Abstract
The complex of two membrane proteins, sensory rhodopsin II (NpSRII) with its cognate transducer (NpHtrII), mediates negative phototaxis in halobacteria N. pharaonis. Upon light activation NpSRII triggers a signal transduction chain homologous to the two-component system in eubacterial chemotaxis. Here we report on crystal structures of the ground and active M-state of the complex in the space group I212121. We demonstrate that the relative orientation of symmetrical parts of the dimer is parallel (“U”-shaped) contrary to the gusset-like (“V”-shaped) form of the previously reported structures of the NpSRII/NpHtrII complex in the space group P21212, although the structures of the monomers taken individually are nearly the same. Computer modeling of the HAMP domain in the obtained “V”- and “U”-shaped structures revealed that only the “U”-shaped conformation allows for tight interactions of the receptor with the HAMP domain. This is in line with existing data and supports biological relevance of the “U” shape in the ground state. We suggest that the “V”-shaped structure may correspond to the active state of the complex and transition from the “U” to the “V”-shape of the receptor-transducer complex can be involved in signal transduction from the receptor to the signaling domain of NpHtrII.
Similar content being viewed by others
Introduction
Archaebacterial photoreceptors mediate phototaxis by regulating cell motility through two-component signaling cascades like those found in chemotaxis signaling chains of enteric bacteria1. The photoreceptor sensory rhodopsin II from N. pharaonis (NpSRII) in complex with its cognate transducer NpHtrII serves as a model system for studying transmembrane signal transfer. This complex displays a 2:2 stoichiometry where two transducers are flanked by two receptors. The long rod-shaped cytoplasmic domain consists of two HAMP domains (present in Histidine kinases, Adenyl cyclases, Methyl-accepting proteins and Phosphatases2,3), which are followed by a methylation domain4 involved in the adaptation processes and a signaling domain which harbors the CheW/CheA binding site5. HAMP subunits contain two amphiphilic helices (AS1 and AS2) joined by a non-helical connector. The first HAMP domain (HAMP1, amino acid residues 83–135) and the second HAMP domain (HAMP2, amino acid residues 157–210) are connected by an α-helical linker (Fig. 1). Despite high abundance of HAMP domains in different classes of proteins, their role and function is still unclear2,6,7. For understanding the signal transfer from the receptor to the transducer it is mandatory to determine the exact structure of the binding surface connecting NpSRII with its cognate transducer. Currently, the structure of the truncated membrane domain has been determined, however for the adjacent proximal region only indirect experimental evidences8,9,10,11 are available. Here, we report on new crystal structures of the ground and active M-state of NpSRII/NpHtrII1–157 complexes, in the space group I212121. Interestingly, the orientation of symmetrical parts of the dimer of two proteins is “U”-shaped contrary to the “V”-shaped quaternary structure reported previously (space group P21212)12. The detailed structure of the retinal-binding pocket remains the same. Our computer modeling data reveals that the HAMP domain in its resting conformation6 acquires complimentary intermolecular interactions with the receptor only in the “U”-shaped conformation. We suggest a biological relevance of this finding and propose that activation of the receptor/transducer complex involves a transition from the “U”- to “V”-shaped quaternary structures.
The dimer of two NpHtrII proteins is flanked by two NpSRII proteins. A-G, TM1 and TM2 are the transmembrane helices. Cytoplasmic part of NpHtrII consists of two HAMP domains (HAMP1 and HAMP2) connected by an α-helical linker (Inter-HAMP) and the kinase control module. Primes denote symmetry mates of the complex.
The previously published X-Ray diffraction data12,13 on the NpSRII/NpHtrII complex only showed electron densities for the transmembrane domain of the transducer but not for its adjacent HAMP domain. These crystals of the wild type NpSRII in complex with the shortened NpHtrII (NpHtrII1–157 and NpHtrII1–114) belonging to the space group P21212 revealed an overall “V”-shaped topology with an opening to the cytosplasmic side. Our homology modeling of the HAMP domain attached to NpHtrII of the “V”-shaped structure did not show any direct contacts that would explain the lack of an electron density in this region. Contrary to these findings, the experiments by various biophysical methods8,9,10,11 demonstrated an intimate interaction of TM2 of the transducer with the E-F loop of the receptor. This discrepancy could be accounted for by the influence of crystallization conditions on the global arrangement of proteins. Previous studies have already shown that modification of crystallization conditions can result in different functional conformations of a protein14.
Results
In an intensive crystallization screening we obtained two new crystal forms displaying space groups with the I212121 and P64 symmetry. Resolution for the both ground and M states was 1.9 Å in the case of the I212121 space group and 2.5 Å in the case of the P64 space group. Since no substantial discrepancies were found between ground state protein structures in both space groups, we will discuss the structure in the I212121 space group as it has higher resolution. When comparing wild type structures solved in the I212121 and previously solved in the P21212 space groups it is seen that the structure of NpSRII is nearly the same (with backbone RMS deviation of 0.22 Å for ordered regions) and is highly similar to the structure of NpSRII when crystallized alone (with backbone RMS deviation of 0.27 Å for ordered regions)15,16,17.
Signal transduction mechanism
The generally accepted mechanism5,18,19,20,21 of the signal transduction in the complex is the following. After isomerization, the retinal looses a proton from the Schiff base, which is then translocated to the Schiff base counterion22 (Asp75 in case of NpSRII). This event leads to the charge rearrangement in the active site of the molecule, disorder of some water molecules of the active site and, therefore, breaking of the hydrogen bonds. This hydrogen bond network integrates helices of resting state NpSRII into a tight bundle and diminishing the strength of this H-bond system results into displacements of the transmembrane helices, in particular F and G, both parallel and perpendicular to the membrane. These shifts change the hydrophobic area of the receptor and have important implications for the signal transduction, as it will be discussed below.
Having two data sets from different space groups (I212121 and P21212), we checked whether different crystal contacts have a structural influence on the retinal-binding site. As can be seen from Fig. 2 there are almost no differences found for the ground state structures (Fig. 2a) as well as for the activated M intermediate (Fig. 2b), indicating that the different crystallization conditions and space groups have little, if any at all, influence on the functionally important regions within NpSRII. This is also substantiated if one analyzes the transition between the ground state and the M state (Fig. 3).
Upon photon absorption by the complex in the space groups I212121 as in P21212, the retinal chromophore switches from all-trans conformation to 13-cis and Lys-Cε simultaneously shifts towards the central water cluster17. The side chain of Asp75 is rotated by about 90° from its ground state position, thereby losing the hydrogen bond to Thr7913,23. Consequently, W3 becomes disordered and disappears in the M state. The pentagon of hydrogen bonds (Asp201-Oδ···W2···W3···Arg72-Nε···W1) does not exist any longer in the M state because molecules W1 and W3 have vanished23. Consequently, helices C and G of NpSRII have more freedom to move independently. The water molecule W2′ is located close to Asp75 and Arg72. Thus, the signal is generated by the retinal isomerization and then propagates to the interface between the receptor and the transducer. The signal transmission is driven by the rearrangement of the hydrogen bond network, mediated by water molecules, which leads to the movement of the receptor helices described above due to the loss of connectivity between C and G helices.
The movement of the helix G of NpSRII initiates the movement of TM2 in NpHtrII. All hydrogen bonds, between the receptor and the transducer, observed for the ground state are still intact in the M state. A piston like movement of the cytoplasmic end of TM2 of about 0.5 Å and rotation of 19° (15° in P21212) are observed and accompanied by alteration in the direction of Tyr199-Asn74 hydrogen bond. The rotation of the transducer’s TM2 has also been observed by EPR spectroscopy5. The similarity of the differences between the ground and the M states in the new type of crystals shows an overall robustness of the signal transduction mechanism in the membrane part of the complex.
Quaternary structure of the complex
Having established that the fine structure of the proteins itself in the complex in the two crystal forms is unperturbed, we analyzed the overall structures in the crystal lattices. It is evident from Fig. 4 that the two receptors are oriented nearly parallel to the transducer and form a “U”-shaped binding crevice in the new crystal form I212121 compared to the “V”-shaped structure in the original P21212 symmetry. The angle between the G helices of NpSRII in the “U”-shape and the “V”-shape is 11°. A single NpSRII monomer of the “U”-shape dimer can be superimposed with the corresponding monomer of the “V”-shape dimer by a rigid body rotation by an angle of 8.5° (see Fig. 4). These large differences between two quaternary structures might account for the altered HAMP domain-NpSRII binding interfaces. Unfortunately, the electron density in this region is too weak in both cases, presumably due to interactions of the proteins in the crystal lattice. However, molecular modeling revealed important details about interactions of the HAMP domain within the complex. We have prepared homology models of the NpHtrII HAMP domains and modeled their possible position relative to the transducer’s transmembrane part and to the receptor NpSRII starting from the known structures of the HAMP domains6,24,25 (PDB codes 1ASW and 3LNR). We found that only the “U”-shape structure allows a close interaction between NpSRII and the HAMP domain, allowing it to be compactly fitted into the “U”-shape topology (Fig. 5). On the other hand, the model based on the “V”-shaped topology does not show any contacts between the HAMP domain and NpSRII, which leads to weaker interactions between these proteins.
Helices G are shown as cylinders. The angle between the two helices G belonging to the same NpSRII molecule in the “U”- and “V” shapes is shown. The axis of rotation of NpSRII as a rigid body during the transition from the “U”- to “V”-shape is shown in red and the corresponding angle is presented. The structure solved in the I212121 space group (“U”-shape, PDB ID 4GY6) is shown in yellow, the structure solved in the P21212 space group (“V”-shape, PDB ID 1H2S) is shown in green. Primes denote symmetry mates. Solid black lines illustrate the approximate membrane boundaries with “IC” designating the intracellular side and “EC” designate the extracellular side.
(A) “V”-shaped NpSRII/NpHtrII complex in P21212 space group. (B) “U”-shaped NpSRII/NpHtrII complex in I212121 space group. Figures on the left show two subsequent crystal layers as seen along the membrane plane. Figures on the right show a single crystal layer as seen perpendicular to the membrane plane. NpSRII molecules are shown in green; parts of NpHtrII molecules resolved in the crystal structure are shown in blue. Models of NpHtrII that continue to the HAMP domain are shown in red. Residues 83–136 in these models are modeled by homology with NMR structure 2ASW. In (A) a big steric clash can be seen between the model of the HAMP domain from one crystal layer and the corresponding NpHtrII molecule from the other crystal layer. There are no observed steric clashes between the model of the HAMP domain from one layer and NpSRII/NpHtrII molecules from the other layer in (B).
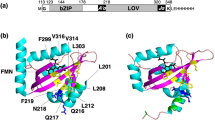
The surface of NpSRII includes a unique group of charged and polar residues at the cytoplasmic ends of helix F (Lys157, Ser158, Arg162, Arg164, and Asn165) and G (Asp214) (Fig. 6). For Ser154 to Lys157 an interaction with the transducer has been proven by ssNMR experiments26. According to our model, this patch of amino acids could interact with Asp85, Ser91, Asp115, Glu116 and Asp119 on TM2 via electrostatic interactions. Furthermore, Lys21 of TM1 and Asp214 of the helix G of the receptor could form a salt bridge and might contribute to the overall stability of the complex conveyed by the HAMP domain-receptor interaction. The negatively charged cytoplasmic domain Gly101-Asp-Gly-Asp-Leu-Asp106 of TM2 of NpHtrII, which is conserved among various species27 and was suggested to interact with NpSRII previously28, apparently has no possible interactions with the receptor in our model (Fig. 6). We should also note that these results well explain the spectroscopic data that indicates that the EF loop interacts with the transducer8,9,26.
Interacting patches in the receptor and the transducer are shown and the G-D-G-D-L-D patch of the transducer proposed previously in literature as interacting with NpSRII. Positive charges are mapped in blue and negative are mapped in red. The expected salt bridges are: Asp214 (NpSRII) to Lys21 (NpHtrII), Lys157 (NpSRII) to Asp119 and Glu116 (NpHtrII).
Normal mode analysis
To further support our hypothesis that signal transduction may involve transition from the “U” to “V”-shaped complex structure, we have constructed an elastic network model (ENM)29 and performed a normal mode analysis (NMA) for the both “U” and “V”-shape structures as the starting models using the rotation translation block (RTB) approach30. The first two normal modes in both cases corresponded to the relative rotation of the subunits in the plane of the membrane. The third mode in corresponded to the “U”-to-“V” transition observed in the crystal structures. To see the effect of a lipid membrane, we repeated the NMA analysis for the NpSRII complex in the “U”-state inserted into a POPC bilayer consisting of 256 lipid molecules. We prepared the initial model using CHARMM-GUI31,32. Then, we equilibrated the bilayer for 1 ns using GROMACS v. 5.033. The protein structure was fixed during the equilibration. In this case, we did not identify a single mode responsible for the “U”-to-“V” transition. However, using a linear combination of ten lowest modes, we could find a motion that reduced the overall RMSD of the transition from 2.9 Å to 1.0 Å.
These finding show that the direction of the U-to-V transition coincides with one or several of temperature fluctuations in the complex, thereby requiring minimal energy to take place. We also clearly observe a piston-like shift of both transmembrane helices of the transducer with respect to NpSRII, which occurs concomitantly with the rotation of the receptors (Fig. 7). When starting from the “U”-shape structure, the complex transitions into a “V”-shape-like model, which closely resembles the crystal structure of the active state (overall RMSD 1.04 Å, also see Supplementary Movie 1).
Retinal isomerization leads to breaking of the receptor into two domains that move in the opposite directions perpendicular to the membrane (1); shift of its transmembrane helices relative to each other creates a hydrophobic mismatch that has to be compensated through a tilt rotation of NpSRII (2) around the intracellular point of contact between NpSRII and NpHtrII (3); this motion levers TM1 and TM2 of the conjugated NpHtrII for a piston-like shift towards the cytosol (4), which concomitantly drags the helices of the other protomer, reversing the mechanism in the symmetric counterpart of the receptor (5) and (6). Solid black lines illustrate the approximate membrane boundaries with “IC” designating the intracellular side and “EC” designate the extracellular side.
Discussion
Multiple experimental studies (calorimetric10, solution NMR11, EPR experiments34) have long predicted the tight binding interface between the transducer and the receptor. In the previously determined structure of the complex (space group P21212) quaternary structure has a “V”-shape and there was no possibility of interaction between the NpHtrII’s HAMP1 domain and NpSRII. Here, we have presented the structure in the space group I212121, where the whole complex is more compact (quaternary structure has a “U”-shape) and the receptor can extensively interact with the transducer’s HAMP domain.
In contrast to the ground state, in the active state of the complex, the interaction between NpSRII and NpHtrII is considerably weakened (dissociation constants are 0.10 μM and 15 μM correspondingly)35,36. This effect can originate from a perturbation of the linker region. However, it cannot be caused by the transmembrane part of the proteins, since the interactions (especially, the hydrogen bonds interactions) are nearly the same in both states. Interestingly, the HAMP domain itself (NpHtrIIG83-Q149) interacts with the receptor in the ground state, but not in the M state11. Since the HAMP domain is sterically quite constrained hampering conformational changes in the “U”-shape, it may be that a transition from “U” to “V”-shape is involved in the signal propagation.
We have recently demonstrated via molecular dynamics study that the HAMP domain adopts an asymmetric conformation in its resting state, in which the protomers are longitudinally shifted by 1.3 Å with respect to each other presumably causing a deviation of the cytoplasmic part of NpHtrII relative to its transmembrane part6. Thus, it may happen that the “U”- and “V”–shaped complexes just correspond to the symmetric and asymmetric states of the HAMP domains and, therefore, correspond to different functional states, namely the ground and the active state. We suggest that the “U”-shape corresponds to a symmetric state of the HAMP domain, where it is stabilized in the rotated conformation via the aforementioned electrostatic interactions with the receptor. In this state, the HAMP domain adopts a compact conformation which has been correlated with the active state of the complex37,38. This conformational state is made possible by a high flexibility of the linker, especially in the region of Gly83-Gly84 amino acid repeat. This hypothesis is supported by another well-established fact: the G83F mutation completely inhibits signal transduction39. Indeed, replacing a small glycine by a considerably larger phenylalanine would limit the ability of the HAMP domain to rotate relative to the membrane part of the transducer and thus forbids the ground state. In line with this reasoning, our data on the crystal structure of the NpSRII/NpHtrII1–135-G83F mutant demonstrate different crystal packing. Although the crystallization conditions were the same as for the wild type “U”-shaped complex in the P64 space group, in this case the space group is P21212 giving the “V”-shape structure similar to those observed before12.
Another evidence in favor of the functional significance of the “U”-shaped membrane protein NpSRII/NpHtrII complex was published recently40. The authors studied how photoexcitation of NpSRII affects the structures of the complex. They conducted two series of 90 nsec molecular dynamics simulations of the complex linked with a modeled HAMP domain in the lipid bilayer using the published “V”-shaped structures of the ground and the M intermediate states12,13 as starting models. The most significant finding was that the orientations of the two NpSRII molecules changed in going from the ground state to the M state: their ‘closed’ ground state can be correlated to the “U”-shaped topology introduced in this paper, whereas their more flexible M state might represent the “V”-shape.
Finally, a very similar transition has been observed in one of the histidine kinases, PhoQ, via disulfide crosslinking41. The PhoQ sensor belongs to the family of two-component signal transduction systems. Similarly to NpHtrII, it contains a HAMP domain and has an analogous TM domain42. TM domain, HAMP and periplasmic domains of PhoQ were investigated using disulfide-scanning mutagenesis. The obtained crosslinking data were best explained by the two-state modeling approach than the one-state modeling, where the signal is suggested to be transferred through scissor-like transition from one state to the other.
Thus, it seems that the mechanism suggested here does not appear to be unique for NpSRII/NpHtrII complex, but rather might be widespread among two-component signaling systems in general. It also does not contradict with the existing models of signal transduction through the HAMP domains. Molecular interactions between NpSRII and HAMP can transition the latter from the compact state to the splayed state as reported before7. Other models like partially disordered HAMP as in the dynamic bundle model43 or alternating dynamics of NPHtrII subdomains38 deal with the signal transfer from the membrane proximal region to the tip of the NPHtrII cytoplasmic and can be incorporated into our model of signal transfer from the membrane domain to the first HAMP domain (HAMP1).
Concluding Remarks
Summarizing, the “U”- and “V”-shaped conformations of the NpSRII/NpHtrII complex may be crucial for the mechanism of signal propagation spanning the membrane domain and feeding into the HAMP domain. This work also represents one more evidence that different crystallization conditions and/or different crystal packing may force the protein to adopt different conformations corresponding to discrete functional states. Obviously, it is not a completely new observation; it was shown that the extracellular domain of metabotropic glutamate receptor can adopt an active-like conformation even in the absence of the ligand14. However, to the best of our knowledge, this is the first such demonstration for the case of membrane proteins where the different space groups could potentially trap different functional states of the transmembrane part of proteins and which might be important for signal transfer in arrays of receptors as it was proposed for chemoreceptors44.
Methods
Protein preparation
The NpSRII and NpHtrII proteins were produced in E. coli, purified, reconstituted into a functional complex, and resolubilized as described before13,45.
Crystallization
We used a new in meso46 approach to obtain crystals. The complex in a crystallization buffer (150 mM NaCl, 25 mM Na/K phosphate, pH 5.1, n-octyl-b-D-glucopyranoside) was added to the lipid phase, formed on base of monooleoyl (Nu-Chek Prep). Best crystals were grown at 22 °C using 1 M Na/K phosphate, pH 5.6 and trehalose as a precipitant.
Trapping of the M intermediate, data collection and refinement
The optimal procedure of the M state trapping was as described before15. X-ray diffraction data were collected at beamline ID14-1 of the European Synchrotron Radiation Facility (ESRF), Grenoble, France. Radiation dose was the same for the ground and M state and kept under reasonable limit to avoid influence of the radiation damage on the M state structure47. Data was integrated using MOSFILM48 and scaled with SCALA49 from the CCP4 program suite50. Molecular replacement was performed using MOLREP51 for a polyalanine model (from Protein Data Bank accession number 1H2S) and gave a unique solution. Structure refinement was done with Phenix52. The occupancy determination of the M state and the structure refinement was performed as before13.
Modeling of the HAMP domain
In order to better understand the influence of the crystal packing on the NpHtrII structure, we modeled its possible continuation to the first HAMP domain (residues 83–136) in two space groups I212121 and P21212, which is not observed in the crystal (Fig. 5). Based on the previous modeling studies for this region based on homology with the available HAMP domain structures6,53, we assumed that the linker between the helix TM2 and the HAMP domain adopts an alpha-helical conformation. The HAMP domain was modeled by homology (based on PDB code 2ASW). We optimized the models using a symmetry module from SAMSON modeling package54. Details of the modeling are provided in SI. We used PyMOL to calculate vacuum electrostatics potentials on molecular surfaces and to produce illustrations for this report55.
Additional Information
Accession codes: Coordinates and structure factors have been deposited in the Protein Data Bank with the following accession numbers: 5JJE (NpSRII/HtrII-157 ground state), 5JJJ (NpSRII/HtrII-135 ground state), 5JJF (the M state of NpSRII/HtrII-157), 5JJN (NpSRII/HtrIIG83F-135).
How to cite this article: Ishchenko, A. et al. New Insights on Signal Propagation by Sensory Rhodopsin II/Transducer Complex. Sci. Rep. 7, 41811; doi: 10.1038/srep41811 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Hoff, W. D., Jung, K. H. & Spudich, J. L. Molecular mechanism of photosignaling by archaeal sensory rhodopsins. Annu. Rev. Biophys. Biomol. Struct. 26, 223–58 (1997).
Ferris, H. U. et al. The mechanisms of HAMP-mediated signaling in transmembrane receptors. Structure 19, 378–85 (2011).
Wang, J., Sasaki, J., Tsai, A.-L. & Spudich, J. L. HAMP Domain Signal Relay Mechanism in a Sensory Rhodopsin-Transducer Complex. J. Biol. Chem. 287, 21316–25 (2012).
Koch, M. K., Staudinger, W. F., Siedler, F. & Oesterhelt, D. Physiological sites of deamidation and methyl esterification in sensory transducers of Halobacterium salinarum. J. Mol. Biol. 380, 285–302 (2008).
Wegener, A. A., Klare, J. P., Engelhard, M. & Steinhoff, H. J. Structural insights into the early steps of receptor-transducer signal transfer in archaeal phototaxis. EMBO J. 20, 5312–9 (2001).
Gushchin, I. Y., Gordeliy, V. I. & Grudinin, S. Role of the HAMP Domain Region of Sensory Rhodopsin Transducers in Signal Transduction. Biochemistry 50, 574–580 (2010).
Airola, M. V., Watts, K. J., Bilwes, A. M. & Crane, B. R. Structure of concatenated HAMP domains provides a mechanism for signal transduction. Structure 18, 436–48 (2010).
Wegener, A. A., Chizhov, I., Engelhard, M. & Steinhoff, H. J. Time-resolved detection of transient movement of helix F in spin-labelled pharaonis sensory rhodopsin II. J. Mol. Biol. 301, 881–91 (2000).
Yang, C.-S., Sineshchekov, O., Spudich, E. N. & Spudich, J. L. The Cytoplasmic Membrane-proximal Domain of the HtrII Transducer Interacts with the E-F Loop of Photoactivated Natronomonas pharaonis Sensory Rhodopsin II. J. Biol. Chem. 279, 42970–42976 (2004).
Hippler-Mreyen, S. Probing the Sensory Rhodopsin II Binding Domain of its Cognate Transducer by Calorimetry and Electrophysiology. J. Mol. Biol. 330, 1203–1213 (2003).
Sudo, Y. et al. Linker region of a halobacterial transducer protein interacts directly with its sensor retinal protein. Biochemistry 44, 6144–52 (2005).
Gordeliy, V. I. et al. Molecular basis of transmembrane signalling by sensory rhodopsin II-transducer complex. Nature 419, 484–7 (2002).
Moukhametzianov, R. et al. Development of the signal in sensory rhodopsin and its transfer to the cognate transducer. Nature 440, 115–9 (2006).
Kunishima, N. et al. Structural basis of glutamate recognition by a dimeric metabotropic glutamate receptor. Nature 407, 971–7 (2000).
Gushchin, I. et al. Active state of sensory rhodopsin II: structural determinants for signal transfer and proton pumping. J. Mol. Biol. 412, 591–600 (2011).
Royant, A. et al. X-ray structure of sensory rhodopsin II at 2.1-A resolution. Proc. Natl. Acad. Sci. USA 98, 10131–6 (2001).
Luecke, H., Schobert, B., Lanyi, J. K., Spudich, E. N. & Spudich, J. L. Crystal structure of sensory rhodopsin II at 2.4 angstroms: insights into color tuning and transducer interaction. Science 293, 1499–503 (2001).
Klare, J. P., Bordignon, E., Engelhard, M. & Steinhoff, H.-J. Sensory rhodopsin II and bacteriorhodopsin: Light activated helix F movement. Photochem. Photobiol. Sci. 3, 543 (2004).
Spudich, J. L., Sineshchekov, O. A. & Govorunova, E. G. Mechanism divergence in microbial rhodopsins. Biochim. Biophys. Acta - Bioenerg. 1837, 546–552 (2014).
Inoue, K., Tsukamoto, T. & Sudo, Y. Molecular and evolutionary aspects of microbial sensory rhodopsins. Biochim. Biophys. Acta - Bioenerg. 1837, 562–577 (2014).
Klare, J. P., Bordignon, E., Engelhard, M. & Steinhoff, H.-J. Transmembrane signal transduction in archaeal phototaxis: the sensory rhodopsin II-transducer complex studied by electron paramagnetic resonance spectroscopy. Eur. J. Cell Biol. 90, 731–9 (2011).
Spudich, E. N., Zhang, W., Alam, M. & Spudich, J. L. Constitutive signaling by the phototaxis receptor sensory rhodopsin II from disruption of its protonated Schiff base-Asp-73 interhelical salt bridge. Proc. Natl. Acad. Sci. USA 94, 4960–5 (1997).
Edman, K. et al. Early Structural Rearrangements in the Photocycle of an Integral Membrane Sensory Receptor. Structure 10, 473–482 (2002).
Hulko, M. et al. The HAMP domain structure implies helix rotation in transmembrane signaling. Cell 126, 929–40 (2006).
Nishikata, K., Fuchigami, S., Ikeguchi, M. & Kidera, A. Molecular modeling of the HAMP domain of sensory rhodopsin II transducer from Natronomonas pharaonis. Biophysics (Oxf). 6, 27–36 (2010).
Etzkorn, M. et al. Complex formation and light activation in membrane-embedded sensory rhodopsin II as seen by solid-state NMR spectroscopy. Structure 18, 293–300 (2010).
Seidel, R. et al. The primary structure of sensory rhodopsin II: a member of an additional retinal protein subgroup is coexpressed with its transducer, the halobacterial transducer of rhodopsin II. Proc. Natl. Acad. Sci. USA 92, 3036–40 (1995).
Pebay-Peyroula, E., Royant, A., Landau, E. M. & Navarro, J. Structural basis for sensory rhodopsin function. Biochim. Biophys. Acta 1565, 196–205 (2002).
Tirion, M. Large Amplitude Elastic Motions in Proteins from a Single-Parameter, Atomic Analysis. Phys. Rev. Lett. 77, 1905–1908 (1996).
Tama, F., Gadea, F. X., Marques, O. & Sanejouand, Y. H. Building-block approach for determining low-frequency normal modes of macromolecules. Proteins 41, 1–7 (2000).
Lee, J. et al. CHARMM-GUI Input Generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM Simulations Using the CHARMM36 Additive Force Field. J. Chem. Theory Comput. 12, 405–413 (2016).
Jo, S., Kim, T. & Im, W. Automated Builder and Database of Protein/Membrane Complexes for Molecular Dynamics Simulations. PLoS One 2, e880 (2007).
Abraham, M. J. et al. GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1, 19–25 (2015).
Bordignon, E. et al. Structural Analysis of a HAMP Domain. J. Biol. Chem. 280, 38767–38775 (2005).
Sudo, Y., Iwamoto, M., Shimono, K. & Kamo, N. Pharaonis phoborhodopsin binds to its cognate truncated transducer even in the presence of a detergent with a 1:1 stoichiometry. Photochem. Photobiol. 74, 489–94 (2001).
Sudo, Y., Yamabi, M., Iwamoto, M., Shimono, K. & Kamo, N. Interaction of Natronobacterium pharaonis phoborhodopsin (sensory rhodopsin II) with its cognate transducer probed by increase in the thermal stability. Photochem. Photobiol. 78, 511–6 (2003).
Doebber, M. et al. Salt-driven Equilibrium between Two Conformations in the HAMP Domain from Natronomonas pharaonis: THE LANGUAGE OF SIGNAL TRANSFER? J. Biol. Chem. 283, 28691–28701 (2008).
Orekhov, P. S. et al. Signaling and Adaptation Modulate the Dynamics of the Photosensoric Complex of Natronomonas pharaonis. PLOS Comput. Biol. 11, e1004561 (2015).
Yang, C. S. & Spudich, J. L. Light-induced structural changes occur in the transmembrane helices of the Natronobacterium pharaonis HtrII transducer. Biochemistry 40, 14207–14 (2001).
Nishikata, K., Ikeguchi, M. & Kidera, A. Comparative Simulations of the Ground State and the M-Intermediate State of the Sensory Rhodopsin II-Transducer Complex with a HAMP Domain Model. Biochemistry 51, 5958–5966 (2012).
Molnar, K. S. et al. Cys-scanning disulfide crosslinking and bayesian modeling probe the transmembrane signaling mechanism of the histidine kinase, PhoQ. Structure 22, 1239–51 (2014).
Goldberg, S. D., Clinthorne, G. D., Goulian, M. & DeGrado, W. F. Transmembrane polar interactions are required for signaling in the Escherichia coli sensor kinase PhoQ. Proc. Natl. Acad. Sci. USA 107, 8141–6 (2010).
Schultz, J. E. & Natarajan, J. Regulated unfolding: a basic principle of intraprotein signaling in modular proteins. Trends Biochem. Sci. 38, 538–45 (2013).
Parkinson, J. S., Hazelbauer, G. L. & Falke, J. J. Signaling and sensory adaptation in Escherichia coli chemoreceptors: 2015 update. Trends Microbiol. 23, 257–266 (2015).
Hohenfeld, I. P., Wegener, A. A. & Engelhard, M. Purification of histidine tagged bacteriorhodopsin, pharaonis halorhodopsin and pharaonis sensory rhodopsin II functionally expressed in Escherichia coli. FEBS Lett. 442, 198–202 (1999).
Caffrey, M. & Cherezov, V. Crystallizing membrane proteins using lipidic mesophases. Nat. Protoc. 4, 706–31 (2009).
Borshchevskiy, V. I., Round, E. S., Popov, A. N., Büldt, G. & Gordeliy, V. I. X-ray-radiation-induced changes in bacteriorhodopsin structure. J. Mol. Biol. 409, 813–25 (2011).
Leslie, A. G. W. The integration of macromolecular diffraction data. Acta Crystallogr. D. Biol. Crystallogr. 62, 48–57 (2006).
Evans, P. Scaling and assessment of data quality. Acta Crystallogr. D. Biol. Crystallogr. 62, 72–82 (2006).
Collaborative Computational Project, N. 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D. Biol. Crystallogr. 50, 760–3 (1994).
Vagin, A. & Teplyakov, A. MOLREP : an Automated Program for Molecular Replacement. J. Appl. Crystallogr. 30, 1022–1025 (1997).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D. Biol. Crystallogr. 66, 213–21 (2010).
Hartmann, M. D. et al. A soluble mutant of the transmembrane receptor Af1503 features strong changes in coiled-coil periodicity. J. Struct. Biol. 186, 357–366 (2014).
Grudinin, S. & Redon, S. Practical modeling of molecular systems with symmetries. J. Comput. Chem. 31, 1799–1814 (2010).
DeLano, W. L. The PyMOL Molecular Graphics System, Version 1.2r3pre, Schrödinger, LLC.
Acknowledgements
We are thankful to E. Pebay-Peyroula for the support of this project and C. Baeken for the assistance with protein preparation. We acknowledge Structural Biology Group of European Synchrotron Radiation Facility for granting access to the synchrotron beamlines. A.I. wishes to thank Stephane Redon for hosting at INRIA NANO-D team where symmetry modeling initiated. The work was supported by the program “Chaires d’excellence” edition 2008 of ANR France, CEA(IBS) – HGF(FZJ) STC 5.1 specific agreement. The molecular dynamics simulations were performed using the JURECA supercomputer at the Research Center Jülich. The work was supported by the Russian Science Foundation research project 14-14-00995. The work used the platforms of the Grenoble Instruct Centre (ISBG; UMS 3518 CNRS-CEA-UJF-EMBL) with support from FRISBI (ANR-10-INSB-05-02) and GRAL (ANR-10-LABX-49-01) within the Grenoble Partnership for Structural Biology (PSB).
Author information
Authors and Affiliations
Contributions
A.I. expressed, purified, and crystallized the proteins, collected and processed data, refined and analyzed the structures and made a major contribution to the preparation of the manuscript draft; E.R. contributed to initial protein production, purification, crystallization and data collection; V.B. took part in data collection, processing and analysis; S.G. performed computer modeling and wrote the manuscript; I.G. contributed to computer modeling and wrote the manuscript; J.K. purified NpHtrII at the initial stage of the project; A.R., V.P. and P.U. contributed to the nanovolume crystallization of the proteins and crystal harvesting; T.B. cloned DNA plasmids; M.E. participated in initiating the project and contributed to the manuscript text; G.B. participated in initiating the project and contributed to the manuscript text; V.G. designed the overall strategy, managed the project and oversaw manuscript preparation; he also obtained the first crystals, collected the data and performed the initial data analysis.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Ishchenko, A., Round, E., Borshchevskiy, V. et al. New Insights on Signal Propagation by Sensory Rhodopsin II/Transducer Complex. Sci Rep 7, 41811 (2017). https://doi.org/10.1038/srep41811
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep41811
This article is cited by
-
Molecular model of a sensor of two-component signaling system
Scientific Reports (2021)
-
The Blue-Green Sensory Rhodopsin SRM from Haloarcula marismortui Attenuates Both Phototactic Responses Mediated by Sensory Rhodopsin I and II in Halobacterium salinarum
Scientific Reports (2019)
-
Photoreaction pathways and photointermediates of retinal-binding photoreceptor proteins as revealed by in situ photoirradiation solid-state NMR spectroscopy
Biophysical Reviews (2019)
-
Structural insights into the signalling mechanisms of two-component systems
Nature Reviews Microbiology (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.