Abstract
Acamprosate supports abstinence in some alcohol-dependent subjects, yet predictors of response are unknown. To identify response biomarkers, we investigated associations of abstinence length with polymorphisms in candidate genes in glycine and glutamate neurotransmission pathways and genes previously implicated in acamprosate response. Association analyses were conducted in the discovery sample of 225 alcohol-dependent subjects treated with acamprosate for 3 months in community-based treatment programs in the United States. Data from 110 alcohol-dependent males treated with acamprosate in the study PREDICT were used for replication of the top association findings. Statistical models were adjusted for relevant covariates, including recruitment site and baseline clinical variables associated with response. In the discovery sample, shorter abstinence was associated with increased intensity of alcohol craving and lower number of days between the last drink and initiation of acamprosate treatment. After adjustment for covariates, length of abstinence was associated with the GRIN2B rs2058878 (P=4.6 × 10−5). In the replication sample, shorter abstinence was associated with increased craving, increased depressive mood score and higher alcohol consumption. Association of abstinence length with GRIN2B rs2058878 was marginally significant (P=0.0675); as in the discovery sample, the minor A allele was associated with longer abstinence. Furthermore, rs2300272, which is in strong linkage disequilibrium with rs2058878, was also associated with abstinence length (P=0.049). This is the first report of a replicated association of genetic markers with the length of abstinence in acamprosate-treated alcoholics. Investigation of the underlying mechanisms of this association and its usefulness for individualized treatment selection should follow.
Similar content being viewed by others
Introduction
The economic costs associated with alcohol use and misuse are staggering, and evidence-based treatment strategies are needed.1,2 Meta-analyses favor acamprosate for its ability to support abstinence,3, 4, 5 which is the most stable type of remission in alcohol-dependent subjects.6 Only a fraction of those treated respond to acamprosate.7 Clinical characteristics, including the number of sober days before initiation of acamprosate treatment, were shown to be associated with abstinence length.8, 9, 10 However, no biomarkers allowing reliable identification of potential responders to acamprosate treatment are currently known. It is expected that a pharmacogenomic approach may lead to the discovery of such biomarkers, enabling individualized recommendations for treatment selection and improved treatment outcomes.11,12
Acamprosate shares structural similarities with glycine and glutamate13 and is thought to work by counteracting the ‘relief craving’ associated with increased glutamate levels in alcoholics with a history of withdrawal.14,15 Several genes, including GATA4, PER2 and SLC29A1, were associated with response to acamprosate treatment in human and animal studies.16, 17, 18 Recent findings also indicate that glycine, which activates N-methyl-D-aspartate (NMDA) and glycine receptors,19,20 may have an important role in alcohol use disorders and treatment response.21, 22, 23, 24, 25 Therefore, in an effort to identify genetic markers associated with abstinence in alcohol-dependent human subjects receiving acamprosate treatment, we investigated sequence variation in genes involved in composition of the glycine and NMDA receptors, glycine and glutamate reuptake, synthesis and metabolism, along with sequence variation in candidate genes previously reported to be associated with acamprosate response in human and animal studies.
To make study findings practically meaningful, we considered several potentially important issues. First, we conducted our study in samples reflecting the population where results will be applied, that is, alcohol-dependent subjects treated in community-based programs. Second, we monitored non-genetic (that is, clinical and demographic) patient characteristics and accounted for relevant covariates and potential confounders in the genetic analyses. Third, to ensure that findings were relevant and readily translated into clinical practice, we utilized diagnostic tools and outcome measures used in community-based treatment programs. In accordance with these considerations, we conducted an open label, naturalistic prospective study investigating genetic markers associated with abstinence in alcohol-dependent subjects treated with acamprosate in community-based programs. Replication analyses were performed using data from a previously conducted study of acamprosate (PREDICT26).
We identified genetic markers associated with the length of sobriety during acamprosate treatment. This is an important step toward the development of personalized treatment recommendations for patients with alcohol use disorders, as genetic markers may be used for selection of patients who have the highest probability of responding to acamprosate treatment. We think that with the limited number of antidipsotropic medications and lack of uniformity in treatment response, the development of such recommendations is of special importance for improvement of treatment outcomes in patients with alcohol use disorders.
Materials and methods
Discovery study
Subjects and recruitment sites
This study was approved by the Institutional Review Board of the Mayo Clinic Rochester and Mayo Clinic Health System, and was conducted according to the Code of Ethics of the World Medical Association (Declaration of Helsinki). All participants signed informed consent approved by the Mayo Clinic Institutional Review Board.
Both men and women between the ages of 18 and 80 with a primary diagnosis of current alcohol dependence based on DSM-IV-TR criteria with the last drink 5 or more days before enrollment were included in the study. We excluded subjects unable to provide informed consent; those unable to speak English; those with psychotic disorders or unstable psychiatric or medical conditions (see Supplementary Materials); women who were pregnant, lactating or planning to become pregnant; subjects taking disulfiram; and those allergic to acamprosate.
Participants were recruited from community-based residential and outpatient treatment programs affiliated with Mayo Clinic in Rochester, Minnesota, and the Mayo Clinic Health System sites in Austin, Minnesota, Albert Lea, Minnesota, and La Crosse, Wisconsin (Supplementary Table S1). In addition, self-referred participants residing in communities adjacent to referral sites not enrolled in treatment programs but interested in taking acamprosate were recruited and included in the analyses as a separate ‘study site.’ A description of programs and the number of subjects recruited at each site is presented in Supplementary Materials.
Assessment
A detailed description of assessments is presented in Supplementary Table S2 and summarized below. The lifetime presence of alcohol dependence and comorbid disorders was assessed using the semistructured interview PRISM.27 Time Line Follow-Back28 was used to assess alcohol consumption before and during treatment. Craving intensity was assessed using Penn Alcohol Craving Scale (PACS).29,30 Association of alcohol use with positive or negative emotional states was assessed using the Inventory of Drug Taking Situations.31,32 The intensity of depressive symptoms and anxiety were assessed using a depression scale from the patient health questionnaire,33 and the Generalized Anxiety Disorder Assessment.34 Alcoholic Anonymous attendance monitoring35 was used to estimate utilization of support networks.
Treatment outcomes including abstinence (defined as time between initiation of acamprosate treatment and first alcohol use) or alcohol use were assessed with self-report (Timeline Follow-Back) and from available medical records during follow-up visits. Gamma-glutamyl transpeptidase measurements were used to assess the accuracy of self-reported sobriety. Compliance with acamprosate was assessed by pill count. The use of other medications was also monitored and considered as potential covariates in the analyses.
Selection of candidate genetic targets in the discovery study
Candidate genes (Supplementary Table S3) were selected based on the following considerations. First, we used a pathway-based approach to systematically investigate genes encoding enzymes involved in glycine metabolism, glycine transporters and subunits of glycine receptors and the NMDA receptor, which is known to be involved in glycine effects.19,36 Second, because acamprosate action is thought to be associated with increased brain glutamate levels,14,15 we included genes associated with glutamate reuptake, synthesis and degradation. Third, we included candidate genes reported to be associated with acamprosate treatment outcomes in human or animal studies16, 17, 18 and genes associated with alcoholism treatment response.37, 38, 39
We used SNPPicker40 for selection of tag single-nucleotide polymorphisms (SNPs) to attain comprehensive coverage of each candidate gene. In addition, 28 SNPs were selected from a panel of ancestry informative markers41 to verify self-reported ancestry. The list of 548 candidate SNPs included in the study is presented in Supplementary Table S4.
Genotyping and quality control
Genotyping was conducted using Illumina Golden Gate custom panel of 576 single-nucleotide variants.42 Of the 576 SNPs included in the genotyping panel (548 candidate SNPs, 28 ancestry informative SNPs), 19 failed and 11 had minor allele frequencies <2% and were excluded from analysis. Of the 433 subjects who started acamprosate treatment and were genotyped, four were excluded from analyses because of low call rates (<90%). As part of genotype quality control, 18 subjects were genotyped in duplicate and a CEPH trio was genotyped eight times. No discordant genotype calls were observed for these replicated samples. Hardy–Weinberg equilibrium was evaluated; however, no SNPs were removed (Supplementary Table S4). To verify self-reported race, STRUCTURE v2.3.3 (Stanford University, Stanford, CA, USA) was used to estimate ancestry of individuals based on the 28 ancestry informative markers. Four self-reported European Americans were removed as they appeared to have >50% non-European ancestry.
Data analyses
Time-to-event (first alcohol use) survival analysis methods were used to examine the association of clinical and genetic markers with treatment outcomes. Demographic and baseline clinical characteristics were first evaluated for associations with length of abstinence using univariate Cox proportional hazard models. As we found an association of treatment outcome with recruitment site, likely reflecting differences in populational characteristics and non-pharmacological treatment components across sites, we controlled for recruitment site as a covariate in all subsequent analyses. Backward stepwise variable selection with covariates that were univariately related to response (P<0.05) was performed to identify other relevant covariates and potential confounders for the pharmacogenetic analyses. On the basis of the results of this selection process, all genetic analyses were adjusted for enrollment site, days since last drink at baseline and baseline PACS.
Replication study
Study sample
To replicate top association findings in the discovery sample, we conducted similar analyses in a subset of participants from the study PREDICT, a double-blind randomized controlled trial that compared treatment outcomes including length of abstinence among alcohol-dependent subjects of German descent recruited from inpatient facilities and treated with acamprosate, naltrexone or placebo for 3 months.26 Clinical and genetic data from 110 males treated with acamprosate who were also included in a genome-wide association study of alcohol dependence43 and, thus, had genotype data available, were included in the replication analyses. The diagnostic assessments and instruments used in the study PREDICT were described previously.26
Genotyping, selection of candidate genetic targets and data analysis in the replication sample
A subset of PREDICT study participants was genotyped using Illumina Human-Hap 550, Illumina Human 610 and Illumina Human 660w quad BeadChips (Illuminia, San Diego, CA, USA) for a genome-wide association study of alcohol dependence.43,44 We selected four top SNPs associated with treatment outcome in the discovery sample (P<0.001) for replication. Three of those SNPs were not genotyped in the replication sample. Therefore, we imputed those SNPs using the ShapeIT tool, v1 (Christchurch, New Zealand) for phasing and IMPUTE (v2.3, Oxford, UK) for imputation.45,46 The European sample from 1000 genomes data was used as the reference panel, and the three imputed SNPs had dosage R2>0.99.
Analysis of our Replication Set data followed the same approach as the Discovery Set analysis. Specifically, Cox proportional hazard models were used to examine the association of length of abstinence with clinical variables and candidate SNPs. Backward stepwise variable selection was again used to identify covariates and potential confounders for the genetic analyses. Genetic association analyses were then performed for SNP genotypes coded as the minor allele count.
Results
Clinical and demographic variables associated with abstinence in the discovery sample
Of the 443 subjects who started acamprosate, 225 European American subjects with available 3-month outcome data (any alcohol use N=93; abstainers N=132) were included in the analyses. The length of abstinence was significantly associated with the recruitment site as well as several clinical variables (Table 1). Increased craving and depression scores (measured by PACS and patient health questionnaire 9, respectively) were associated with shorter abstinence. Longer abstinence was associated with increased number of days since last drink before initiation of acamprosate, increased attendance of Alcoholics Anonymous meetings, having an Alcoholics Anonymous sponsor and attendance at counseling sessions. However, after adjustment for study site, only the baseline PACS score and the number of days between the last drink and initiation of acamprosate treatment remained strongly associated with treatment outcome (P<0.0001 and P=0.0002, respectively). Using a backward stepwise variable selection process, covariates (including study site, PACS score and the number of days between last drink and initiation of acamprosate treatment) were selected for adjustment in pharmacogenomic analyses.
The observed changes in plasma gamma-glutamyl transpeptidase levels between baseline and 3-month follow-up were consistent with self-reported abstinence and alcohol use. As shown in Supplementary Figure S1, gamma-glutamyl transpeptidase levels were elevated at baseline and decreased markedly during treatment in abstainers and non-abstainers. At the 3-month follow-up, average gamma-glutamyl transpeptidase levels were significantly lower and closer to the normative range in abstainers compared with non-abstainers (P<0.001).
Single SNP association with treatment outcomes in the discovery sample
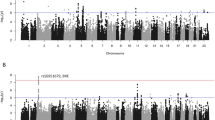
A complete list of candidate SNPs and their association with the length of abstinence is presented in Supplementary Table S4, and summarized in Figure 1. The top results (P<0.001) are also presented in Table 2. The strongest association finding, minor GRIN2B rs2058878 A allele, remains significantly associated with longer abstinence after Bonferroni correction for the number of SNPs included in the analyses (P=4.6 × 10−5, corrected P=0.024). As the replication sample included only alcohol-dependent males, we also explored the association of the top four SNPs with treatment outcome in a subsample of males. As shown in Table 2, the evidence for association with treatment outcome was stronger for minor GLRB rs17035723 A allele in the male subset (hazard ratio=2.88, P=0.98 × 10−5). The hazard ratios of the three GRIN2B SNPs, including rs2058878, were also higher in males; however, P-values reflecting their association with treatment outcome were less significant, possibly because of the decreased sample size in the subsample of males.
Single-nucleotide polymorphism (SNP) association with length of abstinence in the Discovery Sample. Results adjusted for recruitment site, Penn Alcohol Craving Scale and number of days between the last drink and initiation of acamprosate treatment. Dotted line represents the Bonferroni-corrected alpha level of 0.05/518=9.65E−05. Dashed line represents nominal significance of P=0.05.
Clinical and demographic variables associated with abstinence in the replication sample
As shown in Table 3, the replication sample included only alcohol-dependent males and was characterized by higher alcohol consumption compared with the discovery sample. Similar to the discovery sample, increased craving (measured by the Obsessive Compulsive Drinking Scale) was associated with shorter abstinence. Shorter abstinence was also associated with higher alcohol consumption, increased depressive symptoms (measured by the Beck Depression Inventory) and younger age at onset of alcohol dependence. A final multivariable model of potentially relevant covariates (selected using the same process as in the discovery data set) included Beck Depression Inventory score, number of drinks per drinking day and age of onset of alcoholism; all SNP association analyses were adjusted for these covariates.
SNP associations with treatment outcomes in the replication sample
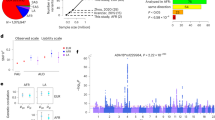
As presented in Table 4, the association of the imputed rs2058878 genotypes with abstinence length in the replication sample was marginally significant (hazard ratio=0.72, P=0.068). As in the discovery sample, the minor A allele was associated with lower risk of relapse (longer abstinence). To gain additional insights into this association signal, we used the available genome-wide SNP data from the replication sample to search for nearby SNPs that were in high linkage disequilibrium (⩾0.9) with rs2058878. We identified the GRIN2B SNP rs2300272 (minor allele frequency=0.48, linkage disequilibrium=0.9 with rs2058878), which was genotyped in the replication sample. Analysis of this SNP revealed association of the minor rs2300272 G allele with shorter abstinence in the replication sample (hazard ratio=1.43, P=0.049).
The association of minor alleles of GRIN2B rs2160734 and rs11612353 (proxy for rs2160733) SNPs with abstinence length was in a reverse direction compared with the discovery sample. The minor GLRB rs11100096 G allele (proxy for rs17035723 A) showed association with treatment outcome in the same direction as the discovery sample; however, the association was not statistically significant.
Discussion
Our findings indicate that the minor GRIN2B rs2058878 A allele is associated with longer abstinence during the first 3 months of acamprosate treatment. This association was replicated in an independent sample with marginally significant evidence of association. Furthermore, the replication sample provided significant evidence for association of abstinence length with rs2300272, which is in high linkage disequilibrium with rs2058878. This is the first report showing the association of these genetic markers with the length of abstinence in alcohol-dependent subjects. It provides a compelling reason to investigate these markers as potential pharmacogenetic predictors of acamprosate treatment response.
GRIN2B rs2058878 was selected as a tag SNP to investigate the association of treatment outcome with corresponding gene segments. It is possible that other rarer variants located in the gene segment ‘tagged’ by rs2058878 and rs2300272 are responsible for physiological mechanisms contributing to the reported associations and a search for such functional variants is necessary. It is also possible that rs2058878 or rs2300272 may have a functional role associated with the expression and/or function of GRIN2B. Indeed, rs2058878 is located in an intronic region where enrichment of histone 3 lysine 4 mono-methylation and histone 3 lysine 27 acetylation was found in several data sets of the UCSC ENCODE database.47 These marks indicate the presence of transcriptional enhancers,37,48 suggesting a role for the region around rs2058878 in the regulation of GRIN2B expression. Interestingly, rs2058878 A allele is part of a canonical E-box sequence identified as a binding site for the transcription factors NeuroD and NeuroG in mouse and Xenopus laevis.49 Review of the Human Brain Transcriptome database (http://hbatlas.org/pages/hbtd) indicates that expression of the GRIN2B gene in human brain areas seems to inversely correlate with expression of the NeuroD and NeuroG transcription factors. These factors are known to have an active role during neural development in humans50 and to act both as transcriptional activators as well as repressors.51,52 However, the predicted consensus site is abolished in the rs2058878 T allele (which is associated with shorter abstinence). Thus, altered transcription factor binding to the rs2058878-containing region and consequent changes in GRIN2B expression could contribute to the association between rs2058878 and duration of abstinence.
The GRIN2B gene encodes the GluN2B subunit of NMDA receptor. Our finding of association between GRIN2B variants and the length of abstinence in alcohol-dependent human subjects treated with acamprosate is in line with experimental evidence associating increased GluN2B subunit expression with chronic ethanol treatment and development of physical dependence.53 Moreover, recent findings indicate that the effects of acamprosate may be related to the Ca2+ ion.54 Providing that Ca2+ influx in neurons is controlled by NMDA receptor, it is of special interest to investigate the effects of rs2058878 or rs2300272 on Ca2+ influx.
Evidence also indicates that GluN2B-containing NMDA receptors are activated by glycine19,20 and are primarily responsible for its intracellular effects36,55 and related behavioral phenotypes, including alcohol intake.56 Moreover, GluN2B-containing NMDA receptors are functionally important for long-term depression.20 Consequently, altered production of GluN2B-containing NMDA receptors may disrupt the balance between long-term depression and long-term potentiation, which are fundamental in brain physiology. Therefore, understanding the effects of rs2058878 and rs2300272 in human alcohol use disorders and animal models of alcohol-related phenotypes is of special interest, especially in the context of experimental evidence suggesting that glycine may have an important role in alcohol dependence and acamprosate response.21,22,57
In addition to investigating genetic variant effects, clinical and demographic variations between study subjects and recruitment sites were investigated as potential predictors of treatment outcomes in our study. Several clinical variables (for example, craving intensity, depressive symptoms and the length of sobriety before initiation of acamprosate treatment) and non-pharmacological treatment components (for example, attendance at Alcoholics Anonymous meetings and having an Alcoholics Anonymous sponsor) were found to be associated with abstinence length. The predictive role of these factors should be accounted for in the pharmacogenomic studies and further investigated in prospective clinical trials. Our findings indicate that tools routinely used for assessment of patients treated in community-based programs, including PACS and Timeline Follow-Back, can be useful for these purposes.
Results of this study should be considered in the context of the following limitations. We have not used a placebo arm in the discovery sample, which limits our ability to differentiate association findings related to acamprosate effects from those associated with sobriety independent from acamprosate effects. Prospective placebo controlled studies are needed to validate the role of our top SNPs as potential pharmacogenomic markers of abstinence length associated with or independent from acamprosate effects.
Analyses presented here are limited to the first 3 months of acamprosate treatment and, therefore, may be relevant only during early stages of abstinence. It is possible that genetic and clinical markers associated with early relapse may be different from those associated with later relapse. This possibility will be investigated in the ongoing analyses focused on genetic and clinical predictors of sobriety during the subsequent 3 months of acamprosate treatment in our study cohort.
Finally, both discovery and replication samples were relatively small, which limited power to discover all meaningful associations. However, analyses resulted in potentially important association findings.
In conclusion, our findings indicate that GRIN2B rs2058878 and rs2300272 SNPs are associated with abstinence length during the first 3 months of acamprosate treatment. These findings support experimental evidence implicating NMDA receptors in the treatment effects of acamprosate. Future studies should prospectively investigate the potential role of these SNPs as biomarkers of abstinence length in treatment-seeking alcoholics and determine the physiological and molecular mechanisms underlying these association findings.
References
Bouchery EE, Harwood HJ, Sacks JJ, Simon CJ, Brewer RD . Economic costs of excessive alcohol consumption in the U.S., 2006. Am J Prev Med 2011; 41: 516–524.
Collins PY, Patel V, Joestl SS, March D, Insel TR, Daar AS et al. Grand challenges in global mental health. Nature 2011; 475: 27–30.
Bouza C, Angeles M, Munoz A, Amate JM . Efficacy and safety of naltrexone and acamprosate in the treatment of alcohol dependence: a systematic review. Addiction 2004; 99: 811–828.
Jonas DE, Amick HR, Feltner C, Bobashev G, Thomas K, Wines R et al. Pharmacotherapy for adults with alcohol use disorders in outpatient settings: a systematic review and meta-analysis. JAMA 2014; 311: 1889–1900.
Rosner S, Leucht S, Lehert P, Soyka M . Acamprosate supports abstinence, naltrexone prevents excessive drinking: evidence from a meta-analysis with unreported outcomes. J Psychopharmacol 2008; 22: 11–23.
Dawson DA, Goldstein RB, Grant BF . Rates and correlates of relapse among individuals in remission from DSM-IV alcohol dependence: a 3-year follow-up. Alcohol Clin Exp Res 2007; 31: 2036–2045.
Rosner S, Hackl-Herrwerth A, Leucht S, Lehert P, Vecchi S, Soyka M . Acamprosate for alcohol dependence. Cochrane Database Syst Rev 2010; CD004332.
Maisel NC, Blodgett JC, Wilbourne PL, Humphreys K, Finney JW . Meta-analysis of naltrexone and acamprosate for treating alcohol use disorders: when are these medications most helpful? Addiction 2013; 108: 275–293.
Mason BJ, Goodman AM, Chabac S, Lehert P . Effect of oral acamprosate on abstinence in patients with alcohol dependence in a double-blind, placebo-controlled trial: the role of patient motivation. J Psychiatr Res 2006; 40: 383–393.
Morley KC, Teesson M, Sannibale C, Baillie A, Haber PS . Clinical predictors of outcome from an Australian pharmacological relapse prevention trial. Alcohol Alcohol. 2010; 45: 520–526.
Kranzler HR, Edenberg HJ . Pharmacogenetics of alcohol and alcohol dependence treatment. Curr Pharm Des 2010; 16: 2141–2148.
Litten RZ, Bradley AM, Moss HB . Alcohol biomarkers in applied settings: recent advances and future research opportunities. Alcohol Clin Exp Res 2010; 34: 955–967.
Chabenat C, Chretien P, Daoust M, Moore N, Andre D, Lhuintre JP et al. Physicochemical, pharmacological and pharmacokinetic study of a new GABAergic compound, calcium acetylhomotaurinate. Methods Find Exp Clin Pharmacol 1988; 10: 311–317.
Dahchour A, De Witte P, Bolo N, Nedelec JF, Muzet M, Durbin P et al. Central effects of acamprosate: part 1. Acamprosate blocks the glutamate increase in the nucleus accumbens microdialysate in ethanol withdrawn rats. Psychiatry Res 1998; 82: 107–114.
Littleton J . Acamprosate in alcohol dependence: how does it work? Addiction 1995; 90: 1179–1188.
Kiefer F, Witt SH, Frank J, Richter A, Treutlein J, Lemenager T et al. Involvement of the atrial natriuretic peptide transcription factor GATA4 in alcohol dependence, relapse risk and treatment response to acamprosate. Pharmacogenomics J 2011; 11: 368–374.
Lee MR, Hinton DJ, Wu J, Mishra PK, Port JD, Macura SI et al. Acamprosate reduces ethanol drinking behaviors and alters the metabolite profile in mice lacking ENT1. Neurosci Lett 2011; 490: 90–95.
Spanagel R, Pendyala G, Abarca C, Zghoul T, Sanchis-Segura C, Magnone MC et al. The clock gene Per2 influences the glutamatergic system and modulates alcohol consumption. Nat Med 2005; 11: 35–42.
Bergeron R, Meyer TM, Coyle JT, Greene RW . Modulation of N-methyl-D-aspartate receptor function by glycine transport. Proc Natl Acad Sci USA 1998; 95: 15730–15734.
Papouin T, Ladepeche L, Ruel J, Sacchi S, Labasque M, Hanini M et al. Synaptic and extrasynaptic NMDA receptors are gated by different endogenous coagonists. Cell 2012; 150: 633–646.
Chau P, Stomberg R, Fagerberg A, Soderpalm B, Ericson M . Glycine receptors involved in acamprosate's modulation of accumbal dopamine levels: an in vivo microdialysis study. Alcohol Clin Exp Res 2010; 34: 32–38.
Lido HH, Marston H, Ericson M, Soderpalm B . The glycine reuptake inhibitor Org24598 and acamprosate reduce ethanol intake in the rat; tolerance development to acamprosate but not to Org24598. Addict Biol 2012; 17: 897–907.
Molander A, Soderpalm B . Accumbal strychnine-sensitive glycine receptors: an access point for ethanol to the brain reward system. Alcohol Clin Exp Res 2005; 29: 27–37.
Molander A, Soderpalm B . Glycine receptors regulate dopamine release in the rat nucleus accumbens. Alcohol Clin Exp Res 2005; 29: 17–26.
Vengeliene V, Leonardi-Essmann F, Sommer WH, Marston HM, Spanagel R . Glycine transporter-1 blockade leads to persistently reduced relapse-like alcohol drinking in rats. Biol Psychiatry 2010; 68: 704–711.
Mann K, Kiefer F, Smolka M, Gann H, Wellek S, Heinz A . Searching for responders to acamprosate and naltrexone in alcoholism treatment: rationale and design of the PREDICT study. Alcohol Clin Exp Res 2009; 33: 674–683.
Hasin DS, Trautman KD, Miele GM, Samet S, Smith M, Endicott J . Psychiatric Research Interview for Substance and Mental Disorders (PRISM): reliability for substance abusers. Am J Psychiatry 1996; 153: 1195–1201.
Sobell L, Sobell M . Timeline follow-back. In: Litten R, Allen J (eds). Measuring Alcohol Consumption. Humana Press: New York, NY, 1992.
Flannery BA, Volpicelli JR, Pettinati HM . Psychometric properties of the Penn Alcohol Craving Scale. Alcohol Clin Exp Res 1999; 23: 1289–1295.
Vander Weg MW, DeBon M, Sherrill-Mittleman D, Klesges RC, Relyea GE . Binge drinking, drinking and driving, and riding with a driver who had been drinking heavily among Air National Guard and Air Force Reserve Personnel. Mil Med 2006; 171: 177–183.
Annis HM, Martin G . Inventory of Drug-Taking Situations. Addiction Research Foundation: Toronto, Canada, 1985.
Annis HM, Turner NE, Sklar SM . Inventory of Drug-Taking Situations: User’s Guide. Addiction Research Foundation, Centre for Addiction and Mental Health: Toronto, Canada, 1997.
Kroenke K, Spitzer RL, Williams JB . The PHQ-9: validity of a brief depression severity measure. J Gen Intern Med 2001; 16: 606–613.
Spitzer RL, Kroenke K, Williams JB, Lowe B . A brief measure for assessing generalized anxiety disorder: the GAD-7. Arch Intern Med 2006; 166: 1092–1097.
Alcoholics Anonymous World Services Inc. Alcoholics Anonymous. www.aa.org, 2014. Accessed 2012.
Cao H, Ren WH, Zhu MY, Zhao ZQ, Zhang YQ . Activation of glycine site and GluN2B subunit of NMDA receptors is necessary for ERK/CREB signaling cascade in rostral anterior cingulate cortex in rats: implications for affective pain. Neurosci Bull 2012; 28: 77–87.
Heilig M, Egli M . Pharmacological treatment of alcohol dependence: target symptoms and target mechanisms. Pharmacol Ther 2006; 111: 855–876.
Kovanen L, Saarikoski ST, Haukka J, Pirkola S, Aromaa A, Lonnqvist J et al. Circadian clock gene polymorphisms in alcohol use disorders and alcohol consumption. Alcohol Alcohol 2010; 45: 303–311.
Lutz UC, Batra A, Kolb W, Machicao F, Maurer S, Kohnke MD . Methylenetetrahydrofolate reductase C677T-polymorphism and its association with alcohol withdrawal seizure. Alcohol Clin Exp Res 2006; 30: 1966–1971.
Sicotte H, Rider DN, Poland GA, Dhiman N, Kocher JP . SNPPicker: high quality tag SNP selection across multiple populations. BMC Bioinformatics 2011; 12: 129.
Kosoy R, Nassir R, Tian C, White PA, Butler LM, Silva G et al. Ancestry informative marker sets for determining continental origin and admixture proportions in common populations in America. Hum Mutat 2009; 30: 69–78.
Oliphant A, Barker DL, Stuelpnagel JR, Chee MS . BeadArray technology: enabling an accurate, cost-effective approach to high-throughput genotyping. Biotechniques 2002; Suppl: 56-58: 60–51.
Treutlein J, Cichon S, Ridinger M, Wodarz N, Soyka M, Zill P et al. Genome-wide association study of alcohol dependence. Arch Gen Psychiatry 2009; 66: 773–784.
Frank J, Cichon S, Treutlein J, Ridinger M, Mattheisen M, Hoffmann P et al. Genome-wide significant association between alcohol dependence and a variant in the ADH gene cluster. Addict Biol 2012; 17: 171–180.
Delaneau O, Marchini J, Zagury JF . A linear complexity phasing method for thousands of genomes. Nat Methods 2012; 9: 179–181.
Howie BN, Donnelly P, Marchini J . A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet 2009; 5: e1000529.
Rosenbloom KR, Sloan CA, Malladi VS, Dreszer TR, Learned K, Kirkup VM et al. ENCODE data in the UCSC Genome Browser: year 5 update. Nucleic Acids Res 2013; 41: D56–D63.
Creyghton MP, Cheng AW, Welstead GG, Kooistra T, Carey BW, Steine EJ et al. Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc Natl Acad Sci USA 2010; 107: 21931–21936.
Seo S, Lim JW, Yellajoshyula D, Chang LW, Kroll KL . Neurogenin and NeuroD direct transcriptional targets and their regulatory enhancers. EMBO J 2007; 26: 5093–5108.
Evsen L, Sugahara S, Uchikawa M, Kondoh H, Wu DK . Progression of neurogenesis in the inner ear requires inhibition of Sox2 transcription by neurogenin1 and neurod1. J Neurosci 2013; 33: 3879–3890.
Itkin-Ansari P, Marcora E, Geron I, Tyrberg B, Demeterco C, Hao E et al. NeuroD1 in the endocrine pancreas: localization and dual function as an activator and repressor. Dev Dyn 2005; 233: 946–953.
Roybon L, Mastracci TL, Ribeiro D, Sussel L, Brundin P, Li JY . GABAergic differentiation induced by Mash1 is compromised by the bHLH proteins Neurogenin2, NeuroD1, and NeuroD2. Cereb Cortex 2010; 20: 1234–1244.
Narita M, Soma M, Mizoguchi H, Tseng LF, Suzuki T . Implications of the NR2B subunit-containing NMDA receptor localized in mouse limbic forebrain in ethanol dependence. Eur J Pharmacol 2000; 401: 191–195.
Spanagel R, Vengeliene V, Jandeleit B, Fischer WN, Grindstaff K, Zhang X et al. Acamprosate produces its anti-relapse effects via calcium. Neuropsychopharmacology 2014; 39: 783–791.
Honer M, Benke D, Laube B, Kuhse J, Heckendorn R, Allgeier H et al. Differentiation of glycine antagonist sites of N-methyl-D-aspartate receptor subtypes. Preferential interaction of CGP 61594 with NR1/2B receptors. J Biol Chem 1998; 273: 11158–11163.
Darstein M, Albrecht C, Lopez-Francos L, Knorle R, Holter SM, Spanagel R et al. Release and accumulation of neurotransmitters in the rat brain: acute effects of ethanol in vitro and effects of long-term voluntary ethanol intake. Alcohol Clin Exp Res 1998; 22: 704–709.
Adermark L, Clarke RB, Olsson T, Hansson E, Soderpalm B, Ericson M . Implications for glycine receptors and astrocytes in ethanol-induced elevation of dopamine levels in the nucleus accumbens. Addict Biol 2011; 16: 43–54.
Acknowledgements
We dedicate this manuscript to the late Dr David Mrazek, the founding director of the Samuel C Johnson Genomics of Addiction program at Mayo and an inspiring leader who relentlessly advocated for the advancement of research in genetic, molecular and biological psychiatry to improve clinical care for alcohol use disorders. We would like to thank Drs M Brunner, M Chauhan, G Melnyk, D Onsrud, B Proctor as well as J Wittkopp, Misty Leidal, BSN, Christine Hanzel, BA, Katrina Schaefer, BA, Evan Loehle-Conger, MA, Sharon Schulz, BSN and Joni Burns-Duenes, BSN for their assistance with patient recruitment and data collection and the Mayo Clinic Psychiatry Information Technology group for their assistance with data management. This publication was supported in part by NIH grants: 1P20AA017830 PIs: Mrazek/Choi (VMK, JMB, JRG, DKH-F, LLL, TDS, MKS, MAF and DSC); RO1 GM28157 PI: Weinshilboum (RMW); U19 GM61388 PI: Weinshilboum (RMW); UL1 TR000135 from the National Center for Advancing Translational Sciences (NCATS) PI: Sundeep Khosla (VMK, JMB, JRG, DKH-F, LLL, TDS, MKS, MAF and DSC). This publication was supported by Mayo Clinic Center for Individualized Medicine (CIM-2012) Grant, PI: Karpyak (VMK), and SC Johnson Genomics of Addictions Program at Mayo Clinic, PI: Choi (VMK, JMB, JRG, DKH-F, LLL, TDS, MKS, MAF and DSC). Study medication (acamprosate) was provided by Forest Pharmaceuticals, which had no other involvement in project planning, conduct, data analyses or preparation of the manuscript. This work was also supported by grant FKZ 01GS08152 from the National Genome Research Network (NGFNplus – see under www.ngfn-alkohol.de) of the German Federal Ministry of Education and Research (BMBF) and by grants FKZ 01GS0117/NGFN and FKZ EB 01011300. KFM was supported by grant 01EB0410 from the Bundesministerium für Bildung und Forschung. MMN received support from the Alfried Krupp von Bohlen und Halbach-Stiftung and is a member of the DFG-funded excellence cluster ImmunoSensation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Translational Psychiatry website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Karpyak, V., Biernacka, J., Geske, J. et al. Genetic markers associated with abstinence length in alcohol-dependent subjects treated with acamprosate. Transl Psychiatry 4, e453 (2014). https://doi.org/10.1038/tp.2014.103
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/tp.2014.103
This article is cited by
-
Managing Pain in Patients With Comorbid Alcohol Use Disorder and Chronic Pain
Current Addiction Reports (2024)
-
TSPAN5 influences serotonin and kynurenine: pharmacogenomic mechanisms related to alcohol use disorder and acamprosate treatment response
Molecular Psychiatry (2021)
-
New steps for treating alcohol use disorder
Psychopharmacology (2018)
-
Metabolomics biomarkers to predict acamprosate treatment response in alcohol-dependent subjects
Scientific Reports (2017)
-
Medikamentöse rückfallprophylaktische Behandlung der Alkoholabhängigkeit
Der Nervenarzt (2017)