Abstract
Human milk contains a diverse array of bioactives and is also a source of bacteria for the developing infant gut. The aim of this study was to characterize the bacterial communities in human milk and infant faeces over the first 3 months of life, in 10 mother-infant pairs. The presence of viable Bifidobacterium and Lactobacillus in human milk was also evaluated. MiSeq sequencing revealed a large diversity of the human milk microbiota, identifying over 207 bacterial genera in milk samples. The phyla Proteobacteria and Firmicutes and the genera Pseudomonas, Staphylococcus and Streptococcus were the predominant bacterial groups. A core of 12 genera represented 81% of the microbiota relative abundance in milk samples at week 1, 3 and 6, decreasing to 73% at week 12. Genera shared between infant faeces and human milk samples accounted for 70–88% of the total relative abundance in infant faecal samples, supporting the hypothesis of vertical transfer of bacteria from milk to the infant gut. In addition, identical strains of Bifidobacterium breve and Lactobacillus plantarum were isolated from the milk and faeces of one mother-infant pair. Vertical transfer of bacteria via breastfeeding may contribute to the initial establishment of the microbiota in the developing infant intestine.
Similar content being viewed by others
Introduction
Early microbial colonisation of the neonatal gut exerts a major effect on host health status1,2. Early colonization is influenced by a number of factors including feeding practice, with the intestinal microbiome composition differing in breast-fed and formula-fed infants3. Breast-fed infants have been reported to have a more stable bacterial population, while the microbiota of formula-fed infants appears to be more diverse, containing higher numbers of E. coli, Clostridium difficile, Enterococcus, Enterobacter and Citrobacter4,5,6. Formula-fed infants have been reported to have higher morbidity and mortality during the first year of life and have an increased risk of developing respiratory and gastrointestinal infections and diseases such as necrotising enterocolitis and sepsis, compared with breast-fed infants7,8.
The health promoting effects of human milk have been linked to the abundance of bioactive molecules present therein, including secretory antibodies, immune cells, antimicrobial proteins such as lactoferrin, CD14 and lysozyme, regulatory cytokines and human milk oligosaccharides (HMOs)9. Though indigestible by the infant, these HMOs exert immunomodulatory effects, stimulate the growth of commensal bacteria, particularly bifidobacteria and are involved in defence against pathogens by preventing adhesion to the gut epithelium during a vulnerable period in life, when the infant’s own defences are immature10.
It has been reported that human milk is in itself a source of commensal bacteria that may colonise the infant gut. Predominant culturable bacterial populations have been identified as Staphylococcus and Streptococcus species, while lactic acid bacteria and bifidobacteria have also been isolated11,12,13. In an earlier culture-independent study, Staphylococcus and Streptococcus dominated the milk microbiota of most mothers, while a more recent study revealed a larger microbial diversity of human milk than originally thought, with more than 700 species identified14,15. Interestingly, the study also reported that the human milk microbiota was compositionally distinct from other body niches and maternal weight and mode of delivery were reported as determinants of composition.
The origin of the human milk microbiome has been the subject of debate. Possible origins include bacteria from maternal skin and the oral cavity of the infant, where it has been demonstrated that during suckling, a high degree of retrograde flow back into the mammary ducts occurs16. More recently, the maternal gut has been suggested as a source, with bacteria entering the mammary glands via the entero-mammary pathway, a route that involves phagocytic dendritic cells penetrating the gut epithelium and trafficking bacteria through the circulatory system17,18,19. This is important since manipulation of the maternal microbiota could be used to promote an optimal human milk microbiome.
Given the significant health effects that breast-feeding exerts on infant health and development, it is important to define the composition of the human milk microbiota. Accordingly, the aim of this study was to characterise the microbiota composition of human milk in ten lactating women, and correlate the findings with gut microbiota composition of their infants, over the first three months of life. Additionally, we assessed the presence of culturable Bifidobacterium and Lactobacillus strains and their recovery from the infant gut.
Results
MiSeq Sequencing of Human Milk and Infant Faeces
MiSeq sequencing of human milk and infant faecal samples yielded a total of 20,206,055 reads, with mean read lengths of 460 bp (330–591 bp). Following quality control, an average of 61,786 (range 34836–109067) and 189,861 (range 98772–327481) reads per sample were obtained for human milk and infant faeces, respectively. Reads were classified into 1313 observed operational taxonomic units (OTUs) for human milk and 264 OTUs for infant faeces at a 3% similarity cut-off. All samples were rarefied to 33,000 sequences to prevent bias due to sampling depth.
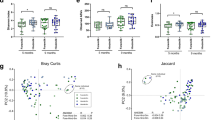
To estimate microbial richness we used the Chao1 richness estimator which revealed significantly higher bacterial richness in milk samples than faecal samples at week 1, 3, 6 and 12 (p < 0.0001, p < 0.0001, p = 0.0002 and p = 0.0003 respectively) (Fig. 1). We applied the Simpson’s index to estimate microbial evenness and this was significantly higher in milk samples at week 1 and 6 (p = 0.002 and p = 0.004). To predict microbial diversity we used the Shannon index, which combines both bacterial richness and evenness, and is more responsive to rare species in terms of species richness than the Simpson’s index which is more sensitive to dominant species. The Shannon index was significantly higher in milk samples at week 1, 3, 6 and 12 (p = 0.0012, p = 0.005, p < 0.0001 and p = 0.004). Over time, there was a significant increase in bacterial richness, Chao1, from week 3 to 6 (p = 0.0462) and a significant decrease in richness from week 6 to 12 (p = 0.0462) in milk samples. The Shannon index demonstrated a significant decrease in diversity in milk samples from week 6 to 12 (p = 0.0482). There were no statistically significant differences in alpha diversity metrics in fecal samples over time.
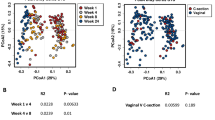
Regarding phyla, at week 1, human milk was dominated by Proteobacteria (41%), Firmicutes (35%) and Bacteroidetes (17%), accounting for 93% of all reads (Fig. 2). Conversely, infant faeces harboured higher relative abundances of Firmicutes (56%) and Actinobacteria (20%) and lower Proteobacteria (21%) and Bacteroidetes (3%). Proteobacteria dominated in milk samples, with the exception of week 3 where Firmicutes showed the highest mean relative abundance (50%).
At genus level, 12 genera appeared to predominate in human milk, as they were detected at a mean relative abundance of ≥1% in at least 90% of samples collected over time (Fig. 3). These genera dominated the community, representing 81% of the relative abundance in human milk samples at week 1 (range 64–91%), 3 (range 64–90%) and 6 (range 61–90%) and 73% of the relative abundance at week 12 (range 50–82%). This core comprised Pseudomonas, Staphylococcus, Streptococcus, Elizabethkingia, Variovorax, Bifidobacterium, Flavobacterium, Lactobacillus, Stenotrophomonas, Brevundimonas, Chryseobacterium and Enterobacter. The remaining sequences mapped to 195 genera, further demonstrating the diversity of the human milk microbiota (Supplementary Table S1). The relative abundances of these genera were individual specific and subject to intra-individual variations over time.
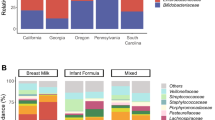
There was little temporal stability in milk samples from most lactating women, as the relative abundance of the bacterial genera present shifted over time (Supplementary Table S1). With the exception of week 3 where Streptococcus had the highest mean relative abundance (32%), Pseudomonas was predominant at all other sampling points detected at 21%, 27% and 19% at week 1, 6 and 12 respectively. By contrast, only 50 genera were detected in infant faecal samples. Pseudomonas was not detected by sequencing and the samples were dominated by Staphylococcus at week 1 (19%), Escherichia-Shigella at week 3 and 12 (17% and 21% respectively) and Veillonella at week 6 (23%) (Fig. 4). When considering genera detected in both human milk and faeces (Table 1), these accounted for 88%, 85%, 88% and 70% of the total reads in infant faecal samples at week 1, 3, 6 and 12 respectively. For human milk, these genera accounted for 37%, 51%, 19% and 27% of the total reads. These genera occurred with varying relative abundances and frequencies. Veillonella and Escherichia/Shigella for example, occurred at a higher mean relative abundance in infant faeces, 14% and 12% respectively, compared to 1% and 0.04% respectively in human milk at week 1. In contrast, Staphylococcus was consistently detected at a high frequency in all samples with a mean relative abundance of 13% and 19% at week 1 in human milk and infant faeces, respectively.
A number of niche-specific genera were exclusive to either human milk or infant faeces. For example, the high relative abundance of Proteobacteria and Bacteroidetes in human milk was largely attributable to the genera Pseudomonas and Variovorax, and Elizabethkingia and Flavobacterium respectively, which were exclusive to human milk. In infant faeces, Eggerthella, which was the only genus not detected in human milk samples, contributed to the higher relative abundance of Actinobacteria.
One subject, M10, reported symptoms of subacute mastitis and withdrew from the study at week 6 following antibiotic administration. Among the milk samples obtained from this subject, the relative abundance of Staphylococcus was higher than the mean relative abundance for healthy subjects, accounting for 73% and 24% of the reads obtained at week 1 and week 3, respectively, compared to 12% and 6% in healthy women. Alpha diversity metrics were also lowest in the mastitic milk, this likely reflects the low abundance of other species in this sample. Also of note was mother-infant pair 9, where Haemophilus was detected at a relative abundance of 24% in the human milk sample and 33% in the infant fecal microbiota at week 1. The mean relative abundance for Haemophilus in other human milk samples at week 1 was 0.1% and 0.5% in infant faeces.
Principal coordinate analysis (PCoA) plots using weighted UniFrac distances demonstrated a clear separation of milk and faecal samples at each week (Supplementary Figure 1). There were no distinct clusters of milk samples over time, although bacterial communities clustered more closely to one another at week 12 than at week 1 (Supplementary Figure 2a). Similarly, in faecal samples no obvious separation was observed between samples at different weeks (Supplementary Figure 2b).
Culture-Dependent Analysis of Human Milk and Infant Faeces
Culture on selective media revealed the presence of presumptive Bifidobacterium and Lactobacillus in the human milk of one mother at 1 × 102 CFU/ml and 3 × 103 CFU/ml respectively at week 3. Bifidobacterium and Lactobacillus were isolated from the corresponding infant fecal sample at 4 × 106 CFU/ml and 7 × 107 CFU/ml, respectively. No culturable Bifidobacterium or Lactobacillus were detected in human milk samples from other mothers in the study. Following enumeration, bacterial isolates from the human milk and the corresponding infant stool sample were subjected to 16S rRNA gene sequencing using species-specific primers. This identified the Bifidobacterium isolates as B. breve and Lactobacillus isolates as L. plantarum in both the human milk and the infants fecal samples. In addition, PFGE analysis of these isolates revealed identical profiles in both the human milk and the infant fecal samples for both B. breve and L. plantarum, which indicated that these isolates belonged to the same strain (Fig. 5).
Pulse-field gel electrophoresis patterns of (a) XbaI-digested genomic DNA of B. breve isolates and (b) ApaI-digested genomic DNA of L. plantarum isolates from human milk and infant faeces. The unedited versions of these images can be found as Supplementary Figures S3 and S4.
Discussion
The benefits of human milk in terms of infant health and development have been well documented1,8,10,12,17,20. Human milk has been recognized as a fundamental source of bioactive components including bacteria that may contribute to neonatal gastrointestinal colonisation and immune development and maturation during the crucial early stages of development8,9,10. Differences have been reported in the microbiota composition of breast-fed infants versus formula-fed with the former suffering from less allergies and gastrointestinal infections3,4. Therefore, the microbiota of breast-fed infants is considered the gold standard in terms of a healthy infant gastrointestinal microbiota. Comprehensively characterising the human milk microbiota is vital for enabling better insight of its significance and activity in relation to the developing infant gut microbiota and health.
In this study, Illumina MiSeq sequencing revealed that 12 genera; Pseudomonas, Staphylococcus, Streptococcus, Elizabethkingia, Variovorax, Bifidobacterium, Flavobacterium, Lactobacillus, Stenotrophomonas, Brevundimonas, Chryseobacterium and Enterobacter dominated the milk of most lactating women, constituting a core milk microbiota. The presence and relative abundances of the remaining 195 genera were unique to individual mothers and subject to variation over time. The core microbiota constituted 81% of the taxa identified at week 1, 3 and 6, however by week 12, this decreased to 73%, suggesting a selected core microbiota drives the early stages but individual-specific taxa become more important in later stages of lactation.
Pseudomonas has been reported as a dominant member of the human milk microbiota in several studies including Ward et al., where it accounted for 61% of the relative abundance from milk samples taken between 9 and 30 days postpartum14,21,22. Similarly Staphylococcus has been found to be a common constituent of the milk microbiota by both culture independent and dependent investigations11,14,15,21,22,23. In a study of breast-fed Swedish infants, both vaginally and caesarean section delivered, 100% of infant feces were colonised by Staphylococcus from day 3 of life. Staphylococcus epidermis in particular appears to have a biological relevance as it has been shown to be the predominant species in human milk and in the faeces of breast-fed infants and is less common in stool of formula-fed infants11,24,25. Genera associated with the oral cavity such as Streptococcus and Veillonella’ reported by Hunt et al. and Cabrera-Rubio et al. were also prevalent in this study14,15. Although not previously reported in the human milk microbiome, Variovorax, strains of which have been isolated from the human oral cavity, were consistently detected in all samples26.
The detection of large proportions of typical inhabitants of the skin and oral microbiota may imply that the origin in this case is secondary contamination. However, anerobic gut-associated populations such as Bacteroides, Blautia, Faecalibacterium, Ruminococcus, Roseburia, Subdoligranulum, Enterococcus and Escherichia-Shigella were also detected here and in other studies15,21. Bifidobacterium was also consistently detected, supporting the findings of Jost et al., and Hunt et al., however Cabrera-Rubio et al. and Ward et al. did not report the presence of Bifidobacterium. These differences are likely due to the method of DNA extraction used, as a bead-beating step was not incorporated in the latter studies. When making comparisons across studies, it is important to note that differences in microbial community composition may also have been affected by factors such as the hypervariable region of the 16S gene examined (we targeted the V3–V4 hypervariable regions of 16S rDNA), geographical differences and the greater sequencing depth achieved using Illumina MiSeq sequencing here compared with Roche 454 pyrosequencing used in other studies. These factors are known to influence diversity and richness estimates and can greatly impact the microbiome of individuals.
When considering the infant fecal microbiota, it was found to be less diverse than that of human milk. The communities were most similar with respect to Staphylococcus which accounted for a mean relative abundance of 15% and 19% at week 1 in human milk and infant faeces, respectively. A number of typically gut-associated genera were common to both human milk and infant faeces including Bifidobacterium, Bacteroides, Enterococcus, Lactobacillus, Clostridium, Coprococcus, Escherichia-Shigella and members of the Lachnospiraceae family. Interestingly, these shared genera accounted for 70–88% of the total reads in infant faecal samples throughout the sampling period. This is in agreement with other studies in mother-infant pairs which have shown that the bacterial composition of the faecal microbiota of the breast-fed infants reflects that found in the breast milk12,24,27,28,29,30. In these studies, the genera Lactobacillus, Staphylococcus, Enterococcus and Bifidobacterium were frequently shared between breast milk and infant faeces. Jimenez et al., in a study with 23 mother-infant pairs identified Staphylococcus as the predominant species in milk and breast-fed infants faeces25. Gronlund et al., reported that maternal breast-milk bifidobacterial counts impacted on the infants’ faecal Bifidobacterium levels27. While Jost et al., identified a number of gut-associated anaerobic genera like Bifidobacterium, Bacteroides and members of the class Clostridia shared between milk and infant faeces30.
Stronger evidence of vertical transfer from mother to infant involves identification of identical strains. Lactobacillus and Bifidobacterium are often considered as members of a healthy microbiota with the predominance of Bifidobacterium in particular appearing to be characteristic of the healthy breast-fed infant and were therefore targeted for strain identification. Despite being detected by MiSeq sequencing, culturable Lactobacillus and Bifidobacterium were found in only one milk sample with a low bacterial count of 3 × 103 CFU/ml and 1 × 102 CFU/ml, respectively. Similarly, Albesharat et al., were unable to isolate any bifidobacteria from human milk samples29. It is unclear whether this was due to their low abundance in human milk, the presence of antimicrobial compounds, the length of storage time of samples prior to culturing and in the case of bifidobacteria, their anaerobicity i.e. viable but not culturable. As MiSeq sequencing does not distinguish between live and dead cells, it is also possible that dead cells are being transferred during feeding which would nonetheless elicit an immune response. Of interest in this case was that these culturable isolates belonged to the same species as those isolated from the corresponding infant fecal samples, namely L. plantarum and B. breve. PFGE analysis confirmed that the same bacterial strain was shared in this mother-infant pair supporting the notion of vertical transfer via human milk. Other studies have also isolated the same Bifidobacterium and Lactobacillus strains from human milk and infant faeces including strains of B. breve and L. plantarum13,28.
A limitation of this study is the relatively small sample size and further investigations in larger populations, including maternal faecal samples are planned to confirm these results and extend the knowledge about the milk microbiome. In summary, our data demonstrate the large diversity of the human milk microbiota with over 207 bacterial genera identified. The relative abundances of these were unique to each mother and subject to variation over time. Coupled with our finding that the same strains of Bifidobacterium and Lactobacillus were found in maternal human milk and corresponding infant fecal samples, these results suggest that there is a microbiota specific for each mother-infant pair that could confer benefits specific to that infant. This is particularly pertinent as commercially available infant formulas and donor milk are sterilized/pasteurised and as such contain little to no microbes. The results of this work also confirmed the presence of microbes typically associated with the gut microbiota in milk samples suggesting strategies to manipulate the maternal gut with bacteria which could confer benefits to the infant may be beneficial. These data emphasise that human milk constitutes a relevant source of a wide range of bacteria for the infant gut and can contribute to infant gut colonisation and therefore to infant health.
Methods
Subjects, Study Design and Sample Collection
This study was approved by the Clinical Research Ethics Committee of the Cork Teaching Hospitals. Parents of infant participants provided written informed consent and all relevant guidelines and regulations were followed. Ten mother-infant pairs were recruited at Cork University Maternity Hospital. Recruits were healthy lactating women and their full-term, healthy breast-fed infants (breast-fed for a minimum of 4 weeks after birth). The clinical characteristics of mother-infant pairs are shown in Table 2. Milk and faecal samples were collected from each mother-infant pair at 1, 3, 6 and 12 weeks. For milk sampling, sterile gloves were worn and the nipple and areola of the human were cleaned with chlorhexidane wipes (Clinell, United Kingdom) prior to manual expression of human milk into a sterile tube. The first few drops (approximately 1 ml) were discarded to prevent chlorhexidine contamination. Faecal samples were also collected into sterile containers and stored at 4 °C until delivery to the laboratory. Aliquots of 1 ml of fresh milk and 1 g faecal sample were subject to culture within 24 hours of donation. Remaining aliquots were immediately frozen at −20 °C for subsequent DNA extraction.
DNA Extraction and MiSeq Sequencing
DNA was purified from stool samples using the QIAmp DNA Stool Mini Kit (Qiagen, UK) according to manufacturer’s instructions with the addition of a bead-beating step (30 s × 3). For milk samples, 1 ml was centrifuged at 5,000 g for 30 minutes. The supernatant was removed and any residue removed with a sterile cotton swab. The pellet was washed twice with sterile PBS and incubated at 55 °C for 15 minutes with 50 mg/ml lysozyme and 5 KU/ml mutanolysin. The remaining steps were performed using the QIAmp DNA Stool Mini Kit according to manufacturer’s instructions with addition of a bead-beating step (30 s × 3). The microbiota composition of the samples was established by amplicon sequencing of a ~380 bp fragment of the V3-V4 hypervariable region of the bacterial 16S rRNA gene following the Illumina 16S Metagenomic Sequencing Library Preperation guide. PCR amplification of V3–V4 region was performed using the forward primer 5′TCGTCGGCAGCGTCAGATGTGTATAAGAGACAGCCTACGGGNGGCWGCAG and reverse primer 5′GTCTCGTGGGCTCGGAGATGTGTATAAGAGACAGGACTACHVGGGTATCTAATCC. Each 25 μl PCR reaction contained 5 ng/μl microbial genomic DNA, 1 μM of each primer and 12.5 μl 2× KAPA HiFi HotStart ReadyMix. The PCR conditions for faecal DNA were as follows: initial denaturation for 5 min at 95 °C; 30 cycles of 15 s at 95 °C, 15 s at 42 °C and 30 s at 72 °C; and 72 °C for 5 min for final extension. The PCR conditions for milk DNA were: initial denaturation for 5 min at 95 °C; 30 cycles of 15 s at 95 °C, 15 s at 57 °C and 30 s at 72 °C; and 72 °C for 5 min for final extension. The Agencourt AMPure XP system (Beckman Coulter, UK) was used to purify the amplicons. A subsequent limited‐cycle amplification step was performed to add multiplexing indices and Illumina sequencing adapters. Amplicons were quantified, normalised and pooled using the Qubit® dsDNA HS Assay Kit (Life Technologies). Library preparation was carried out by GATC Biotech prior to 2 × 300 base-pair (bp) sequencing on the Illumina MiSeq platform.
Sequence and Statistical Analysis
300 bp paired-end reads were assembled using FLASH with parameters of a minimum overlap of 20 bp and a maximum overlap of 120 bp31. The QIIME suite of tools, v1.8.0, was used for further processing of paired-end reads, including quality filtering based on a quality score of >25 and removal of mismatched barcodes and sequences below length thresholds32. Denoising, chimera detection and operational taxonomic unit (OTU) grouping were performed in QIIME using USEARCH v733. Taxonomic ranks were assigned by alignment of OTUs using PyNAST to the SILVA SSURef database release 11134,35. Alpha and beta diversities were generated in QIIME and calculated based on weighted and unweighted Unifrac distance matrices36. Principal coordinate analysis (PCoA) plots were visualised using EMPeror v0.9.3-dev37. To determine any statistically significant differences in microbial diversity between milk and faecal samples, non-parametric Mann-Whitney analysis was completed using Minitab 15 statistical software package. Statistical significance was accepted as p < 0.05, adjusted for ties.
Isolation and Enumeration of Bifidobacterium and Lactobacillus spp
Aliquots of faecal sample, 1 gram, were mixed with 9 ml maximum recovery diluent (Oxoid) to make an initial 10−1 dilution. Serial dilutions were spread-plated onto de Man, Rogosa, Sharpe (MRS) (Difco) agar supplemented with 0.05% L-cysteine hydrochloride (Sigma), 100 μg ml−1 mupirocin (Oxoid) and 50 Units nystatin (Sigma Aldrich) for Bifidobacterium spp. and MRS agar (Difco) supplemented with 50 U ml−1 nystatin for Lactobacillus spp. 1 ml of milk was serially diluted and spread-plated as described above. Agar plates were incubated anaerobically at 37 °C for 72 hours for Bifidobacterium spp. and 5 days for Lactobacillus spp. Bacterial counts were recorded as colony forming units per gram of faeces or per ml of human-milk.
Species Determination by Sanger Sequencing
Bacterial isolates from the Bifidobacterium-selective and Lactobacillus-selective media were grown overnight in MRS broth and genomic DNA extracted using the GenElute™ Bacterial Genomic DNA kit (Sigma) following manufacturer’s instructions. The identity of each putative Bifidobacterium isolate was confirmed by generating a 1.5 kb 16s rRNA gene-internally transcribed spacer (ITS) fragment using the oligonucleotides BIF-specific (5′-GGTGTGAAAGTCCATCGCT-3′) and 23S_bif (5′-GTCTGCCAAGGCATCCACCA-3′)38. Each 50 μl PCR reaction contained 45 μl of Platinum PCR SuperMix High Fidelity (Invitrogen), 1 μl BIF-specific primer (0.15 μM), 1 μl 23S_bif primer (0.15 μM), template DNA and sterile PCR grade water. PCRs were completed under the following conditions: initial denaturation step of 3 min at 94 °C, followed by 35 cycles of 30 s at 94 °C, 30 s at 56.5 °C and 1 min at 72 °C, followed by a single elongation step of 10 min at 72 °C. The resulting amplicons were separated on 1% (w/v) agarose gels and purified using the QIAquick PCR purification kit (Qiagen, United Kingdom). DNA sequencing of both strands was performed by Beckman Coulter (Essex, UK) using the oligonucleotides bif-sec (5′-CATGCCCCTACGTCCAG-3′) and 23S-bif (5′-CAAGGCATCCACCATACGC-3′). Strains were assigned to a particular species following comparison of the 16s rRNA-ITS sequences using NCBI BLAST database (http://www.ncbi.nlm.nih.gov/BLAST/).
Genetic Typing by Pulsed-Field Gel Electrophoresis (PFGE)
High-molecular-weight DNA fragments were isolated from stationary phase cultures by a previously described method39. The restriction enzyme XbaI was used to cleave bifidobacterial chromosomal DNA and ApaI (New England Biolabs, MA, United States) was used for Lactobacillus. The fragments were separated using a contour-clamped homogeneous electric field CHEF-DR III pulsed field system (Bio-Rad Laboratories) with a linear ramp pulse time of 1 to 15 s and 2 to 20 s for bifidobacteria and lactobacilli, respectively. The run time was 18 h at 6 V/cm in a running buffer of 0.5X Tris base-borate-EDTA maintained at a temperature of 12 °C.
Additional Information
How to cite this article: Murphy, K. et al. The Composition of Human Milk and Infant Faecal Microbiota Over the First Three Months of Life: A Pilot Study. Sci. Rep. 7, 40597; doi: 10.1038/srep40597 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Penders, J. et al. Factors influencing the composition of the intestinal microbiota in early infancy. Pediatrics 118, 511–521, doi: 118/2/511 (2006).
Dominguez-Bello, M. G. et al. Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats in newborns. P Natl Acad Sci USA 107, 11971–11975 (2010).
Bezirtzoglou, E., Tsiotsias, A. & Welling, G. W. Microbiota profile in feces of breast- and formula-fed newborns by using fluorescence in situ hybridization (FISH). Anaerobe 17, 478–482, doi: 10.1016/j.anaerobe.2011.03.009 (2011).
Penders, J. et al. Quantification of Bifidobacterium spp., Escherichia coli and Clostridium difficile in faecal samples of breast-fed and formula-fed infants by real-time PCR. Fems Microbiol Lett 243, 141–147, doi: 10.1016/j.femsle.2004.11.052 (2005).
Wang, M. et al. Fecal microbiota composition of breast-fed infants is correlated with human milk oligosaccharides consumed. J Pediatr Gastroenterol Nutr 60, 825–833, doi: 10.1097/MPG.0000000000000752 (2015).
Backhed, F. et al. Dynamics and Stabilization of the Human Gut Microbiome during the First Year of Life. Cell host & microbe 17, 690–703, doi: 10.1016/j.chom.2015.04.004 (2015).
Kramer, M. S. et al. Infant growth and health outcomes associated with 3 compared with 6 mo of exclusive breastfeeding. Am J Clin Nutr 78, 291–295 (2003).
Ladomenou, F., Moschandreas, J., Kafatos, A., Tselentis, Y. & Galanakis, E. Protective effect of exclusive breastfeeding against infections during infancy: a prospective study. Arch Dis Child 95, 1004–1008, doi: 10.1136/adc.2009.169912 (2010).
Newburg, D. S. Innate immunity and human milk. The Journal of nutrition 135, 1308–1312 (2005).
Bode, L. Human milk oligosaccharides: every baby needs a sugar mama. Glycobiology 22, 1147–1162, doi: 10.1093/glycob/cws074 (2012).
Heikkila, M. P. & Saris, P. E. J. Inhibition of Staphylococcus aureus by the commensal bacteria of human milk. J Appl Microbiol 95, 471–478 (2003).
Martin, R. et al. Human milk is a source of lactic acid bacteria for the infant gut. J Pediatr-Us 143, 754–758 (2003).
Martin, R. et al. Lactobacillus salivarius CECT 5713, a potential probiotic strain isolated from infant feces and breast milk of a mother-child pair. Int J Food Microbiol 112, 35–43 (2006).
Hunt, K. M. et al. Characterization of the Diversity and Temporal Stability of Bacterial Communities in Human Milk. Plos One 6 (2011).
Cabrera-Rubio, R. et al. The human milk microbiome changes over lactation and is shaped by maternal weight and mode of delivery. Am J Clin Nutr 96, 544–551 (2012).
Ramsay, D. T., Kent, J. C., Owens, R. A. & Hartmann, P. E. Ultrasound imaging of milk ejection in the breast of lactating women. Pediatrics 113, 361–367 (2004).
Fernández, L. et al. The human milk microbiota: origin and potential roles in health and disease. Pharmacol Res 69, 1–10, doi: 10.1016/j.phrs.2012.09.001 (2013).
Rescigno, M., Rotta, G., Valzasina, B. & Ricciardi-Castagnoli, P. Dendritic cells shuttle microbes across gut epithelial monolayers. Immunobiology 204, 572–581, doi: 10.1078/0171-2985-00094 (2001).
Perez, P. F. et al. Bacterial imprinting of the neonatal immune system: Lessons from maternal cells? Pediatrics 119, E724–E732 (2007).
Jeurink, P. V. et al. Human milk: a source of more life than we imagine. Benef Microbes 4, 17–30 (2013).
Jost, T., Lacroix, C., Braegger, C. & Chassard, C. Assessment of bacterial diversity in breast milk using culture-dependent and culture-independent approaches. Brit J Nutr 110, 1253–1262 (2013).
Ward, T. L., Hosid, S., Ioshikhes, I. & Altosaar, I. Human milk metagenome: a functional capacity analysis. Bmc Microbiol 13, doi: 10.1186/1471-2180-13-116 (2013).
Martin, R. et al. Cultivation-independent assessment of the bacterial diversity of breast milk among healthy women. Res Microbiol 158, 31–37 (2007).
Martin, V. et al. Sharing of Bacterial Strains Between Breast Milk and Infant Feces. J Hum Lact 28, 36–44 (2012).
Jimenez, E. et al. Staphylococcus epidermidis: A differential trait of the fecal microbiota of breast-fed infants. Bmc Microbiol 8 (2008).
Anesti, V. et al. Isolation and molecular detection of methylotrophic bacteria occurring in the human mouth. Environ Microbiol 7, 1227–1238, doi: 10.1111/j.1462-2920.2005.00805.x (2005).
Gronlund, M. M. et al. Maternal breast-milk and intestinal bifidobacteria guide the compositional development of the Bifidobacterium microbiota in infants at risk of allergic disease. Clin Exp Allergy 37, 1764–1772, doi: 10.1111/j.1365-2222.2007.02849.x (2007).
Solis, G., de los Reyes-Gavilan, C. G., Fernandez, N., Margolles, A. & Gueimonde, M. Establishment and development of lactic acid bacteria and bifidobacteria microbiota in breast-milk and the infant gut. Anaerobe 16, 307–310 (2010).
Albesharat, R., Ehrmann, M. A., Korakli, M., Yazaji, S. & Vogel, R. F. Phenotypic and genotypic analyses of lactic acid bacteria in local fermented food, breast milk and faeces of mothers and their babies. Syst Appl Microbiol 34, 148–155 (2011).
Jost, T., Lacroix, C., Braegger, C. P., Rochat, F. & Chassard, C. Vertical mother-neonate transfer of maternal gut bacteria via breastfeeding. Environ Microbiol 16, 2891–2904 (2014).
Magoc, T. & Salzberg, S. L. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963, doi: 10.1093/bioinformatics/btr507 (2011).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nature methods 7, 335–336, doi: 10.1038/nmeth.f.303 (2010).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461, doi: 10.1093/bioinformatics/btq461 (2010).
Caporaso, J. G. et al. PyNAST: a flexible tool for aligning sequences to a template alignment. Bioinformatics 26, 266–267, doi: 10.1093/bioinformatics/btp636 (2010).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic acids research 41, D590–596, doi: 10.1093/nar/gks1219 (2013).
Lozupone, C., Lladser, M. E., Knights, D., Stombaugh, J. & Knight, R. UniFrac: an effective distance metric for microbial community comparison. The ISME journal 5, 169–172, doi: 10.1038/ismej.2010.133 (2011).
Vazquez-Baeza, Y., Pirrung, M., Gonzalez, A. & Knight, R. EMPeror: a tool for visualizing high-throughput microbial community data. GigaScience 2, 16, doi: 10.1186/2047-217X-2-16 (2013).
Turroni, F. et al. Exploring the Diversity of the Bifidobacterial Population in the Human Intestinal Tract. Appl Environ Microb 75, 1534–1545, doi: 10.1128/Aem.02216-08 (2009).
Simpson, P. J., Stanton, C., Fitzgerald, G. F. & Ross, R. P. Genomic diversity and relatedness of bifidobacteria isolated from a porcine cecum. J Bacteriol 185, 2571–2581 (2003).
Acknowledgements
The authors thank the mothers and infants involved in this study for their co-operation and assistance. This work was supported by the Irish Department of Agriculture, Food and Marine funded INFANTMET project, grant number 10/RDT/MFRC/705, and the Science Foundation of Ireland funded APC Microbiome Institute under grant number SFI/12/RC/2273.
Author information
Authors and Affiliations
Contributions
C.S., C.A.R., R.P.R., E.M.D. and P.W.O.T. designed research; K.M., D.C., T.O.C. and C.O.S. conducted research; K.M. analysed data and wrote the paper; C.S. had primary responsibility for final content. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Murphy, K., Curley, D., O’Callaghan, T. et al. The Composition of Human Milk and Infant Faecal Microbiota Over the First Three Months of Life: A Pilot Study. Sci Rep 7, 40597 (2017). https://doi.org/10.1038/srep40597
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep40597
This article is cited by
-
Lactic acid bacteria isolated from women’ breast milk and infants’ faeces have appreciable immunogenic and probiotic potentials against diarrheagenic E. coli strains
BMC Microbiology (2024)
-
Diet at birth is critical for healthy growth, independent of effects on the gut microbiota
Microbiome (2024)
-
Diet–gut microbiome interaction and ferulic acid bioavailability: implications on neurodegenerative disorders
European Journal of Nutrition (2024)
-
Age-specific composition of milk microbiota in Tibetan sheep and goats
Applied Microbiology and Biotechnology (2024)
-
Parental care contributes to vertical transmission of microbes in a skin-feeding and direct-developing caecilian
Animal Microbiome (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.