Abstract
Most inflammatory bowel diseases (IBDs) are classic complex disorders represented by common alleles. Here we aimed to define the genetic architecture of pediatric and adult-onset IBDs for the Polish population. A total of 1495 patients were recruited, including 761 patients with Crohn’s disease (CD; 424 pediatric), 734 patients with ulcerative colitis (UC; 390 pediatric), and 934 healthy controls. Allelotyping employed a pooled-DNA genome-wide association study (GWAS) and was validated by individual genotyping. Whole exome sequencing (WES) was performed on 44 IBD patients diagnosed before 6 years of age, 45 patients diagnosed after 40 years of age, and 18 healthy controls. Altogether, out of 88 selected SNPs, 31 SNPs were replicated for association with IBD. A novel BRD2 (rs1049526) association reached significance of P = 5.2 × 10−11 and odds ratio (OR) = 2.43. Twenty SNPs were shared between pediatric and adult patients; 1 and 7 were unique to adult-onset and pediatric-onset IBD, respectively. WES identified numerous rare and potentially deleterious variants in IBD-associated or innate immunity-associated genes. Deleterious alleles in both groups were over-represented among rare variants in affected children. Our GWAS revealed differences in the polygenic architecture of pediatric- and adult-onset IBD. A significant accumulation of rare and deleterious variants in affected children suggests a contribution by yet unexplained genetic components.
Similar content being viewed by others
Introduction
Inflammatory bowel diseases (IBDs) are chronic disorders with disease onset ranging from early childhood to beyond the sixth decade of life. Childhood-onset IBD represents 10–25% of all IBD cases1,2,3,4. Most IBDs, including Crohn’s disease (CD) and ulcerative colitis (UC), are classic complex disorders5 underlined by genetic variance which may affect the patient’s defense and adaptive mechanisms to environmental factors. Their genetic load is represented by common alleles that, in response to intestinal microbiota, underlie multiple intestinal immunopathological processes3,4. However, a minority of patients with adult-onset IBD and an unknown fraction of pediatric patients, especially those with very early-onset (VEO) disease (<6 years old), may represent monogenic disorders with an IBD-like presentation. These monogenic defects have been found to also alter intestinal immune homeostasis, with defective processing of intracellular bacteria, autophagy, and innate immunity in CD and disruption of the epithelial barrier along with the epithelial response in UC2,6.
Genome-wide association studies (GWASs) provide information on common variants associated with disease susceptibility. The largest genetic association study for IBD included >75,000 patients and controls which uncovered 163 susceptibility loci; 110 of them were shared between CD and UC, whereas 30 and 23 were unique to CD and UC, respectively. Each locus contained an average of five genes7. All but a few loci (e.g., NOD2 and IL23R) exhibit a rather tiny size effect (odds ratio (OR) <1.3)8 and contribute individually to only a small proportion of the expected heritability in IBDs. Multiple known alleles in nearly 200 loci associated with IBD explain only 13.6% and 7.5% of the overall disease variance of CD and UC risk, respectively7.
Pediatric IBDs are typically characterized by a more extensive disease course, a change in disease location over time, and a more frequent positive family history of IBD. In contrast, patients diagnosed between the ages of 20 and 30 years have a relatively less variable phenotype, and those diagnosed after the age of 60 years often have a mild disease severity1,9,10,11. Although the IBD location, progression, and response to therapy depend on the age of onset, multiple determinants of the early age of IBD onset remain largely unknown.
The previously reported pediatric GWASs identified loci that mostly replicated adult-onset IBD studies12. Although underpowered, GWASs carried out exclusively in pediatric patients uncovered 23 of 32 loci previously found in adult-onset CD and 8 of 17 loci previously found in adult-onset UC. Another report14 based on SNPs associated with pediatric-onset IBD suggested a role of NOD2, TNFSF15, POU5F1, and HLA-DRB1*501 in pediatric-onset CD and LAMB1 in pediatric-onset UC. However, few pediatric-onset disease-associated loci have been described, including 20q13, 21q22, and 16p1115,16. As in GWASs of adult-onset IBD, most loci ascertained by pediatric GWASs have a small effect size. Nevertheless, genetic load has been speculated to contribute to IBD etiology and that the phenotype across ages is greater in pediatric-onset than in adult-onset IBD2,17.
To better define risk variants across pediatric- and adult-onset IBDs and identify additional susceptibility loci in the Polish population, the genetic architecture of IBDs was analyzed simultaneously in pediatric and adult cohorts using a pooled-DNA sample-based GWAS to screen for IBD associations. The GWAS findings were further validated using individual DNA samples from enlarged patient cohorts and TaqMan SNP Genotyping Assays. In addition, we searched for rare genetic variants in select sub-groups of patients with VEO and adult-onset IBD using whole exome sequencing (WES).
Materials and Methods
Ethics statement
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. The study was approved by the Ethics Committee of the Medical Center for Postgraduate Education, Warsaw, Poland. Informed consent was obtained from all individual participants included in the study.
Subjects
Between 2010 and 2014, a total of 1495 patients without an IBD family history were recruited at 17 Gastroenterology Departments at different Polish hospitals; 761 were diagnosed with CD and 734 were diagnosed with UC. IBDs were diagnosed in children and adolescents according to the Porto criteria modified in accordance with the recommendations of the European Crohn’s and Colitis Organization (ECCO), and in adults according to ECCO guidelines.
The GWAS cohort comprised 594 patients diagnosed with CD (356 before 17 years of age), 571 patients diagnosed with UC (311 before 17 years of age), and 724 healthy controls. Larger cohorts of cases and controls were enrolled in a replication study, including 761 patients diagnosed with CD (424 before 17 years of age), 734 patients diagnosed with UC (390 before 17 years of age), and 934 healthy controls. Sample sizes and the age distribution of each group are shown in Supplementary Table 1.
See the Supplementary Materials and Methods for detailed description of GWAS, individual genotyping and WES, analysis of mutation, and statistical analysis.
Results
For the association screening, we conducted a cost-effective GWAS. DNA samples from individual patients that passed quality control were equimolarly combined according to patient diagnosis, sex, and age at disease onset to obtain 49 and 30 DNA pools representing 1165 IBD patients and 724 healthy controls, respectively. Of these DNA pools, 9 and 16 represented adult-onset UC (diagnosed ≥17 years of age) and pediatric-onset UC (diagnosed <17 years of age), respectively, and 10 and 14 represented adult- and pediatric-onset CD, respectively. The susceptibility loci for IBD in the Polish population were selected by comparing genome-wide genotypes between healthy controls and patients with IBD (both UC and CD), with UC only, and with CD only. A total of nine separate comparisons were performed for the following groups: adult and pediatric-onset, adult-onset only, and pediatric-onset only. Selected SNPs identified by the GWAS were validated using individual TaqMan SNP genotyping assays for the enlarged cohorts of 761 CD patients, 734 UC patients, and 934 healthy controls.
Association screening and validation assay
A method of selecting loci for validation is crucial for the success of GWASs. In this study, we selected SNPs using three distinct criteria. First, we automatically uncovered 24 SNPs from 20 distinct loci associated at P < 5 × 10−8 (a standard genome-wide significance threshold) in at least one comparison (Supplementary Table 2). Although about half of SNPs selected for chip construction have a MAF >5% in Caucasians, among the 24 SNPs with the highest level of association, 6 were low-frequency variants (MAF = 0.5–5%) and 9 were rare variants (MAF <0.5%). Consequently, out of 20 SNPs subjected to validation (four were not validated for technical reasons), a minor allele was detected for only 7. In line with this observation, among 124 SNPs associated at a P-value between 5 × 10−8 and 10−6, 9 and 94 SNPs were low-frequency and rare variants, respectively, according to the Illumina SNP database18 (Supplementary Table 2). Because rare and low-frequency variants require a very large cohort to achieve a significant association, most of those SNPs were likely false-positive. In the 7 validated findings, 6 SNPs reached the level of significance after multiple test correction, and one SNP was at the nominal level of significance (P < 0.05) (Supplementary Tables 3 and 4). Of these. 5 SNPs were located in close proximity to several other alleles associated in the GWAS with a disease at the same or lower level of significance.
Assuming that an “index SNP” at a given locus is usually not independent of neighboring SNPs, we next focused on loci forming blocks of at least 10 SNPs that remained in strong allele linkage disequilibrium; the distance between two SNPs in the block was less than 30 kb and each SNP associated with a disease was at P < 0.005. Among 72 selected blocks, including 5 loci from the first selection, 55 consisted of at least 10 SNPs associating at P < 0.001 (Supplementary Table 4). Among 67 “index SNPs” from blocks of the second selection, the TaqMan SNP genotyping assays validated association of 62 SNPs at least at the nominal level of significance and OR > 1.2 OR < 0.83 in at least one of nine comparisons, and multiple test correction reduced the number of significant associations to 20 SNPs (Table 1 and Supplementary Table 4). All of the SNPs had the same direction of effect as observed in the GWAS. Altogether, selection of “index SNPs” according to allele linkage disequilibrium reduced the number of false-positive genome-wide associations.
Finally, we selected 14 “index” SNPs from “incomplete” blocks (e.g., composed of a smaller number of SNPs and associating at a P-value between 0.005 and 0.05) because they had associated with IBD in previous reports or represented the potentially interesting regions, such as the major histocompatibility (MHC) region. Genotyping assays validated the association of 5 SNPs at a significant level after multiple test correction and the other 9 SNPs at the nominal level of significance (Supplementary Table 4).
In summary, among 83 SNPs validated for association with the development of CD and/or UC, 31 reached the significance level after multiple test correction, and 52 SNPs were at least at the nominal level of significance. Sixty-five SNPs represented 61 susceptibility loci and 16 represented 7 HLA and 9 non-HLA genes from the MHC region (Supplementary Table 4). For 50 SNPs the minor allele was associated with an increased risk, and for 33 SNPs it had a protective effect. Thirty-four SNPs were shared between CD and UC, whereas 21 and 27 were specific for CD and UC, respectively. All of the shared disease-specific SNPs had the same direction of effect in both types of IBD.
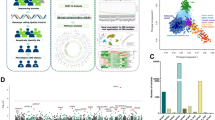
A large number of previously identified loci exhibited a relatively small effect size with an OR < 1.3. In line with this, the genotyping assays revealed a small effect size (1.5 > OR > 1.2 or 0.63 < OR < 0.83) of the association for 53 SNPs, a moderate effect size (2.0 > OR > 1.5 or 0.5 < OR < 0.63) of the association for 22 SNPs and a strong effect size with OR > 2.0 or OR < 0.5 for only eight SNPs. Among the top findings, three SNPs in the NOD2 locus (rs13333062, OR 2.35, P = 3.5 × 10−22, intergenic location; rs6596, OR 3.74, P = 1.7 × 10−54, SNX20 coding location; rs2076756, OR 2.52, P = 8.1 × 10−36, NOD2 intron location) were associated with an increased risk of CD and one SNP in the IL23R locus (rs11209026, OR 0.24, P = 1.9 × 10−6, coding location) was associated with a decreased risk of both UC in adult patients and CD in the two age groups. A new strong association signal found in the MHC region (SNP rs1049526 located in the 3′UTR of BRD2, minimum P = 5.2 × 10−11, OR 2.22–2.70) was associated with an increased risk of IBD, independent of IBD onset (Fig. 1). None of the 15 additional SNPs from this region that were associated with increased or decreased risk reached the effect size of BRD2 rs1049526.
BRD2 rs1049526-centered Manhattan plot of genome-wide associations for Crohn’s disease compared to healthy controls.
The x-axis represents a 2 Mb window with the physical order of the genes. The dashed and dotted horizontal lines indicate the significance threshold of P = 1 × 10−5 and P = 5 × 10−8, respectively.
We then assessed the differences in genetic architecture between pediatric- and adult-onset disease (Table 1). Eleven SNPs were shared between pediatric and adult patients; 5, 5, and 1 were unique to IBD, CD, and UC, respectively. An additional 9 SNPs shared only partly different IBD subtypes between the two patient age groups. Of the remaining associations, one SNPs was unique to the adult-onset population in UC group and 8 SNPs were unique to the pediatric-onset population with 3, 3, and 2 unique to IBD, CD, and UC, respectively. Eleven additional SNPs at the nominal level of significance were shared (completely or partly) between the two patient age groups while 20 and 21 nominally significant SNPs were unique to the adult- or pediatric-onset population, respectively (Supplementary Table 4).
Exome sequencing
GWASs have limited utility in identifying low frequency and rare variants with greater penetrance and true causal genetic variants associated with VEO IBD19. To further search for potential differences in the genetic architecture of VEO and adult-onset IBD, we performed WES analysis for 21 and 22 children diagnosed with CD and UC, respectively, at less than 6 years of age (age range 1–5; median −3) and 23 and 22 patients diagnosed with CD and UC, respectively, after 40 years of age (age range 41–60; median −48.5). In addition, WES was conducted in 18 healthy individuals. Before filtering, sequencing resulted in 150,314 variants discovered: 131,884 SNVs and 18,722 Indels. Each sample had an average 447 Indels and 17,790 SNVs, 640 of which were private variants (sequencing details in Supplementary Table 5).
We identified a total of 2615 rare, homozygous and heterozygous variants (SNPs and Indels), defined by a MAF <2% in the 1 kGP, European-American MAF in the NHLBI Exome Sequencing Project and ExaC, or as novel variants in coding regions of genes selected from an extended list of genes associated with CD or UC20,21. Among these, 1255 variants were categorized as deleterious (Supplementary Table 6) according to the criteria described in the Supplementary Materials and Methods. Analyzing deleterious variant accumulation in genes defined a priori as associated with IBD risk (list provided by Jostins et al.7), we found them to be over-represented among rare variants in VEO IBD patients when compared to both healthy controls and adult IBD patients (Table 2). In contrast, no differences were found in the accumulation of these deleterious variants between adult patients and healthy controls.
Of 2072 rare and novel homozygous and heterozygous variants discovered in coding regions of genes associated with the innate immune system in any of the sequenced exomes, 927 variants were considered deleterious (Supplementary Table 7). As presented in Table 3, these deleterious alleles were also significantly over-represented among rare variants in comparisons between VEO IBD patients and healthy controls, and between VEO and adult IBD patients. They were not over-represented in adult IBD patients compared to healthy controls. Specifically, the significant over-representation of deleterious alleles was observed in children diagnosed with CD compared to healthy controls, and in children with UC compared to affected adults (Table 3).
Of 347 deleterious homozygous variants present in affected children but not in affected adults or healthy controls, 28 variants were located in genes present in the extended list of genes associated with IBD7 (Supplementary Table 8). Six of these variants were found in more than one child (Table 4). Among the remaining 319 variants, 272 (85%) were present in only one individual. None of these variants were present in more than six individuals.
Furthermore, of 53 rare and novel non-synonymous variants (out of which 37 are possibly deleterious with CADD score on PHRED scale above 10) in genes recognized previously as being associated with monogenic IBD (in a list provided by Uhlig et al.5), two homozygote variants were found; NCF4 p.Arg8Trp in one affected adult and WAS p.Glu131Lys (rs146220228, GMAF = 0.0008) in affected child and one adult patient, but not in healthy controls. Forty-six rare variants were present in HLA genes, 16 of which were considered deleterious (Supplementary Table 9), and one homozygous highly deleterious variant (frameshift variant in HLA-DRB1, c.565_566insC) was found in an affected child. Significant over-representation of deleterious alleles among rare variants was not observed for HLA genes (Supplementary Table 10).
Discussion
Association studies
A majority of patients with adult-onset IBD, and possibly a significant proportion of pediatric patients with IBDs, have classic complex (multifactorial) disorders driven by multiple common genetic variants, mostly non-protein-coding SNPs, exhibiting similar small effect sizes3,11,20. However, pediatric-onset IBDs, especially VEO IBDs, may differ from adult IBDs in many aspects, including disease type, disease location, disease behavior, and gender preponderance11,22,23,24. A subset of pediatric patients with a more severe disease course is likely under a higher influence of genetic effect. To discover IBD susceptibility loci specific for pediatric-onset or adult-onset disease, we conducted association studies on adequately powered pediatric and adult cohorts simultaneously after careful selection from the Polish population by experienced gastroenterologists. Consequently, our study thus allowed direct comparisons between healthy controls, adult patients, and pediatric patients.
Association studies can focus on individually selected variants using genotyping assays or on the position of millions of DNA variants using high-throughput technologies. Generally, the greater the sample size of a GWAS, the higher the number of associations that reach the genome-wide significance threshold25. The largest GWAS meta-analyses of IBDs uncovered 163 susceptibility loci, with each locus containing an average of five genes7, whereas in a GWAS limited to hundreds of IBD patients, only a few or no associations at P < 5 × 10−8 were found. Because of the use of statistical, rather than biological, criteria26,27 GWAS may generate both false-positive and false-negative results. Therefore, findings from GWASs are commonly supported by validation and replication studies that use individual genotyping.
The independent and very reliable TaqMan SNP genotyping assays revealed that most associations selected according to the genome-wide significance threshold from our cost-effective GWAS were false-positives. In contrast, of the 72 associations which could be selected according to blocks of SNPs being in strong allele linkage disequilibrium (Supplementary Table 4, lines 3–7 and 11–77), 67 SNPs were validated by individual genotyping; of these 25 and 42 SNPs were at the corrected or the nominal level of significance, respectively. This selection criterion defined a block by a distance of less than 30 kb between each two of at least 10 SNPs associated with a disease at P < 5 × 10−3 and with an “index SNP” in the block at least at P < 10−4, but it did not take into account local probe density (number of probes in a 30,000 base pair window) and the MAF. Thus, although no extensive optimization of the number of required loci, P-value threshold, or window size was performed, the algorithm turned out to be surprisingly effective, possibly because of a high probes density (2,612,357) on the array used. Among verified loci there were many which haven’t passed canonical statistical criterion (examples in Supplementary Figure 1). However, it is not known whether the new associations uncovered in our study were due to DNA pooling for SNP chip hybridization in connection with a novel method for SNP selection or because they are more specific for the Polish population with a relatively higher effect size than commonly found.
Although strict Bonferroni or a similar correction for multiple comparisons is typically required to effectively minimize false positive results, this approach may also reduce the discoverable amount of the heritability by ignoring innumerable truly positive signals27, especially when the correction may involve tests that are correlated25. While the expected number of false-positive results for 83 independent tests with a significance threshold of 0.05 is close to 4, an appropriate correction for multiple comparisons would reduce this to a small fraction of 1. In our validation studies, the multiple test correction reduced the number of associations at the nominal level of significance from 83 to 31, and thus generating a large number of false-negative results and only slightly reducing the expected number of false-positive results. Furthermore, for 55 SNPs the significance threshold was reached in at least 3 out of 9 comparisons that may reduce the likelihood of a false positive for a given SNP. Therefore, we reported both uncorrected and corrected results to balance between false positives and false-negatives. To this end, an informed reader can chose between the lower false positive (Bonferroni correction) and lower false negative (no correction) option.
Altogether, we uncovered much more susceptibility loci than one would expect considering the relatively small sample size. Moreover, although all risk loci with MAF >5% and OR > 1.2 are assumed to already have been identified in IBD patients with European ancestry8, our study replicated only 8 known IBD loci (NOD2, IL23R, IL10, ATG16L1, CYLD, ZMIZ1, SEMA6D, and CCR6) and a few (HLA-DRA, HLA-DQA1) SNPs from the MHC region; all others were newly discovered associations. In line with previous reports, NOD2 SNPs (rs13333062, rs6596, rs2076756) had the strongest effect size in both pediatric- and adult-onset CD (OR 2.3–3.7), but in contrast to other reports28, they did not show protective effects in UC. Among genes with SNPs exhibiting the strongest level of significance in validation genotyping were ATG16L1, VSX2, CYLD, ADAMTS19, NOX3, TFDP1, IL23R, IL10, and BRD2.
The MHC region on chromosome 6p21.3 contains more than 224 genes and is highly polymorphic. Risk variants within the MHC region are known to be associated with more than 100 different autoimmune and infectious diseases, including IBDs2,29,30. Although studies defining the architecture of association and causal alleles from this region are challenging, the high-density mapping of genetic variants from the MHC revealed a relatively equivalent contribution of class I and class II HLA variants to CD risk and HLA class II variation to UC risk30. Our study uncovered the association of 16 SNPs from the MHC region with IBDs (both risk and protective variants) (Supplementary Table 4) and one highly deleterious homozygous variant (HLA-DRB1 c.565_566insC) in an affected child. Thus, we confirmed a significant contribution of the MHC region to IBD risk and uncovered a novel signal, SNP rs1049526, located in the 3′UTR of BRD2, associated with an increased risk of IBD in both pediatric (OR 2.35) and adult (OR 2.66) patients.
BRD2 belongs to the bromodomain and extra-terminal domain (BET) family of chromatin adaptors that control adipogenesis, energy metabolism, and inflammation31. BRD2 directly regulates multiple TH17-associated cytokines, including IL17, IL21, and GMCSF32, and altered TH17 cells mediate autoimmune conditions, including multiple sclerosis, psoriasis, rheumatoid arthritis, and CD32. The mechanisms of BRD2 are in line with observations that many IBD-associated genes are involved in T-cell differentiation, specifically with the IL23 pathway (IL23R, JAK2, STAT3, IL12B, and PTPN2) involved in the maintenance of TH17 cells28,33. However, though GWASs have uncovered associations between BRD2 SNPs and systemic sclerosis34, type I diabetes35,36, multiple sclerosis37, and rheumatoid arthritis38,39, such an association has been not reported previously for IBD. Notably, we did not confirm any association of BRD2 rs1049526 with 458, 133, and 542 patients with primary biliary cirrhosis, primary sclerosing cholangitis, and celiac disease, respectively (data not shown). To this end, we uncovered several novel association signals, among which SNP rs1049526 in BRD2 seems to be particularly interesting. An unresolved question that remains is whether this association is highly IBD-specific only for the Polish population or has been overlooked by other GWASs.
As reported previously, among 32 loci associated with adult-onset CD and 17 loci associated with adult-onset UC, 21 and 8 loci have been replicated in pediatric-onset CD and UC, respectively16. However, only a few loci (including 2q37, 10q22, 16p11, 19q13, 20q13, 21q22, and 22q12) were found to be associated with pediatric IBDs15,16,40. In addition, a potential role of NOD2, TNFSF15, POU5F1, HLADRB1*501, and LAMB1 has been assumed in pediatric-onset CD and UC14. In this study, three NOD2 SNPs had the same strong effect size in both pediatric- and adult-onset CD, and IL23R rs11209026 had a risk effect in both pediatric and adult-onset UC and in pediatric-onset CD. The pediatric-specific ZMIZ1 rs125055041 had a protective effect in pediatric-onset CD and UC as well as adult-onset CD. However, most of the SNPs uncovered in this study differentiated between pediatric- and adult-onset IBD patients.
Exome sequencing
A diverse spectrum of rare genetic disorders with IBD-like phenotypes that result from rare causal alleles confined to exons or exon-intron boundaries3,20,42 can be catalogued by applying targeted exome sequencing or WES. WES interrogates the protein-coding portion of the human genome and, since its introduction, has been shown to be a powerful and cost-effective method for detecting disease variants underlying Mendelian disorders, as well as cataloguing common and rare disease-related genomic alterations43.
Increasing genetic burden is likely associated with an earlier age of IBD onset20. In support of this concept, VEO IBD patients experience distinct and more severe disease phenotypes with a more frequent positive family history3. Consequently, sequencing-based studies conducted in specific subsets of IBD patients (i.e., children younger than 2 years, infantile onset IBD, and familial clusters of affected individuals44) have discovered rare functional variants in genes implicated in the pathogenesis of both VEO (XIAP45, FOXP346, IL10RA, IL10RB, IL1047,48,49, and Il17REL42) and adult-onset disease (GSDMB21). These monogenic defects alter intestinal immune homeostasis via several mechanisms, which were divided by Uhlig et al.5,20 into: (1) disruption of the epithelial barrier and the epithelial response, (2) reduced clearance of bacteria by neutrophil granulocytes and other phagocytes, and (3) altered selection and activation of T-, B-, and Treg-cells. They all account for an unknown, but probably small, fraction of VEO IBD cases.
The discovery of monogenic disorders with IBD-like phenotypes caused by highly penetrant variants and occurring more frequently in VEO patients remains within the possibilities of contemporary genetic research. On the other hand, the identification of rare, potentially pathogenic variants that do not segregate in a strict Mendelian fashion, but contribute individually to disease risk, is challenging42,50,51,52,53. In addition, highly penetrant variants predisposing individuals to complex disorders are likely modified by common variants with low effect size. In this study, we performed WES in 43 children diagnosed with IBD before the age of 6, 45 patients diagnosed after the age of 40, and 18 healthy adults. We identified numerous rare potentially deleterious variants in genes selected from an extended list of IBD-associated genes7,20 and genes associated with the innate immune system54. Both subsets of accumulated variants were over-represented in affected children, but no differences were found between adult patients and healthy controls. Our findings indicate a contribution of these variants to VEO IBD. However, because the effect size for the variants selected by WES is mainly unknown, we cannot speculate on their pathogenicity, and exact determinants of VEO IBD remain to be explained. To what extent the rare non-synonymous variants may account for “the missing heritability” of IBD is also still unknown.
Although a significantly higher aggregation of deleterious variants along large groups of genes was shown in affected children, the lack of clear pathogenic variants in genes associated previously with monogenic IBD suggests that our VEO cases represented mostly a polygenic disease. Among 53 variants in genes recognized previously as being associated with monogenic IBD, only two homozygote variants (NCF4 p.Arg8Trp and WAS p.Glu131Lys) were found in two affected adults, and homozygous WAS p.Glu131Lys was found in one adult and pediatric patient. NCF4 p.Arg8Trp is a new variant, and no clinical data is available for WAS p.Glu131Lys. NCF4 encodes neutrophil cytosolic factor 4. This protein is a regulatory component of the superoxide-producing phagocyte NADPH-oxidase, which is crucial in host defense. Mutations in this gene cause chronic granulomatous disease55, in which the host’s reaction to pathogenic microbes is severely impaired. Up to 40% of patients with chronic granulomatous disease can manifest CD-like symptoms56, and few patients manifest symptoms of UC57. WAS encodes Wiskott-Aldrich Syndrome protein. Wiskott–Aldrich syndrome is a primary immunodeficiency disease in which up to 9% of patients exhibit an IBD-like syndrome58.
To this end, WES allows the cataloguing of genes enriched for rare variants, but more importantly, we need effective methods to investigate the heritability of specific functional alleles in complex disorders. The unresolved question is how to separate the individual causative effects of rare alleles with low/moderate penetrance from the effect of the polygenic burden of common variants. For the discovery of rare incompletely penetrant variants, even large studies are underpowered due to an enormous number of tests needed to establish the significance of the association9.
Summary
To discover differences in the genomic architecture of IBD in Polish patients, we performed both genome-wide association screening and WES simultaneously in pediatric and adult cohorts. A pooled-DNA GWAS approach that efficiently and cost-effectively scanned for common risk alleles59,60,61, together with a new method for identifying SNP associations, appeared to be unexpectedly effective in identifying novel IBD susceptibility loci. Despite “index SNPs” for these loci being selected at P-values much higher than a standard genome-wide significance threshold, most of them were validated in individual genotyping. Our GWAS indicated differences in the polygenic architecture between pediatric- and adult-onset IBD, but a significant accumulation of missense and nonsense rare variants in affected children suggested a contribution of yet unexplained genetic components to VEO IBD. To summarize, Polish pediatric IBD patients exhibited genetically attributable risk that was visibly different from that of adult-onset patients, which may question the previous assumption28 of a close pathogenic relationship between pediatric- and adult-onset IBD.
Additional Information
Accession codes: The datasets used in this GWAS are available in the GEO database under GSE79094. Whole-exome sequencing data (as bam files mapped to hg19 genome assembly) are available in European Nucleotide Archive under accession number PRJEB12993.
How to cite this article: Ostrowski, J. et al. Genetic architecture differences between pediatric and adult-onset inflammatory bowel diseases in the Polish population. Sci. Rep. 6, 39831; doi: 10.1038/srep39831 (2016).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Ruel, J., Ruane, D., Mehandru, S., Gower-Rousseau, C. & Colombel, J.-F. IBD across the age spectrum: is it the same disease ? Nat Rev Gastroenterol Hepatol 11, 88–98 (2014).
Parkes, M., Cortes, A., van Heel, D. A. & Brown, M. A. Genetic insights into common pathways and complex relationships among immune-mediated diseases. Nat. Rev. Genet. 14, 661–673 (2013).
McGovern, D. P. B., Kugathasan, S. & Cho, J. H. Genetics of Inflammatory Bowel Diseases. Gastroenterology 149, 1163–1176.e2 (2015).
de Souza, H. S. P. & Fiocchi, C. Immunopathogenesis of IBD: current state of the art. Nat Rev Gastroenterol Hepatol 13, 13–27 (2016).
Uhlig, H. H. Monogenic diseases associated with intestinal inflammation: implications for the understanding of inflammatory bowel disease. Gut 62, 1795–1805 (2013).
Van Limbergen, J., Radford-Smith, G. & Satsangi, J. Advances in IBD genetics. Nat Rev Gastroenterol Hepatol 11, 372–385 (2014).
Jostins, L. et al. Host-microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124 (2012).
Ellinghaus, D., Bethune, J., Petersen, B.-S. & Franke, A. The genetics of Crohn’s disease and ulcerative colitis—status quo and beyond. Scand. J. Gastroenterol. 50, 13–23 (2015).
Heyman, M. B. et al. Children with early-onset inflammatory bowel disease (IBD): analysis of a pediatric IBD consortium registry. J. Pediatr. 146, 35–40 (2005).
Paul, T. et al. Distinct phenotype of early childhood inflammatory bowel disease. J. Clin. Gastroenterol. 40, 583–586 (2006).
Van Limbergen, J. et al. Definition of phenotypic characteristics of childhood-onset inflammatory bowel disease. Gastroenterology 135, 1114–1122 (2008).
Büning, C. et al. Mutations in the NOD2/CARD15 gene in Crohn’s disease are associated with ileocecal resection and are a risk factor for reoperation. Aliment. Pharmacol. Ther. 19, 1073–1078 (2004).
Maconi, G. et al. CARD15 gene variants and risk of reoperation in Crohn’s disease patients. Am. J. Gastroenterol. 104, 2483–2491 (2009).
Connelly, T. M. et al. Genetic determinants associated with early age of diagnosis of IBD. Dis. Colon Rectum 58, 321–327 (2015).
Kugathasan, S. et al. Loci on 20q13 and 21q22 are associated with pediatric-onset inflammatory bowel disease. Nat. Genet. 40, 1211–1215 (2008).
Imielinski, M. et al. Common variants at five new loci associated with early-onset inflammatory bowel disease. Nat. Genet. 41, 1335–1340 (2009).
Franke, A. et al. Sequence variants in IL10, ARPC2 and multiple other loci contribute to ulcerative colitis susceptibility. Nat. Genet. 40, 1319–1323 (2008).
HumanOmni2.5-8 v1.2 Support Files. Available at: http://support.illumina.com/content/illumina-support/us/en/downloads/humanomni2-5-8-v1-2-product-support-files.html. (Accessed: 25th February 2016).
Cutler, D. J. et al. Dissecting Allele Architecture of Early Onset IBD Using High-Density Genotyping. PLoS ONE 10, e0128074 (2015).
Uhlig, H. H. et al. The diagnostic approach to monogenic very early onset inflammatory bowel disease. Gastroenterology 147, 990–1007.e3 (2014).
Christodoulou, K. et al. Next generation exome sequencing of paediatric inflammatory bowel disease patients identifies rare and novel variants in candidate genes. Gut 62, 977–984 (2013).
Griffiths, A. M. Specificities of inflammatory bowel disease in childhood. Best Pract Res Clin Gastroenterol 18, 509–523 (2004).
Kugathasan, S. & Cohen, S. Searching for new clues in inflammatory bowel disease: tell tales from pediatric IBD natural history studies. Gastroenterology 135, 1038–1041 (2008).
Levine, A. et al. Pediatric onset Crohn’s colitis is characterized by genotype-dependent age-related susceptibility. Inflamm. Bowel Dis. 13, 1509–1515 (2007).
Sham, P. C. & Purcell, S. M. Statistical power and significance testing in large-scale genetic studies. Nat. Rev. Genet. 15, 335–346 (2014).
Sobota, R. S. et al. Addressing population-specific multiple testing burdens in genetic association studies. Ann. Hum. Genet. 79, 136–147 (2015).
Williams, S. M. & Haines, J. L. Correcting away the hidden heritability. Ann. Hum. Genet. 75, 348–350 (2011).
Ek, W. E., D’Amato, M. & Halfvarson, J. The history of genetics in inflammatory bowel disease. Ann Gastroenterol 27, 294–303 (2014).
Shiina, T., Hosomichi, K., Inoko, H. & Kulski, J. K. The HLA genomic loci map: expression, interaction, diversity and disease. J. Hum. Genet. 54, 15–39 (2009).
Goyette, P. et al. High-density mapping of the MHC identifies a shared role for HLA-DRB1*01:03 in inflammatory bowel diseases and heterozygous advantage in ulcerative colitis. Nat. Genet. 47, 172–179 (2015).
Belkina, A. C., Blanton, W. P., Nikolajczyk, B. S. & Denis, G. V. The double bromodomain protein Brd2 promotes B cell expansion and mitogenesis. J. Leukoc. Biol. 95, 451–460 (2014).
Mele, D. A. et al. BET bromodomain inhibition suppresses TH17-mediated pathology. J. Exp. Med. 210, 2181–2190 (2013).
Duerr, R. H. et al. A genome-wide association study identifies IL23R as an inflammatory bowel disease gene. Science 314, 1461–1463 (2006).
Zhou, X. et al. HLA-DPB1 and DPB2 are genetic loci for systemic sclerosis: a genome-wide association study in Koreans with replication in North Americans. Arthritis Rheum. 60, 3807–3814 (2009).
Hakonarson, H. et al. A genome-wide association study identifies KIAA0350 as a type 1 diabetes gene. Nature 448, 591–594 (2007).
Grant, S. F. A. et al. Follow-up analysis of genome-wide association data identifies novel loci for type 1 diabetes. Diabetes 58, 290–295 (2009).
International Multiple Sclerosis Genetics Consortium et al. Risk alleles for multiple sclerosis identified by a genomewide study. N. Engl. J. Med. 357, 851–862 (2007).
Plenge, R. M. et al. Two independent alleles at 6q23 associated with risk of rheumatoid arthritis. Nat. Genet. 39, 1477–1482 (2007).
Plenge, R. M. et al. TRAF1-C5 as a risk locus for rheumatoid arthritis--a genomewide study. N. Engl. J. Med. 357, 1199–1209 (2007).
Amre, D. K. et al. Investigation of reported associations between the 20q13 and 21q22 loci and pediatric-onset Crohn’s disease in Canadian children. Am. J. Gastroenterol. 104, 2824–2828 (2009).
Jakobsen, C. et al. Genetic susceptibility and genotype-phenotype association in 588 Danish children with inflammatory bowel disease. J Crohns Colitis 8, 678–685 (2014).
Sasaki, M. M. et al. Whole-exome Sequence Analysis Implicates Rare Il17REL Variants in Familial and Sporadic Inflammatory Bowel Disease. Inflamm. Bowel Dis. 22, 20–27 (2016).
Auer, P. L. et al. Rare and low-frequency coding variants in CXCR2 and other genes are associated with hematological traits. Nat. Genet. 46, 629–634 (2014).
Stittrich, A. B. et al. Genomic architecture of inflammatory bowel disease in five families with multiple affected individuals. Hum Genome Var 3, 15060 (2016).
Zeissig, Y. et al. XIAP variants in male Crohn’s disease. Gut 64, 66–76 (2015).
Okou, D. T. et al. Exome sequencing identifies a novel FOXP3 mutation in a 2-generation family with inflammatory bowel disease. J. Pediatr. Gastroenterol. Nutr. 58, 561–568 (2014).
Glocker, E.-O. et al. Inflammatory bowel disease and mutations affecting the interleukin-10 receptor. N. Engl. J. Med. 361, 2033–2045 (2009).
Kotlarz, D. et al. Loss of interleukin-10 signaling and infantile inflammatory bowel disease: implications for diagnosis and therapy. Gastroenterology 143, 347–355 (2012).
Moran, C. J. et al. IL-10R polymorphisms are associated with very-early-onset ulcerative colitis. Inflamm. Bowel Dis. 19, 115–123 (2013).
Bonnefond, A. et al. Rare MTNR1B variants impairing melatonin receptor 1B function contribute to type 2 diabetes. Nat. Genet. 44, 297–301 (2012).
Diogo, D. et al. Rare, low-frequency, and common variants in the protein-coding sequence of biological candidate genes from GWASs contribute to risk of rheumatoid arthritis. Am. J. Hum. Genet. 92, 15–27 (2013).
Beaudoin, M. et al. Deep resequencing of GWAS loci identifies rare variants in CARD9, IL23R and RNF186 that are associated with ulcerative colitis. PLoS Genet. 9, e1003723 (2013).
Rivas, M. A. et al. Deep resequencing of GWAS loci identifies independent rare variants associated with inflammatory bowel disease. Nat. Genet. 43, 1066–1073 (2011).
de Bono, B., Gillespie, M., Luo, F. & Gay, N. Innate Immunity Signaling. Reactome - a curated knowledgebase of biological pathways 24, (2008).
Matute, J. D. et al. A new genetic subgroup of chronic granulomatous disease with autosomal recessive mutations in p40phox and selective defects in neutrophil NADPH oxidase activity. Blood 114, 3309–3315 (2009).
van den Berg, J. M. et al. Chronic Granulomatous Disease: The European Experience. Plos One 4, e5234 (2009).
Marciano, B. E. et al. Gastrointestinal Involvement in Chronic Granulomatous Disease. Pediatrics 114, 462–468 (2004).
Catucci, M., Castiello, M. C., Pala, F., Bosticardo, M. & Villa, A. Autoimmunity in Wiskott–Aldrich Syndrome: An Unsolved Enigma. Front Immunol 3, (2012).
Gaj, P. et al. Pooled sample-based GWAS: a cost-effective alternative for identifying colorectal and prostate cancer risk variants in the Polish population. Plos One 7, e35307 (2012).
Gaj, P. et al. Identification of a late onset Alzheimer’s disease candidate risk variant at 9q21.33 in Polish patients. J. Alzheimers Dis. 32, 157–168 (2012).
Paziewska, A. et al. Combination Testing Using a Single MSH5 Variant alongside HLA Haplotypes Improves the Sensitivity of Predicting Coeliac Disease Risk in the Polish Population. Plos One 10, e0139197 (2015).
Acknowledgements
This work was supported by the National Science Centre [2011/02/A/NZ5/00339].
Author information
Authors and Affiliations
Contributions
Conception and design of the study: J.O.; Patients recruitment and clinical data compilation: J.O., I.L., R.T., J.Kierkus, P.S., M.L., G.R., M.Klopocka, G.M., B.I., E.K., K.B.D., J.W., B.K., P.R., U.G.C., P.L., A.J., B.K., T.S., P.A.; DNA isolation, Taqman genotyping, exome sequencing: A.P., F.A., J.Karczmarski, M.D., A.K., A.B., M.P.; GWAS dataset analyses: K.G. and K.P.; Exome-Seq dataset analyses: M.Kulecka; Drafting of the manuscript: J.O., M.Kulecka, K.G., M.M.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Ostrowski, J., Paziewska, A., Lazowska, I. et al. Genetic architecture differences between pediatric and adult-onset inflammatory bowel diseases in the Polish population. Sci Rep 6, 39831 (2016). https://doi.org/10.1038/srep39831
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep39831
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.