Abstract
In this study, RNA sequencing was used to obtain a comprehensive profile of the transcriptomic changes occurring in the mammary gland of lactating sheep suffering from fish oil-induced milk fat depression (FO-MFD). The milk somatic cell transcriptome analysis of four control and four FO-MFD ewes generated an average of 42 million paired-end reads per sample. In both conditions, less than 220 genes constitute approximately 89% of the total counts. These genes, which are considered as core genes, were mainly involved in cytoplasmic ribosomal proteins and electron transport chain pathways. In total, 117 genes were upregulated, and 96 genes were downregulated in FO-MFD samples. Functional analysis of the latter indicated a downregulation of genes involved in the SREBP signaling pathway (e.g., ACACA, ACSL, and ACSS) and Gene Ontology terms related to lipid metabolism and lipid biosynthetic processes. Integrated interpretation of upregulated genes indicated enrichment in genes encoding plasma membrane proteins and proteins regulating protein kinase activity. Overall, our results indicate that FO-MFD is associated with the downregulation of key genes involved in the mammary lipogenesis process. In addition, the results also suggest that this syndrome may be related to upregulation of other genes implicated in signal transduction and codification of transcription factors.
Similar content being viewed by others
Introduction
Sheep milk has a higher content of milk solids than either cow or goat milk, which makes it particularly suitable for the production of cultured dairy products, such as high-quality cheeses. The major proportion of sheep milk fatty acids (FAs) includes saturated FAs (SFA) (>60%), whereas the concentrations of monounsaturated and polyunsaturated FAs (MUFA and PUFA) are much lower (approx. 19–25% and 3–6%, respectively)1,2. Given the beneficial effects of MUFA and PUFA in human health3, different strategies have been developed to modulate the milk FA profile in dairy livestock.
Thus, in dairy sheep, genetic studies have been performed to assess the possible influence of candidate genes implicated in FA desaturation and to identify quantitative trait loci (QTLs) in relation to milk FA composition4,5. In the nutrition field, different feeding strategies have been applied with the aim of increasing milk PUFA rates6,7,8,9,10.
Diet supplementation with marine lipids (namely, fish oil, FO) is an effective tool to modify the milk FA profile by increasing the content of certain bioactive lipids, such as n-3 PUFA and conjugated linoleic acid (CLA)7,8,10. However, this type of supplementation has also been associated with milk fat depression (MFD) syndrome11. In general, this syndrome is characterized by a severe reduction of milk fat content with no changes in milk yield or other milk components12,13. Given that most milk from small ruminants is used for the manufacture of cheeses, MFD hinders the application of this feeding strategy (i.e., the addition of FO to the sheep diet) under practical farm conditions.
The molecular mechanisms underlying marine lipid-induced MFD are not well characterized in ruminants in general and particularly in sheep11,13,14, and MFD nutrigenomic research mainly relies on the evaluation of candidate genes associated with lipid metabolism2,14. However, molecular mechanisms of FO-MFD appear to be complex, and information at the omic level may be very useful to understand these mechanisms. In this regard, next generation RNA sequencing technology (RNA-Seq) offers valuable methodology to unravel previously challenging transcriptomic complexities as it allows complete characterization of the transcriptome and provides the opportunity to quantify transcripts and identify differential regulation between two or more conditions15,16.
Hence, the goal of this study was to elucidate molecular mechanisms underlying FO-induced MFD in dairy sheep using RNA-Seq. To that aim, we compared gene expression profiles of milk somatic cell (MSCs) transcriptomes from control and FO-MFD lactating ewes.
Results
Sequencing and alignment of the ovine MSCs transcriptome
For the four control and four FO-MFD samples sequenced, a total of 42 million paired-end reads per sample were generated on average (mean ± SD = 41,512,563.0 ± 6,396,467.09), with the exception of one control sample that had to be re-sequenced. For this control sample, we acquired two technical replicates of 17,686,309 and 24,006,681 million paired-end reads. Approximately 88.56% of total reads uniquely mapped to the Ovis aries genome (Oar_v3.1).
Power calculations
The power results obtained using the online power analysis tool Scotty (http://scotty.genetics.utah.edu/) are described in Supplementary File S1. According to the parameters indicated in the Material and Methods section, the least expensive experiment to accomplish a differential expression analysis with sufficient power was sequencing five replicates to a depth of 10 million reads aligned to genes per replicate. The most powerful experiment that fits our settings was sequencing 10 replicates to a depth of 40 million reads aligned to genes per replicate. Based on these calculations, the size of our experiment provides adequate power to detect an acceptable proportion of genes differentially expressed under our experimental conditions.
Gene expression level
Gene expression was normalized using the Fragments per Kilobase per Million Mapped Reads (FPKM) method. The total number of expressed genes (genes with at least 0.01 FPKM in one sample15) was 17,134, with an average of 14,195.5 ± 644.51 genes per control sample and 14,422.8 ± 380.09 per FO-MFD sample. According to the FPKM value, expressed genes were classified into low expressed genes (<8 FPKM), middle expressed genes (8–180 FPKM) and highly expressed genes (>180 FKM) (Fig. 1). A total of 219 and 210 genes were identified as highly expressed in the control and FO-MFD samples, respectively. These highly expressed genes were considered ‘core’ genes of the MSCs transcriptome, as they constitute approximately 89% of total FPKM. Among the core genes identified in the two analyzed conditions (control and FO-MFD), a total of 197 genes were identified as highly expressed for both conditions, whereas 22 genes were assigned to the group of highly expressed only in the control and 13 in the FO-MFD samples.
The violin plots are filled in blue (left) and pink (right) for control and fish oil-induced milk fat depression (FO-MFD) samples, respectively. Two transversal black lines separate the three defined groups of expression intensity: low expressed genes (<8 FPKM), middle expressed genes (8–180 FPKM) and highly expressed genes (>180 FPKM).
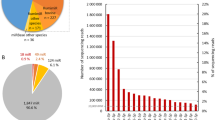
The top 20 genes expressed in the MSCs in the control and FO-MFD supplemented sheep are presented in Fig. 2. These genes constituted on average 80.92% and 81.54% of total FPKM for the control and the FO-MFD samples, respectively. The profile of the top 20 genes was very similar for both conditions, with a considerably increased expression level (involving greater than 20,000 FPKM) for the genes codifying for caseins (CSN1S1, ENSOARG00000010683/CSN1S2, CSN2, CSN3), whey proteins (PAEP and LALBA) and glycosylation-dependent cell adhesion molecule-1 (ENSOARG00000016083/GlyCAM-1) compared with the remaining genes with <10,000 FPKM (Fig. 2).
Graphical representation of the expression levels for the top 20 highly expressed genes in control (blue, left) and FO-MFD (pink, right) ewes are plotted. For each bar graph, Ovis aries gene names (Ovine genome assembly Oar_v3.1) are indicated in the X-axis, and FPKM values are represented in the Y-axis.
To better categorize the core genes, WikiPathways functional enrichment analysis was performed with WebGestalt (Supplementary File S2)17. The top three pathways highlighted by the analysis based on the genes identified as highly expressed in both conditions were cytoplasmic ribosomal proteins (31 genes; padj = 7.75e-57), electron transport chain (9 genes; padj = 4.19e-10) and oxidative stress (3 genes; padj = 0.0025). The 22 genes highly expressed in the control group were enriched in the SREBP signaling pathway (2 genes; padj = 0.0005), and the 13 highly expressed in the FO-MFD sheep in the selenium metabolism and selenoproteins pathway (2 genes; padj = 3.53e-05).
Comparison of transcriptome profiles between control and FO-MFD conditions
Principal component analysis (PCA) performed on the basis of the rlog transformed gene counts revealed a clear difference between the transcriptome profiles of control and FO-MFD ewes (Fig. 3). In the first principal component (PC1), which accumulated 30% of the variance, the control and the FO-MFD gene expression profiles were grouped in distinct clusters. The control samples, which seemed more heterogeneous, fell in the negative direction of the PC1 axis, whereas the FO-MFD samples fall in the positive direction.
PC2 accumulates 19% of the variance. In this component, there were two samples, one control and one FO-MFD sample, which did not cluster with the others likely due to individual variability in the animals.
Differential genetic expression in MSCs between control and FO-MFD conditions
There were 213 differentially expressed genes (DEGs) in the MSC transcriptomes between the control and the FO-MFD sheep. Of these DEGs, 117 genes were upregulated in FO-MFD, and 96 were downregulated in FO-MFD (Supplementary File S3). We performed Gene Ontology (GO) and WikiPathways functional enrichment analyses in the up- and downregulated differentially expressed genes.
Among the genes upregulated in the FO-MFD condition, the significant (padj < 0.05 and a minimum of 5 genes) GO enrichment terms were categorized into 18 functional groups (Supplementary File S4) across the three GO established classes: “biological processes” (3 terms), “molecular function” (6 terms) and “cellular component” (9 terms). The three highest enriched GO terms in each of the categories were: regulation of protein kinase activity (padj = 0.0397), growth factor binding (padj =0.0330) and plasma membrane part (padj = 0.0109) in the biological process, molecular function and cellular component categories, respectively. There were two enriched pathways among the FO-MFD upregulated genes in the WikiPathways analysis (Supplementary File S4?): TGF beta signaling pathway (padj = 3.57e-05; 5 genes) and adipogenesis (padj = 3.57e-05; 5 genes).
Genes downregulated in the FO-MFD condition were significantly clustered (padj < 0.05 and a minimum of 5 genes) in 82 GO functional enrichment terms (Supplementary File S5). Forty terms were significantly enriched in relation to the “biological processes” category, and 22 and 20 GO terms belonged to the “molecular function” and “cellular component” categories, respectively. The highest significantly enriched ‘biological processes’ terms (padj < 1e-05) were associated with lipid metabolisms and lipid biosynthetic processes (Supplementary File S5). The highest enriched GO terms in the “molecular function” and “cellular component” categories were ligase activity, forming carbon-sulfur bonds (padj = 1.81e-07) and cytoplasm (padj = 1.54e-07). The WikiPathways analysis of the 96 downregulated genes for the FO-MFD samples identified one significantly enriched pathway (Supplementary File S5): SREBP signaling (padj = 8.99e-09; 6 genes).
Quantitative Real-Time PCR (qRT-PCR) validation
For the validation of the RNA-Seq results, qRT-PCR analysis was performed in a subset of 12 genes: four upregulated (APP, ATF3, KLF6 and PPT1), four downregulated (ACACA, ACSS1, ACSS2 and LPIN1), and four non-differentially expressed genes (GPAM, GPAT4, SCD and SREBF1). Comparison between fold changes estimated with DESeq2 in the RNA-Seq data and mRNA abundances determined by qRT-PCR exhibited a Pearson correlation coefficient of r = 0.981 (Supplementary File S6). Hence, these 12 genes exhibited similar patterns of mRNA abundance when expression is assessed using these two different technologies.
Discussion
Diet supplementation with FO is an effective strategy to modulate ewe milk fatty acid composition towards a healthier profile8,10. However, the associated MFD constrains the dairy sheep industry due to the concomitant reduction of milk fat concentration. During recent years, RNA-Seq has been progressively used in livestock functional genomics as a revolutionary technology to attempt comprehensive transcriptome analyses that can be useful to detect genetic variants or identify potential biomarkers associated with phenotypes of economic interest18. In this context, the transcriptome analysis presented herein provides a thorough view of DEGs, biological processes and pathways associated with FO-MFD. This knowledge may clarify the molecular mechanisms underlying this syndrome in dairy sheep.
RNA-Seq stranded sequencing was performed in nine RNA samples extracted from MSCs from four FO-MFD and four control lactating ewes. The number of animals used in the experiment was limited due to ethical considerations that recommend following the principle of animal reduction. However, according to the power estimations, the use of four replicates per condition sequenced to a minimum depth per sample of 30 million paired-end reads is adequate for evaluation of the DEGs between the control and FO-MFD ewes (Supplementary File S1).
MSCs are a representative source of lactating sheep mammary gland19. The top-20 expressed genes in the MSCs in this study were consistent with other studies performed on lactating mammary gland, with genes codifying for caseins, whey proteins and GLYCAM1 at the top of the list20,21,22. Moreover, the distribution of genes according to their expression intensity (Fig. 1) was also very similar to that reported in the previously mentioned studies20,21,22, with a small number of genes contributing to a large fraction of the total RNA. This distribution highlights the low complexity of the mammary gland transcriptome during lactation compared with other developmental stages of this organ22.
Results from this study demonstrated that less than 220 genes included within the high gene expression group (>180 FPKM) constitute approximately 89% of total FPKM. These highly expressed genes were considered as ‘core’ genes and were divided into the three following groups: (i) genes identified as highly expressed genes in the two analyzed conditions (197), (ii) genes that were only highly expressed in the FO-MFD (13) and (iii) genes that were only highly expressed in the control condition (22). The most abundant transcripts in MSCs identified in both the control and the FO-MFD sheep were those that codify for major milk proteins. According to the additional functional enrichment analysis performed to better understand the biological role of the other core genes, common genes identified in the two conditions as highly expressed genes were involved in cytoplasmic ribosomal proteins and electron transport chain pathways. These genes were implicated in the transduction processes and the generation of the required energy (ATP) for the massive protein production that occurs in the lactating mammary gland. However, although core genes from both FO-MFD and control sheep were associated with protein synthesis, some biological pathways were uniquely enriched within one of the specific conditions. In the FO-MFD animals, the selenium metabolism and selenoproteins pathway were enriched. Although a report in dairy cows23 suggests that dietary supplementation with selenium may contribute to alleviate MFD, results are not conclusive, and caution should be taken before relating the two core genes (SELENBP1 and FOS) implicated in this pathway and MFD mechanisms. The SREBP signaling pathway, a term enriched among core genes in control animals, is related to fatty acid synthesis. Two genes within this pathway are involved in transcription regulation (INSIG1) and intracellular FA transport (DBI)24.
A total of 213 genes exhibit significantly different expression levels between FO-MFD and control. Interestingly, among the downregulated DEGs in FO-MFD animals, the SREBP signaling pathway was functionally enriched. This observation and the fact that this pathway was also enriched for the core genes of control animals suggest that the lack of stimulation of this SREBP signaling pathway may underlie the MFD mechanisms. The sterol regulatory element-binding protein (SREBP) pathway is a route implicated in lipogenesis25,26, in which the sterol regulatory element-binding transcription factors (SREBFs) and their regulatory SREBF cleavage-activating protein (SCAP) are crucial in the regulation of mRNA expression of enzymes related to de novo synthesis of fatty acids and cholesterol25,27. The SREBP signaling pathway has also been proposed to participate in dietary regulation of lipogenic gene expression during lactation, being critical in MFD14,28. In this analysis, the genes encoding neither SREBFs nor the SCAP protein were identified as differentially expressed. This finding would support the lack of a clear downregulation of the SREBF1 gene observed via qRT-PCR analysis of candidate genes in the same samples29 but contrasts with other studies in sheep and cattle reporting a decreased expression of the SREBF1 gene associated with MFD14,30. In any event, our analysis revealed that several lipogenic and cholesterogenic enzymes regulated by SREBFs (LSS, HMGCS1, ACSS1, MVD, ACACA and FDPS) were downregulated in FO-MFD ewes. This downregulation of the SREBFs target genes without changes in the expression levels of the transcription factors themselves may also suggest an effect at the post-transcriptional instead of at the mRNA level.
Consistent with the WikiPathways analysis, the two highest enriched GO terms among the downregulated genes in FO-MFD animals were also related to lipid metabolism. More precisely, the highest enriched terms were lipid biosynthetic process and lipid metabolic process. However, these GO terms were too general to allow us to draw conclusions regarding metabolic pathways downregulated in the MFD. This finding adds to the complexity of the milk fat synthesis in the mammary gland, with the implication of a complex network of genes24. In this regard, in the hierarchical distribution of GO terms, which encompass the most general to the most specific terms, we tried to identify those specific GO terms that are related to more precise biological processes within the mammary gland lipid metabolism. Hence, we identified the following GO terms: acyl-CoA biosynthetic process (ACSL1, ACSS1, ACSS2, ELOVL6 and ACACA), triglyceride biosynthetic process (ACSL1, PNPLA3, AGPAT2, ELOVL6, ACACA and LPIN1), fatty acid metabolic process (FADS2, ACSL1, ACSS1, AACS, ELOVL6, ACACA and LPIN1) and cholesterol biosynthetic process (LSS, HMGCS1, G6PD, MVD and FDPS). This finding would support the influence of these downregulated biological processes in mammary gland milk fat synthesis and their relation with MFD. Milk fat consists essentially of triglycerides (TAGs) arising from uptake from blood circulating FAs or de novo synthesis31,32 plus a small amount of cholesterol, phospholipids and free FAs32. Our results revealed the downregulation of key genes involved in both de novo synthesis and uptake processes. These genes were associated with the activation of acetoacetate to acetoacetyl-CoA (AACS), activation of FA with CoA (ACSL1, ACSS1, ACSS2), de novo synthesis (ACACA, ELOVL6), desaturation (FADS2) and synthesis of TAGs (AGPAT2, LPIN1), which suggests that the reduction of milk fat content was related to their downregulation. A custom scheme summarizing the principal findings about the downregulated genes in FO-MFD lactating mammary gland is presented in Fig. 4. Some of these genes, specifically ACSS1, ACACA, and LPIN1, were confirmed by qRT-PCR in a previous study on candidate genes using not only the same RNA samples but also additional samples of mammary gland that were biopsied from the same animals29. That study also identified a significant difference in the expression levels of two genes involved in lipid metabolism (FASN and INSIG1) that in our RNA-Seq analysis showed a p-value < 0.05 but did not pass the multiple-adjustment threshold padj < 0.05. Furthermore, although the cholesterol content was not analyzed herein, several genes related to its synthesis were downregulated (LSS, HMGCS1, G6PD, MVD, FDPS), which might allow one to presume a reduction in its content in milk from FO-MFD animals. Although this hypothesis might be supported by a report in cattle suggesting that plant oils inducing MFD would reduce milk cholesterol levels33, caution should be taken when comparing different ruminant species in terms of MFD because ewes, in contrast to cows, are not sensitive to the negative effects of plant oils on milk fat concentration2,6,13.
Principal WikiPathways (top-left) and GO-term results (top-right) from the functional enrichment analysis of downregulated genes in the FO-MFD samples are indicated in green boxes. Blue arrows denote downregulation. Red boxes contain significantly downregulated genes (black) and their function (red). Blue boxes indicate potential products of the proteins codified by the downregulated genes.
The WikiPathways analysis of the upregulated genes in the FO-MFD ewes identified as enriched two pathways: adipogenesis and TGF-beta signaling. The expression of genes involved in the adipogenesis pathway in the analyzed MSCs could be due to the close functional interrelation between the mammary epithelial-stroma and the surrounding adipocytes, which is known as mammary fat pad34. Regarding the TGF-beta signaling pathway, which is implicated in immune response modulation35, a report in mice demonstrated that diet supplementation with FO was associated with increased expression of TGF-beta36, which would merit further research.
Among the results from the GO functional enrichment analysis in the FO-MFD upregulated genes, the large number of genes clustered within GO terms related to the plasma membrane is worthy of mention as it may suggest that dietary FO has an effect on the plasma membrane composition in MSCs. In addition, n-3 PUFA seems to influence lipid raft formation and function in immune cells37. Lipid rafts are combinations of cholesterol, glycosphingolipids and protein receptors organized in glycolipoprotein microdomains that are involved in signal transduction through the activation of signaling pathways involving kinases and phosphatases38. Within our GO term results, we have functionally enriched the biological process regulation of protein kinase activity and some genes clustered in this category that may be related to FO-MFD, such as APP, DUPS1, FGFR1 or ATF3. The interaction of APP, which codifies for amyloid precursor protein, with SREBP1 in the Golgi reticulum has been reported in neurons to inhibit the release of mature SREBP1 (a SREBFs) and its nuclear translocation39. Hence, we hypothesize that if this interaction also occurred in mammary epithelial cells, the target enzymes of the SREBP signaling pathway would be inhibited without downregulation of SREBP1 gene mRNA levels, which is consistent with our results. Thus, the upregulation of the APP gene in the FO-MFD sheep could be a key point in MFD. The DUSP1 gene codifies for dual specificity phosphatase 1 and is upregulated by dietary PUFAs40. The DUSP1 protein is implicated in the inactivation of the mitogen-activated protein kinases (MAPKs) ERK, p38 and JNK41. These kinases phosphorylate SREBFs, enhancing their transcriptional activity40. Thus, the inhibition of MAPKs through DUPS1 decreases the transcriptional activity of SREBFs and therefore lipogenesis. FGFR1 codifies for the fibroblast growth factor receptor 1 and binds to circulating fibroblast growth factor 21 (FGF21)42, which is a potent regulator of metabolism43,44. FGFR21 acts as an inhibitor of the SREBP signaling pathway45, and its expression is modified under different dietary conditions46,47,48. However, it has been reported that the mammary gland is not one of the target tissues of FGF2149. Moreover, to induce FGF21 signaling, FGFR1 must be associated with the co-receptor proteins Klotho or β-Klotho50,51, codified by KL and KLB. In our study, KL was not differentially expressed, and KLB exhibited null expression levels. ATF3 encodes the activating transcription factor 3, a transcriptional repressor. This gene is upregulated under extracellular and intracellular stress conditions52,53. Moreover, ATF3 is a target gene in the regulation of inflammation, immunity and lipid metabolism54,55,56, which are biological processes influenced by n-3 PUFA37,57,58. Finally, within the upregulated genes of the FO-MFD condition, we identified other transcription factors, such as Kruppel-like factors 6 and 11 (KLF6 and KLF11). Contrary to our expectations, KLF6 seems to promote PPARα59, which would enhance SREBP transcriptional activity60. However, consistent with our results, KLF11 was upregulated in a study on the transcriptome adaptation of the bovine mammary gland to diets rich in unsaturated FAs30.
Conclusions
This study presents for the first time in the scientific literature a comprehensive profile of the transcriptomic changes occurring in the mammary gland of lactating sheep suffering from FO-induced MFD. Our results indicate that the reduction in milk fat is associated with the downregulation of key genes involved in the lipogenesis. Moreover, the study suggests that some pathways leading to lipid metabolism downregulation could be related to the upregulation of genes involved in signal transduction and codifying for transcription factors. Overall, these results represent a key application of RNA-Seq technology in ruminant nutrigenomics and provide a great advance in the understanding of functional genomic mechanisms associated with marine lipid-induced MFD in dairy sheep. Our results include a wide list of genes that have not been traditionally analyzed in candidate gene studies in ruminants but may be key regulators in the MFD process and very useful for further research in this field.
Methods
Animals, diets and experimental design
Details of experimental procedures are described in Toral et al. (2016)29. Briefly, eight lactating Assaf ewes at mid lactation (62 ± 2.5 days in milk at the beginning of the assay; parity = 2 ± 0.3) were used. The ewes were individually housed and divided in 2 groups (n = 4) that were assigned to one of 2 dietary treatments: no lipid supplementation (Control) or supplementation with 24 g of fish oil (Afampes 121 DHA; Afamsa, Mos, Spain)/kg of diet dry matter (FO). The basal diet consisted of a total mixed ration based on alfalfa hay (40%) and concentrates (60%). The oil was weekly mixed with the ration, which included molasses to reduce selection of components. All ewes were fed the control diet for a 21-d adaptation period and then both experimental treatments for 40 more days. Samplings for this study were performed at the end of the experiment, when ewes on the FO treatment exhibited a stable decrease in milk fat concentration (monitored daily by infrared spectrophotometric analysis of raw milk samples; ISO 9622:1999). Diets were offered ad libitum, and refusals were removed and weighed each morning. Animals were milked twice daily in a single side milking parlor (DeLaval, Madrid, Spain).
As detailed in the companion study29, feed intake was similar in both groups (on average, 2.72 kg/d), but ewes fed FO exhibited a MFD characterized by a significantly reduced milk fat concentration (in %, 5.87 vs. 4.19, SED = 0.262). On the other hand, milk production was increased in FO-MFD compared with control (1.92 vs 1.47 kg/d respectively, SED = 0.124). Milk FA composition was also different depending on the diet, and details about FA profiles in control and FO-MFD sheep are reported in Toral et al.29.
Collection of milk samples and RNA extraction
Milk samples for RNA extraction were collected on days 38 and 39 on treatments. As previous explained by Suárez-Vega et al.61, to maximize the concentration of mammary epithelial cells within MSCs, the collection was performed approx. 60–70 min after milking and 10 minutes after the injection of oxytocin (5 IU/animal; Facilpart, Laboratorios SYVA, León, Spain). Udders were cleaned with water and soap and then disinfected with povidone iodine. Nipples were also washed with RNAseZap (Ambion, Austin, TX). Individual samples were obtained by hand-milking each half of the mammary gland into an RNAse-free 50-mL tube (2 samples/ewe), which was covered with a sterile gauze to filter the milk. All protocols involving animals were approved by the Animal Welfare Committee of the Instituto de Ganadería de Montaña, the Spanish National Research Council (CSIC) and the Junta de Castilla y León (Spain), following proceedings described in Spanish and EU legislations (R.D. 53/2013, and Council Directive 2010/63/EU). All animals used in this study were handled in strict accordance with good clinical practices, and all efforts were made to minimize suffering.
Samples were held in ice and transferred immediately to the laboratory. Total RNA from MSCs was extracted from 50 mL of fresh milk20,61. Briefly, MSCs were pelleted by centrifugation at 650 × g for 10 min at 4 °C in the presence of a final concentration of 0.5 mM of EDTA. The cell pellet was washed with 10 mL of PBS (pH 7.2 and 0.5 mM of EDTA) followed by an additional centrifugation at 650 × g for 10 min at 4 °C. Washing and centrifugation procedures were repeated twice using 2 and 1.5 mL of the same PBS solution. Then, total RNA was extracted and purified from the milk cell pellet with 500 μL of TRIzol (Invitrogen, Carlsbad, CA).
RNA sequencing
The integrity of the RNA was assessed by the Agilent 2100 Bioanalyzer device (Agilent Technologies, Santa Clara, CA, USA). For each animal, samples with the greatest RNA integrity value (RIN) were used in subsequent analyses. The mean RIN of the selected samples was 8.0 ± 0.13 (range 6.8–8.4). Stranded paired-end libraries with fragments of 300 bp were prepared using the TruSeq Stranded Total RNA Library Prep Kit (Illumina, San Diego, CA, USA). The fragments were sequenced on an Illumina Hi-Seq 2000 sequencer (Fasteris SA, Plan-les-Ouates, Switzerland) according to the manufacturer’s instructions at CNAG (Centro Nacional de Análisis Genómico, Barcelona, Spain), generating stranded paired-end reads of 75 bp. Eight samples from four control and four FO-MFD ewes were sequenced to a minimum depth of 30 million paired-end reads. One of the control samples, which did not reach the minimum depth fixed, was re-sequenced.
Quality control, mapping and quantification
The quality of the raw sequencing data was assessed using FastQC62. STAR aligner v2.4.063 was used to map the reads against the ovine genome assembly v.3.1. (Oar_v3.1; ftp://ftp.ensembl.org/pub/release-83/fasta/ovis_aries/).
For the quantification of gene expression, the annotation of the Oar_v3.1 (ftp://ftp.ensembl.org/pub/release-83/gtf/ovis_aries/) sheep assembly was used as reference. The HTSeq-count v0.6.164 was used to count the number of reads mapped to each gene. Cuffnorm and Cuffquant from Cufflinks package65 were used to obtain normalized FPKM reads.
Power calculations
The web-based software Scotty (http://scotty.genetics.utah.edu/) was used to estimate the power of detection of DE genes of the RNA-Seq experiment. For these calculations, we uploaded the quantification file obtained with HTSeq-count v0.6.164 and fixed the following parameters: a cost per replicate of 200 US Dollars (USD), a cost per million reads aligned to genes of 100 USD, an alignment rate of 85%, a maximum of 10 replicates per condition, a read depth between 10 and 40 million, a maximum cost of the experiment of 100,000 USD, a 50% of differential expressed genes detected with a fold change of 2 and a p-value of 0.05 and a minimum of 50% of genes with at least 50% of maximum power.
Differential expression analysis
Differential gene expression analysis between the milk transcriptomes from FO-MFD and control groups was performed using the DESeq2 (v1.12.4)66 in the R statistical software program (v3.3.1). First, we used DESeq2 to collapse the technical replicates generated for one of the control samples. Then, the program normalized gene count data correcting for library size and RNA composition bias and estimated the dispersion across biological replicates. Finally, pairwise comparison of expression was made between the FO-MFD and the control group for every gene based on a negative binomial model. Fold changes and their associated p-values were obtained. After this step, DESeq2 performs independent filtering, which removes genes that have low counts with the aim of improving the detection power by making the multiple testing adjustment of the p-values less severe. P-values from the subset of genes that pass independent filtering step are adjusted by Benjamini-Hochberg’s approach to correct for multiple testing. Genes with a padj < 0.05 were assigned as differentially expressed. Moreover, DEseq2 was used to perform a principal component analysis (PCA) to cluster samples based on gene expression data.
Gene functional classification: GO term enrichment analysis
The web-based Gene Set Analysis Toolkit (WebGestalt)17 was used to perform a Gene-Ontology (GO) enrichment and WikiPathways analysis. Enriched terms were considered statistically significant when padj was <0.05, and a minimum of five genes were grouped for each significant term. GO terms were categorized in three major functional groups: biological process, molecular function and cellular component.
Quantitative Real-time PCR (qRT-PCR)
To validate the results from the differential expression analysis performed with DESeq2, the mRNA abundances of a subset of 12 genes (4 upregulated, 4 downregulated and 4 non-differentially expressed) were analyzed by qRT-PCR as described in the companion study29 using the primer sequences shown in Supplementary File S7.
Additional Information
How to cite this article: Suárez-Vega, A. et al. Elucidating fish oil-induced milk fat depression in dairy sheep: Milk somatic cell transcriptome analysis. Sci. Rep. 7, 45905; doi: 10.1038/srep45905 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Ramos, M. & Juarez, M. Sheep Milk in Encyclopedia of Dairy Sciences (ed. Fuquay, J. W., Fox, P. F. & McSweeney, P. L. ) Vol. 3, 494–502 (Academic Press, 2011).
Shingfield, K. J., Bonnet, M. & Scollan, N. D. Recent developments in altering the fatty acid composition of ruminant-derived foods. Animal 7 Suppl 1, 132–62 (2013).
Wang, D. D. et al. Association of specific dietary fats with total and cause-specific mortality. JAMA Intern. Med. 176, 1134 (2016).
García-Fernández, M., Gutiérrez-Gil, B., Garcia-Gámez, E. & Arranz, J. J. Identification of single nucleotide polymorphisms in the ovine acetyl-CoA carboxylase-alpha gene. Small Rumin. Res. 90, 34–40 (2010).
García-Fernández, M. et al. Detection of quantitative trait loci affecting the milk fatty acid profile on sheep chromosome 22: role of the stearoyl-CoA desaturase gene in Spanish Churra sheep. J. Dairy Sci. 93, 348–57 (2010).
Castro-Carrera, T. et al. Dietary sunflower oil modulates milk fatty acid composition without major changes in adipose and mammary tissue fatty acid profile or related gene mRNA abundance in sheep. Animal 9, 582–91 (2015).
Reynolds, C. K., Cannon, V. L. & Loerch, S. C. Effects of forage source and supplementation with soybean and marine algal oil on milk fatty acid composition of ewes. Anim. Feed Sci. Technol. 131, 333–357 (2006).
Toral, P. G., Hervás, G., Carreño, D. & Frutos, P. Does supplemental 18:0 alleviate fish oil-induced milk fat depression in dairy ewes? J. Dairy Sci. 99, 1133–44 (2016).
Toral, P. G., Hervás, G., Belenguer, A., Bichi, E. & Frutos, P. Effect of the inclusion of quebracho tannins in a diet rich in linoleic acid on milk fatty acid composition in dairy ewes. J. Dairy Sci. 96, 431–9 (2013).
Toral, P. G. et al. Changes in milk fatty acid profile and animal performance in response to fish oil supplementation, alone or in combination with sunflower oil, in dairy ewes. J. Dairy Sci. 93, 1604–1615 (2010).
Bichi, E., Frutos, P., Toral, P. G., Hervás, G. & Loor, J. J. Dietary marine algae and its influence on tissue gene network expression during milk fat depression in dairy ewes. Anim. Feed Sci. Technol. 186, 36–44 (2013).
Bauman, D. & Griinari, J. Regulation and nutritional manipulation of milk fat: low-fat milk syndrome. Livest. Prod. Sci. 70, 15–29 (2001).
Shingfield, K. J., Bernard, L., Leroux, C. & Chilliard, Y. Role of trans fatty acids in the nutritional regulation of mammary lipogenesis in ruminants. Animal 4, 1140–66 (2010).
Carreño, D., Hervás, G., Toral, P. G., Castro-Carrera, T. & Frutos, P. Fish oil-induced milk fat depression and associated downregulation of mammary lipogenic genes in dairy ewes. J. Dairy Sci. 99, 7971–81 (2016).
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 5, 621–628 (2008).
Wang, Z., Gerstein, M. & Snyder, M. RNA-Seq: a revolutionary tool for transcriptomics. Nat. Rev. Genet. 10, 57–63 (2009).
Wang, J., Duncan, D., Shi, Z. & Zhang, B. WEB-based GEne SeT AnaLysis Toolkit (WebGestalt): update 2013. Nucleic Acids Res. 41, W77–83 (2013).
Wickramasinghe, S., Cánovas, A., Rincón, G. & Medrano, J. F. RNA-Sequencing: A tool to explore new frontiers in animal genetics. Livest. Sci. 166, 206–216 (2014).
Canovas, A. et al. Comparison of five different RNA sources to examine the lactating bovine mammary gland transcriptome using RNA-Sequencing. Sci. Rep. 4, 5297 (2014).
Wickramasinghe, S., Rincon, G., Islas-Trejo, A. & Medrano, J. F. Transcriptional profiling of bovine milk using RNA sequencing. BMC Genomics 13, 45 (2012).
Suárez-Vega, A. et al. Characterization and Comparative Analysis of the Milk Transcriptome in Two Dairy Sheep Breeds using RNA Sequencing. Sci. Rep. 5, 18399 (2015).
Li, C. et al. RNA-Seq reveals 10 novel promising candidate genes affecting milk protein concentration in the Chinese Holstein population. Sci. Rep. 6, 26813 (2016).
Liu, Z. L., Yang, D. P., Chen, P., Dong, W. X. & Wang, D. M. Supplementation with Selenium and Vitamin E Improves Milk Fat Depression and Fatty Acid Composition in Dairy Cows Fed Fat Diet. Asian-Australasian J. Anim. Sci. 21, 838–844 (2008).
Bionaz, M. & Loor, J. J. Gene networks driving bovine milk fat synthesis during the lactation cycle. BMC Genomics 9, 366 (2008).
Rudolph, M. C. et al. Sterol regulatory element binding protein and dietary lipid regulation of fatty acid synthesis in the mammary epithelium. Am. J. Physiol. Endocrinol. Metab. 299 (2010).
Xu, H. F. et al. Overexpression of SREBP1 (sterol regulatory element binding protein 1) promotes de novo fatty acid synthesis and triacylglycerol accumulation in goat mammary epithelial cells. J. Dairy Sci. 99, 783–795 (2016).
Horton, J. D. et al. Combined analysis of oligonucleotide microarray data from transgenic and knockout mice identifies direct SREBP target genes. Proc. Natl. Acad. Sci. USA 100, 12027–32 (2003).
Bionaz, M., Osorio, J. & Loor, J. J. Nutrigenomics in dairy cows: Nutrients, transcription factors, and techniques. J. Anim. Sci. 93, 5531 (2015).
Toral, P. G., Hervás, G., Suárez-Vega, A., Arranz, J. J. & Frutos, P. Isolation of RNA from milk somatic cells as an alternative to biopsies of mammary tissue for nutrigenomic studies in dairy ewes. J. Dairy Sci. 99, 8461–71 (2016).
Ibeagha-Awemu, E. M. et al. Transcriptome adaptation of the bovine mammary gland to diets rich in unsaturated fatty acids shows greater impact of linseed oil over safflower oil on gene expression and metabolic pathways. BMC Genomics 17, 104 (2016).
Bauman, D. E. & Griinari, J. M. Nutritional regulation of milk fat synthesis. Annu. Rev. Nutr. 23, 203–27 (2003).
Garton, G. A. The composition and biosynthesis of milk lipids. J. Lipid Res. 4, 237–54 (1963).
Altenhofer, C. et al. Effects of rapeseed and soybean oil dietary supplementation on bovine fat metabolism, fatty acid composition and cholesterol levels in milk. J. Dairy Res. 81, 120–8 (2014).
Hovey, R. C. & Aimo, L. Diverse and active roles for adipocytes during mammary gland growth and function. J. Mammary Gland Biol. Neoplasia 15, 279–90 (2010).
Li, M. O., Wan, Y. Y., Sanjabi, S., Robertson, A. K. & Flavell, R. A. Transforming growth factor-beta regulation of immune responses. Annu. Rev. Immunol. 24, 99–146 (2006).
Fernandes, G., Bysani, C., Venkatraman, J. T., Tomar, V. & Zhao, W. Increased TGF-beta and decreased oncogene expression by omega-3 fatty acids in the spleen delays onset of autoimmune disease in B/W mice. J. Immunol. 152, 5979–87 (1994).
Calder, P. C. Mechanisms of action of (n-3) fatty acids. J. Nutr. 142, 592S–599S (2012).
Simons, K. & Toomre, D. Lipid rafts and signal transduction. Nat. Rev. Mol. Cell Biol. 1, 31–39 (2000).
Pierrot, N. et al. Amyloid precursor protein controls cholesterol turnover needed for neuronal activity. EMBO Mol. Med. 5, 608–25 (2013).
Kotzka, J. et al. Preventing phosphorylation of sterol regulatory element-binding protein 1a by MAP-Kinases protects mice from fatty liver and visceral obesity. PLoS One 7, e32609 (2012).
Engelbrecht, A.-M. et al. Long-chain polyunsaturated fatty acids protect the heart against ischemia/reperfusion-induced injury via a MAPK dependent pathway. J. Mol. Cell. Cardiol. 39, 940–954 (2005).
Yang, C. et al. Differential specificity of endocrine FGF19 and FGF21 to FGFR1 and FGFR4 in complex with KLB. PLoS One 7, e33870 (2012).
Murata, Y., Konishi, M. & Itoh, N. FGF21 as an endocrine regulator in lipid metabolism: from molecular evolution to physiology and pathophysiology. J. Nutr. Metab. 2011, 981315 (2011).
Woo, Y. C., Xu, A., Wang, Y. & Lam, K. S. L. Fibroblast Growth Factor 21 as an emerging metabolic regulator: clinical perspectives. Clin. Endocrinol. (Oxf). 78, 489–496 (2013).
Zhang, Y. et al. The link between fibroblast growth factor 21 and sterol regulatory element binding protein 1c during lipogenesis in hepatocytes. Mol. Cell. Endocrinol. 342, 41–7 (2011).
Yu, J. et al. Conjugated linoleic acid induces hepatic expression of fibroblast growth factor 21 through PPAR-α. Br. J. Nutr. 107, 461–465 (2012).
Akbar, H., Batistel, F., Drackley, J. K. & Loor, J. J. Alterations in Hepatic FGF21, Co-Regulated Genes, and Upstream Metabolic Genes in Response to Nutrition, Ketosis and Inflammation in Peripartal Holstein Cows. PLoS One 10, e0139963 (2015).
Eissa, L. A., Abdel-Rahman, N. & Eraky, S. M. Effects of omega-3 fatty acids and pioglitazone combination on insulin resistance through fibroblast growth factor 21 in type 2 diabetes mellitus. Egypt. J. Basic Appl. Sci. 2, 75–86 (2015).
Schoenberg, K. M. et al. Plasma FGF21 Is Elevated by the Intense Lipid Mobilization of Lactation. Endocrinology 152, 4652–4661 (2011).
Itoh, N. Hormone-like (endocrine) Fgfs: their evolutionary history and roles in development, metabolism, and disease. Cell Tissue Res. 342, 1–11 (2010).
Long, Y. C. & Kharitonenkov, A. Hormone-like fibroblast growth factors and metabolic regulation. Biochim. Biophys. Acta 1812, 791–795 (2011).
Hai, T., Wolfgang, C. D., Marsee, D. K., Allen, A. E. & Sivaprasad, U. ATF3 and stress responses. Gene Expr. 7, 321–35 (1999).
Hartman, M. G. et al. Role for activating transcription factor 3 in stress-induced beta-cell apoptosis. Mol. Cell. Biol. 24, 5721–32 (2004).
Zmuda, E. J. et al. The roles of ATF3, an adaptive-response gene, in high-fat-diet-induced diabetes and pancreatic beta-cell dysfunction. Mol. Endocrinol. 24, 1423–33 (2010).
Rynes, J. et al. Activating transcription factor 3 regulates immune and metabolic homeostasis. Mol. Cell. Biol. 32, 3949–62 (2012).
Kwon, J.-W. et al. Activating transcription factor 3 represses inflammatory responses by binding to the p65 subunit of NF-κB. Sci. Rep. 5, 14470 (2015).
Sampath, H. & Ntambi, J. M. Polyunsaturated Fatty Acid Regulation of Gene Expression. Nutr. Rev. 62 (2004).
Tai, C. C. & Ding, S. T. N-3 polyunsaturated fatty acids regulate lipid metabolism through several inflammation mediators: mechanisms and implications for obesity prevention. J. Nutr. Biochem. 21, 357–63 (2010).
Bechmann, L. P. et al. Post-transcriptional activation of PPAR alpha by KLF6 in hepatic steatosis. J. Hepatol. 58, 1000–6 (2013).
Pawlak, M., Lefebvre, P. & Staels, B. Molecular mechanism of PPARα action and its impact on lipid metabolism, inflammation and fibrosis in non-alcoholic fatty liver disease. J. Hepatol. 62, 720–733 (2015).
Suárez-Vega, A. et al. Comprehensive RNA-Seq profiling to evaluate lactating sheep mammary gland transcriptome. Sci. Data 3, 160051 (2016).
Andrews, S. Babraham Bioinformatics - FastQC A Quality Control tool for High Throughput Sequence Data. http://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (2012).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Anders, S., Pyl, P. T. & Huber, W. HTSeq - A Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2014).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Acknowledgements
This work was supported by the Spanish Ministry of Economy and Competitiveness (MINECO; AGL2014-54587-R and AGL2015-66035-R projects, co-funded by the European Regional Development Fund). P.G.T. and B.G.G. are funded by two postdoctoral research contracts from MINECO (Juan de la Cierva -JCI-2012-11836- and Ramón y Cajal -RYC-2012-10230 programs, respectively).
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: P.F., G.H. P.G.T. and J.J.A. Performed the experiments: P.G.T., G.H. and P.F. Analyzed the data: A.S.V. and J.J.A. Wrote the paper: A.S.V. and B.G.G. Provided advice and critically reviewed the manuscript: P.F., G.H., J.J.A. and P.G.T. All authors approved the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Suárez-Vega, A., Toral, P., Gutiérrez-Gil, B. et al. Elucidating fish oil-induced milk fat depression in dairy sheep: Milk somatic cell transcriptome analysis. Sci Rep 7, 45905 (2017). https://doi.org/10.1038/srep45905
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep45905
This article is cited by
-
Maternal supplementation with fish oil modulates inflammation-related MicroRNAs and genes in suckling lambs
Tropical Animal Health and Production (2020)
-
Conjugated linoleic acid (CLA)-induced milk fat depression: application of RNA-Seq technology to elucidate mammary gene regulation in dairy ewes
Scientific Reports (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.