Abstract
Pseudomonas aeruginosa move across surfaces by using multiple Type IV Pili (TFP), motorized appendages capable of force generation via linear extension/retraction cycles, to generate surface motions collectively known as twitching motility. Pseudomonas cells arrive at a surface with low levels of piliation and TFP activity, which both progressively increase as the cells sense the presence of a surface. At present, it is not clear how twitching motility emerges from these initial minimal conditions. Here, we build a simple model for TFP-driven surface motility without complications from viscous and solid friction on surfaces. We discover the unanticipated structural requirement that TFP motors need to have a minimal amount of effective angular rigidity in order for cells to perform the various classes of experimentally-observed motions. Moreover, a surprisingly small number of TFP are needed to recapitulate movement signatures associated with twitching: Two TFP can already produce movements reminiscent of recently observed slingshot type motion. Interestingly, jerky slingshot motions characteristic of twitching motility comprise the transition region between different types of observed crawling behavior in the dynamical phase diagram, such as self-trapped localized motion, 2-D diffusive exploration, and super-diffusive persistent motion.
Similar content being viewed by others
Introduction
Type IV Pili (TFP) are nanomotor/motility appendages capable of generating forces in the 100 pN range. These filaments, which function via extension-attachment-retraction cycles, are the strongest linear molecular motors known to date1,2,3,4. TFP are responsible for a rich diversity of bacterial phenomena. P. aeruginosa translocate across surfaces by deploying multiple TFP, resulting in complex motions collectively known as twitching motility5,6,7,8,9,10,11,12,13. TFP also play key roles in bacteria-bacteria interactions and bacterial-surface interactions important to pathogenesis14,15,16,17,18,19,20,21,22,23,24,25,26. TFP driven twitching motility is complex, and can be influenced by many factors, including substrate topography, fluidity and stiffness, and also by surface coating, secretion of biological molecules and oxygen levels27,28,29,30,31,32. Recent work on Neisseria gonorrhoeae suggests that multiple TFP nanomotors can collectively function via a tug-of-war type mechanism33. Moreover, it was shown that Neisseria gonorrhoeae transition between fast and slow motion modes, triggered by oxygen levels and proton motive force32,34,35. In P. aeruginosa. the interactions between TFP and bacterial exopolysaccharides deposited on the surface can guide the organization of microcolonies36.
Analysis of high speed movies of P. aeruginosa reveals that they switch between fast and slow modes, where the fast modes are relatively rare, of short duration, and associated with rotations of the cell (“slingshot”)37. In similar systems it was also shown that within the same population of P. aeruginosa. other distinct dynamical behaviors were observed: while some of the bacteria were consistently performing a super-diffusion motion, others were trapped with a sub-diffusion motility16,17. The relations between these movements and the slingshot-like motion in P. aeruginosa are not known at present.
One poorly understood phenomenon is the emergence of complex twitching motility itself in early surface attached bacterial cells. Pseudomonas cells arrive at a surface with low levels of piliation and TFP activity, which both progressively increase as cells sense the presence of a surface24,25,38,39. How do motility phenomena emerge from these minimal under-piliated conditions, and what are the associated requirements on the TFP nanomotors themselves? The problem is compounded by the general lack of systematic experimental data. Although direct observations of TFP have been made in pioneering experiments using Fluorescence Microscopy by Skerker and Berg in 20015, and later by Zaburdaev et al.30, at present we do not have the full dynamical history of every TFP in a given cell, which makes quantitative analysis difficult.
Here, we establish a multiscale mathematical model of twitching motility bridging the scales from single pilus dynamics and the coordination of the molecular motors to macroscopic properties of bacterial trajectories. We compare the model predictions to experimental trajectories for single cells on glass surface. We do not consider the complications from viscous and solid friction on surfaces in order to asses the minimum degree of complexity needed to reproduce the characteristic motility modes and the slingshot-like jumps observed in the experiments. We find that the main order parameters controlling the type of motility are the average number of surface bound pili and the angular spread of TFP orientation. As schematically displayed in the dynamic phase diagram in Fig. 1, a polarized TFP distribution results in persistent motion, while a wide-spread distribution can lead to persistent, diffusive or trapped behavior. These qualitative observations are robust, although the exact location of the boundaries between the different modes is likely to depend on the properties of the surface, especially in case of nonlinear friction effects such as shear thinning.
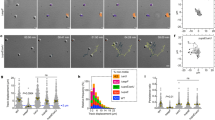
Motility regimes observed at different values of the mean number of surface-bound pili and the angular spread of pili from the cell surface: Diffusive exploration with small number of bound pili pointing into different directions; Trapped localized motion with many bound pili stuck in a tug-of-war situation; and Directionally persistent trajectories with TFP localized at the cell poles and mostly pointing into the same direction. Typical simulation trajectories displayed are 1000 s long and the scale bar is 10 μm. The region where the sling shot jumps are emerging is marked with green.
Slingshot events in the simulations are fast and occur during a small fraction of total time engaged in surface motility (in agreement with experiments). Interestingly, the range of parameters where they are observed (marked green in Fig. 1), is close to the intersection of the three observed crawling modes. This suggests that the slingshots - in addition to facilitating efficient motion on EPS covered surfaces - play an unanticipated important role in mediating changes in motility strategies in response to environmental conditions.
We found that the slingshot jumps can emerge with as little as two TFP bound to the surface, when they are nearly anti-parallel and under high tension (“high-tension antiparallel configuration”, HTAPC) and one of them is suddenly released. With more than two bound pili the motion becomes less jerky, which is consistent with recent findings of Maier et al.31,33, where they modified number of pili on bacteria and observed increased correlation times with increased pili number. However, even in configurations with two bound TFP, not all the sudden jumps in the model resemble the slingshots observed in experiments where the sudden change in speed is correlated with the angular reorientation: we found such correlation only when TFP were very flexible for rotation around the point where they emanate from the surface of the cell. This suggests that the angular flexibility of the motor is a crucial microscopic feature enabling the P. aeruginosa slingshot behaviour.
Model
We designed a multiscale model of P. aeruginosa twitching motility linking the microscopic pili-based mechanisms with the macroscopic properties of bacterial trajectories. The cell is treated as a rigid rod-like body with 12 pili (the number is chosen in accordance with available experiments5,40) emanating from one of its poles5,40,41,42. The pili point to various directions, with equilibrium angles ϕeq that are randomly chosen from a Gaussian distribution with variance ν:  . TFP length changes in time: it is governed by the action of the molecular motors that add or remove pilin monomers resulting in TFP elongation or retraction. The pili are attached to the motor at the cell surface via a flexible hook, while their other end is either free or bound to the surface in a non-specific way (Fig. 2a). In the experiments3, it was established that surface-bound pili predominantly retract - bound pili polymerization was observed only under high tension. However, free pili were observed to equally elongate and retract5,34,43. In line with these observations, we assume that each pilus can be in one of the two modes and that the probability of switching between the elongation and retraction depends on the tension in the filament, which is schematically illustrated in Fig. 2b and detailed in the Methods section. The free pili retract or elongate with a rate 0.8 μm/s based on Skerker and Berg5 and the bound TFP retract with a rate 0.7 μm/s34. Here, we assume that this is also their elongation rate. TFP can shrink to zero length and disappear, however, the total number of pili is conserved and such events are followed by a “birth” of a new pilus in a random direction chosen from the distribution (in another system, of N. gonorrhoeae, it was suggested that there is orientational memory in new pili distribution for explaining directional twitching of bacteria that are assumed to be of spherical symmetry).
. TFP length changes in time: it is governed by the action of the molecular motors that add or remove pilin monomers resulting in TFP elongation or retraction. The pili are attached to the motor at the cell surface via a flexible hook, while their other end is either free or bound to the surface in a non-specific way (Fig. 2a). In the experiments3, it was established that surface-bound pili predominantly retract - bound pili polymerization was observed only under high tension. However, free pili were observed to equally elongate and retract5,34,43. In line with these observations, we assume that each pilus can be in one of the two modes and that the probability of switching between the elongation and retraction depends on the tension in the filament, which is schematically illustrated in Fig. 2b and detailed in the Methods section. The free pili retract or elongate with a rate 0.8 μm/s based on Skerker and Berg5 and the bound TFP retract with a rate 0.7 μm/s34. Here, we assume that this is also their elongation rate. TFP can shrink to zero length and disappear, however, the total number of pili is conserved and such events are followed by a “birth” of a new pilus in a random direction chosen from the distribution (in another system, of N. gonorrhoeae, it was suggested that there is orientational memory in new pili distribution for explaining directional twitching of bacteria that are assumed to be of spherical symmetry).
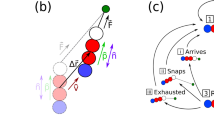
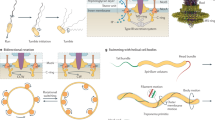
(a) Examples of processes that bacteria can undergo in a KMC simulation step (black line: surface bound pilus, grey line: free pilus). Below is a sketch of a pilus eminating from the cell pole with the visualization of the two elastic terms. (b) The energy landscapes illustrating how the switching rate from a depolymerizing (dep) to a polymerizing (pol) state depends on the stretching tension (top) and how the pulling force on the pilus increases the rate of release from the surface KRel.
TFP are modelled as elastic filaments (Fig. 2a). The elastic energy of a filament with a temporary equilibrium length Leq(t) and actual length Lp(t) oriented in a direction ϕ relative to the cell axes is

where ks = κsLeq/kBT is the linear spring constant that can be estimated from the persistence length of TFP measured in experiments2,5: κs = 10 μN. The filaments bending flexibility was not explicitly considered but is effectively incorporated in the free TFP length. Since bound pili are typically under high tension, significantly bent configurations are unlikely. ϕeq is the orientation of the filament at birth, and ka the angular spring constant measuring the effective flexibility for the filaments to deviate from ϕeq. The values of the effective rotational flexibility ka, as well as the variance of pili distribution ν have not been determined experimentally and are treated as free parameter in our model.
Bacterial trajectories are simulated using the Kinetic Monte Carlo algorithm (KMC)44 considering the following possible processes at each time step (illustrated in Fig. 2a): TFP elongation, retraction, switching between the two, pili rotation, attachment to and release from the surface, and creation of a new pilus (correlated with annihilation of an existing one). At each simulation step, the rate constant for each process is determined and a process is selected with probability proportional to it. The sum of all rate constants determines the lifetime of the present state and thereby the simulation time step44. The chosen processes in the KMC step typically result in stretching or compression of surface bound TFP. After each such process we perform Molecular Dynamics based equilibration of the cell position and orientation in order to minimize the tension created by performing the KMC step. In this way we associate single TFP processes with the motion of the cell and introduce cooperativity among TFP. For each set of the parameter values we run several thousands of single-cell trajectories, 1100 seconds long, and average the results collecting data after 100 seconds. These computer-generated trajectories are then compared to the experimental ones - focusing on statistical properties such as the shape of bacterial speed distribution P(u/u*) and trajectories’ angular persistence.
Results and Discussion
We performed experiments using advanced tracking methods with a time resolution down to 0.1 s. In the numerical simulations we performed a similar number of simulation runs with the same resolution. The details of the experimental and modeling approaches are described in the Materials and Methods section. Quantitative comparison of the results enables us to determine the values of physical parameters that are difficult to assess by direct experiments and apply the additional insight obtained through the modeling to elucidate the microscopic mechanisms of motility.
Slingshot bacteria preform two-legged motility
The motility at large is most notably affected by the number of pili that are typically bound to the surface at the same time. From each simulated trajectory we can define the average such number, 〈NB〉. The value 〈NB〉 ≈ 1 describes a one-leg motility where a single motor action is responsible for cell propulsion, while for larger values  , the propulsion is a result of coordination between the motors pulling on pili and is frequently oppressed due to emergence of tug-of-war configurations. Bacteria speed distribution is thus - not surprisingly - qualitatively different for bacteria with different 〈NB〉. The absolute speed of bacteria in experiment and simulations differs mainly due to the simplicity of the model, e.g. not considering the surface friction. However, by rescaling the speed with the most probable non-zero speed u*, it is still possible to quantitatively compare simlations and experiments. Figure 3a shows three distributions of the rescalled speed, P(u/u*), from the simulations (black) compared to the experimental data (orange). A measure for the discrepancies between the model and experiments,
, the propulsion is a result of coordination between the motors pulling on pili and is frequently oppressed due to emergence of tug-of-war configurations. Bacteria speed distribution is thus - not surprisingly - qualitatively different for bacteria with different 〈NB〉. The absolute speed of bacteria in experiment and simulations differs mainly due to the simplicity of the model, e.g. not considering the surface friction. However, by rescaling the speed with the most probable non-zero speed u*, it is still possible to quantitatively compare simlations and experiments. Figure 3a shows three distributions of the rescalled speed, P(u/u*), from the simulations (black) compared to the experimental data (orange). A measure for the discrepancies between the model and experiments,  , where N is the number of bins used for comparison, is plotted in Fig. 3b showing a clear minimum at 〈NB〉 ≈ 2. Therefore, the best matching trajectories are those where the cells mostly perform two-legged crawling. Figure 3a demonstrates that in case of roughly one-legged crawling, the TFP are often unattached exerting no pulling force on the cell, which results in frequent resting periods and a pronounced peak at zero speed. For higher values of 〈NB〉 there can be frustration due to several bound TFP resulting in trapped configurations with small steps and slow localized speed. Also, in this case TFP length is short (as explained below), resulting in small steps and overall low speed of motion.
, where N is the number of bins used for comparison, is plotted in Fig. 3b showing a clear minimum at 〈NB〉 ≈ 2. Therefore, the best matching trajectories are those where the cells mostly perform two-legged crawling. Figure 3a demonstrates that in case of roughly one-legged crawling, the TFP are often unattached exerting no pulling force on the cell, which results in frequent resting periods and a pronounced peak at zero speed. For higher values of 〈NB〉 there can be frustration due to several bound TFP resulting in trapped configurations with small steps and slow localized speed. Also, in this case TFP length is short (as explained below), resulting in small steps and overall low speed of motion.
(a) Scaled speed distribution P(u/u*) of slingshot bacteria from the experiments (orange) and simulations (black). (b) The discrepancy between the experiments and simulations, χ2, as a function of 〈NB〉. The best match between the simulations and the experiment is achieved when χ2 is minimal, i.e. 〈NB〉 ≈ 2. (c) The mean length of TFP in the simulation, 〈Lp〉, as a function of 〈NB〉. When bound, TFP predominantly retract and do not elongate affecting their average length.
Since the rates of TFP elongation and retraction depend on whether they are bound to the surface or not, the order parameter 〈NB〉 also controls the average length of pili filaments for a given trajectory, as depicted in Fig. 3c. The average length for optimal conditions with 〈NB〉 ≈ 2 is in sub-micron range, which is not in contradiction with the available experimental observations5,41.
The number of bound pili 〈NB〉 depends predominantly on the rates of TFP surface attachment KAd and release KRel, therefore on the chemical properties of pili and surfaces. We have scanned a large parameter space of the KRel and KAd values (14 orders of magnitude of their ratio) and and found that everal combinations of these rates can result in a given 〈NB〉 with an interesting feedback mechanism that results in oscillatory dependence (see Figure S1 on the Online Supplementary Information for a detailed discussion). Comparing simulations and experiments, the best matching speed distribution was obtained with a combination of values: KRel = 130 s−1 and KAd = 5 s−1. These values were used in the rest of the text unless stated otherwise. However, the physical interpretation of these numerical values is difficult since the friction has not been explicitely considered within our model.
Role of tension: slingshot mechanism
Jin et al.37 showed that in the sling-shot motion bacteria change between slow and fast modes, and that while the slow motion is directional, the fast mode (jumps) is often associated with substantial rotation. After obtaining long-time spatially resolved trajectories from both, the experiments and the simulations, we decomposed the trajectories into slow and fast modes of motion in a similar manner as has been done in ref. 37. Jumps are rare events, however, due to the substantial change of the cell position and direction during a jump, they make an important impact on the overall motion. In the simulations we typically observe clearly distinguishable slow and fast modes with an order of magnitude difference in the speed (Fig. 4). As assumed in ref. 37, the fast modes are correlated with the release of a bound pilus. However, only the release events associated with high tension in the filaments lead to fast jumps.
(a) Part of a trajectory from computer simulations. Red dots indicate pilus-release events, blue arrows point on fast jumps, with above-threshold velocity of 0.07 μm/s. Jumps always occur after a release, however not all the release event result in a jump. (b) Time evolution of the x-coordinate and its time derivative. The parameters used here are KRel = 130 s−1, KAd = 5 s−1, ka = 10−14 and ν = π/8.
The tension-mediated cooperative action of TFP is illustrated in Fig. 5 where crawling sequences were isolated from the full trajectories. Each point on the diagram represents a translocation event in the simulation. The instantaneous speed u during the translocation is plotted against the tension in the filaments F/ks. The distinct types of two-legged motion are illustrated above: if picked at random, a likely configuration is that on the left-hand side with two bound TFP pointing into different directions. In such a state the two bound TFP retract by depolymerization which can potentially increase their tension. Assuming that pili cell-attachments are flexible, allowing for free rotation of TFP around them (see Fig. 6), the accompanying stress in the filaments can be relaxed by rotating relative to the cell axis, which results in slow and persistent motion of the cell with relatively small tension in the filaments. Bacteria spend most of the time in this mode. If a pilus is released in such a state, no jump is observed, since there is not enough tension in the filaments. If, however, the cell persists long enough in the slow mode, it moves down the lower branch of points on the speed-tension plot on Fig. 5, reaching an almost antiparallel pili configuration, and then it becomes difficult to relax the tension in the filaments by re-orientation - as the TFP are now puling into opposite directions. This (at the right-hand side of the lower branch) is the so-called high tension antiparallel configuration, HTAPC, indicative of the obstructed motion phase. At this stage, the probability for a pilus release increases - leading to the fast jump, i.e. hopping to the upper branch of the plot in Fig. 5.
The correlation of bacterial instantaneous crawling speed u and the tension in the filaments F/ks for ‘two-legged crawling’ with rotationally flexible TFP. The TFP configuration in various parts of the diagram are illustrated above: If pili are not antiparallel, they can relax the tension by reorientation and the cell motion is slow and persistent. In the HTAPC the tension cannot be relaxed resulting in either obstructed tug-of-war configuration or a fast ‘sling shot’ jump if one of the pili is released. The parameter values are KRel = 130 s−1, KAd = 5 s−1 (corresponding to 〈NB〉 = 1.7), ka = 10−14 and ν = π/8.
(a) Distribution P(θ) of the angular reorientation θ in the fast modes as a function of motor flexibility ka for three values of variance ν: π/20, π/4, and 10π. The pronounced peaks at π/2 are only observed at intermediate ν and low ka. (b) definition of θ, the angle between cell axis and velocity. (c) Comparison of the distribution P(θ) observed in the experiments and in simulations with ν = π/4 and ka = 10−11. (d) The discrepancy χ2 between the experimental observations and simulations in the speed distributions for the fast mode is shown at constant ν = π/4 as a function of the motor flexibility ka (left), and at constant ka = 10−13 as a function of the variance ν (right). In all of the plots KRel = 130 s−1 and KAd = 5 s−1, corresponding to 〈NB〉 ≈ 1.7.
Flexibility-driven twitching
After a jump, the cell axis is oriented approximately in the direction of the remaining bound pilus. Further on, we argue that the reorientation crucially depends on the two key parameters of our study: the rotational flexibility of the motors, ka, and the variance of the distribution of pili equilibrium directions, ν. The evidence for this claim, as provided by simulations, is presented in Fig. 6a where the frequency of jump orientation is plotted as a function of the angular flexibility ka at three different values of the variance ν. The magnitude of reorientation during the sling shot is given by the angle θ between the cell axis and velocity, as in Jin et al.37 (Fig. 6b). In case of very stiff attachment, the angles are more or less uniformly distributed (see large ka values in Fig. 6a) regardless of ν. This is most intuitive as in the case of rotationally stiff attachments high tension could develop in configurations at which the two bound TFP are not necessarily anti parallel, leading to wide distribution of jump angles. For flexible-enough rotations we either observed forward facing jumps with angles around θ = 0 for small values of the variance ν (left), jumps associated with π/2 reorientation (middle) in case of moderately distributed angles, or uniform distribution when ν is large (right). Figure S2(a–c) on the Supplementary Information illustrates typical relocation pathways of bacteria with flexible TFP attachment that lead to the above reorientations. In Fig. 6c a comparison to the experiments is shown for selected values of ν and ka in which the slingshot phenomenon is nicely observed with π/2 reorientation. The discrepancy between the experimental and theoretical distribution of jump reorientation is shown in Fig. 6d, at a fixed value of ν and varying angular elasticity ka, and vice versa. It is evident that only in a narrow regime of parameters, i.e. ka ≤ 10−9 and π/8 ≤ ν ≤ π/3, we observe a quantitative agreement between the experiments and simulations at which slingshot occurs. We also found the best correlation between experiments and simulations in this parameter regime when examining reorientation in the slow motion mode, see Figure S3 in the Supplementary Information. Our findings thus indicate that the angular stiffness must be orders of magnitude smaller than the elastic modulus ks of the TFP filaments in order to enable slingshot jumps with roughly π/2 reorientation.
The reduced angular stiffness in our model can be related to a Young modulus E of the material from which the attachment structure is made of: E = kakBT/L3, where kBT is the thermal energy and L the relevant length scale. Values ka ≤ 10−9 translate to elastic moduli below 1 Pa. This is an unusually small value: For comparison, the elastic moduli of rods made of common flexible materials such as protein fibrils, rubber or bamboo are typically between M Pa and G Pa. However, the fact that the TFP apparatus is anchored on two deformable fluid membranes45 may allow them to exhibit effectively more angular flexibility than that expected from a consideration of only geometry and elasticity of the cellular attachment structure.
Interestingly, we found that the role of motor flexibility does not end in allowing for tension relaxation by reorientation of TFP and approaching HTAPC. Rather, it also controls the orientation of the cell relative to the anti-parallel pili axis just before a jump event, whereby governing the angle θ. The additional insight obtained from modeling allows us to discuss this claim in more detail. Once a cell is stuck in the unfavourable HTAPC, the opposing forces are balanced resulting in vanishing net torque on the cell. At this point rotation of the cell body relative to the bound TFP axis becomes governed by the elasticity: the cell acts like an overdamped harmonic pendulum oscillating around the pili motor positions with a frequency  − until a pilus is released from the surface and the cell reorients into the direction of the remaining bound TFP. The time scale τ should be compared to the mean lifetime of bound TFP, τB, which depends on the detachment probability. At ka ≈ 10−9, our simulations show that τ ≈ 1 s and τB ≈ 0.05 s. For larger values of ka, the two time scales become comparable. Therefore, in the flexible case the jump reorientation is practical identical to the angle with which the cell entered the obstructed phase, while for orientationally stiff motors the cell has enough time to reorient and vary its angle relative to the TFP axis within the obstructed phase (see Figure S2(d) on the Online Supplementary Information). The exact value of the crossover angular elasticity
− until a pilus is released from the surface and the cell reorients into the direction of the remaining bound TFP. The time scale τ should be compared to the mean lifetime of bound TFP, τB, which depends on the detachment probability. At ka ≈ 10−9, our simulations show that τ ≈ 1 s and τB ≈ 0.05 s. For larger values of ka, the two time scales become comparable. Therefore, in the flexible case the jump reorientation is practical identical to the angle with which the cell entered the obstructed phase, while for orientationally stiff motors the cell has enough time to reorient and vary its angle relative to the TFP axis within the obstructed phase (see Figure S2(d) on the Online Supplementary Information). The exact value of the crossover angular elasticity  further depends on effects not considered here, e.g. friction between the cells and surfaces due to nonspecific interactions. Still, our conclusions should qualitatively remain correct.
further depends on effects not considered here, e.g. friction between the cells and surfaces due to nonspecific interactions. Still, our conclusions should qualitatively remain correct.
Another observation to note is that the initial angle with which the cell enters the obstructed phase, depends on the variance ν: if the motors are broadly distributed on the surface of the cell, the initial angles are already smeared out and the jump reorientation angles will be uniformly distributed no matter how flexible the motors are. On the other hand, if the pili angles are very focused, a typical jump will have a forward direction, θ ≈ 0 (see Fig. 2 in the Online Supporting Information). It is only in the intermediate case (roughly, π/6 ≤ ν ≤ π/3) that we did observe sling shots with about π/2 reorientation.
The above discussion is focused on the typical case with two bound pili. When more than two bind the surface, due to geometrical restrictions, the efficiency of rotation in relaxing the tension becomes limited and the tension in the filaments builds up faster. As a result, highly stretched configurations are observed in a large span of pili orientations and the jumps are not associated predominantly with π/2 re-orientations (regardless of ka and ν), as shown in Fig. 7.
Frequency of the fast mode reorientation angles θ at optimal sling shot parameters ν = π/4 and ka = 10−11: The events are separated into those involving two (a) or three (b) TFP - clearly demonstrating that the experimentally observed slingshots result from two-pili processes. (c): Simulation snapshot showing a configuration of high tension with three bound pili. In such cases the tension in the filaments cannot be relaxed by reorientation and thus prior to the release the TFP orientations are broadly distributed - in the depicted snapshot the release event resulted in a forward jump. The scale bar at the bottom is 0.2 μm.
Crawling modes
Based on the above knowledge regarding the high flexibility of TFP attachment and the important role of pili tension, in this section we zoom out from the specific slingshot phenomenon to a wide scan of parameter space for exploring general features of the twitching motion. We systematically explored the characteristics of the single cell motility upon variation of 〈NB〉 (by changing KAd and KRel) and ν. In Fig. 1 we show a schematic dynamic phase diagram, which is based on several thousand simulation trajectories where we evaluated the mean square displacement (MSD) with the same protocol as in recent experiments16,17. We observe three distinct distinct types of trajectories: diffusive, trapped and persistent. The super-diffusive motion is defined here by MSD power larger than 1.1, in the diffusive phase the power is 1 ± 0.03, and the trapped (sub-diffusive mode) is with power less than 0.9. The MSD curves are shown in the Supplementary information (Figure S5), where we also list the exponents and the relevant simulation parameters (Table S1).
Not surprisingly, a very localized distribution of TFP angles results in a persistent super-diffusive crawling mode, as the bound TFP pull the cell forward in a cooperative manner, while at a larger values of the spread ν the type of motion depends on 〈NB〉: Trajectories with 〈NB〉 larger than two are mostly trapped due to the opposing forces pulling in different directions. The diffusive motion appears in a relatively narrow range  and is governed by single TFP retractions in arbitrary directions. Further decrease to 〈NB〉 ≈ 1 leads to super diffusive motion due to pili reorientation in the direction of the cell axis causing an effective narrowing of the angular distribution (see Figure S4 of the Supplementary Information for more details). The motion in this case is directional but slow due to the high frequency of periods with no attached TFP.
and is governed by single TFP retractions in arbitrary directions. Further decrease to 〈NB〉 ≈ 1 leads to super diffusive motion due to pili reorientation in the direction of the cell axis causing an effective narrowing of the angular distribution (see Figure S4 of the Supplementary Information for more details). The motion in this case is directional but slow due to the high frequency of periods with no attached TFP.
From Fig. 1 we see that bacteria with sufficiently broad angular distribution of TFP (delocalized distribution of molecular motors) can exchange between three motion phases by altering the number of bound pili - indicating a potential ability to adapt the surface exploration strategy according to their needs (see Figure S4 in the Supporting Information). On the other hand, hyper-piliated bacteria with  could trigger the transition from super- to sub-diffusive mode by delocalizing the molecular motors - thereby increasing the angular spread of TFP. It is interesting to observe that also the slingshot jumps, despite being rare, play a role in enabling bacteria to switch between the motion phases. The slingshots are observed in the persistent part of the phase diagram with 〈NB〉 ≈ 2 and ν ≈ π/5, which is just below the crossroads between the crawling modes (green region in Fig. 1) and emerge from the antiparallel configuration (HTAPC), which effectively broadens the angular TFP distribution (see S6 in the online Supporting Information). Therefore, the slingshots are correlated with switching from persistent to diffusive or trapped motility modes.
could trigger the transition from super- to sub-diffusive mode by delocalizing the molecular motors - thereby increasing the angular spread of TFP. It is interesting to observe that also the slingshot jumps, despite being rare, play a role in enabling bacteria to switch between the motion phases. The slingshots are observed in the persistent part of the phase diagram with 〈NB〉 ≈ 2 and ν ≈ π/5, which is just below the crossroads between the crawling modes (green region in Fig. 1) and emerge from the antiparallel configuration (HTAPC), which effectively broadens the angular TFP distribution (see S6 in the online Supporting Information). Therefore, the slingshots are correlated with switching from persistent to diffusive or trapped motility modes.
Conclusions
We have designed a coarse-grained model to simulate early stage TFP motility of P. aeruginosa on surfaces and quantitatively compared the model predictions to the experiments performed on the cells tracked by high-speed video microscopy. Full quantitative comparison is not the goal since we do not account for interactions with EPS or solid friction. However, our single-pilus-resolved model allowed us to see how complex motility behavior reminiscent of twitching can result from a remarkably small number (as little as 2) of pili simultaneously bound to the surface. We argue that this condition is commonly met in the early stages of bacterial surface adaptation when the cells are underpiliated. Thus twitching bacteria should emerge from the group once single cells within a group exhibit two or more attached TFP. In later stages of adaptation, with multiple pili per cell, such a scenario can still occur if individual TFP attach weakly to the surface - similar to the low-affinity, high-avidity mechanism of surface attachment that has recently been proposed by Dutcher et al.12 as key to mediating bacterial surface interactions.
The elastic properties of the TFP filaments are key to understanding their collective dynamics12. In our model the slingshot jumps only appear if TFP are able to rotate almost freely around their attachment points on the cells indicating that the elastic penalty for stretching TFP has to be several orders of magnitude larger than that for their rotation. Such effective angular flexibility in the coarse-grained model could be explained either by assuming lateral mobility or sufficient structural flexibility of the molecular motors. Recent detailed analysis of the molecular structure of the motor45 suggests that the motors are unlikely to be laterally mobile, while the presence of flexible linkers in the structure45 makes the assumption of the structural flexibility giving rise to the emergent slingshots plausible. This unanticipated role of flexibility is reminiscent of the buckling instability of the flagellar hook that has been proposed as the key mechanism for bacterial tumbling46,47.
The dynamical phase diagram in Fig. 1 is helpful for visualizing the relations between different motility behaviors in early stage of P. aeruginosa surface adaptation. Our findings point to the ability of bacteria to switch between surface exploration modes by tuning the distribution/orientation of molecular motors on the cell or the number of surface-attached TFP. We predict that a large number of attached TFP will result in a predominantly sub-diffusive motion - qualitatively consistent with previous studies of Deziel et al.48 who found that hyper-piliated mutants exhibit trapped motion. Moreover, as already argued by Weiss40, bacteria may naturally be able to change the number of pili in order to adapt to environmental conditions. The ability to tune the exploration strategy on a single cell level should naturally result in differentiation of bacterial colonies. Indeed, subpopulations exhibiting sub- and super-diffusive behavior have been observed in P. aeruginosa colonies16,17,28. In those experiments, we expect behavior to be influenced by secreted exopoplysaccharides such as Pel and Psl. However, it is interesting that we can see both of these regimes in the phase diagram of a minimal system without the effect of EPS.
Materials and Methods
Experiments
Flow cells (Denmark Technical University) used here were prepared and sterilized using a standard protocol. After injection of bacterial cultures, the flow cell was typically placed for 15 mins, allowing the attaching of cells to coverslip, afterward, unattached cells were washed out. These surface associated cells were further cultured at 30.0 ± 0.1 °C by flowing (3.0 ml/h) FAB mediums. Meanwhile, an inverted microscope (Olympus IX81) equipped with a 100× oil objective was applied to continuously monitor the surface motility of single cells. The dataset, typically containing 20,000 bright-field images, was acquired within 10 fps in first 6 hrs using a sCMOS camera (Andor Neo). These experiments were repeated multiple times in each identical conditions, thus over 100 datasets (>2 · 106 images) were acquired and analyzed in total, in which the 1800 × 1800 16-bit greyscale images were firstly converted to binary images for detection of bacterial cell with a standard image processing algorithm coding by MATLAB, and the x − y positions of cell centroids were subsequently determined and linked individually over time by using an efficient particle tracking algorithm we recently developed16,17,37. As already observed in ref. 37, the motion can be categorized into two modes: the slow persistent mode with the mean speed below 0.07 μm/s and the fast mode associated to sudden change of the direction.
Modeling tension-dependent rate constants
Clausen et al.34 showed that under low stretching force bound TFP predominantly retract, while increasing the force enhances the probability of TFP elongation. To model the switching dynamics between these two states, we assume an energy landscape at which the two modes are represented by minima separated by a barrier (Fig. 2b). The energy level corresponding to the retracting mode is lower than the elongation level and does not depend on the force, while the elongation level linearly decreases with the stretching force up to the stall force of the pilus motor, Fstall = 100 pN2,3. The transition rates, in agreement with experimental findings34, are assumed to be

The probability for pilus release is assumed to increase with the force (see illustration in Fig. 2b). :

with KRel the release constant of the TFP from the surface. The adhesion rate, rAd = KAd is a force-independent parameter.
Additional Information
How to cite this article: Brill-Karniely, Y. et al. Emergence of complex behavior in pili-based motility in early stages of P. aeruginosa surface adaptation. Sci. Rep. 7, 45467; doi: 10.1038/srep45467 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Merz, A. J., So, M. & Sheetz, M. P. Pilus retraction powers bacterial twitching motility. Nature 407, 98–102 (2000).
Maier, B. et al. Single pilus motor forces exceed 100 pN. Proc. Natl. Acad. Sci. USA 99, 16012–16017 (2002).
Clausen, M., Jakovljevic, V., Sogaard-Andersen, L. & Maier, B. High-force generation is a conserved property of type IV pilus systems. J. Bacteriol. 191, 4633–4638 (2009).
Baker, J. L., Biais, N. & Tama, F. Steered Molecular Dynamics Simulations of a Type IV Pilus Probe Initial Stages of a Force-Induced Conformational Transition. PLoS Comput. Biol. 9, e1003032 (2013).
Skerker, J. M. & Berg, H. C. Direct observation of extension and retraction of type IV pili. Proc. Natl. Acad. Sci. USA 98, 6901–6904 (2001).
Bradley, D. E. Function of Pseudomonas-Aeruginosa Pao Polar Pili - Twitching Motility. Can. J. Microbiol. 26, 146–154 (1980).
Henrichsen, J. Twitching Motility. Annu. Rev. Microbiol. 37, 81–93 (1983).
Burrows, L. L. Weapons of mass retraction. Mol. Microbiol. 57, 878–888 (2005).
Kaiser, D. Bacterial motility: How do pili pull? Curr. Biol. 10, R777–R780 (2000).
Mattick, J. S. Type IV pili and twitching motility. Annu. Rev. Microbiol. 56, 289–314 (2002).
Merz, A. J. & Forest, K. T. Bacterial surface motility: slime trails, grappling hooks and nozzles. Curr. Biol. 12, R297–R303 (2002).
Lu, S. et al. Nanoscale Pulling of Type IV Pili Reveals Their Flexibility and Adhesion to Surfaces over Extended Lengths of the Pili. Biophys. J. 108, 2865–2875 (2015).
Strom, M. S. & Lory, S. Structure-function and biogenesis of the type IV pili. Annu. Rev. Microbiol. 47, 565–596 (1993).
Burrows, L. L. Pseudomonas aeruginosa Twitching Motility: Type IV Pili in Action. Annu. Rev. Microbiol. 66, 493–520 (2012).
O’Toole, G. A. & Kolter, R. Flagellar and twitching motility are necessary for Pseudomonas aeruginosa biofilm development. Mol. Microbiol. 30, 295–304 (1998).
Conrad, J. C. et al. Flagella and Pili-Mediated Near-Surface Single-Cell Motility Mechanisms in P. aeruginosa. Biophys. J. 100, 1608–1616 (2011).
Gibiansky, M. L. et al. Bacteria Use Type IV Pili to Walk Upright and Detach from Surfaces. Science 330, 197–U50 (2010).
Chiang, P. & Burrows, L. L. Biofilm formation by hyper-piliated mutants of Pseudomonas aeruginosa. J. Bacteriol. 185, 2374–2378 (2003).
Klausen, M. et al. Biofilm formation by Pseudomonas aeruginosa wild type, flagella and type IV pili mutants. Mol. Microbiol. 48, 1511–1524 (2003).
Singh, P. K., Parsek, M. R., Greenberg, E. P. & Welsh, M. J. A component of innate immunity prevents bacterial biofilm development. Nature 417, 552–555 (2002).
Semmler, A. B., Whitchurch, C. B. & Mattick, J. S. A re-examination of twitching motility in Pseudomonas aeruginosa. Microbiology+ 145, 2863–2873 (1999).
Conrad, J. C. Physics of bacterial near-surface motility using flagella and type IV pili: implications for biofilm formation. Res. Microbiol. 163, 619–629(2012).
Wang, S., Parsek, M. R., Wozniak, D. J. & Ma, L. Z. A spider web strategy of type IV pili-mediated migration to build a fibre-like Psl polysaccharide matrix in Pseudomonas aeruginosa biofilms. Environ. Microbiol. 15, 2238–2253 (3013).
Persat, A., Inclan, Y. F., Engel, J. N., Stone, H. A. & Gitai Z. Type IV pili mechanochemically regulate virulence factors in Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 24, 7563–7568 (2015).
Persat, A. et al. The Mechanical World of Bacteria. Cell 161, 988–997 (2015).
Maier, B. & Wong, G. C. L. How Bacteria Use Type IV Pili Machinery on Surfaces. Trends in Microbiology 23, 775–788 (2015).
Maier, B. The bacterial type IV pilus system – a tunable molecular motor. Soft Matter 9, 5667–5671 (2013).
Zhang, R., Ni, L., Jin, Z., Li, J. & Jin, F. Bacteria slingshot more on soft surfaces. Nature Communications 5, 5541 (2014).
Meel, C., Kouzel, N., Oldewurtel, E. R. & Maier, B. Three-Dimensional Obstacles for Bacterial Surface Motility. Small 8, 530–534 (2012).
Zaburdaev, V. et al. Uncovering the Mechanism of Trapping and Cell Orientation during Neisseria gonorrhoeae Twitching Motility. Biophysical Journal 107, 1523–1531 (2014).
Holz, C. et al. Multiple Pilus Motors Cooperate for Persistent Bacterial Movement in Two Dimensions. Phys. Rev. Lett. 104, 178104 (2010).
Kurre, R. & Maier, B. Oxygen Depletion Triggers Switching between Discrete Speed Modes of Gonococcal Type IV Pili. Biophysical Journal 102, 2556–2563 (2012).
Marathe, R. et al. Bacterial twitching motility is coordinated by a two-dimensional tug-of-war with directional memory. Nature Communications 5, 3759 (2014).
Clausen, M., Koomey, M. & Maier, B. Dynamics of Type IV Pili Is Controlled by Switching Between Multiple States. Biophys. J. 96, 1169–1177 (2009).
Kurre, R., Kouzel, N., Ramakrishnan, K., Oldewurtel, E. R. & Maier, B. Speed switching of gonococcal surface motility correlates with proton motive force. PLoS One 8, e67718 (2013).
Zhao, K. et al. Psl trails guide exploration and microcolony formation in Pseudomonas aeruginosa biofilms. Nature 497, 388–391 (2013).
Jin, F., Conrad, J. C., Gibiansky, M. L. & Wong, G. C. L. Bacteria use type-IV pili to slingshot on surfaces. Proc. Natl. Acad. Sci. USA 108, 12617–12622 (2011).
OToole, G. A. & Wong, G. C. L. Sensational Biofilms: Surface Sensing in Bacteria. Current Opinion in Microbiology 30, 139–146 (2016).
Luo, Y. et al. A Hierarchical Cascade of Second Messengers Regulates Pseudomonas aeruginosa Surface Behaviors. mBio 6, 1–11 (2015).
Weiss, R. L. The structure and occurrence of pili (fimbriae). on Pseudomonas aeruginosa. J. Gen. Microbiol. 67, 135–143 (1971).
Rheinlaender, J., Grabner, A., Ott, L., Burkovski, A. & Schaffer, T. E. Contour and persistence length of Corynebacterium diphtheriae pili by atomic force microscopy. Eur. Biophys. J. 41, 561–570 (2012).
Nudleman, E., Wall, D. & Kaiser, D. Polar assembly of the type IV pilus secretin in Myxococcus xanthus. Mol. Microbiol. 60, 16–29 (2006).
Maier, B., Koomey, M. & Sheetz, M. P. A force-dependent switch reverses type IV pilus retraction. Proc. Natl. Acad. Sci. USA 101, 10961–10966 (2004).
Gillespie, D. T. Exact Stochastic Simulation of Coupled Chemical-Reactions. J. Phys. Chem. 81, 2340–2361 (1977).
Chang, Y.-W. et al. Architecture of the type IVa pilus machine. Science 351, 1165 (2016).
Son, K., Guasto, J. S. & Stocker, R. Bacteria can exploit a flagellar buckling instability to change direction. Nature Physics 9, 494–498 (2013).
Berg, H. C. Turning failure into function. Nature Physics 9, 460–461 (2013).
Deziel, E., Comeau, Y. & Villemur, R. Initiation of Biofilm Formation by Pseudomonas aeruginosa 57RP Correlates with Emergence of Hyperpiliated and Highly Adherent Phenotypic Variants Deficient in Swimming, Swarming, and Twitching Motilities. J. Bacteriol. 183, 1195–1204 (2001).
Acknowledgements
We acknowledge Francisco J. Martinez-Veracoechea for enlightening discussions and his contribution to developing the model. This work was supported by European Union through FP7 grants ARG-ERC-COLSTRUCTION 227758 and ITN-COMPLOIDS 234810 and Horizon 2020 grant ETN-NANOTRANS 674979, by the National Science Foundation of China (21274141 and 21474098), by the Aspen Center for Physics and Kavli Institute for Theoretical Physics (National Science Foundation grants PHY-1066293 and PHY11-25915, respectively). GW acknowledges the support from the Human Frontiers Science Program (RGP0061/2013) and NSF DMR-1411329.
Author information
Authors and Affiliations
Contributions
J.D., D.F., G.C.L.W. and Y.B.-K. conceived the work, Y.B.-K. performed computer simulations, F.J. performed the experiments, Y.B.-K., J.D., D.F. and G.C.L.W. analysed the data, discussed the results and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Brill-Karniely, Y., Jin, F., Wong, G. et al. Emergence of complex behavior in pili-based motility in early stages of P. aeruginosa surface adaptation. Sci Rep 7, 45467 (2017). https://doi.org/10.1038/srep45467
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep45467
This article is cited by
-
Modelling bacterial twitching in fluid flows: a CFD-DEM approach
Scientific Reports (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.