Abstract
Excessive accumulation of white adipose tissue (WAT) is a hallmark of obesity. The expansion of WAT in obesity involves proliferation and differentiation of adipose precursors, however, the underlying molecular mechanisms remain unclear. Here, we used an unbiased transcriptomics approach to identify the earliest molecular underpinnings occuring in adipose precursors following a brief HFD in mice. Our analysis identifies Heme Oxygenase-1 (HO-1) as strongly and selectively being upregulated in the adipose precursor fraction of WAT, upon high-fat diet (HFD) feeding. Specific deletion of HO-1 in adipose precursors of Hmox1fl/flPdgfraCre mice enhanced HFD-dependent visceral adipose precursor proliferation and differentiation. Mechanistically, HO-1 reduces HFD-induced AKT2 phosphorylation via ROS thresholding in mitochondria to reduce visceral adipose precursor proliferation. HO-1 influences adipogenesis in a cell-autonomous way by regulating events early in adipogenesis, during the process of mitotic clonal expansion, upstream of Cebpα and PPARγ. Similar effects on human preadipocyte proliferation and differentiation in vitro were observed upon modulation of HO-1 expression. This collectively renders HO-1 as an essential factor linking extrinsic factors (HFD) with inhibition of specific downstream molecular mediators (ROS & AKT2), resulting in diminished adipogenesis that may contribute to hyperplastic adipose tissue expansion.
Similar content being viewed by others
Introduction
White adipose tissue (WAT) has a remarkable capacity to expand or remodel in order to meet the energy demands of the organism. In the face of caloric excess, WAT expands through the enlargement of existing white adipocytes (hypertrophy) as well as by recruitment of new fat cells (hyperplasia)1,2,3. Visceral adipocyte hypertrophy is detrimental for metabolic health in humans/mice4,5,6,7. However, increased fat tissue expansion resulting from adipocyte hyperplasia produces less pronounced impairments in glucose tolerance than a similar magnitude of obesity resulting from adipocyte hypertrophy8. Accordingly, adequate adipogenesis throughout the process of adipose expansion is a necessity to maintain metabolic homeostasis in “metabolically healthy obese humans/mice”, whereas excessive hypertrophy in the absence of the generation of new, metabolically healthy adipocytes is associated with the pathophysiology of obesity-related disease, such as diabetes and cardiovascular disease3,9,10,11,12. In obese humans, hyperplastic adipose tissue (many small adipocytes) is associated with better glucose, insulin and lipid profiles compared with adipose hypertrophy (i.e. few large adipocytes)13. In addition, there is a decreased preadipocyte frequency in visceral adipose tissue from type 2 diabetes mellitus subjects14. Thus, understanding the molecular and cellular mechanisms that regulate adipose homeostasis represents a promising strategy for identifying novel therapeutic opportunities to fight obesity-related complications.
Adipogenesis occurs in two steps: ‘commitment’ of adipose precursors (APs) to a preadipocyte fate and ‘terminal differentiation’ describing the process by which the preadipocyte acquires the characteristics of the mature adipocyte1,2. The majority of studies on adipocyte precursors and adipogenic differentiation had been performed in vitro, primarily using human and murine cell lines15,16. While these studies have provided valuable insights, most notably unraveling the adipogenic transcription cascade, when and where specific factors activate adipose precursors in vivo cannot be determined in the in vitro system17.
Lineage tracing studies have demonstrated that most adipose precursors within WAT are of non-endothelial and non-hematopoietic origin (CD31− and CD45−, respectively) and express surface cell markers including CD34, CD29, as well as Platelet-derived growth factor receptor alpha (Pdgfra)17,18,19,20. These studies have shown that HFD-induced adipose tissue hyperplasia is restricted to visceral fat19,21,22,23. However, lineage tracing cannot reveal details about physiological signals and molecular mechanisms underlying the response of APs to various (patho-)physiological stimuli.
To reveal early molecular targets in APs following overnutrition, we fed mice a HFD for three days and analyzed the transcriptome of APs isolated from subcutaneous (sc) and visceral (vi) WAT. Interestingly, this approach revealed HO-1 being upregulated by HFD in APs, but not other cellular constituents of adipose tissue. Based on our previous work showing that HO-1 is a conserved pro-inflammatory mediator necessary for the adverse metabolic effects of obesity24, we here investigate the role of HO-1 on adipogenesis. In our previous work we deleted HO-1 in five metabolic tissues; interestingly, in three of these - beta-cells, adipose tissue (using aP2-Cre) and muscle - presence or absence of HO-1 appeared negligible after HFD feeding. Intriguingly though, liver and macrophage HO-1 knockout mice exhibit an exquisitely healthy metabolic profile, however, a role of HO-1 in adipogenesis was not addressed at this time, due to a lack of suitable Cre lines to delete HO-1 in adipose precursors. Although various aP2-Cre lines have been used to target adipose tissue and were the only ones available; the lack of specificity of these lines for adipocytes as well as expression of aP2 during terminal differentiation25 made this model unattractive for studying adipogenesis. We chose to readdress a role of HO-1 in adipogenesis, as HO-1 top-scored in our adipose precursor array screen, and genetic tools to target adipose precursors have now been characterized19. We conditionally deleted HO-1 in early (Pdgfra-Cre) as well as late (AdipoQ-Cre) adipogenesis to distinguish whether observed phenotypes are the result of gene function within adipocyte precursors (APs) or mature adipocytes. Deletion of HO-1 in AP cells using Pdgfra-Cre enhanced viAP proliferation and differentiation. Mechanistically, we demonstrate that deficiency of HO-1 in viAPs raised reactive oxygen species (ROS) levels and promoted proliferation and differentiation via increasing Akt2 signaling.
Results
HFD feeding targets HO-1 in adipose precursors
Short periods of consumption of a HFD (<1 wk) lead to rapid visceral-specific expansion of adipocyte precursors which is phenotypically similar to the accumulation of abdominal WAT in men19. However, the nutrient-associated genetic factors regulating this process in vivo are largely unknown. We used an unbiased transcriptomics approach to identify the earliest molecular underpinnings occuring in APs following a brief HFD in mice. Body weight as well as fat pad weight significantly increased in wildtype mice fed a HFD for three days (Figure S1A). We enriched adipose precursors by bead-purification19,26,27. High expression of the adipose precursor marker Pdgfra selectively in APs, paralleled by low CD68 and AdipoQ expression demonstrated successful enrichment, in addition to separation from Lin+ (CD31+CD45+Ter119+) cells and mature adipocytes (which show low Pdgfra expression) (Figure S1B). Microarray analysis was performed on AP cells lysed directly after isolation to avoid cellular phenotype alterations occuring in culture28. By comparing standard diet (SD)- with HFD-fed mice, microarray analysis revealed regulation of a greater number of genes in visceral APs (viAP) as compared to subcutaneous APs (scAP) (Fig. 1A and B). In viAPs from HFD fed mice 496 genes were regulated, of which 443 were induced and 53 repressed more than 2.0-fold, respectively (Fig. 1B and Table S1). In scAPs from HFD fed mice 172 genes were regulated, of which 149 were induced and 23 repressed more than 2.0-fold, respectively (Fig. 1B and Table S2). Of note, we could verify previous findings of several genes that are differentially expressed between sc and vi APs (in SD fed mice)29 including Tbx15, Sfrp2, Hoxc10, Ccl8, Mmp3 and others.
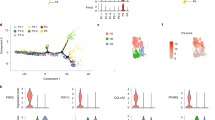
All visceral and subcutaneous APs were isolated from mice on a SD or HFD for 3 days via Sca-1 bead pull down (see ‘Experimental Procedures’). (A) Dot plot displaying diet-induced changes in gene expression in APs. Purple, green and red spots represent genes significantly differentially expressed greater than 2-fold (P < 0.05; n = 6 mice, pooled into two replicates per group). Grey dots indicate genes that are not statistically significantly enriched. (B) Venn diagram illustrating overlap between gene signatures derived from expression analysis of AP fractions. (C) GO analysis using DAVID80 of scAP and viAP molecular signatures. Notable terms are highlighted; (see also Supplemental Tables S3 + S4). P-values are derived from Fisher’s exact test and pass Benjamini-Hochberg corrected P < 0.05. (D) GSEA analysis of AP fractions. Analyzed gene sets include GO, KEGG and Reactome-derived cell cycle pathways as well as custom-curated genesets from a 3T3-L1 differentiation time course31 (see ‘Experimental Procedures’ for details). (E) Upper panel: Network plots for genes comprising the ‚Leading edge‘ of cell cycle from the KEGG, GO and Reactome compendium. Lower panel: Network plots for the 66 genes comprising a core HFD-associated AP gene signature. Blue node color represents downregulation and red node color represents upregulation of gene expression in APs. The node size is associated with the gene’s co-expression in the entire data set. The edge (line) thickness is linked to the gene’s connectivity (co-expression within the module). (F) Heatmap of top 10 genes regulated by HFD common to APs from scWAT and viWAT. Each column represents pooled cells from three mice. Blue color represents downregulation and red color represents upregulation of gene expression. (G) HO-1 mRNA expression in scWAT or viWAT fractions (n = 6). (H) Western blot of protein lysates from APs enriched from SVF of scWAT or viWAT. Each lane represents pooled cells from 2 mice. (I) Quantification of western blots in (H) showing HO-1 protein levels normalized to β-actin. (J) Whole mount immunofluorescence staining for HO-1 and CD24 in viWAT. Arrowheads indicate CD24+ cells coexpressing HO-1. Bar = 20 μm.
The HFD fed mice gene-signature in viAPs was characterized by significant enrichment of cell cycle/cell proliferation pathways, based on (i) viAP Ki67 gene upregulation (Fig. 1A), (representing proliferative capacity)19,30; (ii) DAVID classification (Fig. 1C, top panel and Table S3); (iii) GSEA analysis using curated ‘Cell cycle’ pathways from Gene Ontology, Reactome as well as KEGG. GSEA-enrichment of custom-built gene sets comprising two distinct stages of in vitro (3T3-L1) preadipocyte differentiation31: Whereas the gene set ‘terminal differentiation’ as a group exhibited less remarkable regulation, the gene set ‘preadipocyte proliferation’ showed marked upregulation specifically in viAPs from HFD-fed mice (Fig. 1D). (iv) Enrichment of a gene coexpression network representing the common genes from the GSEA-‚ ‘Leading Edge’ in viAPs (Fig. 1E, upper panel, and Table S5).
HFD feeding targets genes in scAPs associated with regulation of cytoarchitecture and/or angiogenesis (Fig. 1C, lower panel and Table S4) including Thsp132, Timp133 and Flt-1. Of the 172 genes whose expression is regulated by more than 2-fold in scAPs, 66 of these were also changed in viAPs (Fig. 1B and Tables S1 and S2). Thus, these 66 genes define a core HFD-induced molecular signature common to both primary visceral and subcutaneous APs, shown in Fig. 1E, lower panel, and Tables S1 and S2. The gene most highly induced in our ‘HFD-signature’ was HO-1 (Fig. 1E, lower panel & 1 F marked in bold letters). Importantly, HFD failed to upregulate HO-1 in Lin+ cells as well as in mature adipocytes (Fig. 1G). Significant elevation of HO-1 protein levels, detected by western blotting in isolated APs from scWAT and viWAT from mice fed either SD or HFD for three days (Fig. 1H and I), confirmed our observations at mRNA level (Fig. 1G). Enzymatic digestion of WAT followed by bead-separation may induce ‘unspecific’ gene expression in adipose cells34 and interfere with HO-1 expression in our APs. Thus, to exclude that experimental manipulation caused HO-1 expression, we performed histological staining of HO-1 in WAT of SD versus HFD fed mice. In SD fed mice, HO-1 staining was barely detectable, however, HFD feeding for three days strongly increased HO-1 expression. Importantly, HFD-induced HO-1 expression was found in cells expressing the adipose precursor marker CD24 (Fig. 1J). Serum of mice exposed to HFD for three days contains elevated levels of non-esterified free fatty acids (NEFAs)35. Therefore, we assessed whether fatty acids are responsible for HO-1 induction in APs. Indeed, incubating these cells with palmitic acid resulted in strong upregulation of HO-1 mRNA in a time-dependent manner (Figure S1C), in contrast to linoleic and oleic acid (Figure S1D). Further, HO-1 upregulation was dependent on the transcription factor Nrf2, since palmitic acid failed to induce HO-1 mRNA in APs from Nrf2 knockout mice (Figure S1E). Thus, these data strongly support our notion that HO-1 is part of an early molecular mechanism determining AP activation during HFD feeding. Together, our data show that HFD feeding triggers depot-specific-, as well as depot-independent gene expression signatures in APs in vivo, and that viAPs express a proliferation network which is absent in scAPs. Among the top-regulated genes in APs from both fat depots we found HO-1, upregulated by HFD almost exclusively in the AP-fraction of WAT, but not Lin+ or mature adipocyte cells, suggesting that HO-1 may transduce HFD effects in APs in vivo.
HO-1 blocks HFD-induced visceral adipose precursor proliferation
To test whether HO-1 affects the proliferation gene network in adipose precursors in vivo, we generated a conditional HO-1 knockout with the Pdgfrα promoter, which targets adipocyte precursors, but not liver or muscle17,19,22,36. We found that full excision of the HO-1 gene is achieved in APs from both scWAT and viWAT, while Lin+ cells still contain the full-length Hmox1 gene (Fig. 2A). Transgenic Hmox1fl/flPdgfraCre mice were born at the expected mendeleian frequency, with no observed signs of abnormality, illness or increased mortality and exhibited normal body weight at six weeks of age (Figure S2A). HO-1 levels in APs from unchallenged, naive mice are barely detectable as seen in Fig. 1G–I, so to confirm efficient HO-1 protein deletion in APs, the bead purified APs were further stimulated with the strong HO-1 inducer, hemin. Consistently, hemin treatment resulted in reduced heme oxygenase activity in APs derived from Hmox1fl/flPdgfraCre mice compared with Hmox1fl/fl littermates (Fig. 2B). In vivo, an initial burst of AP proliferation precedes differentiation into mature adipocytes. Thus, we focused our analysis on the Lin−:CD29+:CD34+:Pdgfrα+ population of stromal cells within WAT as candidate adipose precursors and assessed their proliferation by intracellular staining for the cell proliferation antigen Ki67 (Figure S2B). On standard diet, flow cytometry analysis of APs freshly isolated from adipose SVF showed that Ki67 values were similar in APs from Hmox1fl/flPdgfraCre and Hmox1fl/fl control mice. However, when these mice were challenged with a HFD for a short period of time (3 days), we found a significant increase in AP proliferation in viWAT of Hmox1fl/flPdgfraCre mice compared to HFD-fed controls (Hmox1fl/fl) (Fig. 2C and D, right panel), despite similar food intake (Figure S2C), whereas APs derived from scWAT, proliferation levels were similar (Fig. 2D, left panel). To eliminate the possibility that HO-1 from mature adipocytes could have an effect on AP proliferation, we generated mice in which the Hmox1 gene was selectively deleted in differentiated adipocytes (designated Hmox1fl/flAdipoQCre, Fig. 2E). Consistent with its primary role in APs, deletion of HO-1 in mature adipocytes did not affect AP proliferation (Fig. 2F).
(A) PCR analysis of DNA isolated from bead-purified APs and Lin+ cells from the viWAT and scWAT depots of Hmox1fl/flPdgfraCre mice. (B) HO-1 activity in APs, challenged with hemin for 16 hrs (n = 5). (C) Exemplary flow cytometry plots of Ki67 staining in APs obtained from the SVF of scWAT or viWAT from Hmox1fl/fl and Hmox1fl/flPdgfraCre mice after feeding a HFD for 3 days. (D) Quantification of Ki67 in APs obtained from Hmox1fl/fl and Hmox1fl/flPdgfraCre mice (n = 6). (E) PCR analysis of DNA isolated from AP cells from the viWAT and scWAT depots of Hmox1fl/flAdipoQCre mice. (F) Quantification of Ki67 in APs obtained from Hmox1fl/fl and Hmox1fl/flAdipoQCre mice. n = 6 per group. Results are mean ± SEM. *p < 0.05, **p < 0.01, ***p < 0.001.
HO-1 inhibits adipogenesis in primary adipose precursor cells isolated from mice fed a brief HFD
Bead-purified APs from Hmox1fl/flPdgfraCre and Hmox1fl/fl control mice were induced to differentiate into mature adipocytes by treatment with an adipogenic induction cocktail containing dexamethasone, IBMX, TZD and insulin. APs derived from sc- and viWAT from unchallenged naive (SD-fed) Hmox1fl/flPdgfraCre and Hmox1fl/fl mice exhibited identical adipogenic capacity, as measured with Oil Red-O staining for triglycerides (Fig. 3A, left panel & Fig. 3B). An expected result as HO-1 is largely absent in APs from unchallenged naive mice, as shown in Fig. 1H. Therefore, we fed Hmox1fl/flPdgfraCre and Hmox1fl/fl control mice a HFD for three days prior to AP isolation, resulting in strong HO-1 induction in APs (Fig. 1G–I). Under those obesogenic conditions, treatment with an adipogenic cocktail resulted in more efficient adipogenesis in APs isolated from HFD-fed Hmox1fl/flPdgfraCre mice as compared with APs isolated from Hmox1fl/fl control mice (Fig. 3A, right panel & Fig. 3B). Noteworthy, not only the number of lipid accumulating cells, but also the intracellular lipid content was higher when viAPs were differentiated from Hmox1fl/flPdgfraCre mice as compared with Hmox1fl/fl control mice (Fig. 3C).
All visceral and subcutaneous APs were isolated from mice on a SD or HFD for 3 days via Sca-1 bead pull down (see ‘Experimental Procedures’). (A,B) Oil-Red O staining of scAP or viAP cell cultures derived from Hmox1fl/fl and Hmox1fl/flPdgfraCre mice 7 days after the induction of adipocyte differentiation (n = 3). (C) Flow cytometry for viAP cell cultures derived from Hmox1fl/fl and Hmox1fl/flPdgfraCre mice Bodipy lipid-stained 4 days after the induction of adipocyte differentiation, measured as mean fluorescence intensities (n = 3). Results are mean ± SEM. *p < 0.05, **p < 0.01, ***p < 0.001.
HO-1 inhibits early events during adipogenesis in a cell-autonomous manner
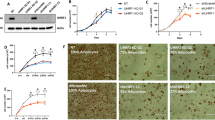
Besides HO-1 induction, HFD feeding will have ‘off target’ effects, therefore we turned to 3T3-L1 cells (which endogenously express HO-1), and generated gain- as well as loss of function cells. Cells with reduced HO-1 demonstrated enhanced adipogenic potential, including greater lipid accumulation (Fig. 4A) and increased expression of adipocyte marker genes such as PPARγ, CEBPα, FABP4 and Adiponectin (Fig. 4B). Overexpression of HO-1 in these cells markedly blocked adipogenesis, as shown by reduced Oil-red O staining of neutral lipids (Fig. 4C). Adipocyte markers were significantly decreased in HO-1-overexpressing cells (Fig. 4D), further confirming an antiadipogenic effect of HO-1. We applied microarray experiments comparing control (MSCV) versus HO-1 (MSCV-HO1) over-expressing cells treated with an adipogenic cocktail (Fig. 4E). Pathway analysis of genes affected by HO-1 expression (Fig. 4F) demonstrates an inhibitory role of HO-1 on cell cycle as well as proliferation-related events (Fig. 4G). To test whether HO-1 affects early clonal expansion, which is a prerequisite for 3T3-L1 adipocyte differentiation37, we performed BrdU-incorporation assays in transgenic 3T3-L1 cell lines treated with growth medium or an adipogenic medium. Treatment of control cells (LMP) with an adipogenic medium increased BrdU incorporation, an effect that was significantly enhanced in HO-1 knockdown cells (miHO1) (Fig. 4H). Accordingly, HO-1 overexpression (MSCV-HO1) resulted in significantly reduced BrdU incorporation as compared with control cells (MSCV) following treatment with an adipogenic medium (Fig. 4I). Within 48 hrs of adipogenic stimulation, growth-arrested preadipocytes re-enter the cell cycle and undergo two rounds of proliferation, a process termed ‘mitotic clonal expansion’ (MCE), ultimately leading to the upregulation of PPARγ and CEBPα37. While HO-1 overexpression before adipogenic induction reduced the accumulation of intracellular TGs as well as expression of PPARg, Cebpa and AdipoQ, (Fig. 4C,D), HO-1 overexpression after MCE had no effect on adipogenic gene expression on day 7 as well as TG accumulation (Fig. 4J). We conclude that HFD feeding induces HO-1 in APs and HO-1 cell-autonomously inhibits their subsequent differentiation into mature adipocytes via interfering with early adipose precursor activation (i.e. proliferation), upstream of Cebpα and PPARγ.
(A,C) Left panels: 3T3-L1 preadipocytes were transduced with (A) a retrovirus expressing a shRNAmir specific for HO-1 (miHO-1) or ctrl LMP vector (LMP), or with (C) a retrovirus expressing HO-1 or empty MSCV vector, and protein lysates were prepared for Western blotting at confluence. β-actin demonstrates equal protein loading. Middle and right panels: Oil-red O staining was performed on day 7 after induction of adipocyte differentiation. (B,D) Q-PCR analysis of adipocyte genes PPARγ, CEBPα, Adiponectin and aP2 were analyzed with Q-PCR on days 0, 1, 2, 3 and 4 after induction of adipocyte differentiation. (E) Microarray analysis of control (MSCV) or HO-1 overexpressing (MSCV-HO1) 3T3-L1 cells 8 hrs after addition of an adipogenic cocktail. (F) Heatmap of HO-1 independent (Cluster 1 +2) and HO-1 dependent (Clusters 3 + 4) genes following addition of an adipogenic cocktail (n = 2 biological replicates). (G) Pathway analysis of HO-1 dependent genes (Clusters 3 + 4). (H,I) BrdU incorporation into HO-1 gain- and loss of function 3T3-L1 cells 24 and 48 hrs after addition of an adipogenic medium (n = 4). (J) 3T3-L1 cells were induced to differentiate into mature adipocytes. 48 hrs later, cells were infected with a control (AdLacZ) or HO-1 overexpressing (AdHO-1) adenovirus and differentiation was continued for another 5 days followed by Oil red-O staining. Results are mean ± SEM. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
HO-1 prevents HFD-induced Akt2 signaling in visceral adipose precursor cells
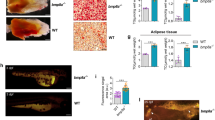
We next pondered on how HO-1 blocks AP proliferation. Phosphorylation of Akt2 has been shown to mediate the effects of HFD on AP proliferation19. HFD feeding for three days elevated phosphorylated AKT2 at S474 in bead-isolated, uncultured viAPs in our control (Hmox1fl/fl) mice, as described previously19; and more importantly, pAKT2 levels were higher in viAPs of Hmox1fl/flPdgfraCre mice compared to Hmox1fl/fl littermate controls (Fig. 5A and B). We have previously shown that HO-1 triggers mitochondrial changes that contribute to the “pre”-programming of cellular function in macrophages and hepatocytes24. We therefore investigated if HO-1 exerts similar effects in APs, linking adipose precursor HO-1 with the well documented role of mitochondrial metabolism in adipogenesis38. Intriguingly, when assessing mitochondrial activity in naive viAPs, we found clearly elevated basal, spare as well as maximal respiratory capacity in the HO-1-deficient cells (Fig. 5C,D). Recent studies have suggested that mild ROS elevations activate mitochondrial respiration39. If ROS balance was altering oxidative capacity (Fig. 5C), mitochondrial function in HO-1 knockout cells should be particularly sensitive to antioxidants. In line with this idea, 1 hr preadministration of the ROS quencher N-acetyl-cysteine (NAC), reverted HO-1- deficient mitochondrial functional parameters back to control levels (Fig. 5E,F). Further, ROS levels were significantly increased in viAPs derived from Hmox1fl/flPdgfraCre mice based on elevated measures of MitoSOX fluorescence, a probe that selectively detects mitochondrial superoxide and nitroblue tetrazolium (NBT) reduction staining (Fig. 5G,H), suggesting that HO-1 regulates acute ROS thresholding of mitochondrial respiration in APs. Finally, we also tested viAPs from Hmox1fl/flAdipoQCre mice to exclude any possible influence of HO-1 from mature adipocytes. As expected loss of HO-1 in adipocytes had no effect on mitochondrial activity in APs (Fig. 5I,J). Mechanistically, our data suggest that HO-1 continuously limits activation of key signaling systems (PI3K-AKT2), controlling adipose precursor activation through an acute and ROS-dependent mechanism. Taken together, these data indicate that visceral APs activated very early after high-fat diet feeding, undergo adipogenesis more efficiently when HO-1 levels are reduced, facilitating ROS thresholding of mitochondrial respiration upstream of AKT2. We further tested if these mice exhibit any metabolic abnormalities after an extended HFD feeding period of 8 weeks. Intriguingly, fasting blood glucose and insulin as well as Homeostasis model assessment of insulin resistance (HOMA-IR) levels were lower in Hmox1fl/flPdgfraCre as compared to Hmox1fl/fl control mice (Fig. 6A–C), indicating improved metabolic health in the HFD-treated Hmox1fl/flPdgfraCre mice. Serum levels of free fatty acids were significantly lower in HFD-fed Hmox1fl/flPdgfraCre mice compared to their control littermates (Fig. 6D). Irrespective of diet, Hmox1fl/flPdgfraCre and Hmox1fl/fl littermates gained weight to a similar extent, and displayed similar fat pad weights (Figure S3A–D). Histological examination of scWAT and viWAT of Hmox1fl/flPdgfraCre mice revealed a trend towards higher number of small adipocytes in viWAT from Hmox1fl/flPdgfraCre mice (Figure S3E). To explain the metabolic improvements observed in Hmox1fl/flPdgfraCre mice we investigated adipose tissue fibrosis and ex-vivo lipolysis in viWAT. Total collagen content as well as trichrome staining demonstrated reduced tissue fibrosis in viWAT of obese HFD-fed Hmox1fl/flPdgfraCre mice compared with Hmox1fl/fl control mice (Fig. 6E,F). Because serum fatty acid levels can be a reflection of adipose tissue lipolysis40, ex vivo lipolysis of viWAT was analyzed. In fact, viWAT explants from obese HFD-fed Hmox1fl/flPdgfraCre mice showed reduced isoprenaline stimulated lipolysis when compared with Hmox1fl/fl control mice (Fig. 6G). Together, our data show that obese Hmox1fl/flPdgfraCre mice lacking HO-1 specifically in adipose precursors display reduced markers of metabolic disease, associated with a decrease in lipolysis and adipose tissue fibrosis.
(A) pAKT2(S474) and total AKT2 levels were measured by immunoblotting of bead purified viAPs from Hmox1fl/fl and Hmox1fl/flPdgfraCre mice fed either SD or HFD for 3 days. A representative western blot is shown. The experiment was repeated at least three times. (B) Quantification of the pAKT2(S474)/AKT2 ratio. (C,D) Oxygen consumption rates (OCR) and mitochondrial function of viAPs derived from Hmox1fl/fl and Hmox1fl/flPdgfraCre mice (n = 4). (E,F) Oxygen consumption rates (OCR) and mitochondrial function of viAPs derived from Hmox1fl/fl and Hmox1fl/flPdgfraCre mice. N-acetyl-cysteine (NAC, 10 μM) was added 1 hr before measurements (n = 4). (G) Flow cytometric quantitation of mitoSOX-stained viAPs derived from Hmox1fl/fl and Hmox1fl/flPdgfraCre mice (n = 6). (H) NBT reduction staining of viAPs derived from Hmox1fl/fl and Hmox1fl/flPdgfraCre mice, after 48 hrs in culture (n = 4). (I–J) Oxygen consumption rates (OCR) and mitochondrial function of viAPs derived from Hmox1fl/fl and Hmox1fl/flAdipoQCre mice (n = 4). Results are mean ± SEM. *p < 0.05, **p < 0.01, ***p < 0.001.
All mice (Hmox1fl/fl and Hmox1fl/flPdgfraCre34) were kept on HFD for 8 weeks. (A–D) Fasting circulating levels of glucose (A) and insulin (B) in obese mice. (C) HOMA-IR (n = 10–12). (D) Serum free fatty acids (NEFA) in obese mice (n = 10–12). (E) viWAT collagen content of Hmox1fl/fl and Hmox1fl/flPdgfraCre mice (n = 6). (F) Trichrome staining of visceral adipose tissue of Hmox1fl/fl and Hmox1fl/flPdgfraCre mice. Bar = 50 μm. (G) Isoprenaline stimulated lipolysis in viWAT pieces from Hmox1fl/fl and Hmox1fl/flPdgfraCre mice (n = 6). Results are mean ± SEM. *p < 0.05, **p < 0.01, ***p < 0.001.
HO-1 impairs human adipocyte proliferation and differentiation
To investigate if HO-1 exerts similar effects on human adipogenesis, we used primary adipose stromal cells (hASC) isolated from lipoaspirates41. In order to manipulate HO-1 levels, we selected an overexpression strategy based on our observations that endogenous HO-1 levels in hASC are low. Transient HO-1 overexpression was achieved by transduction of hASC with an adenovirus carrying HO-1 expression construct (Fig. 7A)24. We determined if HO-1 affects hASC proliferation using an in vitro BrdU cell proliferation assay. Following plating of equivalent numbers of adenovirally-transduced hASC, AdLacZ-transduced cells readily incorporated BrdU 24 and 48 hrs later, an effect that was reduced when HO-1 was overexpressed (Fig. 7B). When induced to differentiate into mature adipocytes, control- (AdLacZ) transduced hASC showed prominent lipid accumulation as revealed by Oil-Red O staining. However, when hASC were instead transduced with an HO-1 adenovirus (AdHO-1), adipocyte differentiation was markedly reduced (Fig. 7C). Adipocyte markers were significantly decreased in HO-1-overexpressing cells, evaluated at day 10 of differentiation (Fig. 7D). Thus, elevated HO-1 levels act anti-proliferative and anti-adipogenic in murine and human adipose precursor cells.
All hASC were transduced with an Adenovirus for LacZ (AdLacZ) or HO-1 (AdHO-1) and experiments started 48 hrs later. (A) Western blot analysis of HO-1 protein expression in human ASC. β-actin demonstrates equal protein loading. (B) Cell proliferation analysis in human ASC. (C) Human ASC transduced with an Adenovirus for LacZ (AdLacZ) or HO-1 (AdHO-1) were induced to differentiate into mature adipocytes. Oil-red O staining was performed on day 10 after induction of adipocyte differentiation. (D) Q-PCR analysis of mature adipocyte genes LPL and RBP4 analyzed with Q-PCR 10 days after induction of adipocyte differentiation. Results are mean ± SEM. *p < 0.05, **p < 0.01, ***p < 0.001.
Discussion
In this study, we aimed to identify novel HFD target genes involved in the activation of adipose precursor cells, on the basis of (1) a hitherto unknown function in adipocyte biology, (2) a robust induction upon HFD feeding in APs in vivo, and (3) the potential to act upstream of the adipogenic master regulators Cebpα and PPARγ. Using these criteria, we identified among the top-most regulated genes, HO-1. Conditional HO-1 knockout mouse models using Pdgfra-Cre as well as AdipoQ-Cre reveal a novel role for HO-1 in pathways that govern obesogenic adipocyte proliferation and differentiation.
Although APs from different WAT depots have distinct gene signatures29,42,43,44,45, it is not clear if these cells also differ in their response to environmental cues, such as hormones or overfeeding. Our microarray screen using purified murine sc- and viAPs19,26,27 extends on these studies providing potential molecular underpinnings of hyperplastic fat expansion, some of which have a documented role in adipogenesis, mostly via facilitation of clonal expansion, including Cyclins -D1 & -D3, -E1 & -E2 as well as Cdk-2 and -446,47,48. Some of the genes targeted by HFD in scAPs are (dys)regulated in obesity and/or have a documented role in regulation of extracellular matrix (ECM) architecture and angiogenesis in adipose tissue such as Tsp132, Timp133 or Flt149. Maintaining a high degree of flexibility of the ECM allows the AT to expand in a healthy manner, without adverse metabolic consequences1,6,50,51. Further studies are needed to investigate if these genes play a role in the postulated function of APs as being sensors of overnutrition52,53.
Pdgfra+ APs responded to nutrient overload in WAT with proliferation as early as 24 hours following nutrient overload17,18,21,52,53. Biochemical data using purified APs show that HO-1 is a HFD target gene in APs from both depots, making HO-1 a candidate gene with the potential to regulate the ‘adipogenic’ response. Mice fed a HFD for three days display elevated NEFAs35, and we identified palmitic acid as such a HFD-elicited signal responsible for upregulation of HO-1 in APs, dependent on Nrf2. The expression of several cell cycle regulators identified in our screen were enhanced in viAPs from our AP-specific HO-1 knockout mice suggesting a direct role of HO-1 on obesogenic AP activation. Enhanced Ki67 staining, as shown by FACS analysis in Pdgfra+ APs from HO-1 knockout mice, further support a role of HO-1 in AP proliferation at the onset of obesity. BrdU-uptake assays in 3T3-L1 cells, engineered for HO-1 gain- and loss of function, further support an inhibitory role of HO-1 on adipogenesis-induced AP proliferation and positions HO-1 in the ‘commitment’ phase of adipogenesis. The depot-specific action of HO-1 on visceral, but not subcutaneous AP proliferation may likely be explained by the recent observations that obesogenic AP proliferation is restricted to this compartment19,23,54,55. Future studies will identify potential effects of HFD-induced HO-1 in subcutaneous APs.
In addition to enhanced AP proliferation, AP differentiation was also enhanced in HO-1 deficient APs (Fig. 3). A reduction in adipogenesis was observed solely when APs were isolated from HFD-fed mice, but not from chow fed animals, a phenomenon that may be explained by the finding that HO-1 levels are (too) low in APs from chow-fed mice (Fig. 1H). APs activated during the first week on HFD, undergo adipogenesis to form new adipocytes over a period of several weeks19,23,43. Remarkably, adipocyte size distribution and viWAT weight remained unaffected. Further studies utilizing longer HFD durations and examination of the development of other WAT depots such as retroperitoneal and mesenteric WAT56 as well as the use of alternative genetic tools to mark adipose precursors (e.g. Pdgfrb-rtTA; TRE-Cre “MuralChaser mouse”57) are needed. The formation of adipocytes during development can vary greatly from the route by which new adipocytes are recruited in response to overnutrition. Interestingly, both, Hmox1fl/flPdgfraCre and Hmox1fl/fl control mice mice fed a standard diet display no differences in adipocyte differentiation and/or recruitment of APs. This suggests that HO-1 plays a negligible role during physiologic WAT development. The most obvious explanation for this finding may be that HO-1 is largely absent in APs from ‘unstressed’ animals (Fig. 1H). In stark contrast, HO-1 is markedly induced in stressed animals, fed a HFD for even a short period of three days, resulting in a consecutive block of adipose precursor cell proliferation and differentiation. Forced overexpression of HO-1 in 3T3-L1 and human scWAT lipoaspirates strongly support the cell-autonomous effects of HO-1 on adipogenesis. Intriguingly, HO-1 & Akt2 may be functionally linked as modulation of both, HO-1 and Akt2 signaling affects adipose precursor proliferation in the setting of overnutrition, yet dispensable for normal development of WAT. Our data showing enhanced phosphorylation of Akt2 in HO-1 deficient cells (Fig. 5A and B) would support this notion and place HO-1 upstream of Akt2. We cannot exclude that other pathways more upstream of Akt2 are also co-effected by HO-1, which needs to be evaluated by future studies, but for now we focused on Akt2 because of its critical role in AP proliferation19. Importantly, the anti-proliferative and anti-adipogenic effect of HO-1 was also observed in primary human preadipocytes which could potentially be important for humans, considering that fat cell numbers increase, also in humans, when overfed for a short period of time58. Pharmacologic HO-1 induction has provided conflicting data on the role of HO-1 in adipocyte differentiation59,60,61,62. “Off-target” effects of pharmacologic HO-1 inducers (discussed in ref. 24), and the use of bone marrow- as opposed to adipose tissue derived mesenchymal stem cells in these studies may explain the discrepancies, considering the cell-autonomous differences between WAT- and bone marrow derived ‘preadipocytes’63,64. Mechanistically, HO-1 acts upstream of Cebpα and PPARγ, as overexpression of HO-1 in 3T3-L1 cells which require MCE to trigger the terminal differentiation transcription cascade, blocks their differentiation only when overexpressed before MCE occurs (Fig. 4C vs J). Our data suggest a trigger-like role for HO-1 in adipogenesis by compromising mitochondria-derived ROS signaling, a scenario shared by macrophages and hepatocytes24. Evidence is accumulating that “low amounts” of ROS are actually fundamental signaling molecules in many cellular processes, promoting stress resistance, and that excessive ROS scavenging may have damaging side effects65. Our findings that elevated mitochondrial activity/ROS levels in HO-1 deficient APs display enhanced proliferation and differentiation capacities are consistent with ROS facilitating fat cell formation38,66,67,68. Here, we add HO-1 as a new upstream building block of such redox-dependent signal “preconditioning” in vivo and suggest a model whereby Akt2 acts as a transducer of HO-1 dependent ROS levels to support AP proliferation. This collectively renders HO-1 as an essential factor linking extrinsic factors (HFD) with specific molecular mediators, resulting in diminished adipogenesis.
Our goal was to evaluate functional effects that occur when activated APs have differentiated into mature adipocytes, a process that requires up to 7 weeks to complete19,23,55, before metaflammation occurs69. Hmox1fl/flPdgfraCre mice fed a HFD for 8 weeks displayed reduced levels of several metabolic parameters (Fig. 6A–D), despite similar fat pad weights. Several possible mechanisms may exist: (i) reduced lipolysis in viWAT examined ex vivo (Fig. 6G), (ii) reduced ECM deposition (Fig. 6E,F), (iii) enhanced lipid filling capacity of activated APs from Hmox1fl/flPdgfraCre mice over the 8 week HFD time course. It is well established that adipose tissue fibrosis as well as elevated FFAs cause systemic metabolic alterations6,70. Our data are in line with a previous report linking reduced lipolysis and free fatty acids with improved glucose and insulin metabolism in adipose tissue GSα-deficient mice, despite similar adiposity71. We recently demonstrated that macrophage as well as hepatocyte HO-1 is a major driver of metaflammation24. Our results are in line with these studies and place adipose precursor HO-1 upstream, and show that HO-1 induction in APs happens very early in a cascade leading to metabolic dysfunction. In this study, we demonstrate that HO-1 regulates the HFD-induced activation of visceral adipose tissue. In the absence of adipose precursor HO-1, proliferation of viAPs in HFD-fed mice is increased, resulting in a clear improvement of WAT function and important metabolic parameters.
Methods
Animals
Animal procedures were approved by the Austrian Ministry for Science and Research and the experiments were carried out in accordance with the approved guidelines and regulations. All mice used for these studies were on the C57BL/6 J genetic background. Hmox1fl/fl mice contain LoxP sites flanking exon 2 of the HO-1 gene locus and were described previously24. Elimination of exon2 leads to a consecutive frameshift in exon 3, an early stop codon, thus resulting in a truncated peptide consisting of 9 amino acids24. PdgfraCre (stock no. 013148) and AdipoQCre (stock no. 010803) mice were purchased from Jackson Laboratories. Nrf2-deficient mice72 were from the same genetic background, which were kindly provided by Masayuki Yamamoto, Department of Medical Biochemistry, Tohoku University Graduate School of Medicine. Mice were males 6–8 weeks of age at the start of experiments. In this study, viWAT refers to the perigonadal visceral adipose tissue in mice. scWAT refers to inguinal subcutaneous adipose tissue in mice. For some experiments, BrdU was administered in the drinking water at 0.8 mg/ml for experiments lasting one week. High-fat diet is from Research Diets (D12492). Standard diet is from Harlan Laboratories (2018S).
Food intake
For food intake experiments, mice were individually caged and food intake was measured by weighing the food pellets prior and after the feeding period. Kilocalories of food intake were normalized to the average body weight of the mouse at the beginning and end of the 3-day period.
Isolation of human preadipocytes
Human SAT was obtained from healthy individuals undergoing lipoaspiration as described41. A total of 5 female donors were used throughout the study and the experiments were performed in accordance with the relevant guidelines and regulations by ethical committee of the Medical University of Vienna ethics committee and the General Hospital of Vienna (protocol ID EK no. 1115/2010). Informed consent was obtained from all subjects.
Heme Oxygenase activity
Cells (approximately 3*106) treated for 16 h with 20 μM hemin or vehicle alone (DMSO) were homogenized in 200 μl of a buffer containing 300 mM sucrose, 20 mM TRIS and 2 mM EDTA at a pH of 7.4 by repeated freezing/unfreezing. 90 μl of the homogenate was added to a reaction mixture containing 500 nmole NADPH in a 100 mM potassium phosphate buffer with 1 mM EDTA (assay buffer pH: 7.4), supplemented with 20 nmoles of hemin. The mixture was incubated under constant agitation at 37 °C in darkness for 30 min. The reaction was stopped by transferring the samples on ice followed by addition of 1/5 volume of saturated KCl. The formed bilirubin was extracted into benzene (4× the assay volume) by vortexing (3 × 30 s) and the organic phase was harvested by centrifugation (250 × g). Bilirubin concentration of the extracts was determined by photo spectroscopy (U-3000, Hitachi, Tokyo, Japan), scanning twice the absorption between 600 and 400 nm using the following settings: slit: 2 nm, 120 nm/min, PMT: autogain, high resolution. The difference in absorption was determined between 450 and 520 nm and the calculation of bilirubin concentration was obtained using a standard calibration curve, generated by adding known amounts of bilirubin to the assay buffer. Protein concentrations of cell homogenates were determined using the Coomassie Brilliant Blue assay (Bradford). HO activity was calculated as nmole bilirubin formed per mg protein per 30 min.
Whole-mount immunofluorescence
Whole-mount visceral fat samples were processed and stained as described73. Briefly, 6–7 week old male C57BL6/J mice (three per group) were fed a SD or HFD for three days. Thereafter, fat samples were minced into 1–2 mm pieces and permeabilized for 30 min at room temperature with 0,5% PBS-T. After 3 washes with 0,05% PBS-T tissue pieces were incubated in 5% normal goat serum (Dako) in PBS to block unspecific binding sites. After blocking, tissues were incubated overnight at 4 °C with primary antibodies diluted in PBS-T (rabbit anti-HO-1; 1:500; ab13243, Abcam and rat anti-CD24; 1:500; Cat.-Nr. 101801, Biolegend). The following secondary antibodies were used for immunofluorescence staining: fluorescein anti-rat IgG (Cat.-Nr. FI-4001, Vector Laboratories), Alexa Fluor-594 goat anti-rabbit IgG (Cat.-Nr. A11037, Life Technologies). For control experiments, the primary antibody was substituted with preimmune serum. Stained tissues were incubated with Celltracker Green Bodipy Dye (Cat.-Nr. C2102, Thermo Scientific) to visualize lipid droplets for 10 minutes at room temperature before mounting in ibiTreat μ-dishes (ibidi GmbH) and subjected to confocal imaging analysis on a LSM 700 Laser Scanning Microscope (Zeiss).
Flow cytometry analysis of cell proliferation
Isolation of adipose tissue stromal cells was performed essentially as described20. Briefly, adipose tissue was excised, minced, and digested in Hank’s Balanced Salt Solution (HBSS) (Sigma no. H8264) containing 3% BSA and 0.8 mg/ml Collagenase Type 2 (Worthington Biochemical; LS004174) for 75 min in a shaking water bath at 37 °C. The mixture was then filtered through a 40 μm filter, and filtered cells were pelleted and washed in HBSS containing 3% BSA. For Ki67 analysis, cells were stained with the following antibodies at room temperature: CD45 PE-Cy7 (Biolegend; clone 30-F11), CD31 PE-Cy7 (Biolegends; clone 390), CD29 Alexa Fluor 700 (BioLegend; clone HMβ1-1), CD34 Alexa Fluor 647 (BioLegend; clone MEC14.7) and CD140a PE (Biolegend; clone APA5). Anti-Ki67 FITC antibody (eBioscience, clone SOLA15) was used overnight at 4 °C. Cells were washed, fixed and permeabilized using the Fixation/Permeabilization kit (eBioscience) as per manufacturer’s protocol. All antibody incubations and washing steps were performed in HBSS with 3% BSA. Following antibody incubation, samples were analyzed on a BD LSR Fortessa analyzer. Data analysis was performed using BD FACS Diva software (BD Biosciences).
Isolation of AP cells by bead separation
sc or viWAT was digested, filtered, and washed as described for flow cytometry analysis. Adipose precursors were then isolated as per the manufacturer’s MACS protocol (Miltenyi) using first negative selection of CD45 and CD31 followed by positive selection for SCA-1. Briefly, cell suspensions were incubated with the Non-Adipocyte Progenitor Depletion Cocktail, washed and passed over a lineage depletion column. The flow-through was collected, washed, and incubated with the Adipocyte Progenitor Isolation Cocktail. Cells were then washed and passed over a lineage selection column. Retained cells were eluted, counted, and either lysed directly into RLT lysis buffer for RNA isolation and subsequent Q-PCR, or RIPA buffer for protein isolation and subsequent western blotting, or cultured as previously described (Ohno et al., 2012). Briefly, APs were expanded in DMEM F12 with 10% fetal bovine serum (GIBCO) in 6 cm tissue culture plates at low confluency until ready for use. Cells were replated at 1 × 105 cells per well in 24-well plates and treated as indicated. After treatment, cells were thoroughly washed, and RNA isolated using RNeasy mini kit (Qiagen) as per manufacturer’s protocol.
AP stimulation with fatty acids
Bead isolated APs from male C57BL/6 J and NRF2 knockout mice were seeded in 24 Well plates and cultured until they reached confluency. Cells were then treated with either palmitic, oleic or linoleic acid at a concentration of 250 μM for 16 hours. Fatty acid/BSA conjugates were prepared by dissolving the fatty acid powder in 0.1 M NaOH at 70 °C to generate a 100 mM stock solution. A 10 mM solution was prepared by mixing the stock solution with a 5% (w/v) free fatty acid free BSA solution in double distilled water. Finally, the 10 mM solution was further diluted in serum free medium to obtain the final concentration of 1.0 mM fatty acid/0.5% BSA.
Adenovirus experiments
Generation of AdHO-1 and AdLacZ was described previously24. In brief, Primary hASC were infected with 30 plaque forming units (pfu)/cell.
Gene expression profiling
Adipose precursor cell screen
Each of the eight microarray samples represent RNA isolated from APs derived from scWAT or viWAT from SD- or HFD fed C57BL/6 J mice (at age 7–8 weeks) representing pooled samples from 3 mice. Data were imported and preprocessed using R and Bioconductor, specifically the ‘oligo’ package version ‘1.34.2’ and ‘pd.mogene.1.0.st.v1’ version ‘3.14.1’. Data were preprocessed using quantile normalization to account for latent batch effects and RMA to summarize probesets. This combination has been shown to effectively reduce batch effects and to provide good sensitivity for the detection of differentially expressed genes74. For each probeset we fit a linear model to the effect of CRE treatment. An empirical bayes procedure was used to exploit information across different probesets by shrinking means and standard deviations toward common values. Non-specific filtering was applied to reduce the number of statistical hypothesis tests. Only probesets with log2 expressions above 7 in at least 3 samples, and a mean absolute deviation in the upper terzile were selected. Among the remaining probesets we only selected those that had a unique Gene-Symbol annotation using Bioconductor package ‘mogene10sttranscriptcluster.db’ version 8.4.0. Statistical significance of the estimated differential effects was assessed using t-tests and corresponding p-values. Multiple testing adjustment was performed using the Benjamini-Hochberg procedure, which provides control of the false discovery rate.
Gene Set Enrichment Analysis (GSEA)
Biological insights concerning the differentially expressed genes were explored via Gene Set Enrichment Analysis (GSEA). The analysis was performed with the GSEA software75 (version 2.1.0) using the c2 (version 5) gene set database. In addition, a 3T3-L1 differentiation time course31 was used to construct gene sets resembling preadipocyte‚ ‘proliferation’ or‚ ‘terminal differentiation’, by assembling differentially expressed genes between day-2 vs. day 0 or day 7 vs. day 2, respectively76. Genes were first ranked based on real value using the weighted signal-to noise metric. P-values and the false discovery rate (FDR) for the enrichment score of each gene set were calculated based on 1000 gene set permutations. Enrichment plots were generated using Cytoscape 3.3.0 and the plugin Enrichment map77.
Oxygen consumption assay and bioenergetic profile
Oxygen consumption rates (OCR) of bead-purified murine APs from viWAT were performed using the XF24 Flux Analyzer (Seahorse Bioscience) as reported previously24. In brief, bead-purified visceral APs were seeded into XF 24-well cell culture microplates. After a 24 hour recovery period cells were washed and a final volume of 630 μl buffer-free Assay Medium (Seahorse Bioscience, supplemented with 5 mM glucose (Sigma) and 1 mM sodium pyruvate (Gibco)) was added to each well prior to the experimental protocol. Cells were then kept in a CO2-free incubator at 37 °C for 1 hour. After instrument calibration, cells were transferred to the XF24 Flux Analyzer to record cellular oxygen consumption rates. Measurements were performed with repetitive cycles of 2 min mixture, 2 min wait and 4 min OCR measurement times. Injected compounds for the mitochondrial stress test were oligomycin (2 μM working concentration) to inhibit ATP synthase, followed by FCCP (1 μM working concentration) to induce mitochondrial uncoupling and rotenone/antimycin A (2 μM working concentration each) to block the mitochondrial respiratory chain. Following the assay, the medium was carefully aspirated and cellular protein content was measured using the Pierce BCA Protein Assay Kit (Thermo Scientific) to adjust for potential differences in cell numbers. For ROS-quenching experiments, APs were incubated with 10 μM buffered N-acetyl-cysteine (NAC) for 1 hour prior to otherwise unchanged experimental procedures.
Western blot analysis
Protein concentration for the resulting lysates was determined using the BCA (bicinchoninic acid) Protein Assay kit from Pierce, and lysates were run on 10% polyacrylamide gels. PVDF membranes (GE healthcare) were incubated with following antibodies: HO-1 (Abcam), β-actin (Sigma), pAkt2-S474 and Akt2 (all Cell Signaling). Hrp-conjugated IgG secondary antibodies were used (Cell Signaling) and blots were developed with ECL Plus Western Blotting Detection System (GE Healthcare). ImageLab software (BioRad) was used for densitometric quantification.
Adipocyte differentiation
At 2 days post-confluency preadipocytes were treated with a differentiation cocktail containing: DMEM, 10% CS, insulin (870 nM), TZD (5 μM), Dex (1 μM), and IBMX (500 μM). On day 3 of differentiation, cells were switched to differentiation cocktail excluding IBMX and Dex for remaining duration of differentiation.
Q-PCR
Q-PCR was performed as previously published41. All Q-PCR data are normalized to amounts of acidic ribosomal phosphoprotein P0 (RPLP0), unless otherwise stated. Primer sequences are summarized in Supplemental Table S6.
Oil-red O staining
Cells were fixed in 10% formalin, thereafter stained for 20 minutes with Oil-RedO (Sigma) working solution (composed of 4 parts water and 6 parts 0.6% Oil-red O dye in isopropanol). Oil-Red O was extracted using 100% isopropanol and absorbance was measured at 490 nm using a spectrophotometer.
Retrovirus preparation and infection
Retrovirus preparation and infection were performed as described78. Briefly, pMSCV, LMP empty vectors, or their derivatives containing specific cDNA or shRNAmir, along with vectors containing reverse transcriptase (gag-pol) and VSV-G-expressing plasmids, were transfected into Phoenix packaging cells with the CellPhect transfection kit (Amersham Biosciences). Viral supernatant was collected 48 hrs after transfection, filtered through 0.45 μm filters, and added to target cells for 12 hrs along with 8 g/ml Polybrene. Cells were selected with 4 g/ml puromycin or 400 g/ml hygromycin to make stable lines and were maintained in media containing appropriate antibiotics.
Construction of transgenic cell lines
Constitutive stable HO-1 knock-down in 3T3-L1 cells was generated by transduction with the microRNA (miRNA) adapted retroviral vector LMP (OpenBiosystems, Huntsville, AL, USA). shRNAmir (microRNA-adapted short hairpin RNA) against murine HO-1 generated in pSM2 vector (Open Biosystems) was subcloned into the LMP vector with XhoI and EcoRI (Invitrogen, Carlsbad, CA) restriction enzymes. Confirmation was verified by restriction site analysis and sequencing. To produce murine stem cell virus (MSCV) particles, HEK293FT cells were transiently co-transfected with a vector containing the viral packaging proteins gag and pol, a vector containing env, and either LMP, or LMP-miHO1 (LMP containing miRNA against human HO-1). Vectors containing gag, env, and pol were kind gifts from Dr. Ewan Rosen (Beth Israel Deaconess Medical Center, Harvard Medical School, Boston, MA, USA). Lipofectamine 2000 (Invitrogen) reagent was used for transfection. 48 hours after transfection, viral supernatants were collected, centrifuged at 1500 rpm for 3 min, filtered through a 0.4 μm filter, supplemented with 8 μg/ml Polybrene (Sigma), and used to infect 3T3-L1 cells. Stable integrants were selected with puromycin (5 μg/ml) over a period of 2 weeks. HO-1 knock-down was verified by western blotting. The coding region of human HO-1 cDNA (a gift of Dr. Matthias Mayerhofer, Department of Laboratory Medicine, Medical University of Vienna, Vienna, Austria) was inserted into the retroviral vector pMSCV-puro (Clontech, Mountain View, CA, USA) at the BamHI and HindIII sites within the MCS. To produce MSCV retrovirus, HEK293FT cells were transiently co-transfected with a vector containing the viral packaging proteins gag and pol, a vector containing env and MSCV or MSCV-HO1. Lipofectamine 2000 (Invitrogen) reagent was used for transfection. 48 hours after transfection, viral supernatants were collected, centrifuged at 1500 rpm for 3 min, filtered through a 0.4 μm filter, supplemented with 8 μg/ml Polybrene (Sigma), and used to infect 3T3-L1 cells. Stable cell pools transduced with either empty plasmid or plasmid carrying HO-1 cDNA were termed MSCV and MSCV-HO1, respectively. HO-1 expression was verified by western blotting.
BrdU cell proliferation assay
BrdU uptake in 3T3-L1 gain/loss of function cells and in human SVFs was determined as previously described41. Briefly, untreated, LMP, miHO-1, MSCV and MSCV HO-1 were plated into 96-well culture plates at a density of 10,000 cells per well. Differentiation was initiated two days after seeding and cells were kept at 37 °C for 10 days. Proliferation was established using the Cell Proliferation ELISA, BrdU kit (Roche) according to manufacturer’s protocol. In brief, 10 μl BrdU labeling solution was added to 100 μl differentiation medium per well. Cells were then incubated at 37 °C for 2 hours, followed by the removal of labeling medium and addition of 200 μl/well FixDenat solution. After a 30 minute incubation time at RT the solution was removed by tapping. Thereafter, 100 μl/well anti-BrdU-POD working solution was added for 90 minutes at RT. Wells were washed 3 times and then 100 μl/well substrate solution was added for 15 minutes at RT. Finally, cells were treated with 25 μl/well 1 M H2SO4 stop solution. Absorbance was measured on a multimode microplate reader (Synergy 2, BioTek Instruments) at 450 nm and a reference wavelength of 690 nm.
Nitroblue tetrazolium (NBT) assay
Bead-purified adipose precursors of 8 to 10-week-old control (Hmox1fl/fl) and HO-1 knockout (Hmox1fl/flPdgfraCre) mice were cultured in 48-well culture plates. After a 48 hour recovery phase cells were incubated with 0.2% NBT (Sigma) in PBS for 2 hours at 37 °C. Reduced NBT was dissolved in 50% acetic acid and absorbance was measured at 560 nm. Results were normalized to individual cell counts.
MitoSOX Red measurement
Visceral APs were bead-purified as previously described and seeded into 24 Well plates. Cells were allowed to adhere for 24 hours and afterwards incubated with 5 μM MitoSOX Red (Molecular Probes; M36008) for 15 min at 37 °C. Cells were washed off with PBS and after two further washing steps cells were resuspended in HBSS containing 3% BSA. Superoxide production was measured as fluorescence intensities at 510/580 nm.
Histology and Adipocyte size
Hematoxylin and eosin (H&E) staining was performed on 5 μm paraffin sections. Adipocyte size distribution was determined by semi-automated morphometry. In brief, 3 fields of view of 3 different sections per animal were quantified. Visceral and subcutaneous fat pads from 4 animals per group were analyzed. Adipocyte size was measured using Adiposoft79.
Adipose tissue collagen assay
Collagen was assessed using the QuickZyme Total Collagen Assay as per manufacturer’s protocol. Briefly, after 8 weeks on high fat diet ~300 mg of visceral adipose tissue was directly incubated in 6 M HCl for 20 hours at 95 °C. Tubes were then cooled to room temperature and centrifuged at 13,000 g for 10 min. 35 μl of the supernatant was added to 75 μl of assay buffer in a microplate and incubated at RT for 20 min. Afterwards 75 μl detection reagent was added to each well and the microplate was incubated for 1 hour at 60 °C. Absorbance was measured at 570 nm and collagen content was calculated using a standard curve.
Ex vivo lipolytic assessment of adipose tissue
After an 8 week long HFD feeding period, visceral adipose tissue was surgically removed and washed in PBS. Fat depots were cut into 20 mg pieces and incubated in DMEM containing 2% fatty-acid-free BSA either with 10 μM isoproterenol (stimulated) or without (basal) for 1 hour at 37 °C. Fat pieces were then transferred into fresh medium and incubated for another hour. Free fatty acid release into the medium was measured using the WAKO NEFA-HR (2) kit as per manufacturer’s protocol.
Trichrome staining
Visceral adipose tissue of obese mice was fixed in 10% formalin for at least 24 hours at 4 °C and afterwards embedded in paraffin. Blocks were cut in 10 μm slices and incubated overnight at 56 °C. Following standard dehydration steps (20 min Xylen, 2 min 100% EtOH, 2 min 95% EtOH, 2 min 70% EtOH) slices were submerged into Postmordanding solution (200 ml saturated picuis acid; 8 g mercuris chloride) for 1 hour. After a warm tap water wash slices were incubated in Verhoff’s stain (120 ml hematoxylin; 13,5 ml ferric chloride; 66,5 ml Weigerts iodine) for 8 min and washed again. Slices were stained for 5 min with Fast yellow (2 g Acid yellow 17; 1 ml glacial HAc; 200 ml dH2O) and washed three times with 0,5% HAc. This was followed by 6 min in Biebrich stain (1,5 g Biebrich scarlet; 1,5 g acid fuchsin; 1,5 g Ponceau 2 R; 1 ml glacial HAc; 200 ml dH2O) and a wash with dH2O. Afterwards slices were incubated for 6 min with Phosphotungstic acid (2%) and washed again with dH2O. Finally, slices were stained with Light green stain (4 g Light green SF; 4 g glacial HAc; 200 ml dH2O) for 10 min and washed three times with 0,5% HAc. Slices were washed extensively in dH2O, and following 2 min in 70% EtOH and 2 min in 100% EtOH they were incubated in butylacetat for 20 min. Eukitt embedded fatpad slices were imaged using an Axioimager M2 (Zeiss).
Statistical Analysis
Data are expressed as mean ± standard error of the mean (SEM). Statistical significance was tested by Student’s t test or 2-way ANOVA where appropriate. Correlations were tested by linear regression. All figures and statistical analysis were generated using Prism 6 (GraphPad). All reported p values are two-tailed. p < 0.05 was considered to indicate statistical significance.
Additional Information
Accession Code: Data sets have been deposited in the Gene expression omnibus archive as series GSE80148. A reviewer accessible link has been created and is available at http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?token=ejgdqouubpirhsv&acc=GSE80148.
How to cite this article: Wagner, G. et al. HO-1 inhibits preadipocyte proliferation and differentiation at the onset of obesity via ROS dependent activation of Akt2. Sci. Rep. 7, 40881; doi: 10.1038/srep40881 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Cristancho, A. G. & Lazar, M. A. Forming functional fat: a growing understanding of adipocyte differentiation. Nat Rev Mol Cell Biol 12, 722–734, doi: 10.1038/nrm3198 (2011).
Rosen, E. D. & Spiegelman, B. M. What we talk about when we talk about fat. Cell 156, 20–44, doi: 10.1016/j.cell.2013.12.012 (2014).
Pellegrinelli, V., Carobbio, S. & Vidal-Puig, A. Adipose tissue plasticity: how fat depots respond differently to pathophysiological cues. Diabetologia, doi: 10.1007/s00125-016-3933-4 (2016).
Spalding, K. L. et al. Dynamics of fat cell turnover in humans. Nature 453, 783–787, doi: 10.1038/nature06902 (2008).
Arner, P. et al. Variations in the size of the major omentum are primarily determined by fat cell number. J Clin Endocrinol Metab 98, E897–901, doi: 10.1210/jc.2012-4106 (2013).
Sun, K., Tordjman, J., Clement, K. & Scherer, P. E. Fibrosis and adipose tissue dysfunction. Cell Metab 18, 470–477, doi: 10.1016/j.cmet.2013.06.016 (2013).
Rutkowski, J. M., Stern, J. H. & Scherer, P. E. The cell biology of fat expansion. J Cell Biol 208, 501–512, doi: 10.1083/jcb.201409063 (2015).
Naaz, A. et al. Loss of cyclin-dependent kinase inhibitors produces adipocyte hyperplasia and obesity. FASEB J 18, 1925–1927, doi: 10.1096/fj.04-2631fje (2004).
Samocha-Bonet, D. et al. Metabolically healthy and unhealthy obese–the 2013 Stock Conference report. Obes Rev 15, 697–708, doi: 10.1111/obr.12199 (2014).
Gustafson, B., Hedjazifar, S., Gogg, S., Hammarstedt, A. & Smith, U. Insulin resistance and impaired adipogenesis. Trends Endocrinol Metab 26, 193–200, doi: 10.1016/j.tem.2015.01.006 (2015).
Heilbronn, L., Smith, S. R. & Ravussin, E. Failure of fat cell proliferation, mitochondrial function and fat oxidation results in ectopic fat storage, insulin resistance and type II diabetes mellitus. Int J Obes Relat Metab Disord 28 Suppl 4, S12–21, doi: 10.1038/sj.ijo.0802853 (2004).
van Tienen, F. H. et al. Preadipocytes of type 2 diabetes subjects display an intrinsic gene expression profile of decreased differentiation capacity. Int J Obes (Lond) 35, 1154–1164, doi: 10.1038/ijo.2010.275 (2011).
Hoffstedt, J. et al. Regional impact of adipose tissue morphology on the metabolic profile in morbid obesity. Diabetologia 53, 2496–2503, doi: 10.1007/s00125-010-1889-3 (2010).
Muir, L. A. et al. Adipose tissue fibrosis, hypertrophy, and hyperplasia: Correlations with diabetes in human obesity. Obesity (Silver Spring) 24, 597–605, doi: 10.1002/oby.21377 (2016).
Green, H. & Meuth, M. An established pre-adipose cell line and its differentiation in culture. Cell 3, 127–133 (1974).
Fischer-Posovszky, P., Newell, F. S., Wabitsch, M. & Tornqvist, H. E. Human SGBS cells - a unique tool for studies of human fat cell biology. Obes Facts 1, 184–189, doi: 10.1159/000145784 (2008).
Berry, R., Jeffery, E. & Rodeheffer, M. S. Weighing in on adipocyte precursors. Cell Metab 19, 8–20, doi: 10.1016/j.cmet.2013.10.003 (2014).
Lee, Y. H., Petkova, A. P., Mottillo, E. P. & Granneman, J. G. In vivo identification of bipotential adipocyte progenitors recruited by beta3-adrenoceptor activation and high-fat feeding. Cell Metab 15, 480–491, doi: 10.1016/j.cmet.2012.03.009 (2012).
Jeffery, E., Church, C. D., Holtrup, B., Colman, L. & Rodeheffer, M. S. Rapid depot-specific activation of adipocyte precursor cells at the onset of obesity. Nat Cell Biol 17, 376–385, doi: 10.1038/ncb3122 (2015).
Church, C. D., Berry, R. & Rodeheffer, M. S. Isolation and study of adipocyte precursors. Methods Enzymol 537, 31–46, doi: 10.1016/B978-0-12-411619-1.00003-3 (2014).
Lee, Y. H., Mottillo, E. P. & Granneman, J. G. Adipose tissue plasticity from WAT to BAT and in between. Biochim Biophys Acta 1842, 358–369, doi: 10.1016/j.bbadis.2013.05.011 (2014).
Berry, R. & Rodeheffer, M. S. Characterization of the adipocyte cellular lineage in vivo . Nat Cell Biol 15, 302–308, doi: 10.1038/ncb2696 (2013).
Wang, Q. A., Tao, C., Gupta, R. K. & Scherer, P. E. Tracking adipogenesis during white adipose tissue development, expansion and regeneration. Nat Med 19, 1338–1344, doi: 10.1038/nm.3324 (2013).
Jais, A. et al. Heme oxygenase-1 drives metaflammation and insulin resistance in mouse and man. Cell 158, 25–40, doi: 10.1016/j.cell.2014.04.043 (2014).
Lee, K. Y. et al. Lessons on conditional gene targeting in mouse adipose tissue. Diabetes 62, 864–874, doi: 10.2337/db12-1089 (2013).
Lee, M. W. et al. Activated type 2 innate lymphoid cells regulate beige fat biogenesis. Cell 160, 74–87, doi: 10.1016/j.cell.2014.12.011 (2015).
Cawthorn, W. P., Scheller, E. L. & MacDougald, O. A. Adipose tissue stem cells meet preadipocyte commitment: going back to the future. J Lipid Res 53, 227–246, doi: 10.1194/jlr.R021089 (2012).
Javazon, E. H., Beggs, K. J. & Flake, A. W. Mesenchymal stem cells: paradoxes of passaging. Exp Hematol 32, 414–425, doi: 10.1016/j.exphem.2004.02.004 (2004).
Macotela, Y. et al. Intrinsic differences in adipocyte precursor cells from different white fat depots. Diabetes 61, 1691–1699, doi: 10.2337/db11-1753 (2012).
Hudak, C. S. et al. Pref-1 marks very early mesenchymal precursors required for adipose tissue development and expansion. Cell Rep 8, 678–687, doi: 10.1016/j.celrep.2014.06.060 (2014).
Mikkelsen, T. S. et al. Comparative epigenomic analysis of murine and human adipogenesis. Cell 143, 156–169, doi: 10.1016/j.cell.2010.09.006 (2010).
Kong, P., Cavalera, M. & Frangogiannis, N. G. The role of thrombospondin (TSP)-1 in obesity and diabetes. Adipocyte 3, 81–84, doi: 10.4161/adip.26990 (2014).
Maquoi, E., Munaut, C., Colige, A., Collen, D. & Lijnen, H. R. Modulation of adipose tissue expression of murine matrix metalloproteinases and their tissue inhibitors with obesity. Diabetes 51, 1093–1101 (2002).
Ruan, H., Zarnowski, M. J., Cushman, S. W. & Lodish, H. F. Standard isolation of primary adipose cells from mouse epididymal fat pads induces inflammatory mediators and down-regulates adipocyte genes. J Biol Chem 278, 47585–47593, doi: 10.1074/jbc.M305257200 (2003).
Jais, A. et al. Myeloid-Cell-Derived VEGF Maintains Brain Glucose Uptake and Limits Cognitive Impairment in Obesity. Cell 165, 882–895, doi: 10.1016/j.cell.2016.03.033 (2016).
Jeffery, E. et al. Characterization of Cre recombinase models for the study of adipose tissue. Adipocyte 3, 206–211, doi: 10.4161/adip.29674 (2014).
Tang, Q. Q., Otto, T. C. & Lane, M. D. Mitotic clonal expansion: a synchronous process required for adipogenesis. Proc Natl Acad Sci USA 100, 44–49, doi: 10.1073/pnas.0137044100 (2003).
Tormos, K. V. et al. Mitochondrial complex III ROS regulate adipocyte differentiation. Cell Metab 14, 537–544, doi: 10.1016/j.cmet.2011.08.007 (2011).
Mailloux, R. J., McBride, S. L. & Harper, M. E. Unearthing the secrets of mitochondrial ROS and glutathione in bioenergetics. Trends Biochem Sci 38, 592–602, doi: 10.1016/j.tibs.2013.09.001 (2013).
Senol-Cosar, O. et al. Tenomodulin promotes human adipocyte differentiation and beneficial visceral adipose tissue expansion. Nat Commun 7, 10686, doi: 10.1038/ncomms10686 (2016).
Lindroos, J. et al. Human but not mouse adipogenesis is critically dependent on LMO3. Cell Metab 18, 62–74, doi: 10.1016/j.cmet.2013.05.020 (2013).
Gupta, R. K. et al. Zfp423 expression identifies committed preadipocytes and localizes to adipose endothelial and perivascular cells. Cell Metab 15, 230–239, doi: 10.1016/j.cmet.2012.01.010 (2012).
Vishvanath, L. et al. Pdgfrbeta(+) Mural Preadipocytes Contribute to Adipocyte Hyperplasia Induced by High-Fat-Diet Feeding and Prolonged Cold Exposure in Adult Mice. Cell Metab 23, 350–359, doi: 10.1016/j.cmet.2015.10.018 (2016).
Tchkonia, T. et al. Mechanisms and metabolic implications of regional differences among fat depots. Cell Metab 17, 644–656, doi: 10.1016/j.cmet.2013.03.008 (2013).
Wu, J. et al. Beige adipocytes are a distinct type of thermogenic fat cell in mouse and human. Cell 150, 366–376, doi: 10.1016/j.cell.2012.05.016 (2012).
Abella, A. et al. Cdk4 promotes adipogenesis through PPARgamma activation. Cell Metab 2, 239–249, doi: 10.1016/j.cmet.2005.09.003 (2005).
Merkestein, M. et al. FTO influences adipogenesis by regulating mitotic clonal expansion. Nat Commun 6, 6792, doi: 10.1038/ncomms7792 (2015).
Lim, G. E. et al. 14-3-3zeta coordinates adipogenesis of visceral fat. Nat Commun 6, 7671, doi: 10.1038/ncomms8671 (2015).
Kamba, T. et al. VEGF-dependent plasticity of fenestrated capillaries in the normal adult microvasculature. Am J Physiol Heart Circ Physiol 290, H560–576, doi: 10.1152/ajpheart.00133.2005 (2006).
Sun, K., Kusminski, C. M. & Scherer, P. E. Adipose tissue remodeling and obesity. J Clin Invest 121, 2094–2101, doi: 10.1172/JCI45887 (2011).
Divoux, A. et al. Fibrosis in human adipose tissue: composition, distribution, and link with lipid metabolism and fat mass loss. Diabetes 59, 2817–2825, doi: 10.2337/db10-0585 (2010).
Lee, Y. H., Petkova, A. P. & Granneman, J. G. Identification of an adipogenic niche for adipose tissue remodeling and restoration. Cell Metab 18, 355–367, doi: 10.1016/j.cmet.2013.08.003 (2013).
Lee, Y. H. & Granneman, J. G. Seeking the source of adipocytes in adult white adipose tissues. Adipocyte 1, 230–236, doi: 10.4161/adip.20804 (2012).
Jeffery, E. et al. The Adipose Tissue Microenvironment Regulates Depot-Specific Adipogenesis in Obesity. Cell Metab 24, 142–150, doi: 10.1016/j.cmet.2016.05.012 (2016).
Wang, Q. A. & Scherer, P. E. The AdipoChaser mouse: A model tracking adipogenesis in vivo . Adipocyte 3, 146–150, doi: 10.4161/adip.27656 (2014).
Gao, H. et al. Early B cell factor 1 regulates adipocyte morphology and lipolysis in white adipose tissue. Cell Metab 19, 981–992, doi: 10.1016/j.cmet.2014.03.032 (2014).
Vishvanath, L. et al. Pdgfrbeta+ Mural Preadipocytes Contribute to Adipocyte Hyperplasia Induced by High-Fat-Diet Feeding and Prolonged Cold Exposure in Adult Mice. Cell Metab 23, 350–359, doi: 10.1016/j.cmet.2015.10.018 (2016).
Tchoukalova, Y. D. et al. Regional differences in cellular mechanisms of adipose tissue gain with overfeeding. Proc Natl Acad Sci USA 107, 18226–18231, doi: 10.1073/pnas.1005259107 (2010).
Choudhary, A. K. et al. Administration of heme arginate ameliorates murine type 2 diabetes independently of heme oxygenase activity. PLoS One 8, e78209, doi: 10.1371/journal.pone.0078209 (2013).
Li, M. et al. Treatment of obese diabetic mice with a heme oxygenase inducer reduces visceral and subcutaneous adiposity, increases adiponectin levels, and improves insulin sensitivity and glucose tolerance. Diabetes 57, 1526–1535, doi: 10.2337/db07-1764 (2008).
Nicolai, A. et al. Heme oxygenase-1 induction remodels adipose tissue and improves insulin sensitivity in obesity-induced diabetic rats. Hypertension 53, 508–515, doi: 10.1161/HYPERTENSIONAHA.108.124701 (2009).
Vanella, L. et al. Increased heme-oxygenase 1 expression in mesenchymal stem cell-derived adipocytes decreases differentiation and lipid accumulation via upregulation of the canonical Wnt signaling cascade. Stem Cell Res Ther 4, 28, doi: 10.1186/scrt176 (2013).
Hsiao, S. T. et al. Comparative analysis of paracrine factor expression in human adult mesenchymal stem cells derived from bone marrow, adipose, and dermal tissue. Stem Cells Dev 21, 2189–2203, doi: 10.1089/scd.2011.0674 (2012).
Berry, R., Rodeheffer, M. S., Rosen, C. J. & Horowitz, M. C. Adipose Tissue Residing Progenitors (Adipocyte Lineage Progenitors and Adipose Derived Stem Cells (ADSC)). Curr Mol Biol Rep 1, 101–109, doi: 10.1007/s40610-015-0018-y (2015).
Ristow, M. Unraveling the truth about antioxidants: mitohormesis explains ROS-induced health benefits. Nat Med 20, 709–711, doi: 10.1038/nm.3624 (2014).
Kanda, Y., Hinata, T., Kang, S. W. & Watanabe, Y. Reactive oxygen species mediate adipocyte differentiation in mesenchymal stem cells. Life Sci 89, 250–258, doi: 10.1016/j.lfs.2011.06.007 (2011).
Chang, Y. C. et al. Deficiency of NPGPx, an oxidative stress sensor, leads to obesity in mice and human. EMBO Mol Med 5, 1165–1179, doi: 10.1002/emmm.201302679 (2013).
Lee, H., Lee, Y. J., Choi, H., Ko, E. H. & Kim, J. W. Reactive oxygen species facilitate adipocyte differentiation by accelerating mitotic clonal expansion. J Biol Chem 284, 10601–10609, doi: 10.1074/jbc.M808742200 (2009).
Nishimura, S. et al. CD8+ effector T cells contribute to macrophage recruitment and adipose tissue inflammation in obesity. Nat Med 15, 914–920, doi: 10.1038/nm.1964 (2009).
Boden, G. Obesity, insulin resistance and free fatty acids. Curr Opin Endocrinol Diabetes Obes 18, 139–143, doi: 10.1097/MED.0b013e3283444b09 (2011).
Li, Y. Q. et al. Gsalpha deficiency in adipose tissue improves glucose metabolism and insulin sensitivity without an effect on body weight. Proc Natl Acad Sci USA 113, 446–451, doi: 10.1073/pnas.1517142113 (2016).
Itoh, K. et al. An Nrf2/small Maf heterodimer mediates the induction of phase II detoxifying enzyme genes through antioxidant response elements. Biochem Biophys Res Commun 236, 313–322 (1997).
Berry, R. et al. Imaging of adipose tissue. Methods Enzymol 537, 47–73, doi: 10.1016/B978-0-12-411619-1.00004-5 (2014).
Klinglmueller, F., Tuechler, T. & Posch, M. Cross-platform comparison of microarray data using order restricted inference. Bioinformatics 27, 953–960, doi: 10.1093/bioinformatics/btr066 (2011).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102, 15545–15550, doi: 10.1073/pnas.0506580102 (2005).
Kang, S. et al. Identification of nuclear hormone receptor pathways causing insulin resistance by transcriptional and epigenomic analysis. Nat Cell Biol 17, 44–56, doi: 10.1038/ncb3080 (2015).
Merico, D., Isserlin, R., Stueker, O., Emili, A. & Bader, G. D. Enrichment map: a network-based method for gene-set enrichment visualization and interpretation. PLoS One 5, e13984, doi: 10.1371/journal.pone.0013984 (2010).
Tauber, S. et al. Transcriptome analysis of human cancer reveals a functional role of heme oxygenase-1 in tumor cell adhesion. Mol Cancer 9, 200, doi: 10.1186/1476-4598-9-200 (2010).
Galarraga, M. et al. Adiposoft: automated software for the analysis of white adipose tissue cellularity in histological sections. J Lipid Res 53, 2791–2796, doi: 10.1194/jlr.D023788 (2012).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4, 44–57, doi: 10.1038/nprot.2008.211 (2009).
Acknowledgements
This work was supported by the Austrian Science Fund (W1205, CCHD PhD program). We thank the following colleagues from the Medical University of Vienna, Vienna, Austria: Markus Jeitler for help with DNA Microarrays; Julian Mohrherr and Sylvia Wolkerstorfer for help with mouse work; Florian Kiefer and Alexander Jais for fruitful discussions and critical reading of the manuscript. The authors are grateful to Masayuki Yamamoto (Department of Medical Biochemistry, Tohoku University Graduate School of Medicine, Japan) for providing Nrf2 deficient mice. The financial support by the Federal Ministry of Science, Research and Economy (BMWFW) and the National Foundation for Research, Technology, and Development is gratefully acknowledged.
Author information
Authors and Affiliations
Contributions
M.B. conceived the project. G.W., J.L.-C., E.E., J.H., K.L., S.R., T.C.Z. and C.D. performed the experiments. G.W., A.S., M.G., T.M., F.K. and M.B. analyzed the data. G.P., H.S.-F., B.H., F.G., J.Y. contributed reagents/materials/tools. R.L. and O.W. provided insightful comments. G.W., H.E. and M.B. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Wagner, G., Lindroos-Christensen, J., Einwallner, E. et al. HO-1 inhibits preadipocyte proliferation and differentiation at the onset of obesity via ROS dependent activation of Akt2. Sci Rep 7, 40881 (2017). https://doi.org/10.1038/srep40881
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep40881
This article is cited by
-
The associations of oxidative stress and inflammatory markers with obesity in Iranian population: MASHAD cohort study
BMC Endocrine Disorders (2024)
-
Associations of dietary, lifestyle, other participant characteristics, and oxidative balance scores with plasma F2-isoprostanes concentrations in a pooled cross-sectional study
European Journal of Nutrition (2022)
-
Heme Oxygenase 1 in Vertebrates: Friend and Foe
Cell Biochemistry and Biophysics (2022)
-
LMO3 reprograms visceral adipocyte metabolism during obesity
Journal of Molecular Medicine (2021)
-
Surgical fat removal exacerbates metabolic disorders but not atherogenesis in LDLR−/− mice fed on high-fat diet
Scientific Reports (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.