Abstract
The autism susceptibility candidate 2 gene (AUTS2) has been associated with multiple neurological diseases including autism spectrum disorders (ASDs). Previous studies showed that AUTS2 has an important neurodevelopmental function and is a suspected master regulator of genes implicated in ASD-related pathways. However, the regulatory role and targets of Auts2 are not well known. Here, by using ChIP-seq (chromatin immunoprecipitation followed by deep sequencing) and RNA-seq on mouse embryonic day 16.5 forebrains, we elucidated the gene regulatory networks of Auts2. We find that the majority of promoters bound by Auts2 belong to genes highly expressed in the developing forebrain, suggesting that Auts2 is involved in transcriptional activation. Auts2 non-promoter-bound regions significantly overlap developing brain-associated enhancer marks and are located near genes involved in neurodevelopment. Auts2-marked sequences are enriched for binding site motifs of neurodevelopmental transcription factors, including Pitx3 and TCF3. In addition, we characterized two functional brain enhancers marked by Auts2 near NRXN1 and ATP2B2, both ASD-implicated genes. Our results implicate Auts2 as an active regulator of important neurodevelopmental genes and pathways and identify novel genomic regions that could be associated with ASD and other neurodevelopmental diseases.
Similar content being viewed by others
Introduction
Autism spectrum disorders (ASDs) are common (1/88 in the United States),1 highly heritable2 pervasive developmental disorders characterized by variable deficits in social communication, language and restrictive and repetitive behaviors.3 AUTS2 has been increasingly implicated as an ASD candidate gene with over 50 unrelated individuals with ASD or ASD-related phenotypes, such as intellectual disability or developmental delay, having structural variants disrupting the AUTS2 region.4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17 AUTS2 structural variants have also been associated with other neurological phenotypes including epilepsy,18 schizoaffective disorder,19 bipolar disorder,20,21 attention deficit-hyperactivity disorder,22 differential processing speed,23 suicidal tendencies under the influence of alcohol24 and dyslexia.10 In addition, a non-coding single-nucleotide polymorphism within AUTS2 (rs6943555) is significantly associated with alcohol consumption.25 AUTS2 has also been implicated in human-specific evolution.26, 27, 28
AUTS2 is a nuclear protein that is expressed in numerous neuronal cell types including glutamatergic neurons (in the cortex, olfactory bulb and hippocampus), GABAergic neurons (Purkinje cells) and tyrosine hydroxylase-positive dopaminergic neurons (substantia nigra and ventral tegmental area).29 In mice, Auts2 is expressed in the developing olfactory bulb, cerebral cortex and cerebellum, and is located in the nuclei of neurons and some neuronal progenitors.29 At embryonic day (E) 16, Auts2 shows strong expression in the cerebral cortex with a gradient of high rostral to low caudal expression.29
The precise function of AUTS2 is not well known; however, zebrafish knockdowns have shown auts2 to be critical in neurodevelopment. auts2 morphant fish display microcephaly with a decrease in neuronal cells in the brain,13,30 which may be caused by failure of cells to differentiate into mature neurons.30 auts2 knockdown also leads to craniofacial abnormalities in zebrafish13 and reduced movement, possibly caused by fewer motor and/or sensory neurons.30 Sequence analysis of AUTS2 identified a predicted PY motif (PPPY) at amino acids 515–519,31 a potential WW-domain-binding region involved in protein–protein interactions. This motif is thought to be involved in the activation of transcription factors, suggesting that AUTS2 may be involved in transcriptional regulation.5
Several proteins are suggested to affect the expression of AUTS2. T-box, brain, 1 (Tbr1), a transcription factor that has been implicated in ASD,32 regulates regional and laminar identity in postmitotic neurons29 and is critical for proper neocortex development,33 interacts with the Auts2 promoter in the developing neocortex.33 Tbr1-deficient mice have decreased levels of reelin,33 a protein necessary for proper neuronal migration in developing brains that can be expressed at decreased levels in individuals with ASD.34 Auts2 expression is reduced in SATB homeobox 2 (Satb2) null mice, with Satb2 being a known regulator of Tbr1 expression in callosal projection neurons.35 Auts2 has been reported to have a 1.33-fold change in cerebellar gene expression in methyl CpG binding protein 2 (Mecp2) null mice, a gene implicated in neurodevelopmental disorders including Rett syndrome and autism.36 AUTS2 is a potential target of GTF2I repeat domain-containing 1 (GTF2IRD1), one of 26 genes deleted in neurodevelopmental disorder Williams–Beuren syndrome.37,38 Zinc-finger matrin-type 3 (Zmat3, also known as wig-1) is a transcription factor regulated by p53 and has an important role in RNA protection and stabilization. wig-1 downregulation leads to a significant reduction in Auts2 mRNA levels in the brains of BACHD mice, a Huntington’s disease mouse model.39
AUTS2 is also thought to interact with other genes and pathways including the notch and ERK signaling pathways, PRC1, and SEMA5A.40,41 Notch signaling has been shown to be involved in neuronal migration through its interaction with Reelin42,43 and AUTS2 expression was found to oscillate in phase with other notch pathway genes.44,45 Polycomb-group repressive complex 1 (PRC1), a polycomb-group gene often involved in transcriptional repression physically interacts with AUTS2, implicating a role for AUTS2 in developmental transcriptional regulation.46 The regulatory pathway of the autism candidate gene semaphorin 5 A (SEMA5A) contains multiple ASD-associated genes, including AUTS2, that overlap rare copy number variations that have been associated with ASD. Twelve regulators of SEMA5A-regulated genes were identified, including AUTS2, suggesting that AUTS2 is a master regulator in ASD-related pathways.47 The Drosophila melanogaster Tay bridge gene, which has a region of homology with AUTS2 (30% identity in the amino acids 1764–2019 with human AUTS2 amino acids 486–782) in the C-terminal region, is a component of the EGFR/Erk signaling pathway that antagonizes EGFR signaling and interacts with Erk.41 This suggests that AUTS2’s normal function in humans may be involved in Erk signaling, having a role in the differentiation and survival of cells during development.41 Given the limited homology between the proteins, it is difficult to deduce any functional conservation.48 However, expression of human AUTS2 in the wing disc was shown to interfere with EGFR signaling, albeit in an opposite manner to Tay. The authors conclude that the effects of AUTS2 on drosophila EGFR signaling are consistent with a role in the regulation of Erk in humans.41 Combined, these interactions implicate AUTS2 in neurodevelopment pathways, including processes important for cell differentiation and ASD.40 However, the actual downstream regulatory targets of AUTS2 still remain largely unknown.
Given AUTS2’s suspected role in neurodevelopment and ASD, and that AUTS2 may be a master regulator in ASD-related pathways,47 we explored the genomic targets of Auts2 in E16.5 mouse forebrains using chromatin immunoprecipitation followed by deep sequencing (ChIP-seq), RNA-seq and zebrafish enhancer assays. We found multiple lines of evidence that Auts2 is associated with promoters and distal enhancers of genes that are active during neurodevelopment. In addition, motif analysis of Auts2-marked sites found enrichment for known motifs involved in neurodevelopment including paired-like homeodomain 3 (Pitx3), transcription factor 3 (TCF3) and forkhead box O3 (FOXO3). Finally, we identified two novel brain enhancers marked by Auts2, which are located near neurexin 1 (NRXN1) and ATPase Ca++ transporting plasma membrane 2 (ATP2B2), two genes implicated in ASD.
Materials and methods
ChIP-seq
Mouse embryos were harvested from timedpregnant CD-1 females (Charles River, Wilmington, MA, USA) at E16.5. The forebrains were dissected in cold phosphate-buffered saline and batches of eight forebrains each were collected in a tube and washed twice with phosphate-buffered saline. Forebrains were cut to <1 mm size and crosslinked with 1% formaldehyde for 10 min. Chromatin from forebrain tissue was isolated and sheared using a Bioruptor (Diagenode, Denville, NJ, USA) and immunoprecipitation was performed using 5 mg of anti-AUTS2 antibody (HPA000390, Sigma Aldrich (St Louis, MO, USA), lot no. A33089; previously verified to be specific for the Auts2 protein).29 The antibody is polyclonal, which can be prone to batch variability and can contain multiple epitopes leading to nonspecific background. We confirmed Auts2 antibody specificity through immunoblot and immunoprecipitation assays. To further validate our results, we performed ChIP-seq with three additional polyclonal Auts2 antibodies: Everest (Upper Heyford, England; catalog no. EB09003), Santa Cruz (Dallas, TX, USA; catalog no. sc-163717) and Abcam (Cambridge, England; catalog no. ab96326). We obtained peaks using the Everest (139 peaks) and Santa Cruz (51 peaks) antibodies but no unique peaks were obtained with the Abcam antibody. Analysis of the overlap of those peaks with the Sigma antibody ChIP-seq peaks found 137 (98.5%) of the Everest peaks and all 51 (100%) of the Santa Cruz peaks to overlap the Sigma peaks (Supplementary Table S1). Furthermore, we performed quantitative PCR to validate the ChIP-seq results and showed specific enrichment for four Everest peaks that overlap Sigma peaks, and no enrichment for four negative controls (regions lacking Auts2 association; data not shown). Chromatin from the same sample was processed for the input control. Illumina libraries were constructed from ChIP and input DNA by the UC Davis Genomics core and sequenced on a HiSeq2500 (Illumina). Auts2 ChIP-seq data from this study are available in SRA (http://www.ncbi.nlm.nih.gov/sra; SRA experiment SRR1292304 (Sigma), SRR1292309 (input), SRR1365079 (Everest), SRR1365081 (Santa Cruz)). ChIP-seq reads were demultiplexed and aligned to the mouse genome (mm9) using Bowtie49 allowing one mismatch per read and retaining only reads with a single reportable alignment (example command: bowtie -p 12 -m 1 -v 1 -S). Resulting SAM files were converted to BAM format for peak calling with MACS2 (version 2.0.10).50 For peak calling, MACS2 extended the reads to 300 bp and kept only peaks where FDR⩽0.01 after normalizing relative to input DNA (example command: macs2 callpeak -f BAM -g mm --keep-dup 1 --nomodel --extsize 300 −q 0.01). A total of 49 364 210 reads were generated with the Sigma Auts2 input and 49 289 630 reads were generated from the input control. For both Auts2 and input, the read length was 50 bp.
RNA-seq
Total RNA was extracted from two replicates of E16.5 mouse forebrain tissue and purified using the RNeasy Maxi Kit (Qiagen, Venlo, Limburg, The Netherlands) and sent to Otogenetics for ribosomal RNA depletion, cDNA production using random primers, library preparation, and pair end sequencing (~100 bp) using Illumina HiSeq2000. Resulting reads were demultiplexed and aligned to the mouse genome (mm9) using TopHat v2.0.7.49,51,52 The two replicates were merged and read counts mapping to each transcript were obtained using HTseq.53 Expression of each transcript was quantified as fragments per kilobase of transcript per million fragments mapped (FPKM) by dividing the total number of reads mapping to each transcript by transcript length, and the total number of reads aligned to the genome divided by one million. The Wilcoxon test from the statistical toolkit R was used to calculate differences in gene expression between genes whose promoter contains an Auts2-marked site and all other genes. RNA-seq data from this study are available in SRA (http://www.ncbi.nlm.nih.gov/sra; SRR experiment SRR1298758 (replicate 1) and SRR1298760 (replicate 2)).
Analyzing the genomic context near Auts2 peaks
Distance from Auts2-marked sites to nearest transcription start site (TSS) was assessed using genomic coordinates downloaded from the UCSC Genome Browser’s mm9 RefSeq Genes track. Histone modification ChIP-seq data from mouse E14.5 whole brain54 were downloaded from UCSC Genome Browser (H3K4me3, H3K27ac and H3K27me3). All data sets were downloaded in October 2013. BedTools’55 intersectBed command was used to identify overlap between this study’s Auts2-marked sites and histone modification peaks from E14.5 mouse whole brain. A single base pair overlap was sufficient to consider two regions overlapping. We observed a minimum of 3 bp, maximum of 1475 bp and a median of 349 bp overlap. Significance was calculated using a permutation p-test (1000 permutations). To determine how many Auts2-marked sites reside within each gene in the mouse genome, genes were defined by transcription start and end sites in mm9 RefSeq. BedTools’ windowBed command (with 1 bp window) was used to count the number of Auts2-marked sites within each gene.
Gene ontology, pathway and motif analysis
Pathway analysis was performed using ingenuity pathway analysis (IPA; Ingenuity Systems, www.ingenuity.com) by inputting a list of genes near Auts2-marked regions (one gene per Auts2-marked site, based on closest distance to TSS, listed in Supplementary Table S2). IPA performs multiple hypothesis test correction using the Benjamini–Hochberg method. Gene ontology analysis was performed using Genomic Regions Enrichment of Annotations Tool (GREAT).56 The Q-value, calculated by GREAT, applies a false discovery rate correction to the binomial raw P-value to correct for multiple hypothesis testing.56 To associate genomic regions with genes in GREAT, we used the ‘basal plus extension’ setting, except when examining promoter marked sites, in which case ‘single nearest gene’ was chosen. Background regions were defined using GREAT’s ‘whole genome’ setting except when examining promoter marked sites, where we defined a promoter-specific background containing all promoter regions (2500 bp upstream and 500 bp downstream of a TSS) plus a window the size of the largest Auts2 ChIP-seq peak (1457 bp). MEME and ChIPmunk57,58 were used to search for de novo motifs within Auts2-marked sites. MEME-ChIP and GOMO59,60 were used to identify experimentally characterized motifs and their gene ontologies. MEME-ChIP and GOMO analyses were performed on 26 March 2014. For MEME-Chip, transcription factor-binding motif input came from JASPAR vertebrates and UniPROBE mouse databases. The expected motif site distribution was set to zero or one occurrence per sequence and motif width was set between 6 and 30. For GOMO, the supported database category was set to multiple species and the database was set to Mus Musculus. The signal threshold was set to q⩽0.05.
Transgenic enhancer assays
Enhancer candidate sequences were selected from Auts2-marked sites based on proximity to the SFARI genes that had a gene score of 1–3 (corresponding to high confidence genes, strong candidates and genes with suggestive evidence),61 evolutionary conservation (sequences showing ⩾70% identity for at least 100 bp) and/or overlap with enhancer-associated histone marks (H3K27ac and H3K4me1), but not promoters (H3K4me3) from whole-brain E14.5 ChIP-seq data sets.54 PCR was carried out on human genomic DNA (Roche) and products were cloned into the E1b-GFP (green-fluorescent protein)-Tol2 enhancer assay vector containing an E1b minimal promoter followed by GFP62 and verified by sequencing. Constructs were injected following standard procedures63,64 into at least 100 zebrafish embryos along with Tol2 mRNA65 to facilitate genomic integration. GFP expression was observed and annotated up to 48 h post fertilization (hpf). An enhancer was considered positive if at least 15% of all fish surviving to 48 hpf showed a consistent expression pattern after subtracting out percentages of tissue expression in fish injected with the empty enhancer vector. For each construct, at least 50 fish were analyzed for GFP expression at 48 hpf. All animal work was approved by the UCSF Institutional Animal Care and Use Committee (protocol number AN100466).
Results
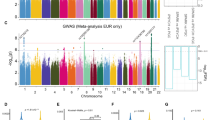
We performed RNA-seq and ChIP-seq using an Auts2 antibody on E16.5 mouse forebrains. E16.5 was chosen because of the reported strong Auts2 expression in the forebrain29 and the established neurogenesis for many relevant brain structures at this time point.66 Through RNA-seq, we identified 8897 transcripts expressed at this time point (quantified as FPKM>0.3). Our Auts2 ChIP-seq found 1930 marked sites, the majority of which (1146=59%) do not overlap gene promoters (2500 bp upstream and 500 bp downstream of a TSS). Nonetheless, the 784 Auts2-marked promoters we detected are significantly more than expected by chance (P<0.001; permutation test), and most promoter peaks (602=31% of all peaks) directly overlap the TSS (Figure 1a).
Analysis of Auts2 ChIP-seq peaks. (a) Distance distribution of the 1930 Auts2-marked sites to the nearest transcription start site (TSS) shows preferential binding near TSSs. Histogram displays bins of 5 kb. (b) FPKM transcript expression scores (FPKM>0.3) for genes whose promoters localized with Auts2 display significantly higher expression than those that do not (P<2.2e−16; Wilcoxon test). (c) Overlaps of Auts2-marked sites with histone modifications show significant localization of Auts2 at promoters (H3K4me3) and active enhancers (H3K27ac; P<0.001; permutation test) but not repressed regions (H3K27me3; P-value =0.081; permutation test). Histone data were acquired from previously reported ChIP-seq for mouse E14.5 whole brain.54
Promoters of actively transcribed neurodevelopmental genes are marked by Auts2
We initially focused on the 784 Auts2-marked sites that reside within promoter regions. These promoter peaks correspond to 776 genes, as a few genes have multiple Auts2-marked sites within their promoter region. Our RNA-seq analysis showed that these genes display significantly higher expression levels at E16.5 than transcripts that were not marked by Auts2 (P<2.2e−16; Wilcoxon test; Figure 1b). Consistent with their association with highly expressed genes, 88% of Auts2 marks at promoters (689/784) are also marked by the active promoter histone modification H3K4me3 in previously published ChIP-seq data generated from E14.5 whole brain54 (P<0.001; permutation test; Figure 1c). These results indicate that the presence of Auts2 at promoters correlates with transcriptional activation.
Using IPA, we comprehensively analyzed pathways and networks of the 776 genes whose promoters overlap Auts2-marked sites, and therefore may be directly regulated by the Auts2 protein. We found that these genes are significantly enriched for diseases and biological functions related to neurodevelopment, including epileptic seizures, disorders of the basal ganglia, migration of neural precursor cells, cell movement of neurons, polarization of neurons and more (Figure 2a, Supplementary Table S2). Interestingly, these genes are also enriched for processes involved in gene expression and cell cycle, including expression of RNA, transcription, proliferation of cells, splicing of RNA, expression of DNA and cell death (Figure 2a).
Pathway analysis and gene ontology of Auts2-marked sites. (a) ingenuity pathway analysis (IPA) pathway analysis of genes whose promoters contain an Auts2-marked site; the figure shows selected significant (after Benjamini–Hochberg correction) neurological, gene expression and cell cycle-related disease and biological functions. (b) GREAT56 gene ontology analysis of non-promoter Auts2-marked sites; the figure shows all significant (after false discovery rate correction) neurological-related mouse phenotypes.
Non-promoter Auts2-marked regions function as enhancers
We next analyzed the 1146 Auts2 ChIP-seq marked sites that did not overlap promoters. Supporting the hypothesis that these distal peaks are functionally important genomic elements, 74% are evolutionarily conserved (phastcons 30-way mammal conserved elements),67 significantly more than expected by chance (P<0.001; permutation test). Further suggesting that Auts2 is primarily an activator, 26% (294/1146) of Auts2 distal peaks overlap previously reported mouse E14.5 whole-brain H3K27ac ChIP-seq peaks,54 an active enhancer mark, which is significantly more than expected by chance (P<0.001; permutation test). In contrast, only 2% (20/1146) overlap E14.5 mouse forebrain H3K27me3 peaks,54 a repressive mark (P=0.081; permutation test; Figure 1c). Five Auts2-marked sites were identified within the Auts2 gene itself, all of which overlap mouse E14.5 forebrain H3K4me1 marks,54 and two that overlap H3K27ac marks,54 suggesting an autoregulatory active role for Auts2. None of the Auts2-marked sites overlap the TSS of any of the transcripts (Supplementary Figure S1). Combined, these results imply that many of the non-promoter Auts2-marked sites likely function as activate regulatory elements.
We investigated the regulatory functions of the 1146 non-promoter Auts2-marked sites using the gene ontology tool GREAT. These Auts2-marked non-promoter sites reside near genes involved in mouse brain development (Figure 2b) including corpus callosum size and neuron number, both of which have been implicated in ASD.68, 69, 70 Taken together, these data support a neurodevelopmental and gene expression role for genes associated with Auts2-marked regulatory regions.
Given our observed correlation between Auts2-marked sites and enhancer marks, we next tested whether these sequences function as enhancers in vivo. Ten Auts2-marked enhancer candidates (AMECs; Supplementary Table S3) were tested for enhancer activity using a zebrafish transgenic enhancer assay. Candidates were selected based on proximity to ASD-related genes, conservation and overlap with enhancer-associated histone modifications.54 Four of the ten candidates were positive enhancers at 24 or 48 hpf (Supplementary Figure S2). AMEC1 lies in an intron of NRXN1 and showed positive enhancer activity in the heart and forebrain (olfactory epithelium) at 48 hpf (Figure 3a, Supplementary Figure S2a). AMEC2, which lies ~56 kb upstream of contactin 4 (CNTN4), displayed enhancer activity in the somitic muscles at 48 hpf (Supplementary Figure S2b). AMEC5, which lies ~571 kb upstream of the RNA-binding protein fox-1 homolog (RBFOX1), had enhancer activity in the notochord at 48 hpf (Supplementary Figure S2c). AMEC8, which lies in an intron of ATP2B2, showed enhancer activity in the midbrain and hindbrain (potentially in the trigeminal sensory neuron) and the spinal cord at 24 hpf, and in somitic muscles at 48 hpf (Figure 3b, Supplementary Figure S2d). These results show that Auts2 occupies functionally active enhancers, some of which drive expression in the developing brain.
Auts2-marked enhancers. (a) A UCSC Genome Browser snapshot of the Nrxn1 locus in mm9, including tracks for the RefSeq gene, Auts2 ChIP-seq, whole-brain E14.5 H3K27ac/H3k4me1/H3kme3 ChIP-seq54 and the Auts2-marked enhancer candidate (AMEC) 1. A representative picture of AMEC1 shows positive enhancer activity in the zebrafish heart and forebrain (olfactory epithelium (red arrow)) at 48 h post fertilization (hpf) is shown below. (b) UCSC browser snapshot of the Atp2b2 region including RefSeq and ChIP-seq tracks. Below, a representative 24 hpf embryo showing enhancer activity of AMEC8 in the midbrain (red arrow) and hindbrain (trigeminal sensory neurons).
Additional genome-wide pathway analyses confirm neurodevelopmental function
We examined the function of nearby genes for all 1930 Auts2-marked sites and the subset of 784 promoter marked sites using GREAT (Supplementary Table S2) and observed an enrichment for gene expression in the mouse cerebral cortex (promoter sites) and in the lower jaw (all marked sites; Supplementary Table S2), which fits with previously reported human phenotypes13 as well as auts2 morpholino knockdown phenotypes of smaller heads13,30 and undersized and reduced jaws.13 GREAT disease ontology enrichment also identified motor neuron disease and hereditary degenerative disease of central nervous system (promoter marked sites; Supplementary Table S2), which also matches the auts2 knockdown phenotype of fewer motor neuron cell bodies in the spinal cord along with improperly angled weaker projections.30
We also performed IPA analysis on all Auts2-marked sites and the subset of 1146 non-promoter Auts2-marked sites using the gene whose TSS was closest to the ChIP-seq peak (Supplementary Table S2). For both data sets, we found significant enrichment of many diseases and functions including multiple neurodevelopmental related categories (for example, migration of neurons, differentiation of neurons, development of the cerebral cortex, disorders of the basal ganglia, Parkinson’s disease, schizophrenia, and so on; Supplementary Table S2). In addition, we observed enrichment for genes involved in the axonal guidance signaling canonical pathway (P=4.6e−9; right-tailed Fisher exact test; for all Auts2-marked sites; Supplementary Figure S3, Supplementary Table S2) and ERK/MAPK signaling canonical pathway (P=6.79e−7; right-tailed Fisher exact test; for all Auts2-marked sites; Supplementary Figure S4, Supplementary Table S2).
Motif analysis identifies transcription factors involved in neuronal development
To identify known motifs present in Auts2-marked sites, we compared the sequences of those sites with position weight matrices of several hundreds of experimentally determined transcription factor-binding sites using MEME-ChIP.59 We analyzed gene ontology terms linked to identified motifs with the gene ontology tool GOMO.60 In Auts2-marked promoter regions, we identified motifs involved in many functions including olfactory receptor activity, translation, transcription, neuron fate commitment and structural constituent of ribosomes (Supplementary Table S4). Among the neuro-associated enriched motifs is a binding motif for Pitx3, a transcriptional regulator involved in the differentiation and maintenance of dopaminergic neurons during development.71 A paralog of PITX3 with a similar binding motif is PITX1, which has been implicated in autism.72 In Auts2-marked non-promoter regions, we identified motifs involved in many functions including calcium ion binding, translation, structural constituent of ribosomes and olfactory receptor activity (Supplementary Table S4). Among the known motifs enriched in non-promoter marked regions is the T-cell acute lymphocyte leukemia/TCF3 heterodimer (TAL1::TCF3). Tcf3 is a transcriptional regulator expressed in the developing cerebral cortex involved in the regulation of cell growth, differentiation and commitment of multiple cell lineages including neurons.73, 74, 75 TAL1 and TCF3 are also associated with acute lymphoblastic leukemia (ALL),76,77 which has been associated with AUTS2.40 In addition enriched is FOXO3, a transcriptional activator involved in neuronal cell death78 (Supplementary Table S4). Using MEME and ChIPmunk, two de novo motif algorithms,57,58 we were only able to identify motifs comprising simple repeats, for instance [CA]ACA[CA]ACA[CA]ACA.
Discussion
Our mouse E16.5 forebrain RNA-seq and ChIP-seq analyses lend further support to a neurodevelopmental role for Auts2. We showed that Auts2 is frequently and significantly localized near promoters of active genes, suggesting that Auts2 is involved in the activation or maintenance of gene expression. IPA and GREAT analyses identified significant associations with neurodevelopmental genes and pathways near Auts2 ChIP-seq peaks. Both GREAT and IPA found highly significant enrichment for genes involved in gene expression and ribosome-related proteins. Interestingly, dynamic regulation of individual ribosomal proteins control gene expression and mammalian development within vertebrate embryos.79 Future experiments are needed to determine whether and how AUTS2 and ribosomes work together to affect neurodevelopment. IPA analysis also identified axonal guidance signaling as a significantly enriched canonical pathway, which includes the SEMA5A gene and supports AUTS2’s role in SEMA5A-related pathways including neurodevelopment and ASD progression.47
Our analyses identified several interesting genes that Auts2 may target. Glycerophosphodiester phosphodiesterase domain containing 1 (Gdpd1) contains an Auts2 mark at its promoter and is highly expressed in our RNA-seq data set (FPKM=2.58). GDPD1 encodes a glycerophosphodiester phosphodiesterase and could be a candidate region linking AUTS2 to alcoholism, given that other classes of phosphodiesterases are involved in alcohol seeking and consumption behaviors.80 GABA B receptor 1 (Gabbr1), which also contains an Auts2-marked site at its promoter and is highly expressed in our data set (FPKM=11.18), is a key component of GABAergic signaling important for synaptic regulation and is implicated in autism, epilepsy and alcohol and cocaine addiction.81, 82, 83 Ubiquitin carboxyl-terminal esterase L1 (Uchl1) is very highly expressed in our data set (FPKM= 43.36) and displays Auts2 localization at its promoter. Uchl1 is expressed specifically in neurons and is a strong Parkinson’s disease susceptibility gene,84 providing insight on a potential new role for AUTS2 in the progression of Parkinson’s disease. Another very highly expressed gene (FPKM=28.08) that has a promoter marked with Auts2 is neruocan (Ncan), a gene thought to be involved in axon guidance85 that is implicated in bipolar disorder and schizophrenia.86,87 These genes represent possible connections between AUTS2 and a wide range of neurological disorders and provide a list of candidate interactions for further functional studies of the role of AUTS2 in neurodevelopment and disease.
In addition to neurodevelopment, AUTS2 may have a role in cancer. Previous studies have linked AUTS2 with ALL,40 matching our finding of enrichment of the TCF3 motif in Auts2-marked sites, as TCF3 is associated with ALL.77 In addition, our GREAT results identified the MSigDB perturbation term ‘genes whose DNA methylation differ between primary ALL cells and peripheral blood samples’ as significant (p=7.82e−6; Binomial test; non-promoter marks).
De novo motif analysis of Auts2-marked regions did not find any non-repetitive novel motifs, and the distribution of known motifs was not centrally enriched within the Auts2-marked regions, supporting the theory that Auts2 acts as a cofactor rather than by binding directly to DNA. AUTS2 lacks identified DNA-binding motifs but contains several predicted protein–protein interaction domains including an SH2, a PY and 13 SH3 domains.29,31,40 More research needs to be performed to determine Auts2-binding partners and how they converge to bind DNA.
Auts2-binding analysis revealed that non-promoter Auts2-marked sites overlap the enhancer mark H3K27ac significantly more than expected by chance, but not with the repressive H3K27me3 mark, suggesting that Auts2 binds to active enhancer regions throughout the genome. Taken together, this suggests that Auts2 also has an activating role in distant gene regulatory elements. We functionally characterized 10 Auts2-marked sequences near genes implicated in ASD using a transgenic zebrafish enhancer assay. Four AMECs showed positive zebrafish enhancer expression, two of which were positive in the brain. Given our small sample size, testing only 10 AMECs and finding 4 to be positive, we cannot definitively conclude that Auts2 can be used as an enhancer mark or that it is directing the activity of these enhancers. AMEC1, which showed positive zebrafish expression in the olfactory epithelium, is in the intron of NRXN1, a gene involved in synapse formation and signaling that has been implicated in ASD and other neurological disorders.88,89 Nrxn1 is also expressed in the olfactory epithelium at E14.5,90 suggesting this enhancer could regulate Nrxn1. Several other lines of evidence, including our motif analysis, known expression patterns of Auts2 in mouse and zebrafish,29,30 and our IPA and GREAT analyses, further support an olfactory role for Auts2. AMEC8 lies in an intron of ATP2B2, which is implicated in ASD, likely because of altered Ca2+ signaling when ATP2B2 is defective.91,92 This sequence showed positive zebrafish enhancer activity in trigeminal sensory neurons matching Atp2b2’'s expression in the mouse trigeminal ganglion.90 AMEC4, which is within AUTS2 (Supplementary Figure S1), did not display enhancer activity. It is possible that this region is not an enhancer, or may be active at time points outside the annotated 24–48 hpf.
Using ChIP-seq and RNA-seq on mouse E16.5 forebrains we identified potential Auts2-regulated regions and found that they mark active regulatory elements. We located 1930 targets of Auts2, ~40% of which lie in promoter regions. Transcripts with Auts2-bound promoters had significantly higher expression levels than ones without Auts2 binding. Both the genes and regulatory sequences that are bound by Auts2 provide distinctive candidate regions to investigate nucleotide variation associated with neurodevelopmental disorders. AUTS2 is emerging as a critical regulator of active neurodevelopmental genes, and future studies such as mouse knockouts could confirm and increase our understanding of the neurodevelopmental function of this gene.
References
Baio J . Prevalence of autism spectrum disorders — autism and developmental disabilities monitoring network, 14 Sites, United States, 2008. Centers Dis Control Prev MMWR Surveill Summ 2012; 61: 1–19.
Risch N, Spiker D, Lotspeich L, Nouri N, Hinds D, Hallmayer J et al. A genomic screen of autism: evidence for a multilocus etiology. Am J Hum Genet 1999; 65: 931.
Geschwind DH . Advances in autism. Annu Rev Med 2009; 60: 367–380.
Pinto D, Pagnamenta AT, Klei L, Anney R, Merico D, Regan R et al. Functional impact of global rare copy number variation in autism spectrum disorders. Nature 2010; 466: 368–372.
Kalscheuer VM, FitzPatrick D, Tommerup N, Bugge M, Niebuhr E, Neumann LM et al. Mutations in autism susceptibility candidate 2 (AUTS2) in patients with mental retardation. Hum Genet 2007; 121: 501–509.
Bakkaloglu B, Roak BJO, Louvi A, Gupta AR, Abelson JF, Morgan TM et al. Molecular cytogenetic analysis and resequencing of contactin associated protein-Like 2 in autism spectrum disorders. Am J Hum Genet 2008; 82: 165–173.
Huang X-L, Zou YS, Maher Ta, Newton S, Milunsky JM . A de novo balanced translocation breakpoint truncating the autism susceptibility candidate 2 (AUTS2) gene in a patient with autism. Am J Med Genet A 2010; 152A: 2112–2114.
Glessner JT, Wang K, Cai G, Korvatska O, Kim CE, Wood S et al. Autism genome-wide copy number variation reveals ubiquitin and neuronal genes. Nature 2009; 459: 569–573.
Ben-David E, Granot-Hershkovitz E, Monderer-Rothkoff G, Lerer E, Levi S, Yaari M et al. Identification of a functional rare variant in autism using genome-wide screen for monoallelic expression. Hum Mol Genet 2011; 20: 3632–3641.
Girirajan S, Brkanac Z, Coe BP, Baker C, Vives L, Vu TH et al. Relative burden of large CNVs on a range of neurodevelopmental phenotypes. PLoS Genet 2011; 7: e1002334.
Talkowski ME, Rosenfeld JA, Blumenthal I, Pillalamarri V, Chiang C, Heilbut A et al. Sequencing chromosomal abnormalities reveals neurodevelopmental loci that confer risk across diagnostic boundaries. Cell 2012; 149: 525–537.
Nagamani SCS, Erez A, Ben-Zeev B, Frydman M, Winter S, Zeller R et al. Detection of copy-number variation in AUTS2 gene by targeted exonic array CGH in patients with developmental delay and autistic spectrum disorders. Eur J Hum Genet 2013; 21: 1–4.
Beunders G, Voorhoeve E, Golzio C, Pardo LM, Rosenfeld Ja, Talkowski ME et al. Exonic deletions in AUTS2 cause a syndromic form of intellectual disability and suggest a critical role for the C terminus. Am J Hum Genet 2013; 92: 210–220.
Girirajan S, Johnson RL, Tassone F, Balciuniene J, Katiyar N, Fox K et al. Global increases in both common and rare copy number load associated with autism. Hum Mol Genet 2013; 22: 2870–2880.
Cuscó I, Medrano A, Gener B . Autism-specific copy number variants further implicate the phosphatidylinositol signaling pathway and the glutamatergic synapse in the etiology of the disorder. Hum Mol Genet 2009; 18: 1795–1804.
Tropeano M, Ahn JW, Dobson RJB, Breen G, Rucker J, Dixit A et al. Male-biased autosomal effect of 16p13.11 copy number variation in neurodevelopmental disorders. PLoS ONE 2013; 8: e61365.
Jolley A, Corbett M, McGregor L, Waters W, Brown S, Nicholl J et al. De novo intragenic deletion of the autism susceptibility candidate 2 (AUTS2) gene in a patient with developmental delay: a case report and literature review. Am J Med Genet A 2013; 2: 1–5.
Mefford HC, Muhle H, Ostertag P, von Spiczak S, Buysse K, Baker C et al. Genome-wide copy number variation in epilepsy: novel susceptibility loci in idiopathic generalized and focal epilepsies. PLoS Genet 2010; 6: e1000962.
Hamshere ML, Green EK, Jones IR, Jones L, Moskvina V, Kirov G et al. Genetic utility of broadly defined bipolar schizoaffective disorder as a diagnostic concept. Br J Psychiatry 2009; 195: 23–29.
Hattori E, Toyota T, Ishitsuka Y, Iwayama Y, Yamada K, Ujike H et al. Preliminary genome-wide association study of bipolar disorder in the Japanese population. Am J Med Genet B Neuropsychiatr Genet 2009; 150B: 1110–1117.
Lee H, Woo H, Greenwood T . A genome-wide association study of seasonal pattern mania identifies NF1A as a possible susceptibility gene for bipolar disorder. J Affect Disord 2012; 145: 200–207.
Elia J, Gai X, Xie HM, Perin JC, Geiger E, Glessner JT et al. Rare structural variants found in attention-deficit hyperactivity disorder are preferentially associated with neurodevelopmental genes. Mol Psychiatry 2010; 15: 637–646.
Luciano M, Hansell N, Lahti J . Whole genome association scan for genetic polymorphisms influencing information processing speed. Biol Psychol 2011; 86: 193–202.
Chojnicka I, Gajos K, Strawa K, Broda G, Fudalej S, Fudalej M et al. Possible association between suicide committed under influence of ethanol and a variant in the AUTS2 gene. PLoS ONE 2013; 8: e57199.
Schumann G, Coin LJ, Lourdusamy A, Charoen P, Berger KH, Stacey D et al. Genome-wide association and genetic functional studies identify autism susceptibility candidate 2 gene (AUTS2) in the regulation of alcohol consumption. Proc Natl Acad Sci USA 2011; 108: 7119–7124.
Pollard KS, Salama SR, Lambert N, Lambot M-A, Coppens S, Pedersen JS et al. An RNA gene expressed during cortical development evolved rapidly in humans. Nature 2006; 443: 167–172.
Prabhakar S, Noonan JP, Pääbo S, Rubin EM . Accelerated evolution of conserved noncoding sequences in humans. Science 2006; 314: 786.
Green RE, Krause J, Briggs AW, Maricic T, Stenzel U, Kircher M et al. A draft sequence of the Neandertal genome. Science 2010; 328: 710–722.
Bedogni F, Hodge RD, Nelson BR, Frederick Ea, Shiba N, Daza Ra et al. Autism susceptibility candidate 2 (Auts2) encodes a nuclear protein expressed in developing brain regions implicated in autism neuropathology. Gene Exp Patterns 2010; 10: 9–15.
Oksenberg N, Stevison L, Wall JD, Ahituv N . Function and regulation of AUTS2, a gene implicated in autism and human evolution. PLoS Genet 2013; 9: e1003221.
Sultana R, Yu C-E, Yu J, Munson J, Chen D, Hua W et al. Identification of a novel gene on chromosome 7q11.2 interrupted by a translocation breakpoint in a pair of autistic twins. Genomics 2002; 80: 129–134.
O’Roak BJ, Vives L, Girirajan S, Karakoc E, Krumm N, Coe BP et al. Sporadic autism exomes reveal a highly interconnected protein network of de novo mutations. Nature 2012; 485: 246–250.
Bedogni F, Hodge RD, Elsen GE, Nelson BR, Daza R a M, Beyer RP et al. Tbr1 regulates regional and laminar identity of postmitotic neurons in developing neocortex. Proc Natl Acad Sci USA 2010; 107: 13129–13134.
Fatemi SH, Snow A V, Stary JM, Araghi-Niknam M, Reutiman TJ, Lee S et al. Reelin signaling is impaired in autism. Biol Psychiatry 2005; 57: 777–787.
Srinivasan K, Leone DP, Bateson RK, Dobreva G, Kohwi Y, Kohwi-Shigematsu T et al. A network of genetic repression and derepression specifies projection fates in the developing neocortex. Proc Natl Acad Sci USA 2012; 109: 19071–19078.
Ben-Shachar S, Chahrour M, Thaller C, Shaw C a, Zoghbi HY . Mouse models of MeCP2 disorders share gene expression changes in the cerebellum and hypothalamus. Hum Mol Genet 2009; 18: 2431–2442.
Young EJ, Lipina T, Tam E, Mandel a, Clapcote SJ, Bechard a R et al. Reduced fear and aggression and altered serotonin metabolism in Gtf2ird1-targeted mice. Genes Brain Behav 2008; 7: 224–234.
O’Leary J, Osborne LR . Global analysis of gene expression in the developing brain of Gtf2ird1 knockout mice. PLoS ONE 2011; 6: e23868.
Sedaghat Y, Mazur C, Sabripour M, Hung G, Monia BP . Genomic analysis of wig-1 pathways. PLoS ONE 2012; 7: e29429.
Oksenberg N, Ahituv N . The role of AUTS2 in neurodevelopment and human evolution. Trends Genet 2013; 29: 1–9.
Molnar C, de Celis JF . Tay Bridge is a negative regulator of EGFR signalling and interacts with Erk and Mkp3 in the Drosophila melanogaster wing. PLoS Genet 2013; 9: e1003982.
Hashimoto-Torii K, Torii M, Sarkisian MR, Bartley CM, Shen J, Radtke F et al. Interaction between Reelin and Notch signaling regulates neuronal migration in the cerebral cortex. Neuron 2008; 60: 273–284.
Wang G-S, Hong C-J, Yen T-Y, Huang H-Y, Ou Y, Huang T-N et al. Transcriptional modification by a CASK-interacting nucleosome assembly protein. Neuron 2004; 42: 113–128.
William D a, Saitta B, Gibson JD, Traas J, Markov V, Gonzalez DM et al. Identification of oscillatory genes in somitogenesis from functional genomic analysis of a human mesenchymal stem cell model. Dev Biol 2007; 305: 172–186.
Dequéant M-L, Glynn E, Gaudenz K, Wahl M, Chen J, Mushegian A et al. A complex oscillating network of signaling genes underlies the mouse segmentation clock. Science 2006; 314: 1595–1598.
Gao Z, Zhang J, Bonasio R, Strino F, Sawai A, Parisi F et al. PCGF homologs, CBX proteins, and RYBP define functionally distinct PRC1 family complexes. Mol Cell 2012; 45: 344–356.
Cheng Y, Quinn JF, Weiss LA . An eQTL mapping approach reveals that rare variants in the SEMA5A regulatory network impact autism risk. Hum Mol Genet 2013; 22: 1–13.
Tian W, Skolnick J . How well is enzyme function conserved as a function of pairwise sequence identity? J Mol Biol 2003; 333: 863–882.
Langmead B, Trapnell C, Pop M, Salzberg SL . Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 2009; 10: R25.
Zhang Y, Liu T, Meyer C a, Eeckhoute J, Johnson DS, Bernstein BE et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol 2008; 9: R137.
Trapnell C, Pachter L, Salzberg SL . TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 2009; 25: 1105–1111.
Kim D, Pertea G, Trapnell C, Pimentel H, Kelley R, Salzberg SL . TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol 2013; 14: R36.
Anders S, McCarthy DJ, Chen Y, Okoniewski M, Smyth GK, Huber W et al. Count-based differential expression analysis of RNA sequencing data using R and Bioconductor. Nat Protoc 2013; 8: 1765–1786.
Shen Y, Yue F, McCleary DF, Ye Z, Edsall L, Kuan S et al. A map of the cis-regulatory sequences in the mouse genome. Nature 2012; 488: 116–120.
Quinlan AR, Hall IM . BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 2010; 26: 841–842.
McLean CY, Bristor D, Hiller M, Clarke SL, Schaar BT, Lowe CB et al. GREAT improves functional interpretation of cis-regulatory regions. Nat Biotechnol 2010; 28: 495–501.
Bailey TL, Gribskov M . Fitting a mixture model by expectation maximization to discover motifs in biopolymers. In: Proceedings of the Second International Conference on Intelligent Systems for Molecular Biology. Menlo Park, California 1994 p 2: 28–36.
Kulakovskiy I V, Boeva V a, Favorov a V, Makeev VJ . Deep and wide digging for binding motifs in ChIP-Seq data. Bioinformatics 2010; 26: 2622–2623.
Machanick P, Bailey TL . MEME-ChIP: motif analysis of large DNA datasets. Bioinformatics 2011; 27: 1696–1697.
Buske F a, Bodén M, Bauer DC, Bailey TL . Assigning roles to DNA regulatory motifs using comparative genomics. Bioinformatics 2010; 26: 860–866.
Abrahams BS, Arking DE, Campbell DB, Mefford HC, Morrow EM, Weiss L a et al. SFARI Gene 2.0: a community-driven knowledgebase for the autism spectrum disorders (ASDs). Mol Autism 2013; 4: 36.
Li Q, Ritter D, Yang N, Dong Z, Li H, Chuang JH et al. A systematic approach to identify functional motifs within vertebrate developmental enhancers. Dev Biol 2010; 337: 484–495.
Westerfield M . The Zebrafish Book. 5th edn, University of Oregon Press: Eugene, Oregon, 2007.
Nusslein-Volhard C, RD . Zebrafish. Oxford University Press: Oxford, 2002.
Kawakami K . Transposon tools and methods in zebrafish. Dev Dyn 2005; 234: 244–254.
Finlay B, Darlington R . Linked regularities in the development and evolution of mammalian brains. Science 1995; 268: 1578–1584.
Siepel A, Bejerano G, Pedersen JS, Hinrichs AS, Hou M, Rosenbloom K et al. Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res 2005; 15: 1034–1050.
Egaas B, Courchesne E, Saitoh O . Reduced size of corpus callosum in autism. Arch Neurol 1995; 52: 794–801.
Lainhart J, Lange N . Increased neuron number and head size in autism. JAMA J Am Med 2011; 306: 2031–2032.
Courchesne E, Mouton PR, Calhoun ME, Semendeferi K, Ahrens-Barbeau C, Hallet MJ et al. Neuron number and size in prefrontal cortex of children with autism. JAMA 2011; 306: 2001–2010.
Chung S, Hedlund E, Hwang M, Kim DW, Shin B-S, Hwang D-Y et al. The homeodomain transcription factor Pitx3 facilitates differentiation of mouse embryonic stem cells into AHD2-expressing dopaminergic neurons. Mol Cell Neurosci 2005; 28: 241–252.
Philippi A, Tores F, Carayol J, Rousseau F, Letexier M, Roschmann E et al. Association of autism with polymorphisms in the paired-like homeodomain transcription factor 1 (PITX1) on chromosome 5q31: a candidate gene analysis. BMC Med Genet 2007; 8: 74.
Slattery C, Ryan MP, McMorrow T . E2A proteins: regulators of cell phenotype in normal physiology and disease. Int J Biochem Cell Biol 2008; 40: 1431–1436.
Massari M, Murre C . Helix-loop-helix proteins: regulators of transcription in eucaryotic organisms. Mol Cell Biol 2000; 20: 429–440.
Gray P a, Fu H, Luo P, Zhao Q, Yu J, Ferrari A et al. Mouse brain organization revealed through direct genome-scale TF expression analysis. Science 2004; 306: 2255–2257.
Kelliher M a, Seldin DC, Leder P . Tal-1 induces T cell acute lymphoblastic leukemia accelerated by casein kinase IIalpha. EMBO J 1996; 15: 5160–5166.
Barber KE, Harrison CJ, Broadfield ZJ, Stewart ARM, Wright SL, Martineau M et al. Molecular Cytogenetic characterization of TCF3 acute lymphoblastic leukemia. Genes Chromosom Cancer 2007; 46: 478–486.
Xie Q, Hao Y, Tao L, Peng S, Rao C, Chen H et al. Lysine methylation of FOXO3 regulates oxidative stress-induced neuronal cell death. EMBO Rep 2012; 13: 371–377.
Kondrashov N, Pusic A, Stumpf CR, Shimizu K, Hsieh AC, Xue S et al. Ribosome-mediated specificity in Hox mRNA translation and vertebrate tissue patterning. Cell 2011; 145: 383–397.
Wen R-T, Zhang M, Qin W-J, Liu Q, Wang W-P, Lawrence AJ et al. The phosphodiesterase-4 (PDE4) inhibitor rolipram decreases ethanol seeking and consumption in alcohol-preferring Fawn-Hooded rats. Alcohol Clin Exp Res 2012; 36: 2157–2167.
Fatemi SH, Folsom TD, Reutiman TJ, Thuras PD . Expression of GABA(B) receptors is altered in brains of subjects with autism. Cerebellum 2009; 8: 64–69.
Xi B, Chen J, Yang L, Wang W, Fu M, Wang C . GABBR1 gene polymorphism (G1465A) is associated with temporal lobe epilepsy. Epilepsy Res 2011; 96: 58–63.
Enoch M-A, Zhou Z, Kimura M, Mash DC, Yuan Q, Goldman D . GABAergic gene expression in postmortem hippocampus from alcoholics and cocaine addicts; corresponding findings in alcohol-naïve P and NP rats. PLoS ONE 2012; 7: e29369.
Maraganore DM, Lesnick TG, Elbaz A, Chartier-Harlin M-C, Gasser T, Krüger R et al. UCHL1 is a Parkinson’s disease susceptibility gene. Ann Neurol 2004; 55: 512–521.
Watanabe E, Aono S, Matsui F, Yamada Y, Naruse I, Oohira a . Distribution of a brain-specific proteoglycan, neurocan, and the corresponding mRNA during the formation of barrels in the rat somatosensory cortex. Eur J Neurosci 1995; 7: 547–554.
Cichon S, Mühleisen TW, Degenhardt F a, Mattheisen M, Miró X, Strohmaier J et al. Genome-wide association study identifies genetic variation in neurocan as a susceptibility factor for bipolar disorder. Am J Hum Genet 2011; 88: 372–381.
Mühleisen TW, Mattheisen M, Strohmaier J, Degenhardt F, Priebe L, Schultz CC et al. Association between schizophrenia and common variation in neurocan (NCAN), a genetic risk factor for bipolar disorder. Schizophr Res 2012; 138: 69–73.
Missler M, Südhof T . Neurexins: three genes and 1001 products. Trends Genet 1998; 14: 20–26.
Ching MSL, Shen Y, Tan W-H, Jeste SS, Morrow EM, Chen X et al. Deletions of NRXN1 (neurexin-1) predispose to a wide spectrum of developmental disorders. Am J Med Genet B Neuropsychiatr Genet 2010; 153B: 937–947.
Diez-Roux G, Banfi S, Sultan M, Geffers L, Anand S, Rozado D et al. A high-resolution anatomical atlas of the transcriptome in the mouse embryo. PLoS Biol 2011; 9: e1000582.
Carayol J, Sacco R, Tores F, Rousseau F, Lewin P, Hager J et al. Converging evidence for an association of ATP2B2 allelic variants with autism in male subjects. Biol Psychiatry 2011; 70: 880–887.
Yang W, Liu J, Zheng F, Jia M, Zhao L, Lu T et al. The evidence for association of ATP2B2 polymorphisms with autism in Chinese Han population. PLoS ONE 2013; 8: e61021.
Acknowledgements
We would like to thank Megan Laurance for her assistance with IPA and Mariel Mckenzie Finucane with her assistance with statistics. This work was supported by a grant from the Simons Foundation, SFARI no. 256769, the National Institute of Neurological Disorders and Stroke grant number 1R01NS079231 and German-Israel Foundation grant number I-2360-203. RYB is supported in part by the Career Integration grant (CIG) number 630849. NA is also supported in part by the National Institute of Diabetes and Digestive and Kidney Diseases grant number 1R01DK090382, the National Human Genome Research Institute grant numbers R01HG005058 and R01HG006768, the National Institute of Child and Human Development grant number R01HD059862, and the National Institute of General Medical Sciences grant number GM61390. KSP and GDEH are also supported by NHLBI grant number HL098179, a gift from the San Simeon Fund, and institutional funds from the J David Gladstone Institutes.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Translational Psychiatry website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Oksenberg, N., Haliburton, G., Eckalbar, W. et al. Genome-wide distribution of Auts2 binding localizes with active neurodevelopmental genes. Transl Psychiatry 4, e431 (2014). https://doi.org/10.1038/tp.2014.78
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/tp.2014.78
This article is cited by
-
Erotomania and phenotypic continuum in a family frameshift variant of AUTS2: a case report and review
BMC Psychiatry (2021)
-
AUTS2 isoforms control neuronal differentiation
Molecular Psychiatry (2021)
-
Neuroplastin deletion in glutamatergic neurons impairs selective brain functions and calcium regulation: implication for cognitive deterioration
Scientific Reports (2017)