Abstract
Mice are commonly used to study the temporomandibular joint (TMJ) and to model human TMJ disease. However, evaluating TMJ pathology in mice using standard histologic methods is time consuming, labor intensive, and dependent upon investigators’ expertise at consistently orienting and sectioning across tiny specimens. We describe a method that uses confocal microscopy to rapidly and reliably assess indicators of mandibular condyle cartilage pathology in mice. We demonstrate the utility of this method for detecting abnormalities in chondrocyte distribution in mice lacking lubricin (Prg4), the major boundary lubricant of articular cartilage. We further show that the method can provide information about recombination sites and efficiency in mandibular cartilage for Cre-driver strains. Because specimen preparation and data acquisition with confocal microscopy are simple and fast, the method can serve as a primary screening tool for TMJ pathology, before proceeding to complicated, time consuming, secondary analyses.
Similar content being viewed by others
Introduction
The temporomandibular joint (TMJ) is a specialized synovium-enclosed articulating structure composed of the mandibular condyle, an interposed fibrocartilaginous disc, and the glenoid fossa1. The TMJ is needed for eating and speaking, making it among the most used joints. Like other synovial joints, the TMJ can be affected by inflammatory, traumatic, infectious, developmental, and neoplastic diseases. In fact, at least one sign of TMJ disease is present in 40 to 75% of US adults2.
Mice and humans share many developmental and molecular pathways, which led to mice being used for modeling human TMJ diseases3. However, methods for evaluating the consequences of TMJ disease in mice are challenging because of the small size of this joint. Therefore, to facilitate TMJ studies in mice, we developed a simple and fast confocal microscopic imaging method for the mandibular condyle. Herein, we describe the imaging method and demonstrate its utility by studying wild-type (WT) and lubricin (Prg4) deficient mice, the latter modeling the human genetic disease Camptodactyly-Arthropathy-Coxa vara- Pericarditis (CACP) syndrome4, and by determining the pattern of Cre-mediated recombination for the AcanCreERt2 mouse strain5. This strain, which has a tamoxifen-inducible Cre-recombinase knocked into the Aggrecan (Acan) locus, is commonly used to study articular cartilage.
Results
Confocal imaging of a mouse mandibular condyle
Each specimen could be loaded and imaged for two fluoroscopic channels in less than 6 minutes. Representative three dimensional (3D) reconstructed confocal images for the middle region of the mandibular cartilage from a 6-week-old WT mouse are shown in Fig. 1. DAPI stained and EdU containing chondrocyte nuclei were reliably detected to a depth of 50 μm beneath the articulating surface. Therefore, measurements of chondrocyte number and condyle volume were performed using a 760 × 760 × 50 μm field. In order to determine the coefficient of variation (CV) for counting the number of chondrocyte nuclei, for measuring cartilage volumes, and for determining the number of cells per unit volume, we imaged condyles from 3 WT mice 10 times each. Every CV was less than 0.11 and most of them were less than 0.05, which led us to conclude that imaging a specimen 3 times would be sufficient to obtain an average value that would fall close to the true value. We then imaged 10 WT 6-week-old male mice. In these genetically identical mice, the standard deviations (SD) for cell number and cartilage volume were ∼14% of the mean, whereas the SDs for the number of cell nuclei/cartilage volume (hereafter termed “cell density”) were ∼7% of the mean for the cartilage volume lying within 10 μm of the articular surface, the “superficial cell density” (Table 1 and Supplementary Table 1). We chose the volume 10 μm closest to the condylar surface based on prior work that showed these cells express Dkk3 whereas deeper cells do not6,7. Thus, as few as 4 mice per group could be used to detect a ≥ 20% difference in superficial cell density between experimental and control animals (Table 1). All subsequent experiments considered measures of cell density, since larger sample sizes would be required to observe significant differences in chondrocyte number or cartilage volume (Supplementary Table 1). The other cell density measure we considered was the number of nuclei in the imaged cartilage volume from 10 μm below the surface to the bottom of the imaged cartilage, the “deeper cell density”.
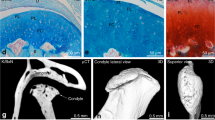
(a) Side view of the condyle specimen holder showing [1] a 0.05 mm diameter stainless steel wire (Ormco, Orange, CA) that has been bent 180° in order to clasp the neck of the condyle and keep it in line with the coracoid process, [2] a plastic stop to keep the condyle from moving, [3] pairs of 0.018 × 0.025 inch rectangular arch wires (Orthopli Corp, Philadelphia, PA) implanted vertically in dental resin to grasp [4] two parallel J120 closed clear power chains (Rocky Mountain Orthodontics, Denver, CO) which keep the ramus in line with the condyle. (b) Top view of the specimen holder demonstrating how the wire has been bent to efficiently insert and clasp the condylar neck. (c) Top view of the specimen holder with the 2 power chains added. (d) Side views indicating how the condyle (stained blue with toluidine) and coracoid processes are oriented to image the condyle’s anterior, middle, and posterior regions. (e) H&E stained sagittal section containing the middle region of the condyle with length and depth of the confocal imaging field obtained by confocal objective indicated. (f) Bird’s eye view of a 3D reconstruction depicting the confocal imaged volume (yellow) of the middle condylar region from a 6-week-old male mouse. (g) Bird’s eye view of a 4-week-old mouse condyle showing centroids of DAPI stained (blue) and EdU containing (pink) chondrocyte nuclei. (h) Sagittal section through the contralateral mouse condyle imaged for DAPI (blue) and EdU (pink) fluorescence, showing that the frequency and distribution of EdU-containing chondrocytes are comparable to that found with confocal imaging.
The Bland-Altman analyses revealed no significant biases in any of the measures used to calculate either the superficial or deep cell densities (Supplementary Figure 2 and Table 2).
Confocal microscopy confirms chondrocyte loss in mice lacking lubricin
It has previously been reported, using standard histology, that lubricin knockout mice develop age-related abnormalities in their TMJs, including widening and flattening of condylar surface with superficial chondrocyte loss1,8,9. Our confocal imaging detected these same changes in Prg4 knockout mice without requiring decalcification, paraffin embedding, sectioning, and staining (Fig. 2).
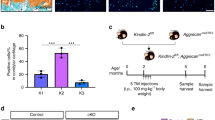
(a) 3D reconstruction depicting nuclear centroids in the middle region of a 2-month-old wild-type mouse (superficial cell nuclei, that is those within 10 μm of the condylar surface, are pseudocolored yellow and deeper cell nuclei are pseudocolored blue). (b) 3D reconstruction of the middle region of a 2-month-old Prg4 knockout (Prg4−/−) mouse with the same pseudocoloring as in panel A. Note there are fewer surface chondrocytes in the knockout mouse condyle. (c) Bar graph showing the mean (+SD) superficial cell densities for the middle region of 2-month-old Wild-type (n = 3) and Prg4−/− (n = 3) mice. (d and e) Similar 3D reconstructions as in panels A and B, but in 6-month-old Wild-type and Prg4−/− mice, respectively; note the condyle is now flattened and widened in the knockout mouse. (f) Bar graph showing the mean (+SD) superficial cell densities for the middle region of 6-month-old Wild-type (n = 4) and Prg4−/− (n = 5) mice. (g) H&E stained coronal section through the middle region of the TMJ from a 6-month-old wild-type mouse. (h) H&E stained coronal section through the middle region of the TMJ from a 6-month-old Prg4−/− mouse. Note the knockout mouse has a thicker fibrocartilaginous disc and less superficial chondrocyte nuclei compared to the wild-type mouse. (i) Toluidine blue stained TMJ section from the knockout mouse confirming that the cartilage surface has become acellular.
Confocal microscopy reveals non-uniform Cre-expression in the TMJ of AcanCreERt2 mice
The AcanCreERt2 mouse strain is frequently used to conditionally recombine genes with floxed alleles in articular cartilage, including the TMJ10. We observed that AcanCreERt2 is not uniformly active throughout the mandibular cartilage. We suspected this after we used the AcanCreERt2 strain to induce diphtheria toxin (DTA) expression in order to kill TMJ chondrocytes. We administered 3 daily doses of tamoxifen to P63 (9-week-old) AcanCreERt2/+; ROSA26l-s-lDTA/+ mice and then evaluated their mandibular condyles at P73 (Fig. 3). In control mice, we observed no difference in chondrocyte density between the anterior, middle, and posterior condylar regions (data not shown). However in the AcanCreERt2/+; ROSA26l-s-lDTA/+ tamoxifen-treated mice, significantly more chondrocyte loss occurred in anterior and middle regions compared to the posterior region (Fig. 3). We repeated the experiment using mice with a floxed fluorescent reporter allele (ROSA26mTmG) and again found more AcanCreERt2 activity in the anterior and middle regions compared to the posterior region and in deeper chondrocytes compared to superficial chondrocytes (Fig. 4). These confocal microscopic observations were confirmed when we examined the condyles with standard histologic methods (Figs 3 and 4).
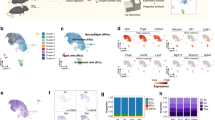
(a) Schematic depicting the experimental timeline for inducing diphtheria toxin (DTA) expression with a tamoxifen-inducible Cre knocked into the Acan locus. Mice were given tamoxifen for 3 consecutive days at P63 (9-weeks-old) and studied at P73. (b) 3D reconstruction depicting nuclear centroids in the middle region of a 10-week-old control (AcanCreERt2/+) mouse, pseudocolored white. (c) 3D reconstruction of the middle region from a 10-week-old DTA (AcanCreERt2/+; ROSA26l-s-lDTA/+) mouse; i.e., a mouse induced to express diphtheria toxin when 9-weeks-old. (d and e) 3D reconstructed images from the anterior (d) and posterior (e) regions of the same DTA mouse. Note there are fewer nuclei in the DTA condyle compared to the control condyle, and fewer nuclei in the anterior and middle regions compared to the posterior region of the DTA condyle. (f) Bar graph showing the mean (+SD) superficial cell densities for the middle region of control mice (n = 6) and the anterior, middle, and posterior regions of DTA mice (n = 4). Note, in control mice the surface cell density is the same for the anterior, middle, and posterior regions (data not shown). In the DTA mice, there is significantly less cell loss in the posterior compared to anterior and middle regions. (g) Bar graph showing the mean (+SD) deeper cell densities for the middle region of control mice (n = 6) and the anterior, middle, and posterior regions of DTA mice (n = 4). Note, deeper cell loss is greater in the anterior and middle regions of the DTA mouse condyle compared to the posterior region. (h and i) H&E stained coronal sections through the anterior, middle, and posterior regions of the TMJs from a control and a DTA mouse, respectively. Note the posterior region of the DTA mouse has less cell loss than the anterior and middle region. Also, note there is more killing of deeper cells compared to superficial cells in anterior and middle regions of the DTA mouse condyle.
(a) Flattened confocal images indicating green fluorescent protein (GFP) expressing chondrocytes in the anterior and posterior regions of a AcanCreERt2/+; ROSA26mTmG/+ mouse that was induced to express membrane bound GFP instead of tomato fluorescent protein when 9 weeks-old. Twenty-five images taken at 2 μm intervals to a depth of 50 μm were flattened for this image. (b) Flattened confocal images representing centroids of GFP expressing chondrocytes that have been pseudocolored yellow in the anterior and posterior regions of the same mouse depicted in panel A; non-GFP expressing chondrocyte nuclei are pseudocolored blue. Note the uniform distribution of nuclei from GFP expressing chondrocytes in the anterior region contrasts with the asymmetric distribution observed in the posterior region. (c) Fluorescence microscopic image of a sagittal section from a similarly treated mouse confirming there are more GFP expressing chondrocytes in the anterior and middle regions of the condyle compared to the posterior region; DAPI-stained nuclei are pseudocolored blue. (d) Bar graph showing the mean (+SD) percentages of GFP expressing superficial cells and deeper cells in the anterior and posterior regions of the mandibular condyle from tamoxifen-treated AcanCreERt2/+; ROSA26mTmG/+ mice (n = 4). Note the fraction of GFP expressing cells is greater in the anterior compared to posterior region and greater among deeper cells compared to superficial cells.
Discussion
The mouse is commonly used to model human TMJ disease3. Mice can also be used to identify diseases that cause TMJ pathology, which may be asymptomatic in patients. For example, lubricin deficiency causes striking and profound TMJ pathology in mice1,8,9, and in fish11, but the clinical literature does not mention TMJ involvement in patients. Even in mice, TMJ pathology may be missed because most scientists lack expertise in evaluating the mouse TMJ. Therefore, to facilitate studies in the mouse, we adapted a confocal imaging technique for studying long bone articular cartilage12. Because the mouse mandibular condyle is small, we designed a simple positioning apparatus to image the condyle’s anterior, middle, and posterior regions (Fig. 1 and Supplementary Figure 1). We showed that specimen processing requires minimal sample handling and that the condyle can be imaged to a depth of 50 μm quickly and reproducibly (Fig. 1, Table 1, and Supplementary Table 1).
Using wild-type mice, we determined that cell density was similar across the anterior, middle, and posterior condylar regions (data not shown), and higher nearer the surface (Supplementary Figure 1). We then compared WT and lubricin (Prg4) knockout mice to show that few mice were needed to detect a significant difference in cell density (Fig. 2). Importantly, we confirmed our confocal imaging results using histologic methods, and showed the latter agreed with results obtained by other investigators1,8,9. Taken together, our findings suggest that confocal microscopy can serve as a rapid screening tool for TMJ disease in mice. Mice found to have condylar abnormalities by confocal microscopy can then have their entire TMJ examined in detail using more extensive, but more costly, histologic methods.
Another use of condylar confocal microscopy will be to determine patterns of Cre-recombinase activity for driver strains. We demonstrated this using the AcanCreERt2 allele, which is used to conditionally alter genes in cartilage. When we administered tamoxifen to 9-week-old mice with an AcanCreERt2 allele and a reporter allele (i.e., DTA or mTmG), we observed significantly more Cre-recombinase activity in the anterior compared to the posterior region of the condyle (Figs 3 and 4). We also observed more Cre-recombinase activity in deeper compared to superficial chondrocytes. These confocal data agree with data for AcanCreERt2 that other investigators obtained using standard histology and LacZ staining10, and are consistent with there being a greater abundance of fibrochondrocytes, compared to hyaline chondrocytes, near the surface and in and posterior region of the condyle. The confocal method extends these earlier findings by quantifying the rate of Cre-recombination (Figs 3 and 4) and by demonstrating that recombination efficiency varies depending upon the reporter allele (Figs 3 and 4). Specifically, we observed AcanCreERt2 was more efficient at recombining the ROSA26l-s-lDTA allele than the ROSA26mTmG allele. The latter allele appears more difficult to Cre-recombine than other ROSA26 alleles13, perhaps because it contains two consecutive highly homologous cDNAs encoding fluorescent reporter proteins.
We are not the first to apply confocal imaging to the mouse mandibular condyle. For example, confocal microscopy has been used to evaluate sites of fluorescent protein expression in transgenic mice7. However, these investigators did not describe standardized methods for orienting their specimens or for collecting quantitative data. We used common dental supplies to build a specimen holder and developed a standard positioning system that enabled us to image the condylar surface quickly and at high resolution. Although there are other 3D imaging strategies for examining mouse cartilage in situ, including μMRI and μCT14, the advantages of confocal microscopy are its higher resolution compared to μMRI and its shorter image acquisition time compared to μMRI and μCT.
A limitation of our imaging method is that it does not examine the fibrocartilaginous disc or the glenoid fossa. Therefore, it will be less sensitive for screening mice whose disorders affect the disc and fossa before they affect the condyle. At present, we do not know when condylar involvement occurs in mouse models of TMJ. In the case of lubricin deficiency, condylar involvement occurs early even though the major sites of Prg4 expression are in the disc and glenoid fossa1. Similarly, when TMJ partial discectomy is performed to induce disease, condylar involvement appears within weeks of the primary injury15. Therefore, we anticipate that our method will be useful for many mouse models in which TMJ disease develops and for other studies of TMJ development, such as the transdifferentiation of condylar chondrocytes into osteoblasts16. Other image quantification strategies17 may also improve the utility of our method. We look forward to other investigators extending our method by adding new fluorescent agents, image analysis algorithms, and mouse models of TMJ disease.
Methods
All studies were performed in an AAALAC accredited facility in accordance with National Institutes of Health and Boston Children’s Hospital guidelines for research involving mice. The Institutional Animal Care and Use Committee at the Boston Children’s Hospital approved these studies.
Experimental animals
WT and ROSA26l-s-IDTA mice on a C57BL/6 J background, ROSA26mTmG mice on a 129 × 1/SvJ background, and AcanCreERt2 mice were obtained from the Jackson Laboratories (Bar Harbor, ME), stock numbers 00664, 009669, and 007576, respectively. Prg4 loss-of-function alleles, Prg4− and Prg4GT, have been previously described4,9; these mice are also available from the Jackson Laboratories (stock numbers 025737 and 025740, respectively).
Cre-recombinase activity was induced in 9-week-old mice with the AcanCreERt2 allele by administering 3 daily intraperitoneal (IP) injections of tamoxifen (150 mg/kg/dose) diluted in corn oil (both from Sigma-Aldrich, St. Louis, MO).
Labeling of dividing cells was performed by administering a single 50 mg/kg dose of 5-ethynyl-2′-deoxyuridine (EdU) IP (Santa Cruz Biotechnology, Dallas, TX).
Experimental animals were killed by CO2 inhalation immediately prior to specimen recovery and processing.
Specimen processing
After sacrifice, the skull and mandible were hemisected. To recover individual mandibular condyles, the overlying muscle was removed, the TMJ capsule opened, and the mandibular condyle was disarticulated from its glenoid fossa and fibrocartilaginous disc. The mandible was then fixed in 4% paraformaldehyde (PFA, Affymetrix, Cleveland, OH) at 4 °C. The contralateral mandible was either disarticulated and fixed or fixed in situ with the other components of the TMJ for whole mount sectioning, depending on the experiment.
Histology
The right PFA-fixed hemi-mandible was decalcified with 14% EDTA (pH 7.5) for up to 14 days, depending upon the animal’s age. Samples were then paraffin embedded so that the condyle could be sectioned in the sagittal or coronal plane. Six μm thick sections were obtained and stained with hematoxylin and eosin (H&E) or toluidine blue using standard methods. Sections from equivalent regions of the TMJ were compared between animals.
For experiments assessing the efficiency of EdU labeling in WT mice and of Cre-mediated recombination in AcanCreERt2/+; ROSA26mTmG/+ mice, after the mandible was decalcified in EDTA, it was equilibrated in 15% followed by 30% sucrose before being embedded in O.C.T. compound (Sakura, Torrance, CA). Eight μm thick frozen sections from WT and AcanCreERt2/+; ROSA26mTmG/+ mice were then obtained in the sagittal plane. For EdU containing sections, the Alexa Fluor 647 Click-iT EdU imaging kit (Invitrogen, Carlsbad, CA) was using following the manufacturer’s directions. All frozen sections were stained by ProLong Gold antifade reagent with DAPI (4′,6-diamidino-2-phenylindole) (Molecular Probes, Eugene, OR) and imaged using a Nikon Eclipse 80i (Nikon, Sendai, Japan) fluorescence microscope with a CoolSNAP HQ2 camera (Photometrics, Tucson, AZ).
Confocal microscopy
The non-decalcified left PFA-fixed hemi-mandible was placed in PBS with DAPI (Thermo-Fisher Scientific, Eugene, OR, 10 μg/ml) at 4 °C for at least 24 hours. For imaging cells that had incorporated EdU during cell division, the fluorophore Alexa 647 (Thermo-Fisher Scientific, Eugene, OR) was covalently linked to EdU-containing DNA as previously described12.
We fabricated a simple specimen holder using commonly available dental equipment (Fig. 1a–c). Condyles were loaded in the specimen holder, submerged in PBS, and imaged at 2 μm intervals to the depth of 50 μm. Three different orientations were tested in order to compare data for the anterior, middle, and posterior regions (Fig. 1d–f). Images were obtained with a Zeiss LSM780 confocal laser scanning (CLS) microscopy system (Carl Zeiss, Germany) with 20x Plan Apochromat NA 0.8 WD 550 mm dry objective, and a scanning speed of 6 as described previously12. When counting chondrocytes, we first determined the depth at which DAPI stained nuclei were clearly visible. When then determined the cartilage volume that extended to this depth. IMARIS software (Bitplane, Switzerland) was then used to generate 3D image reconstructions, calculate chondrocyte number and cartilage volumes, and determine the number of cell nuclei in volumes at different depths beneath the surface as previously described12.
Statistical Analyses
To assess the agreement and correlation of our confocal imaging method, the left and right condyles of 10 mice were measured and analyzed by the Bland-Altman method. The two-sided t-test was used to supplement the agreement between the measurement of left and right condyles. The minimal sample sizes to detect significant differences under the two-sided t-test were estimated assuming equal numbers of mice in case and control groups. Student’s t-test and a linear regression model were applied for comparisons between experimental and control groups. At least 3 animals per group were studied, and p-values < 0.05 were considered statistically significant.
Additional Information
How to cite this article: He, Y. et al. Confocal imaging of mouse mandibular condyle cartilage. Sci. Rep. 7, 43848; doi: 10.1038/srep43848 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Koyama, E. et al. Lubricin is Required for the Structural Integrity and Post-natal Maintenance of TMJ. Journal of dental research 93, 663–670, doi: 10.1177/0022034514535807 (2014).
Scrivani, S. J., Keith, D. A. & Kaban, L. B. Temporomandibular disorders. The New England journal of medicine 359, 2693–2705, doi: 10.1056/NEJMra0802472 (2008).
Suzuki, A. & Iwata, J. Mouse genetic models for temporomandibular joint development and disorders. Oral diseases 22, 33–38, doi: 10.1111/odi.12353 (2016).
Rhee, D. K. et al. The secreted glycoprotein lubricin protects cartilage surfaces and inhibits synovial cell overgrowth. The Journal of clinical investigation 115, 622–631, doi: 10.1172/JCI22263 (2005).
Henry, S. P. et al. Generation of aggrecan-CreERT2 knockin mice for inducible Cre activity in adult cartilage. Genesis 47, 805–814, doi: 10.1002/dvg.20564 (2009).
Kaul, R. et al. The Effect of Altered Loading on Mandibular Condylar Cartilage. PloS one 11, e0160121, doi: 10.1371/journal.pone.0160121 (2016).
Utreja, A. et al. Cell and matrix response of temporomandibular cartilage to mechanical loading. Osteoarthritis and cartilage 24, 335–344, doi: 10.1016/j.joca.2015.08.010 (2016).
Bechtold, T. E. et al. Osteophyte formation and matrix mineralization in a TMJ osteoarthritis mouse model are associated with ectopic hedgehog signaling. Matrix biology: journal of the International Society for Matrix Biology 52–54, 339–354, doi: 10.1016/j.matbio.2016.03.001 (2016).
Hill, A., Duran, J. & Purcell, P. Lubricin protects the temporomandibular joint surfaces from degeneration. PloS one 9, e106497, doi: 10.1371/journal.pone.0106497 (2014).
Jing, J. et al. Osterix couples chondrogenesis and osteogenesis in post-natal condylar growth. Journal of dental research 93, 1014–1021, doi: 10.1177/0022034514549379 (2014).
Askary, A. et al. Ancient origin of lubricated joints in bony vertebrates. eLife 5, doi: 10.7554/eLife.16415 (2016).
Zhang, M. et al. Induced superficial chondrocyte death reduces catabolic cartilage damage in murine posttraumatic osteoarthritis. The Journal of clinical investigation 126, 2893–2902, doi: 10.1172/JCI83676 (2016).
Kozhemyakina, E. et al. Identification of a Prg4-expressing articular cartilage progenitor cell population in mice. Arthritis & rheumatology 67, 1261–1273, doi: 10.1002/art.39030 (2015).
Ruan, M. Z. et al. Quantitative imaging of murine osteoarthritic cartilage by phase-contrast micro-computed tomography. Arthritis and rheumatism 65, 388–396, doi: 10.1002/art.37766 (2013).
Xu, L. et al. Early-onset osteoarthritis of mouse temporomandibular joint induced by partial discectomy. Osteoarthritis and cartilage 17, 917–922, doi: 10.1016/j.joca.2009.01.002 (2009).
Jing, Y. et al. Chondrocytes Directly Transform into Bone Cells in Mandibular Condyle Growth. Journal of dental research 94, 1668–1675, doi: 10.1177/0022034515598135 (2015).
Ren, Y., Lin, S., Jing, Y., Dechow, P. C. & Feng, J. Q. A novel way to statistically analyze morphologic changes in Dmp1-null osteocytes. Connective tissue research 55 Suppl 1, 129–133, doi:10.3109/03008207.2014.923879 (2014).
Acknowledgements
Y.H. is a recipient of a China Scholarship Council (CSC) Scholarship. This work was also supported by NIH RO1 AR050180 (to M.L.W.). The authors thank Ms. Evelyn Flynn for providing histology expertise.
Author information
Authors and Affiliations
Contributions
Y.H., M.Z., and M.L.W. conceived the study. Y.H. and Y.C. designed the apparatus. Y.H. and M.Z. collected data. A.Y.H. performed statistical analyses. Y.H., M.Z., A.Y.H., Y.C., D.B., and M.L.W. reviewed and analyzed the data, contributed to writing the manuscript, gave final approval, and agreed to be responsible for all aspects of this work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
He, Y., Zhang, M., Huang, A. et al. Confocal imaging of mouse mandibular condyle cartilage. Sci Rep 7, 43848 (2017). https://doi.org/10.1038/srep43848
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep43848
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.