Abstract
Base-calling accuracy is crucial for high-throughput DNA sequencing and downstream analysis such as read mapping and genome assembly. Accordingly, we made an endeavor to reduce DNA sequencing errors of Illumina systems by correcting three kinds of crosstalk in the cluster intensity data. We discovered that signal crosstalk between adjacent clusters accounts for a large portion of sequencing errors in Illumina systems, even after correcting color crosstalk caused by the overlap of dye emission spectra and phasing/pre-phasing caused by out-of-step nucleotide synthesis. Interestingly and importantly, spatial crosstalk between adjacent clusters is cluster-specific and often asymmetric, which cannot be corrected by existing deconvolution methods. Therefore, we introduce a novel mathematical method able to estimate and remove spatial crosstalk, thereby reducing base-calling errors by 44–69% at a given mapping rate from Illumina systems. Furthermore, the resolution gained from this work provides new room for higher throughput of DNA sequencing and of general measurement systems using fluorescence-based imaging technology. The resulting base-caller 3Dec is available for academic users at http://github.com/flishwnag/3dec. Not only does it reduce 62.1% errors compared to the standard pipeline, but also its implementation is fast enough for daily sequencing.
Similar content being viewed by others
Introduction
Illumina’s “sequencing-by-synthesis” technology is widely used for DNA sequencing in research and medicine. With this technology, billions of clusters of DNA templates are laid out on a flow cell and extended cycle-by-cycle using reversible terminators. The extended nucleotides of A, C, G, and T of each cycle are identified by their attached fluorescent dyes, which emit light when excited by laser. Each cycle generates four images capturing the emitted lights of the DNA clusters in four wavelength bands. Base-calling aims to reconstruct the underlying sequences from the fluorescence signals. In order to reduce base-calling errors, we need to enhance the signals using different types of data corrections. The first major correction involves removal of color crosstalk caused by the overlapping emission spectra of the four dyes. The principle of optimal color correction was presented1 in the context of capillary sequencing, and a similar scheme was proposed2 for the Illumina platform. The second major correction targets the crosstalk among adjacent cycles which results from out-of-step nucleotide synthesis of some DNA molecules. This phasing/pre-phasing issue has been modeled and effectively managed by various methods, including Illumina’s Bustard software and third-party software programs, such as FreeIbis3 and AYB4.
In this study, we focused on the crosstalk among spatially adjacent clusters. Ideally, the intensities of each cluster, as defined by image processing, correspond to just one DNA sequence. However, the observed intensities often contain signals from neighboring DNA sequences, other than the nominal one, and, hence, are not pure (see Fig. 1a). We showed that a large portion of base-calling errors were caused by this “spatial crosstalk”. Moreover, spatial crosstalk is cluster-specific and often asymmetric.
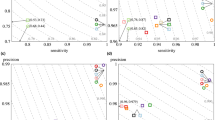
“C1” and “C2” are two neighbor clusters from tile s_4_1113 in the HiSeq2000 dataset. Panel (a,c): fluorescence signals (Y-axis) versus the cycle (X-axis): upper – “cluster 1”; lower – “cluster 2”. Panel (a) shows the signals before correction (Color crosstalk and phasing/pre-phasing has been removed using 3Dec, and the raw intensities are shown in Supplementary Fig. S4). Obviously, the signals of “Cluster 1” are contaminated by those of “Cluster 2”, and in the 13th cycle, the base is mistakenly called as T (Panel a), instead of A (Panel c). Panel (c) shows the signals after correction. Panel (b,d): the corresponding scatter plots of the intensity signals in (a) and (c). In each scatter plot, the x- and y-values of each point represent the signals of “cluster 2” and “cluster 1” in a specific cycle and channel, respectively. A point is marked “ × ” if the corresponding nucleotide in this cycle and channel appears in “cluster 1”, but not in “cluster 2”; it is marked “ + ” if the nucleotide in this cycle and channel appears in “cluster 2”, but not in “cluster 1”; it is marked “*” or “o” if the nucleotide appears in both or in neither, respectively. In the absence of spatial crosstalk (Panel (d)), the straight line passing through the centers of the clouds marked respectively by “o” and by “ + ” would be parallel to the x-axis. However, this straight line tilts upward in panel (b), indicating crosstalk from Cluster 2 to Cluster 1. The slope of this line corresponds to the spatial crosstalk coefficients from Sequence 2 to Cluster 1, as defined in Method, Equation (3).
To address the raised challenge, we developed an adaptive decorrelation method that estimates the extent of spatial crosstalk for each pair of adjacent clusters. The estimated spatial cross-talk matrix consisting of all pairwise coefficients is then used to correct the signals. Based on the method, we developed the base-calling software 3Dec to increase the sequencing accuracy for the Illumina platforms. The software 3Dec is available to academic users at http://github.com/flishwnag/3dec.
Results
Spatial crosstalk and its cluster-specific pattern
A large portion of base-calling errors were caused by “spatial crosstalk”. Specifically, we calculated the base-calling correlations between adjacent clusters, making use of the reads from bacteriophage PhiX174, which are often included as quality control in Illumina sequencers. The reads of three tiles of bacteriophage PhiX174 from GAII, HiSeq2000 and MiSeq, respectively, were mapped to the consensus reference genome using Bowtie25 (see Method). In all three tiles, the depths of mapped reads were more than 400x; thus, mismatches from the consensus were regarded as base-calling errors. Next, we calculated the percentage of mistaken calls identical to the bases of their nearest neighbor clusters of the same cycle. As a comparison, we also computed the percentage of mistaken calls identical to a random base in the tile of the same cycle (Fig. 2). As expected, the latter calculation, as shown in black lines, was approximately 25%, while the former calculation, as shown in red lines, ranged from 35% to 60% in the three datasets. The margins beyond the random setting imply that about 13% ( ) to 47% of base-calling errors are caused by spatial crosstalk from the nearest neighbors. The current Illumina pipeline uses a “chastity filter” to detect clusters with high impurity and discards them in base calling2. Without the “chastity filter”, spatial crosstalk would be even more serious in the original data.
) to 47% of base-calling errors are caused by spatial crosstalk from the nearest neighbors. The current Illumina pipeline uses a “chastity filter” to detect clusters with high impurity and discards them in base calling2. Without the “chastity filter”, spatial crosstalk would be even more serious in the original data.
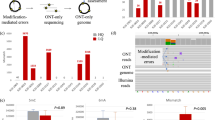
Among all base-calling errors from a tile of bacteriophage PhiX174, we calculated the percentage of calls that were identical to the bases of their nearest neighbor clusters of the same cycle. The percentages (Y axis) stratified by each cycle (X axis) are shown by the red lines in panel (a–c). As a comparison, the black lines show the percentages of mistaken calls identical to random calls of the same cycle. (a): data from the GAII tile s_4_21 without pass of any filter. (b): data from the HiSeq2000 tile s_4_1113 with the pass of the chastity filter. (c): data from the MiSeq tile s_4_1101 with the pass of both the chastity filter and the quality control (QC) filter. Under the random setting, the percentages are around 25% in all three cases, while the observed percentages in the red lines range from 35% to 60%. The margins beyond the random setting are thus caused by the spatial interference between adjacent clusters. (d) Illustration of spatial crosstalk cluster specificity, as indicated by a small region in tile “s_4_1113”. The X-axis and Y-axis are the coordinates of the clusters’ positions. Each black ‘ + ’ corresponds to a cluster, and each red-head, blue-tail arrow represents a spatial crosstalk coefficient larger than 0.05 from the tail to the head (see Method).
The novel decorrelation method
To address the challenge raised by cluster-specific spatial crosstalk, we developed an adaptive decorrelation. The method assumes that each cluster has one nominal DNA sequence fragment to be identified and that the fluorescence signals of each cluster are a weighted sum of the signals of the involved DNA fragments, whose weights are to be estimated. We refer to the weights as “spatial crosstalk coefficients” from the involved sequences. Without loss of generality, we first consider a pair of clusters indexed by i and j. The spatial crosstalk from Fragment j to Cluster i is measured by cij, and the crosstalk in the reverse direction is measured by cji (Method, Equation (3)). We note that cij and cji are specific to the cluster pair, but not necessarily identical.
The estimation of spatial crosstalk coefficients is motivated by the pairwise scatter plot of the signals. Figure 1b shows the scatter plot of the example given in Fig. 1a, in which four clouds are observed. More precisely, if during one cycle, one kind of nucleotide, say A, appears in DNA Fragment 2, but not in Fragment 1, then the corresponding point is marked “+”, and all these points form a cloud (see Fig. 1 for the meanings of other clouds). The slope of the line that passes through the origin and the center of Cloud “+” corresponds to cij(i = 1, j = 2), which is the crosstalk coefficient from Fragment 2 to Cluster 1. It obviously tilts upward, indicating the presence of crosstalk from Fragment 2 to Cluster 1. Based on this visualization, we define an objective function, and its minimizer gives an estimate of the crosstalk coefficient cij (See Method). The biases and standard errors of the crosstalk estimates were assessed by a bootstrap method (see Supplementary Fig. S2 and Supplementary Table S1 for some results).
The general spatial crosstalk model for a tile of intensities is as follows. The intensities of all the clusters correspond to a linear transformation of the intensities of all DNA fragments. This linear transformation can be represented by a spatial crosstalk matrix whose diagonal elements are ones and whose off-diagonal coefficient of Row i, Column j represents the crosstalk coefficient from Fragment j to Cluster i. To a good approximation, we can estimate each coefficient using the intensities of Cluster i and j. Since only the coefficients corresponding to spatially neighboring pairs are nonzero, the matrix is sparse. Our decorrelation of the spatial crosstalk is thus the multiplication of the inverse of the crosstalk matrix to the cluster signals.
The distribution of spatial crosstalk coefficients
Spatial crosstalk occurred frequently when the distance between cluster pairs was less than 3 pixels for the HiSeq2000 dataset. The detailed distributions of the estimated spatial crosstalk coefficients stratified by the distances between the involved cluster pairs can be found in Supplementary Figure S1. Obviously, the closer two clusters are, the more spatial crosstalk is likely to occur. Furthermore, we can define the total crosstalk into cluster i by summing up all  , over all j such that i ≠ j. It is shown in Supplementary Figure S3 that the error rate goes up as the total crosstalk effect increases.
, over all j such that i ≠ j. It is shown in Supplementary Figure S3 that the error rate goes up as the total crosstalk effect increases.
Cluster-specificity of spatial crosstalk
The extent of spatial crosstalk cannot be quantified by classical deconvolution models that assume a fixed point spread function. We illustrated this by the detailed spatial crosstalk pattern of a small region of the HiSeq2000 tile “s_4_1113” in Fig. 2d, where the arrows representing the spatial crosstalk coefficients are asymmetric. Spatial crosstalk occurs quite often among closely located clusters, but its pattern is unpredictable.
To quantify this, we selected several 2-D vectors, as shown in the first column of Supplementary Table S2. Then we calculated the coordinate differences from each cluster to their nearest neighbor clusters. The precision of the cluster coordinates was taken to be 0.1 pixel. All cluster pairs whose coordinate differences were equal to a preselected vector formed a group. Thus, the cluster pairs in such group were roughly of the same distance and orientation. We then calculated the variations of the spatial crosstalk coefficients between these cluster pairs, and the results are shown in the third and fourth column of Supplementary Table S3. The variations of the coefficients of a given distance and orientation are several-fold larger than the averages of their standard errors. This indicates that the coefficients cannot be determined by the distances and orientations of the cluster pairs; in other words, spatial crosstalk is cluster-specific and that its removal requires an adaptive correction method.
Software
Based on the above method, we developed a base-calling software program, termed 3Dec, including the corrections of color crosstalk, cycle crosstalk (otherwise known as phasing and pre-phasing), and spatial crosstalk. The pipeline of 3Dec is shown in Fig. 3. Its input is a CIF file defined from the image processing. After correcting color and cycle crosstalk (Step 1), 3Dec estimates each non-zero coefficient in the spatial crosstalk matrix (Step 2), then corrects spatial crosstalk (Step 3). The resulting intensities are cleaner and are used for base-calling in Step 4.
The input data of 3Dec are CIF files that include the florescence intensity values of each cluster in each cycle and each channel. Step 1: the software corrects color crosstalk and cycle crosstalk (aka. Phasing/pre-phasing). Step 2: Using the corrected data from Step 1, 3Dec estimates each non-zero coefficient in the spatial crosstalk matrix. Step 3: the pipeline of 3Dec corrects spatial crosstalk by multiplying the inverse of the spatial crosstalk matrix from Step 2 to the corrected intensities from Step 1. The resulting intensities are cleaner and are then used for base-calling in Step 4. On the other hand, in the alternative pipeline, the multiplication of the inverse of the spatial crosstalk matrix from Step 2 is applied to the raw data instead. Thus the resulting cluster intensities can be used by other base callers to improve accuracy.
Alternatively, 3Dec has a separate module of spatial crosstalk correction (See Fig. 3). The original cluster intensity data are taken as input by the module, and the corrected intensities can be applied interchangeably with other base-callers, such as Bustard and AYB (hereinafter termed 3Dec + Bustard and 3Dec + AYB in this paper). That is, the alternative pipeline 3Dec + AYB and 3Dec + Bustard carry out spatial crosstalk correction by the 3Dec module and base calling by AYB and Bustard, respectively. Thus the gains in their base-calling accuracy compared to AYB and Bustard represents the improvement of our decorrelation method of spatial crosstalk.
Software comparison
We compared the mapping rate, PF (perfectly matched) rate and error rate on various methods (see Method for details). The comparison results are shown in Table 1, in which three kinds of error rates are presented. The first one is the traditional error rate defined as the sum of the edit distances6 divided by the total base account of all mapped reads4,7. However, this measure may underestimate the performance of a better base-calling method. Under the same mapping criterion, the total numbers of mapped reads could be larger for a better base caller. Thus, when more low-quality reads were successfully mapped, more errors would be included too. Therefore, we report two other measures of error rates. The second one calculates rates by counting errors in a common subset of reads for each dataset. That is, at a given threshold on the edit distances, we obtained the set of reads that were successfully mapped to the reference for each base-calling method. Then we picked the subset of reads that are common to all methods. And the error rate was calculated based on this subset. The third one counts errors in those mapped reads obtained at a given mapping rate. For each given mapping rate, reads with the smallest edit distance to the consensus reference were selected for the calculation. Consequently equal number of reads were selected for each method, although the subsets of reads could differ from one base-caller to another. See details in Method, and also Supplementary Note for the differences among the three definitions of the error rates.
Figure 4a shows the error rates versus the mapping rates for the HiSeq2000 dataset. Zero error rate corresponds to the case of perfect match to the consensus (PF). Based on cluster intensity files corrected for spatial crosstalk by 3Dec and files without correction, Bustard and AYB, respectively, increased PF rates from 84.16% and 89.32% to 88.00% and 91.72% and reduced error rates at the 97.50% mapping rate from 0.406% and 0.244% to 0.227% and 0.137% (see Table 1 for more details). The error rates were reduced by 44.1% and 44.9%.
3Dec + AYB and 3Dec + Bustard signify spatial crosstalk decorrelation by 3Dec and base calling by AYB and Bustard, respectively. Panel (a,b): the error rates (Y-axis) versus the mapping rates (X-axis) on HiSeq2000 and GAII datasets, respectively. The error rates were calculated as follows. First, we mapped all reads to the consensus reference; then we sorted the mapped reads in the descending order of their edit distances to the reference; finally, we selected the top K reads for the error rate calculation, where K equals the number of total reads multiplied by a given mapping rate. (c) For the phage tile of HiSeq2000 s_4_1113, the red line and blue line show the percentages (Y axis) of mistaken calls identical to the bases of their nearest neighbor clusters stratified by each cycle (X axis), using respectively the original and the 3Dec-corrected cluster intensities. The black line, which shows the percentages of mistaken calls identical to random calls of the same cycle, serves as the baseline. The consistency between the blue and black lines indicates that almost no spatial crosstalk exists after the decorrelation. (d) The rates of perfect mapped reads stratified by the cluster chastity values. The x-axis represents the average chastity of the first 25 cycles of a cluster, and the y-axis represents the percentage of PF reads. 3Dec improves sequencing data quality more prominently in low-chastity clusters.
We also compared the base-calling accuracies by the results of DNA assembly. That is, we assembled the Phix174 genome by SparseAssembler8 for sequencing reads obtained by different base-calling methods. We generated random samples according to a specific coverage (5x, 10x and 20x), and assembled the genome based on each random sample. The N50s and maximum contig lengths reported in Table 2 were the averages of 100 random samples at each coverage. 3Dec + AYB achieved the best N50 and maximum contig length while 3Dec gave comparable results.
Speed of computation
Computationally, the pipeline of 3Dec in Fig. 3 is more efficient, while almost reaches the highest accuracy achieved by the alternative pipeline 3Dec + AYB in Fig. 3. On our ThinkPad W530 mobile workstation, 3Dec processed the HiSeq2000 data in 740 seconds while reducing 62.1% errors compared to Bustard. The pipeline of 3Dec is more than five times as fast as 3Dec + AYB (see the last column of Table 1). The runtime for an entire Hiseq flowcell (8 lanes X 200 M clusters, 101-cycle, pair-end) would take ~1.2 days on a 32GB RAM, 32-thread workstation. The current implementation of 3Dec is thus fast enough for daily sequencing.
The chastity score and base-calling accuracy
The Illumina pipeline uses the chastity score as a measure of cluster quality. In Fig. 4d, we show the improvement of 3Dec in terms of PF rates stratified by chastity scores. The lower chastity scores, the more improvement. Earlier it was shown that the improvement of base-calling accuracy is primarily achieved by the correction of spatial crosstalk. This implies that spatial crosstalk, to a great extent, accounts for the low chastity scores. To test the effectiveness of 3Dec in this regard, we re-extracted the cluster intensity files from the GAII image data but without eliminating any low chastity clusters deliberately. Figure 4b shows the error rates versus the mapping rates of the base-calling results. Based on the cluster intensity files corrected for spatial crosstalk by 3Dec, AYB increases PF rates from 91.03% to 93.61%, while reduces the error rates by 69.0% at the 95.53% mapping rate (see Fig. 4b and Table 1).
Justification of the adaptive decorrelation method
The effectiveness of decorrelation can be examined from two perspectives. First, we can check the pairwise scatter plot as shown in Fig. 1. After the decorrelation, the line representing the spatial crosstalk in the scatter plot of Fig. 1d becomes parallel to the x-axis, and the corrected intensities are shown in Fig. 1c. Second, we can calculate the base-calling correlations between adjacent clusters using the 3Dec base-calling results, as shown in Fig. 2b. When we did this, most correlations were eliminated, as shown in Fig. 4c.
Phred quality score
Currently, the quality scores of 3Dec + Bustard and 3Dec + AYB can be defined by Bustard and AYB respectively, and the development of a quality score module specific for 3Dec is still ongoing.
Discussion
In this article, we have reported, for the first time, an adaptive decorrelation that can correct the cluster-specific spatial crosstalk found in Illumina sequencing intensity files. The crosstalk was evidenced by the base-calling correlations between adjacent clusters. The new method can be used in several ways to improve Illumina technology. First, we can remove the base-calling errors caused by spatial crosstalk using the current cluster intensity files provided by the sequencers. Particularly, the method can be applied to the MiSeq model, which is the only NGS system approved by U.S. Food and Drug Administration (FDA) for clinical use9. Second, we can increase sequencing throughput by either increasing cluster densities or lowering the threshold of the chastity filter in the Illumina pipeline. The potentially more serious spatial crosstalk is then corrected by the proposed decorrelation method. This strategy can be tested on Illumina non-patterned sequencing platforms, such as HiSeq2000/2500 and MiSeq. Third, this study presents the decorrelation by the settings of non-patterned flow cells. Recently, Illumina released a new kind of sequencer using patterned flow cells characterized by uniform clustering. Yet quite a portion of clusters are polyclonal and discarded. It is anticipated that the proposed decorrelation method will be able to save some clusters by separating the different DNA clones for further base calling.
Imaging coupled with fluorescence technology is widely used in many high-throughput measurement systems of molecular biology. The proposed adaptive decorrelation provides a general tool to enhance resolution and remove crosstalk among neighboring target molecules.
Method
Model
In the standard Illumina pipeline, the raw image files generated by scanning each flow cell surface go through a series of processing steps, including template generation, registration, and intensity extraction. The outputs of image processing are cluster intensity files (CIF), which store the fluorescence intensity values for each cluster across all cycles and across all four wavelength channels. To reduce the file size, Illumina further divides each flow cell into tiles. For the sake of simplicity, we hereinafter present the decorrelation of the spatial crosstalk with respect to each tile. Assume a tile contains m clusters of n sequencing cycles. We denote the intensity signals of the  cluster by an n × 4 matrix Oi, whose (p, q) entry Oi(p, q) represents the signal in the
cluster by an n × 4 matrix Oi, whose (p, q) entry Oi(p, q) represents the signal in the  cycle and the
cycle and the  channel. We also assume that each cluster has one nominal DNA sequence fragment to be identified from the intensity data. The sequence is represented by an n × 4 matrix Si whose (p, q) entry Si(p, q) is one if the
channel. We also assume that each cluster has one nominal DNA sequence fragment to be identified from the intensity data. The sequence is represented by an n × 4 matrix Si whose (p, q) entry Si(p, q) is one if the  base of this sequence is of type q and zero otherwise, where the nucleotides A, G, C, T are encoded by type 1, 2, 3, 4, respectively.
base of this sequence is of type q and zero otherwise, where the nucleotides A, G, C, T are encoded by type 1, 2, 3, 4, respectively.
As discussed in Results and the literature1,2,4,7, spatial crosstalk, cycle crosstalk (also known as phasing and pre-phasing), and color crosstalk can be approximated by linear transformations. Accordingly, we use an m × m matrix C, an n × n matrix P and an 4 × 4 matrix M to represent the spatial, cycle, and the color crosstalk, respectively. Generally speaking, the non-diagonal element in Row i and Column j of each matrix represents the crosstalk from the j-th component to the i-th component. In the context of spatial crosstalk, C(i, j) measures the portion of the signals of the nominal Sequence j that leaks into the signals of Cluster i. We refer to it as the spatial crosstalk coefficient from Sequence j to Cluster i. In addition, we assume that C(i, i) =1. Thus, for any cluster i = 1, 2, …m, we have:

where λi is the average signal strength of Cluster i, MT is the transpose of M, cij is the (i, j) entry of the spatial crosstalk matrix C, N is an n × 4 matrix representing the background noise for all the clusters, and εi, an n × 4 matrix, is the error term.
We built this model to describe the phenomenon of spatial crosstalk along with color crosstalk and phasing/pre-phasing. If spatial crosstalk did not exist, matrix C would become an identity matrix, and the model in Equation (1) would be reduced to the model used in Bustard (Equation (2) in the paper by Massingham and Goldman4). In fact, we can use existing methods to estimate matrices M and P, followed by correction of color crosstalk and phasing/pre-phasing. Notably, these corrections do not interfere with the spatial crosstalk effect in the signals because they transform the signals uniformly across clusters, whereas the spatial crosstalk pattern remains unchanged. The above model uses the coefficient cij to describe the spatial crosstalk from Sequence j to Cluster i. Importantly, cji which is the spatial crosstalk coefficient from Sequence i to Cluster j, is not necessarily the same as cij, which is the coefficient from Sequence j to Cluster i. We estimate each coefficient cij based solely on the signals of cluster i and j, with no need for additional information, except for neighboring clusters. In other words, the estimation is adaptive to each pair of clusters. The method includes the following major steps:
-
1
Correct color crosstalk, phasing and background noise;
-
2
Estimate spatial crosstalk coefficients;
-
3
Correct spatial crosstalk;
-
4
Call the bases.
In the next sections, we present the details that explain each step above.
Correction of phasing, color crosstalk and background noise
In our software, we first ignore the low chastity clusters and robustly estimate matrices M and P using the iterative weighted least square method similar to that in AYB4. Let Ti be  , where P, M, N are plugged in by their estimates, respectively. Thus Ti is the signal of Cluster i after correction. According to Equation (1), Ti satisfies
, where P, M, N are plugged in by their estimates, respectively. Thus Ti is the signal of Cluster i after correction. According to Equation (1), Ti satisfies

where  are the error terms in the model.
are the error terms in the model.
Estimation of the spatial crosstalk matrix
We take a pairwise strategy to estimate matrix C. Namely, to a good approximation, we focus on the signals of cluster i and j when estimating the pair of crosstalk coefficients cij and cji. The pairwise strategy significantly reduces the computational complexity. On the other hand, even though possible higher-order interactions are ignored in one round of correction, we could improve the estimate by iterations if necessary. Consequently, we consider a simplified version of Equation (2), as follows:

where ηi and ηj are the error terms. In this model, both the crosstalk coefficients (cij, cji) and the nominal signal (Si, Sj) are unknown to us. Thus it is not a standard inversion but a blind inversion problem10. Heuristically, we want to estimate cij based on the information in the pairwise scatter plot shown in Fig. 1b, in which the slope roughly corresponds to the crosstalk coefficient cij (i = 1, j = 2). Notice that

When t = cij, the second term disappears, namely, the signals Ti − tTj contain no spatial crosstalk from Sequence j. Once we have a fair impurity measure that can approximately remove the effect of the first term, then the minimizer of the impurity of Ti − tTj would give an estimate of cij.
Ideally, in the signals of a DNA sequence without any kind of crosstalk, only the value corresponding to the nucleotide base would be significantly non-zero in each cycle. If, in the same cycle, the values in the other three channels were non-zero, then they would be the result of measurement errors or spatial crosstalk. It should be recalled that color crosstalk and phasing have already been corrected in the last step. Thus, we take the sum of the non-zero values in the three non-principal channels as the impurity:

The value of f(·) measures the extent of spatial crosstalk from one specific adjacent sequence across all cycles. Because Ti − tTj contains the least spatial crosstalk when t = cij, the spatial crosstalk coefficient can be estimated by minimizing the plug-in impurity measure f(Ti − tTj). Returning to Fig. 1b, those points marked in “×” and “*” correspond to the signals with nucleotide bases in cluster i ( = 1), and their effect is made approximately null by the impurity measure. Those points marked in “+” correspond to the signals with nucleotide bases in Cluster j( = 2), but not Cluster i(= 1), and the possible crosstalk is estimated by the regression of Ti on Tj.
When the signal-to-noise ratio of Ti is low, we found that the variation of the estimates of cij is large. To avoid overcorrection of spatial crosstalk, we impose some penalty on t. This leads to our estimator, as follows:

where u is a positive penalty constant. We make a few notes here. First, we take the sum of absolute values, rather than the sum of squares, in the impurity measure. Consequently, the more robust method, least absolute deviations (LAD), is used for estimation. Second, in implementation, we apply to each cycle a weight, which is set to be inversely proportional to the variance of each cycle to reduce the variation of the estimate.
Overall, we estimate the spatial crosstalk matrix C as follows:  ; For any pair of Cluster i and j such that i ≠ j, if Cluster j is located near Cluster i, we estimate cij by minimizing Equation (5) using the bisection method11, and cji can be estimated similarly; otherwise cij = cji = 0 because significant spatial crosstalk exists only between adjacent clusters. The algorithm of finding clusters’ neighbors is described in Supplementary Note and Supplementary Figure S5.
; For any pair of Cluster i and j such that i ≠ j, if Cluster j is located near Cluster i, we estimate cij by minimizing Equation (5) using the bisection method11, and cji can be estimated similarly; otherwise cij = cji = 0 because significant spatial crosstalk exists only between adjacent clusters. The algorithm of finding clusters’ neighbors is described in Supplementary Note and Supplementary Figure S5.
Correction of spatial crosstalk
At this point, we assume that color crosstalk and phasing have been corrected and that the spatial crosstalk matrix has been estimated, as shown above. Leaving out the errors in Equation (2), for any Cycle p and Channel q, we have:

Then  can be calculated by solving the above linear equations. Because most elements in C are zeros, a linear equation solver for sparse matrices is used12,13 for the calculation. To reduce the size of the linear equation and take advantage of parallel computation, we divide the clusters into blocks based on their locations. Then we carry out crosstalk estimation followed by correction separately for each block.
can be calculated by solving the above linear equations. Because most elements in C are zeros, a linear equation solver for sparse matrices is used12,13 for the calculation. To reduce the size of the linear equation and take advantage of parallel computation, we divide the clusters into blocks based on their locations. Then we carry out crosstalk estimation followed by correction separately for each block.
For those who prefer to use existing base-calling methods, the spatial crosstalk correction is applied directly to the original cluster intensity data. Leaving out the errors, we see that the second part of equation (1) is equivalent to:

In most cases, the background noise has been subtracted when generating the cluster intensity files; therefore, N(p, q) is close to zero. Ignoring the background noise in the model, we calculate  by solving the sparse linear equations. The corrected cluster intensity signals
by solving the sparse linear equations. The corrected cluster intensity signals  are in the standard CIF format and can thus be used by any existing base callers.
are in the standard CIF format and can thus be used by any existing base callers.
Base calling
Based on the corrected signals in Equation (6), we call bases as the types with the largest signals in each cycle.
Data
The datasets used in this article are listed below.
-
Illumina GAII phiX174. This dataset contains the sequencing images of 5 tiles, and the cluster fluorescence intensities were generated using our own tool (unpublished) from the images. The image data were distributed with SWIFT2 and were downloaded at http://sgenomics.org/swift/paperdataset.html. It has about 100,000 single-end reads of 37 sequencing cycles in each tile. The generated cluster intensity files and cluster location files can be obtained at http://www.statsci.amss.ac.cn/yjscy/yjy/ll/201508/t20150828_305802.html.
-
Illumina HiSeq2000 phiX174. This dataset contains the cluster intensity data for 3 tiles. It was distributed along with BlindCall7 and downloaded at http://www.cbcb.umd.edu/~hcorrada/secgen/. Each tile has about 1,900,000 single-end reads of 101 sequencing cycles.
-
Illumina MiSeq phiX174. This was a public dataset in Illumina Basespace and can be downloaded at https://basespace.illumina.com/s/flRIEtUDLckW. In this article, we took the first end of tile s_1_1101, which contains 650,582 sequences of 250 cycles.
Base-calling correlations between adjacent clusters
The sequences of the first two datasets (Illumina GAII phiX174; Illumina HiSeq2000 phiX174) were identified using the AYB base caller (Version 2.10, downloaded at https://github.com/timmassingham/AYB2/), and the sequences of the other dataset were extracted from the associated FASTQ files. We calculated spatial correlations of base calling for three tiles: s_4_21 from the first dataset, s_4_1113 from the second, and s_1_1101 from the third. In all three cases, the depths of mapped reads were more than 400X at each position, making it possible to define a reliable consensus sequence for each tile. We first used Bowtie25 (version 2.2.5, using the default option of “–sensitive”) to map the reads to the reference (NC_001422.1). Then the consensus was defined as the nucleotide occurring the most frequently at each bp. For each cluster, the nearest neighbor is determined based on cluster coordinates under Euclidian distance measure. Called bases different from those in the consensus references were regarded as base-calling errors, and deletions from the references, which were about 0.51–1.8% of the total differences, were neglected. Bases are ignored if themselves, the neighbors or the random ones include “N”; reads are ignored if more than half of the bases are identical to the neighbors or to the random ones. The results of spatial correlations were shown in Fig. 2a–c.
Software comparison
The cluster fluorescence intensities of the Illumina GAII phiX174 and Illumina HiSeq2000 phiX174 datasets were used for the comparison. We ran the software on a ThinkPad W530 mobile workstation with 24GB memory and the Intel Core i7-3820QM CPU. We ran the standard sequencing software Bustard (included in the Illumina Off-line Base-caller, version 1.9.4) and AYB to call bases using both original and 3Dec-corrected cluster intensity data. We also ran base calling using only 3Dec modules. After base calling, we used Bowtie2 to map the called sequences to the consensus references (see the above paragraph for mapping details). The comparison results are shown in Table 1, in which three kinds of error rates are presented. In the first definition of error rates, we counted errors in mapped reads obtained by using the same default mapping criterion in Bowtie2 across different base-calling methods. In the second definition, we calculated error rates for each method based on a common subset of reads. That is, at a given threshold on the edit distances, we found the set of reads that were successfully mapped to the reference for each base-calling method (one SAM file for each method). Then we picked the subset of reads that are common to all methods and did the calculation. In the third definition, we counted errors in those selected reads at a given mapping rate, which is 97.5% for the HiSeq2000 dataset in Table 1. Reads with the smallest edit distances to the consensus reference were selected at each given mapping rate. Specifically, we selected reads by the following steps: first, we mapped all reads to the consensus reference; then we sorted the mapped reads in the descending order of their edit distances to the reference; finally, we selected the top reads according to a given mapping rate, which equals the number of selected reads divided by the number of total reads.
Accuracy assessment of the estimation
We used the following method to estimate the biases and variances of the estimates of spatial crosstalk coefficients. We randomly selected a block (Cluster No. 283,639 to No. 346,710) in HiSeq2000 tile s_4_1113 for this assessment. The Moving Block Bootstrap14 (MBB) re-sampling technique was used to evaluate estimation accuracy because serial correlations exist between adjacent sequencing cycles, and MBB fits this situation15. The re-sampling procedure was as follows. After correcting color crosstalk and phasing, we calculated the weights used in the estimation of the spatial crosstalk coefficients based on all the data from the tile. Then we performed resampling 30 times for the selected clusters, as follows. Each time, we generated 20 random seed integers uniformly distributed in the range  and formed a group consisting of the 5 or 6 consecutive cycles following each seed integer. In total, we had 20 groups, which, as a pool, represented one set of the resampled data. Next, the spatial crosstalk coefficients were estimated for each set of the resampled data. The standard errors were evaluated based on the bootstrap estimates. The bootstrap accuracy assessment results are shown in Supplementary Figure S2. The same bootstrap technique was used in Supplementary Table S1 to show the significance of the crosstalk coefficients and in Supplementary Table S2 to show the phenomenon of cluster specificity.
and formed a group consisting of the 5 or 6 consecutive cycles following each seed integer. In total, we had 20 groups, which, as a pool, represented one set of the resampled data. Next, the spatial crosstalk coefficients were estimated for each set of the resampled data. The standard errors were evaluated based on the bootstrap estimates. The bootstrap accuracy assessment results are shown in Supplementary Figure S2. The same bootstrap technique was used in Supplementary Table S1 to show the significance of the crosstalk coefficients and in Supplementary Table S2 to show the phenomenon of cluster specificity.
Additional Information
How to cite this article: Wang, B. et al. An adaptive decorrelation method removes Illumina DNA base-calling errors caused by crosstalk between adjacent clusters. Sci. Rep. 7, 41348; doi: 10.1038/srep41348 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Li, L. & Speed, T. P. An estimate of the crosstalk matrix in four-dye fluorescence-based DNA sequencing. Electrophoresis 20, 1433–1442, doi: 10.1002/(SICI)1522-2683(19990601)20:7<1433::AID-ELPS1433>3.0.CO;2-0 (1999).
Whiteford, N. et al. Swift: primary data analysis for the Illumina Solexa sequencing platform. Bioinformatics 25, 2194–2199, doi: 10.1093/bioinformatics/btp383 (2009).
Renaud, G., Kircher, M., Stenzel, U. & Kelso, J. freeIbis: an efficient basecaller with calibrated quality scores for Illumina sequencers. Bioinformatics 29, 1208–1209, doi: 10.1093/bioinformatics/btt117 (2013).
Massingham, T. & Goldman, N. All Your Base: a fast and accurate probabilistic approach to base calling. Genome Biol 13, R13, doi: 10.1186/gb-2012-13-2-r13 (2012).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat Methods 9, 357–359, doi: 10.1038/nmeth.1923 (2012).
Navarro, G. A guided tour to approximate string matching. Acm Comput Surv 33, 31–88, doi: 10.1145/375360.375365 (2001).
Ye, C., Hsiao, C. & Corrada Bravo, H. BlindCall: ultra-fast base-calling of high-throughput sequencing data by blind deconvolution. Bioinformatics 30, 1214–1219, doi: 10.1093/bioinformatics/btu010 (2014).
Ye, C., Ma, Z. S., Cannon, C. H., Pop, M. & Yu, D. W. SparseAssembler: de novo Assembly with the Sparse de Bruijn Graph. arXiv preprint arXiv:1106.2603 (2011).
Sheridan, C. Milestone approval lifts Illumina’s NGS from research into clinic. Nat Biotechnol 32, 111–112, doi: 10.1038/nbt0214-111 (2014).
Li, L. Blind Inversion Needs Distribution (BIND): General Notion and Case Studies. (Institute of Mathematical Statistics, 2003).
Burden, R. L. & Faires, J. D. Numerical analysis. 3rd edn, (Prindle, Weber & Schmidt, 1985).
Li, X. Y. S. An overview of SuperLU: Algorithms, implementation, and user interface. Acm T Math Software 31, 302–325, doi: 10.1145/1089014.1089017 (2005).
Demmel, J. W., Eisenstat, S. C., Gilbert, J. R., Li, X. Y. S. & Liu, J. W. H. A supernodal approach to sparse partial pivoting. Siam J Matrix Anal A 20, 720–755, doi: 10.1137/S0895479895291765 (1999).
Kunsch, H. R. The Jackknife and the Bootstrap for General Stationary Observations. Ann Stat 17, 1217–1241, doi: 10.1214/aos/1176347265 (1989).
Lahiri, S. N. Theoretical comparisons of block bootstrap methods. Ann Stat 27, 386–404, doi: 10.1214/aos/1018031117 (1999).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant No. 91530105, 91130008, No. 11571349, and No.11201460), the Strategic Priority Research Program of the Chinese Academy of Sciences (Grant No. XDB13040600), the National Center for Mathematics and Interdisciplinary Sciences of the CAS, and the Key Laboratory of Systems and Control of the CAS. Lei M Li’s research was also supported by the Program of One hundred talented people, CAS. Lin Wan’s research is also supported by the Youth Innovation Promotion Association of the CAS.
Author information
Authors and Affiliations
Contributions
B.W. derived the model, designed the method, carried out the data analysis, wrote the software, and drafted the manuscript. L.W. conceived the project, and participated in method design, data analysis and writing. A.W. participated in sequence alignment and detection of base-calling errors. L.M.L. conceived and designed the study, and drafted the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Wang, B., Wan, L., Wang, A. et al. An adaptive decorrelation method removes Illumina DNA base-calling errors caused by crosstalk between adjacent clusters. Sci Rep 7, 41348 (2017). https://doi.org/10.1038/srep41348
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep41348
This article is cited by
-
Systematic evaluation of error rates and causes in short samples in next-generation sequencing
Scientific Reports (2018)
-
Estimating Phred scores of Illumina base calls by logistic regression and sparse modeling
BMC Bioinformatics (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.