Abstract
Planococcus is a Gram-positive halotolerant bacterial genus in the phylum Firmicutes, commonly found in various habitats in Antarctica. Quorum quenching (QQ) is the disruption of bacterial cell-to-cell communication (known as quorum sensing), which has previously been described in mesophilic bacteria. This study demonstrated the QQ activity of a psychrotolerant strain, Planococcus versutus strain L10.15T, isolated from a soil sample obtained near an elephant seal wallow in Antarctica. Whole genome analysis of this bacterial strain revealed the presence of an N-acyl homoserine lactonase, an enzyme that hydrolyzes the ester bond of the homoserine lactone of N-acyl homoserine lactone (AHLs). Heterologous gene expression in E. coli confirmed its functions for hydrolysis of AHLs, and the gene was designated as aidP (autoinducer degrading gene from P lanococcus sp.). The low temperature activity of this enzyme suggested that it is a novel and uncharacterized class of AHL lactonase. This study is the first report on QQ activity of bacteria isolated from the polar regions.
Similar content being viewed by others
Introduction
Quorum sensing (QS) or bacterial cell-to-cell communication has become a focus of research due to its high potential as a novel application to prevent the onset of bacterial infections and reduce the current over-use of antibiotics which itself is a selective pressure leading to increased antibiotic resistance1. Bacteria communicate with each other to control numerous phenotypic characteristics, such as the production of virulence factors2, antibiotic biosynthesis3, and biofilm differentiation4. In nature, QS could be highly advantageous particularly in the contexts of symbioses and niche adaptation, and for facilitating population migration towards/away from favourable/unfavourable conditions in their local environment5. Antarctica provides some of the most challenging environments on Earth for life6,7. Metagenomic analysis of Antarctic soil has revealed that Antarctic microbial communities are more complex and higher diversity than previously thought8. The presence of QS genes in Antarctic soil, together with antibiotic, biofilm formation, virulence and other toxic compound resistance genes, suggests that QS provides these bacteria with a competitive advantage in hostile Antarctic environments9.
The disruption of QS signals, termed quorum quenching (QQ), was first described by Dong et al.10 in Bacillus sp. However, QQ activity in extremophiles is not well studied, and has only been characterized in detail in a thermophile, Geobacillus kaustophilus11. To our knowledge, there is no report or characterization of QQ activity in bacteria isolated from the polar regions. With the ability to disrupt intercellular communication, QQ potentially makes an important contribution to competitive ability.
Planococcus is a member of the family Planococcaceae, a group of halotolerant Gram-positive bacteria12 commonly found in various habitats in Antarctica13,14. In this study, we report QQ activity in Antarctic Planococcus versutus strain L10.15T, that was capable of inactivating synthetic N-acyl homoserine lactones (AHLs) with acyl side chain lengths C4-C12, and functioning at a temperature as low as 4 °C. The gene responsible for the QQ activity of P. versutus L10.15T was identified and confirmed for its function in an expression study. The cold-active characteristic of the enzyme coded by this gene suggested that it belonged to a novel class of N-acyl homoserine lactonase, and we therefore term the gene as ‘autoinducer degrading gene from P lanococcus sp.’ (aidP). The discovery of the AidP protein provides the foundation for new routes of prospecting QQ enzymes for industrial applications.
Results
Isolation and molecular identification of QQ bacteria
The enrichment KG medium became turbid after one week of incubation, suggesting that the bacteria were able to utilise the synthetic AHL (C6-HSL) as sole carbon source15. The bacteria were then screened for QQ activity in an AHL inactivation assay using the CV026 biosensor. Amongst the 15 bacterial isolates obtained, strain L10.15T demonstrated strong QQ activity and was selected for further analysis. Web-based similarity comparison against the GenBank database suggested that strain L10.15T belonged to the genus Planococcus, and this was confirmed by phylogenetic analysis. Phylogenetic analysis of 16S rRNA gene sequences indicated that the strain’s closest relatives were P. halocryophilus, P. kocurii and P. donghaensis. Polyphasic taxonomic study confirmed that strain L10.15 represents a new taxon within Planococcus, which we designated as P. versutus L10.15T16.
Degradation of AHLs
QQ activity of P. versutus L10.15T was verified using synthetic AHL (C6-HSL) screened with biosensor CV026. Synthetic C6-HSL was selected for initial screening since it was used as the sole source of carbon in the enrichment medium. Strain L10.15T cells degraded 100 μM of C6-HSL in 100 μl of cell suspension within 24 h (Supplementary Fig. S1). As AHLs will undergo lactonolysis under alkaline conditions17, turnover of AHLs by alkaline lactonolysis was ruled out as no change in pH values was observed in the reaction mixtures after 24 h (data not shown).
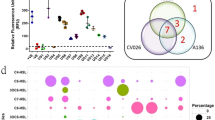
Because biosensor CV026 is only applicable for detection of short chain AHLs, rapid resolution liquid chromatography (RRLC) (Fig. 1) was used to confirm the QQ activity of strain L10.15T. A range of AHLs (C4-C13-HSL, with or without substituted groups) was tested. The results indicated that P. versutus L10.15T degraded all the AHLs tested and exhibited high activities toward most, including those with 3-hydroxy or 3-oxo substitutions and un-substituted homoserine lactones. The substrate specificity was significantly affected by the length of the acyl acid chain, with the estimated relative activity of strain L10.15T showing a gradual decease toward AHLs with longer chain lengths. The strain had low QQ activity toward C13-HSL (data not shown). No AHL degradation was observed in all control AHL degradation assays repeated with E. coli DH5α cells and PBS.
Determination of AHL-lactonase activity via acidification assay
In order to demonstrate that P. versutus L10.15T produced an AHL-degrading enzyme, the bacterial suspension was boiled at 95 °C for 30 min to denature any enzyme present before addition to the synthetic AHLs. The boiled cell suspension no longer possessed QQ activity (data not shown), indicating that the activity was most probably catalyzed by an enzyme. An acidification assay17 was conducted to re-lactonise the opened ring of AHL lactones caused by the AHL-lactonase. A proportion of the AHL amounts degraded by strain L10.15T was restored after acidification, with C4-HSL being restored up to 30% (Supplementary Fig. S2), further supporting that the QQ activity of P. versutus L10.15T may be due to an AHL-lactonase.
Complete genome sequencing and assembly of strain L10.15
Sequenced data of P. versutus L10.15T comprised a total of 135,526 polymerase reads (Reads N50: 9333 bp) which, upon filtering, resulted in a total of 167,910 subreads (Reads N50: 7768 bp). The subreads were subjected to de novo assembly using Hierarchical genome assembly process (HGAP) algorithm version 2, which constructed a final assembly (mean coverage: 219.3 times) of three circular contigs with sizes of 3.2 Mb, 70.7 kb and 9.8 kb, respectively. All contigs contained self-overlapping ends, indicating their circularity. The coordinates of the overlapping ends were estimated using the gepard dotplot tool and were subsequently trimmed to generate blunt-ended circular contigs.
BLASTn searches against the NCBI non-redundant nucleotide database provided a preliminary identity assignment for each contig, with the largest contig being determined as a chromosomal contig and the two smaller contigs as being extrachromosomal contigs, and most likely representing plasmids due to sequence similarity to deposited plasmid sequences (Supplementary Tables S1–S3). Based on these results, designations of pPS15-1 and pPS15-2 were given to the 70.7 kb and 9.8 kb contigs, respectively.
Further support for the identity of pPS15-1 and pPS15-2 was provided by the identification of genes associated with the replication module of plasmids. For example, in pPS15-1, a putative parA (WP_007723335.1), and in its opposite transcriptional orientation, 2 CDSs (WP_049694985.1 and WP_049694987.1) were identified, genes which encode putative proteins with sequence similarities to various plasmid replication proteins. In pPS15-2 a putative RepB family plasmid replication initiator protein gene (WP_049694148.1), a common occurrence among the plasmids of cold-active bacteria, was identified18.
The chromosomal origin of the 3.2 Mb contig was further established through the identification of a 9.8 kb section which contains a 708 bp putative oriC region (9 DnaA boxes) (Supplementary Figure S3) flanked by a gene cluster which comprised rmpH (WP_049693545.1), rnpA (WP_049693546.1), dnaA (WP_049693544.1), dnaN (WP_049693543.1), yaaA (WP_049693542.1), recF (WP_049693541.1), gyrB (WP_049693540.1), and gyrA (WP_049693539.1). The chromosome was rearranged to begin with the putative dnaA CDS (1 to 1344 nt) and to end with the putative oriC region (3248918 to 3249625 nt). In order to obtain an in silico evaluation of the degree of completeness of the chromosome, a Z-curve was then plotted using the rearranged chromosome. From the plot, an inverted V-shape was observed for both AT- and GC- disparity curves, where the maximum (estimated location of putative replication terminus (terC) site) and the minimum (corresponding with the location of predicted oriC site) were situated at diametrically opposite locations (Supplementary Fig. S3). The presence of both the origin and terminus of replication provided an indication of the completeness of the chromosome.
Overview of general genome features of strain L10.15T
The assembled genome of strain L10.15T comprised three circular replicons with a mol% G + C content of 39.4%, which is consistent with the reported GC contents of P. donghaensis and P. kocurii19,20. A total of 3153 coding sequences (CDS), 9 rRNA operons and 71 tRNAs were predicted to be encoded by the genome (Table 1). The NCBI genome accession number for the chromosome is CP016540, for pPS15-1 is CP016541.2 and for pPS15-2 is CP016542.2.
Identification of Planococcus versutus L10.15T AHL-lactonase gene
Analysis of the annotated genome from the RAST server revealed the presence of a candidate QQ gene (annotated as an N-acyl homoserine lactonase). The 282 amino acid sequence was subsequently subjected to BLASTp search against the NCBI non-redundant protein sequences database. The comparison result revealed that the predicted proteome of the CDS showed this protein was annotated as an MBL fold metallo-hydrolase. BLASTp search also revealed a MBL fold metallo-hydrolases with 98% identity in P. antarticus, P. faecalis and P. kocurii, indicating that the gene is conserved across several Planococcus species. BLASTp search also indicated that genes with lower identity (<85%) were detected in Bacillus, Jeotgalibacillus, Panaenibacillus, Lysinibacillus and other genera from the Order Bacillales. However, only one AHL lactonase gene21 (from Lysinibacillus sp. Gs50) with identity 81% has been characterized. The BLASTp also detected several putative conserved domains (Supplementary Fig. S4) including AHL lactonase MBL-fold (Table 2).
When the amino acid sequence of aidP was compared with those of the known AiiA-type lactonases (Fig. 2), the zinc-binding motifs HXHXDH~H, which are commonly conserved sequences of the metallo-β-lactamase superfamily, were found in both the known AiiA-type lactonases and in aidP (117HLHLDH122 ~ H219). The multiple alignment analysis of aidP and other AiiA-type lactonases also indicates (1) the active-site residue of the AHL lactonase enzyme, Tyr222, Leu120, Asp121, which are known to be conversed in most of the AiiA-type lactonases; (2) the amino acid residue that interacts with the primary ligand, H265 and D219; and (3) a non-conservative residue in the N-binding region, G235, was also detected. The catalytic mechanism involved in the active-site residue, and also the possible product-bound mechanism, are considered below.
The amino acid sequence of AidP (Genbank accession number WP_049694637.1) was compared with those AiiA from Bacillus sp.240B1 (accession number AAF62398), AiiB from Agrobacterium tumefaciens C58 (accession number AAK91031), AhlD from Arthrobacter sp. strain IBN110 (accession number AAP57766.1), AttM from A. tumefaciens strain A6 (accession number AAD43990), AidC from Chryseobacterium sp. strain StRB126 (accession number BAM 28988.1), and QlcA from the soil metagenome (accession number ABV58973.1). Sequences were aligned using the ClustalW program and shaded using the Genedoc program (http://www.nrbsc.org/gfx/genedoc/). The zinc-binding motif is boxed with rectangles. The amino acid residues that are essential for enzymatic activity of the known AHL lactonase are indicated by red arrows.
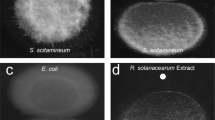
To confirm whether this candidate gene is responsible for the QQ activity in P. versutus L10.15T, it was ligated with expression vector pET-100 and cloned into E. coli BL21 StarTM. Recombinant E. coli BL21 StarTM cells harboring the plasmids were screened for QQ activity with 10 μM C6-HSL using RRLC as described before. E. coli BL21 StarTM containing the plasmid ligated with the candidate gene showed significant AHL-degrading activity against C6-HSL (Fig. 3). To further confirm the AHL degradation gene is an AHL lactonase gene, the AHL residue was analyzed by liquid chromatography-mass spectrometry (LC-MS). This analysis indicates retention time of C6-HSL was shifted from 1.45 min (Fig. 4a) to 0.94 min (Fig. 4b). ESI-MS analysis of C6-HSL with retention time 1.45 min exhibits a strong quasimolecule (M-H) ion at mass-to-charge ratio (m/z) of 200.0 (Fig. 4c). However, ESI-MS analysis of the AHL residue revealed a product with m/z of 218.0 (Fig. 4d). This result indicates that the C6-HSL experiences a mass increase of 18, corresponding to a water molecule, further suggesting that the candidate gene produced an enzyme that hydrolyzes the ester bond of the homoserine lactone ring of C6-HSL. This QQ gene, which may represent a novel class of AHL lactonase, was termed aidP (autoinducer degrading gene from P lanococcus sp.).
AidP is a new member of the AiiA-type AHL-lactonase
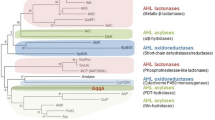
To evaluate the novelty of AidP in the AHL-lactonase family, we used the amino acid sequence of aidP to determine the phylogenetic relationship between AidP and known AHL lactonases from various bacteria using the neighbour-joining method (Fig. 5). The AHL lactonase can be grouped into the metallo-β-lactamase superfamily, α/β-hydrolase-fold family, and the phosphotriesterase (PTE) family. Among these groups, the metallo-β-lactamase superfamily was well characterized from various bacteria. The amino acid sequence of aidP also clustered into the metallo-β-lactamase superfamily. However, it clustered distantly from the other lactonase clusters represented by six other AiiA-type lactonases. The aidP gene has low similarity to each of the known AiiA-type AHL lactonases - 26.2% identity across the entire length of AiiA from Bacillus sp. 240B122, 24.9% identity across AttM from Agrobacterium tumefaciens strain C5823, 27.5% across AhlD from Arthrobacter sp. strain IBN11024, 29% identity across AidC from Chryseobacterium sp. strain StRB12625 and 15.1% identity across QlcA from the soil metagenome26 and 41.5% across AiiB from Agrobacterium tumefaciens27. A phylogenetic tree of the highest similarity amino acid sequences with aidP from NCBI BLASTp search was also constructed (Fig. 6). AidP and aidP-homologous gene from Planococcus species were clustered in one clade.
Phylogenetic tree based on amino acid sequences of AidP, AiiB, AhlD, AiiA, AttM, AidC, QlcA, AidH (accession number ACZ73823.1), and AidH from Ochrobactrum sp. strain T63, AiiM from Microbacterium testaceum strain StLB037 (accession number BAH97082.2), and QsdA from Rhodococcus erythropolisstrain W2 (accession number AAT06802.1). The dendrogram was constructed by the neighbour-joining method using MEGA 6 software. The scale bar represents 0.2 substitutions per amino acid position.
Discussion
By enrichment targeting QQ bacteria using the protocol described by Chan et al.15 at a modified lower growth temperature of 4 °C, we successfully isolated P. versutus L10.15T, a novel psychrotolerant QQ bacterium. This strain had the ability to selectively degrade AHLs with carbon side chain lengths between C4 and C12.
We infer that the QQ activity shown by strain L10.15 was achieved via an AHL lactonase enzyme, a class of QQ enzyme that hydrolyzes the homoserine lactone ring by cleaving its ester bond. This led us to hypothesize that P. versutus L10.15T possesses an AHL lactonase gene similar to the AiiA gene, an AHL lactonase gene identified from Bacillus sp.22. RAST identified the QQ gene and revealed it to be an N-acyl homoserine lactonase. The BLASTp result indicated that the gene is annotated as an MBL fold hydrolase, and revealed that several other Planococcus species possess the MBL fold metallo-hydrolase gene with high similarity (98%) to that found in the current study. Other hits from BLASTp included genes with lower identity (85% or below) from various taxa in the order Bacillales, all of which were annotated as MBL fold metallo-hydrolases. Even though BLASTp identified several genes as AHL lactonase (Fig. 6), only AHL lactonase from Lysinibacillus sp. Gs50 has been characterized and confirmed in terms of its activity21. The QQ gene (designated as aidP) identified in P. versutus L10.15T is, therefore, yet to be characterized, and could be a novel class of AHL lactonase enzyme from the metallo-β-lactamase family. This inference was further supported by the AHL lactonase MBL fold domain detected by BLASTp search. The domain was detected as a “specific hit”, indicating that the sequence has a high confidence level of the inferred function with the domain model, the Bacillus thuringiensis AHL lactonase (AiiA) (accession number: cd07729), with high similarity (e-value: 3.82 × 10−78). CDD also classified that this domain belongs to the metallo-hydrolase-like MBL fold superfamily.
The AHL lactonases from the MBL superfamily are known to have a signature zinc-binding motif HXHXDH ~ H. Crystallographic studies of AiiA28,29, AiiB30 and AidC31, have also revealed many other amino acid residues that are functionally crucial. These amino acid residues play critical functions in the catalytic mechanism. In this study, we also identify a number of functionally-important amino acid residues that were previously reported in the crystallographic studies of AiiA-type enzymes. A mutagenesis study of the active site of AiiA revealed that tyrosine (Y194) and aspartic acid (D108) residues are directly involved in the catalytic mechanism. Multiple alignment analysis of aidP and other AHL lactonases from the MBL superfamily revealed the presence of both amino acids (Y222 and D121) in aidP. The tyrosine residue provides stabilization for the substrate, while aspartic acid acts as a proton shuttle that tightens the active site, interacting with the hydroxyl leaving group of the product30,32. Asp219 (D219) of aidP, which is homologous with D191 of AiiA and D213 in AiiB, is important in the formation of zinc bridging. G235, which is homologous with Gly207 of AiiA, was also detected in AidP; mutation of this amino acid residue which is located in the N-acyl binding region will cause a significant decrease in the activity of AiiA.
In order to determine if aidP is conserved in the genus Planococcus, all available genome data of Planococcus sp. were subjected to RAST analysis. ‘N-acyl homoserine lactonase’ was found in the three closest available relatives of P. versutus L10.15T, these being the type species of P. antarcticus, P. faecalis and P. kocurii, all with high similarity (98%) to aidP. This indicates that these species carry a homologous aidP gene. Both P. antarcticus and P. faecalis were also isolated from Antarctica and, even though P. kocurii was not isolated from Antarctica, ANI analysis indicated that P. faecalis and P. kocurii have high OrthoANI33 values (98.2%) and may belong to same taxon34 (Supplementary Figure S5). We did not identify N-acyl homoserine lactonase or other QQ genes in genomes of the other Planococcus spp. As aidP and aidP-homologous genes have mostly been detected in the genome of Planococcus strains originating from Antarctica, we speculate that this may indicate the importance of QQ activity in members of Planococcus as a strategy for competition in harsh soil environments of the continent.
In addition to enhancing competitive ability in the Antarctic environment, the cold-active QQ enzymes of P. versutus L10.15T could give additional advantages to this strain. Previously, P. rifientoensis has been reported to promote the growth of plants35, even though a complete genome analysis of P. rifientoensis identified no gene sequence coding for N-acyl homoserine lactonase36. As QQ enzymes produced by bacteria have been shown to effectively prevent the plant pathogen, Pectobacterium sp., from causing soft root disease on plant tubers9, we hypothesise a capability of P. versutus L10.15T to promote plant growth, and a potential for it to act as a biocontrol agent in horticulture.
Using QQ as biocontrol strategy will minimize the selective pressure imposed on the targeted pathogens, and thus may reduce the development of resistance. The search for more effective QQ agents is important, as the resistance of bacteria to available antibiotics is increasing, and the development of alternative strategies against pathogens is critical1,36. It has been recognized that polar and other extreme environments are potentially important sources of novel and industrially important enzymes37. In this context, the discovery of psychrotolerant QQ bacteria with potential for use as biocontrol, remediation or growth promoting agents would be advantageous. In combination with the possible positive influence of nitrogen-fixing ability on plant growth, we suggest that P. versutus L10.15T could be a beneficial bacterium to explore for applications in agriculture in cold environments.
Materials and Methods
Sampling location
The soil sample (upper 6 cm depth) was collected using a sterile sampling tube at the edge of an elephant seal (Mirounga leonina) wallow area on Lagoon Island (Ryder Bay, Adelaide Island, west of the Antarctic Peninsula; 67° 35.689′S 068° 14.495′′E). The soil sample was collected as part of a survey of bacterial communities at locations around Ryder bay (Supplementary Fig. S6).
Bacterial isolation
For enrichment and isolation of QQ bacteria, a basal medium was used as previously described15. Briefly, 1 g of soil sample and 5 ml sterile medium including 100 μg C6-HSL were added to a sterile 50 ml plastic tube and incubated at 4 °C with shaking (150 rpm). After 1 week of incubation, 100 μl of the bacterial suspension was inoculated into fresh, sterile medium supplemented with C6-HSL. This step was repeated three times and finally, 100 μl of bacterial suspension was plated onto LB agar in order to obtain pure colonies.
Bacterial strains, growth media, and culture conditions
The biosensor C. violaceum CV026 was used to screen for presence of AHL in the AHL inactivation assays38. E. coli DH5α was used as the host for cloning the 16S rDNA gene and as negative control for the AHL inactivation assay, and was grown at 37 °C in LB broth to achieve the desired OD600 of 1.0. Strain L10.15 was cultured at 4 °C in LB broth.
Molecular identification and phylogenetic analysis
P. versutus L10.15T DNA was extracted using the QIAamp DNA mini kit (Qiagen, Germany). Universal primers 27F39 and 1525R40 were used for 16S ribosomal RNA (rRNA) gene amplification. PCR products were ligated into pGEM-T following the manufacturer’s protocol (Promega, USA). DNA sequencing was performed by routine automated methods in which standard T7 forward and SP6 reverse primers were used. Phylogenetic and molecular evolutionary relationships were analysed using MEGA version 641. Phylogenetic analyses of 16S rRNA were carried out using the 16S rRNA sequences of Planococcus type species. Alignment was performed using the MUSCLE algorithm42 and the phylogenies were constructed using the default settings of the neighbour-joining algorithm.
AHL inactivation assay
To assay the ability to inactivate AHLs, synthetic AHL was dispensed into 1.5 ml sterile tubes and solvent evaporated, after which 100 μl of either P. versutus L10.15T or E. coli strain DH5α cells were added to rehydrate the dried AHL (100 μM final concentration). These cell suspension mixtures were then incubated at 4 °C (except for E. coli DH5α at 37 °C) with shaking (220 rpm). Aliquots were withdrawn at 0 h and 24 h. The reaction was stopped by heating as described previously37. After cooling, the reaction mix (10 μl) was spotted onto agar overlaid with biosensor CV026 followed by overnight incubation at 28 °C. Control experiments were carried out using extraction buffer (PBS 10 mM, pH 6.5) and E. coli DH5α.
Detection of AHL-degrading activity of P. versutus L10.15T and acidification assay
The residue AHL from the assay was analysed by rapid resolution liquid chromatography (RRLC) 6400 series (Agilent, USA) using an Agilent Poroshell 120 EC-C18 column (4.6 mm × 100 mm, 2.7 μm particle size) with the elution procedure consisting of an isocratic profile of acetonitrile/water (35:65, v/v), and a constant flow rate of 0.7 mL/min, with UV detection set at 205 nm. The assay was carried out as described above, except the residue AHLs were extracted twice using an equal volume of ethyl acetate after 0 h, 24 h or 48 h of incubation, followed by evaporation to dryness. AHL extracts were reconstituted in HPLC grade acetonitrile before being subjected to RRLC analysis. For identification of AHL lactonase activity, the method of Yates et al.17 was followed. Samples withdrawn from the AHL inactivation assay were divided into two aliquots of equal volume (50 μl), and one was acidified with 100 mM HCL to pH 2.0 to induce recyclization of the lactone ring, while the other untreated aliquot was used as control. The acidified sample was incubated at 4 °C for 48 h before adjusting to pH 6.0 with MOPS buffer (0.5 M, pH 7.5).
Whole genome sequencing and functional gene annotation
Genomic DNA of P. versutus L10.15T was isolated using the MasterPure™ Gram-positive DNA purification kit (Epicentre Technologies) following the manufacturer’s recommended protocol. DNA quality was confirmed using a Nanodrop spectrophotometer (Thermo Scientific, USA) and Qubit® 2.0 Fluorometer (Life Technologies, USA) before being constructed into a 20 kb SMRTbell template library. Pacific Biosciences (PacBio) RSII sequencing platform was used to perform whole genome sequencing using C4 chemistry in a single molecule real time (SMRT) cell. The reads were de novo assembled using hierarchical genome assembly process (HGAP) algorithm version 2 into a circular contig. Genome annotation was performed using the NCBI Prokaryotic Genome Annotation Pipeline (PGAP) version 2.5 and Rapid Annotation using Subsystem Technology (RAST) version 3.042,43,44,45.
Cloning of candidate QQ gene and confirmation of QQ activity of the candidate gene
The candidate QQ gene from P. versutus L10.15T was amplified by PCR using the designed primer pair forward (5′-CACCATGACTGGTATTATCAAGCC) and reverse (5′- TTATTCGTAGTATCCTTCAGTCGACT) and KAPA HiFi polymerase (Kapa Biosystem, USA). PCR was performed using the following cycling parameters: 95 °C for 30 s, 62 °C for 30 s, and 72 °C for 1 min, for 35 cycles. The amplicons were purified using the AMPure XP- PCR purification system (Beckman Coulter, USA). The purified PCR product was cloned into the expression vector (pET-200) and transformed into E. coli BL21 StarTM. Next, the pET-AidP was amplified in the BL21 StarTM host, and extracted using the Qiagen Plasmid Mini kit (Qiagen, Germany). pET-AidP was then transformed into expression host E. coli BL21 StarTM. The E. coli BL21 StarTM with pET-AidP plasmid was induced by IPTG (0.4 mM) and its QQ activity was assessed as described above, except that a temperature of 16 °C was used. The AHL (C6-HSL with a final concentration of 2 mM) residues were subjected to liquid chromatography-mass spectrometry (LC-MS) analysis. Briefly, the samples were analyzed by LC using an Agilent Poroshell 120 EC-C18 column (4.6 mm × 100 mm, 2.7 μm particle size) with a mobile phase of acetonitrile-water (0.1% formid acid; a linear gradient [v/v] of acetonitrile from 5 to 95% over 1.5 min at a flow rate of 1.8 ml min−1. The separated fractions were further analyzed by electrospray ionization mass spectrometry (ESI-MS).
Conclusions
We confirmed the in vitro QQ activity of P. versutus L10.15T that produced a lactonase that degraded AHL molecules at low (4 °C) temperature, providing a new source for the isolation of these industrially important enzymes from Antarctic microbiota. P. versutus L10.15T showed broad AHL degradation activity. Genomic analysis enabled the identification of the AHL lactonase gene, which belongs to a novel class of AHL lactonase gene. Further study is required to fully characterize this enzyme.
Nucleotide sequence accession number
The complete genome sequence of P. versutus L10.15T has been deposited in GenBank under the accession number CP016540.
Additional Information
How to cite this article: See-Too, W. S. et al. AidP, a novel N-Acyl homoserine lactonase gene from Antarctic Planococcus sp. Sci. Rep. 7, 42968; doi: 10.1038/srep42968 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Rasmussen, T. B. & Givskov, M. Quorum-sensing inhibitors as anti-pathogenic drugs. Int. J. Med. Microbiol. 296, 149–161 (2006).
Jones, S. et al. The lux autoinducer regulates the production of exoenzyme virulence determinants in Erwinia carotovora and Pseudomonas aeruginosa . The EMBO Journal 12, 2477 (1993).
Pierson, L., Keppenne, V. D. & Wood, D. W. Phenazine antibiotic biosynthesis in Pseudomonas aureofaciens 30–84 is regulated by PhzR in response to cell density. J. Bacteriol. 176, 3966–3974 (1994).
Davies, D. G. et al. The involvement of cell-to-cell signals in the development of a bacterial biofilm. Science 280, 295–298 (1998).
Williams, P., Winzer, K., Chan, W. C. & Camara, M. Look who’s talking: communication and quorum sensing in the bacterial world. Philos. Trans. R. Soc. Lond. B Biol. Sci. 362, 1119–1134 (2007).
Peck, L. S., Convey, P. & Barnes, D. K. A. Environmental constraints on life histories in Antarctic ecosystems: tempos, timings and predictability. Biol. Rev. Camb. Philos. Soc. 81, 75–109 (2006).
Convey, P. et al. The spatial structure of Antarctic biodiversity. Ecol. Monogr. 84, 203–244 (2014).
Lee, C. K., Barbier, B. A., Bottos, E. M., McDonald, I. R. & Cary, S. C. The inter-valley soil comparative survey: the ecology of Dry Valley edaphic microbial communities. The ISME Journal 6, 1046–1057 (2012).
Pearce, D. A. et al. Metagenomic analysis of a southern maritime Antarctic soil. Front. Microbiol. 3, 403 (2012).
Dong, Y.-H., Xu, J.-L., Li, X.-Z. & Zhang, L.-H. AiiA, an enzyme that inactivates the acylhomoserine lactone quorum-sensing signal and attenuates the virulence of Erwinia carotovora . Proc. Natl Acad. Sci. USA 97, 3526–3531 (2000).
Chow, J. Y. et al. Directed evolution of a thermostable quorum-quenching lactonase from the amidohydrolase superfamily. J. Biol. Chem. 285, 40911–40920 (2010).
Romano, I., Giordano, A., Lama, L., Nicolaus, B. & Gambacorta, A. Planococcus rifietensis sp. nov, isolated from algal mat collected from a sulfurous spring in Campania (Italy). Syst. Appl. Microbiol. 26, 357–366 (2003).
Olivera, N. L., Sequeiros, C. & Nievas, M. L. Diversity and enzyme properties of protease-producing bacteria isolated from sub-Antarctic sediments of Isla de Los Estados, Argentina. Extremophiles 11, 517–526 (2007).
Miller, K. J. & Leschine, S. B. A halotolerant Planococcus from Antarctic Dry Valley soil. Curr. Microbiol. 11, 205–209 (1984).
Chan, K.-G., Yin, W.-F., Sam, C.-K. & Koh, C.-L. A novel medium for the isolation of N-acylhomoserine lactone-degrading bacteria. J. Ind. Microbiol. Biot. 36, 247–251 (2009).
See-Too, W.-S. et al. Planococcus versutus L10.15T, isolated from Antarctic soil. Int. J. Syst. Evol. Microbiol, doi: 10.1099/ijsem.0.001721 (2016).
Yates, E. A. et al. N-acylhomoserine lactones undergo lactonolysis in a pH-, temperature-, and acyl chain length-dependent manner during growth of Yersinia pseudotuberculosis and Pseudomonas aeruginosa . Infect. Immun. 70, 5635–5646 (2002).
Dziewit, L. & Bartosik, D. Plasmids of psychrophilic and psychrotolerant bacteria and their role in adaptation to cold environments. Front. Microbiol. 5, 596 (2014).
Pearson, M. D. & Noller, H. F. The draft genome of Planococcus donghaensis MPA1U2 reveals nonsporulation pathways controlled by a conserved Spo0A regulon. J. Bacteriol. 193, 6106 (2011).
See-Too, W.-S. et al. De novo assembly of complete genome sequence of Planococcus kocurii ATCC 43650T. a potential plant growth promoting bacterium. Mar. Genomics 28, 33–35 (2016).
Garge, S.-S. & Nerurkar, A.-S. Attenuation of quorum sensing regulated virulence of Pectobacterium carotovorum subsp. carotovorum through an AHL lactonase produced by Lysinibacillus sp. Gs50. Plos One 11, e0167344 (2016).
Wang, L.-H., Weng, L.-X., Dong, Y.-H. & Zhang, L.-H. Specificity and enzyme kinetics of the quorum-quenching N-acyl homoserine lactone lactonase (AHL-lactonase). J. Biol. Chem. 279, 13645–13651 (2004).
Zhang, H.-B., Wang, L.-H. & Zhang, L.-H. Genetic control of quorum-sensing signal turnover in Agrobacterium tumefaciens . Proc. Natl Acad. Sci. USA 99, 4638–4643 (2002).
Park, S.-Y. et al. AhlD, an N-acylhomoserine lactonase in Arthrobacter sp., and predicted homologues in other bacteria. Microbiology 149, 1541–1550 (2003).
Wang, W.-Z., Morohoshi, T., Someya, N. & Ikeda, T. AidC, a novel N-acylhomoserine lactonase from the potato root-associated Cytophaga-Flavobacteria-Bacteroides (CFB) group bacterium Chryseobacterium sp. strain StRB126. Appl. Environ. Microbiol. 78, 7985–7992 (2012).
Riaz, K. et al. A metagenomic analysis of soil bacteria extends the diversity of quorum‐quenching lactonases. Environ. Microbiol. 10, 560–570 (2008).
Carlier, A. et al. The Ti plasmid of Agrobacterium tumefaciens harbors an attM- paralogous gene, aiiB, also encoding N-acyl homoserine lactonase activity. Appl. Environ. Microbiol. 69, 4989–4993 (2003).
Liu, D. et al. Mechanism of the quorum-quenching lactonase (AiiA) from Bacillus thuringiensis. 1. Product-bound structures. Biochemistry 47, 7706–7714 (2008).
Momb, J. et al. Mechanism of the quorum-quenching lactonase (AiiA) from Bacillus thuringiensis. 2. Substrate modeling and active site mutations. Biochemistry 47, 7715–25 (2008).
Liu, D. et al. Structure and specificity of a quorum-quenching lactonase (AiiB) from Agrobacterium tumefaciens . Biochemistry 46, 11789–11799 (2007).
Mascarenhas, R. et al. Structural and biochemical characterization of AidC, a quorum-quenching lactonase with atypical selectivity. Biochemistry 54, 4342–4353 (2015).
Liu, D. et al. Three-dimensional structure of the quorum-quenching N-acyl homoserine lactone hydrolase from Bacillus thuringiensis . Proc. Natl Acad. Sci. USA 102, 11882–11887 (2005).
Lee, I., Kim, Y. O., Park, S. C. & Chun, J. OrthoANI: an improved algorithm and software for calculating average nucleotide identity. Int. J. Syst. Evol. Microbiol. 66, 1100–1103 (2015).
Goris, J. et al. DNA–DNA hybridization values and their relationship to whole-genome sequence similarities. Int. J. Syst. Evol. Microbiol. 57, 81–91 (2007).
Rajput, L., Imran, A., Mubeen, F. & Hafeez, F. Y. Salt-tolerant PGPR strain Planococcus rifietoensis promotes the growth and yield of wheat (Triticum aestivum L.) cultivated in saline soil. Pak. J. Bot. 45, 1955–1962 (2013).
See-Too, W.-S. et al. Complete genome of Planococcus rifietoensis M8T, a halotolerant and potentially plant growth promoting bacterium. J. Biotechnol. 221, 114–115 (2016).
Nichols, D. et al. Developments with Antarctic microorganisms: culture collections, bioactivity screening, taxonomy, PUFA production and cold-adapted enzymes. Curr. Opin. Biotechnol. 10, 240–246 (1999).
McClean, K. H. et al. Quorum sensing and Chromobacterium violaceum: exploitation of violacein production and inhibition for the detection of N-acylhomoserine lactones. Microbiology 143, 3703–3711 (1997).
Weisburg, W. G., Barns, S. M., Pelletier, D. A. & Lane, D. J. 16S ribosomal DNA amplification for phylogenetic study. J. Bacteriol. 173, 697–703 (1991).
Dewhirst, F. E., Paster, B. J. & Bright, P. L. Chromobacterium, Eikenella, Kingella, Neisseria, Simonsiella, and Vitreoscilla species comprise a major branch of the beta group Proteobacteria by 16S ribosomal ribonucleic acid sequence comparison: transfer of Eikenella and Simonsiella to the family Neisseriaceae (emend.). Int. J. Syst. Evol. Microbiol. 39, 258–266 (1989).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 30, 2725–2729 (2013).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Aziz, R. K. et al. The RAST Server: rapid annotations using subsystems technology. BMC Genomics 9, 1 (2008).
Overbeek, R. et al. The SEED and the Rapid Annotation of microbial genomes using Subsystems Technology (RAST). Nucleic Acids Res. 42, D206–D214 (2014).
Brettin, T. et al. RASTtk: a modular and extensible implementation of the RAST algorithm for building custom annotation pipelines and annotating batches of genomes. Sci. Rep. 5, doi: 10.1038/srep08365 (2015).
Acknowledgements
This work was supported by the University of Malaya (UM) High Impact Research Grants (UM-MOHE HIR Grant UM.C/625/1/HIR/MOHE/CHAN/14/1, Grant No. H-50001-A000027; UM-MOHE HIR Grant UM.C/625/1/HIR/MOHE/CHAN/01, Grant No. A-000001-50001) awarded to Kok-Gan Chan. Wah-Seng See-Too thanks the Bright Sparks Program of University of Malaya for scholarship support. Robson Ee was supported by PPP grant (PG084-2015B) from the University of Malaya, and Yan Lue Lim by PPP grant (PG133-2016A). Fieldwork in Antarctica was supported by the British Antarctic Survey. Peter Convey is supported by NERC core funding to the BAS ‘Biodiversity, Evolution and Adaptation’ Team. This paper also contributes to the Scientific Committee on Antarctic Research international programme ‘Antarctic Thresholds – Ecosystem Resilience and Adaptation’ (AnT-ERA).
Author information
Authors and Affiliations
Contributions
W.S.S.T. conceived the study, acquired the data, performed statistical analyses and drafted the manuscript. R.E., Y.L.L., P.C., D.P., Y.F.Y. and K.G.C. contributed to the manuscript. All authors reviewed the manuscript and provided meaningful intellectual contributions to the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
See-Too, W., Ee, R., Lim, YL. et al. AidP, a novel N-Acyl homoserine lactonase gene from Antarctic Planococcus sp.. Sci Rep 7, 42968 (2017). https://doi.org/10.1038/srep42968
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep42968
This article is cited by
-
Efficacy of ceftiofur N-acyl homoserine lactonase niosome in the treatment of multi-resistant Klebsiella pneumoniae in broilers
Veterinary Research Communications (2023)
-
Pseudomonas aeruginosa quorum sensing inhibition by clinical isolate Delftia tsuruhatensis 11304: involvement of N-octadecanoylhomoserine lactones
Scientific Reports (2019)
-
Characterization of a novel N-acylhomoserine lactonase, AidP, from Antarctic Planococcus sp.
Microbial Cell Factories (2018)
-
Characterization of AiiK, an AHL lactonase, from Kurthia huakui LAM0618T and its application in quorum quenching on Pseudomonas aeruginosa PAO1
Scientific Reports (2018)
-
Complete genome of Arthrobacter alpinus strain R3.8, bioremediation potential unraveled with genomic analysis
Standards in Genomic Sciences (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.