griffith D et al. Primary and secondary kinase genotypes correlate with the biological and clinical activity of sunitinib in imatinib-resistant gastrointestinal stromal tumor. J Clin Oncol 2008; 26: 5352–5359.
Wagle N, Emery C, Berger MF, Davis MJ, Sawyer A, Pochanard P et al. Dissecting therapeutic resistance to RAF inhibition in melanoma by tumor genomic profiling. J Clin Oncol 2011; 29: 3085–3096.
Timmermann B, Kerick M, Roehr C, Fischer A, Isau M, Boerno ST et al. Somatic mutation profiles of MSI and MSS colorectal cancer identified by whole exome next generation sequencing and bioinformatics analysis. PLoS One 2010; 5: e15661.
Acknowledgements
We thank Dr Sabino Ciavarella for critically revising the manuscript, Dr Giorgia Ludovici for excellent technical assistance and A.R.B.O. Financial support for this work was provided by a research grant from the Italian Association of Cancer Research (AIRC IG11647) to F.S. and grant MERIT RBNE08NKH7 to F.G.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Savonarola, A., Palmirotta, R., Guadagni, F. et al. Pharmacogenetics and pharmacogenomics: role of mutational analysis in anti-cancer targeted therapy. Pharmacogenomics J 12, 277–286 (2012). https://doi.org/10.1038/tpj.2012.28
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/tpj.2012.28
Keywords
This article is cited by
-
BI-847325, a selective dual MEK and Aurora kinases inhibitor, reduces aggressive behavior of anaplastic thyroid carcinoma on an in vitro three-dimensional culture
Cancer Cell International (2022)
-
The pharmacogenomics of osteosarcoma
The Pharmacogenomics Journal (2017)
-
Association of ABCB1 and ABCG2 single nucleotide polymorphisms with clinical findings and response to chemotherapy treatments in Kurdish patients with breast cancer
Tumor Biology (2016)
-
Single nucleotide polymorphism detection by optical DNA-based sensing coupled with whole genomic amplification
Analytical and Bioanalytical Chemistry (2013)