Abstract
Decoding the patterns of miRNA regulation in diseases are important to properly realize its potential in diagnostic, prog- nostic, and therapeutic applications. Only a handful of studies computationally predict possible miRNA-miRNA interactions; hence, such interactions require a thorough investigation to understand their role in disease progression. In this paper, we design a novel computational pipeline to predict the common signature/core sets of miRNA-miRNA interactions for different diseases using network inference algorithms on the miRNA-disease expression profiles; the individual predictions of these algorithms were then merged using a consensus-based approach to predict miRNA-miRNA associations. We next selected the miRNA-miRNA associations across particular diseases to generate the corresponding disease-specific miRNA-interaction networks. Next, graph intersection analysis was performed on these networks for multiple diseases to identify the common signature/core sets of miRNA interactions. We applied this pipeline to identify the common signature of miRNA-miRNA inter- actions for cancers. The identified signatures when validated using a manual literature search from PubMed Central and the PhenomiR database, show strong relevance with the respective cancers, providing an indirect proof of the high accuracy of our methodology. We developed miRsig, an online tool for analysis and visualization of the disease-specific signature/core miRNA-miRNA interactions, available at: http://bnet.egr.vcu.edu/miRsig.
Similar content being viewed by others
Introduction
MicroRNAs (miRNAs) are non-coding RNAs of ~22 nucleotides in length that inhibit gene expression at the post transcriptional level by binding to the 3′ UTR region of target mRNAs through complementary base pairing1. However, a couple of studies have instead reported an activation of target gene expression as well2,3. By virtue of this gene regulation mechanism, miRNAs play a critical role in several biological processes4 and patho-physiological conditions, including cancers5. The role of miRNA regulations in diseases have been widely recorded6, however the precise patterns through which a miRNA regulates a certain disease(s) are still elusive. For example, it is not yet clear how a miRNA’s up/down regulation directly or indirectly affects a disease’s progression or repression because of the many intermediate factors involved. Thus, predicting and identifying miRNA-disease associations has been a primary research area for several groups. Moreover, the multi-level interactions of miRNAs in cancer-like multi-factorial diseases are more complex due to the possibility of several types of interactions, such as, the classical miRNA-mRNA, miRNA-environmental factors, miRNA- transcription factors-miRNA7, and our newly hypothesized direct miRNA-miRNA interactions without any intermediate linkers (e.g., transcription factors)8. However, till date, no experimental proof of direct miRNA-miRNA interactions exists except, a single study reported in mouse9.
Although, the precise patterns or the reasons behind miRNAs’ deregulation in cancers are not fully understood, it has been found that miRNAs tend to work together in groups10, as evidenced in certain diseases11. Such co-ordinated regulation, comprising mutual co-targeting and co-regulation, as well as miRNA regulation by other miRNAs are reported in many disease conditions, including various cancers10. To elucidate the miRNA-disease associations at the regulome level, we earlier developed the miRegulome database and corresponding analytic tools12. Furthermore, in cancers it has been observed that groups of miRNAs, known as superfamilies, express consistently across several cancers and may act as drivers of tumorigenesis, where few key miRNAs direct the global miRNA expression patterns13. Identification and existence of such groups or super-families of miRNAs obviously leads to the intuition, that the therapeutic suppression or expression of any one of the miRNAs in the family, would compensate for the other participants of the family13. Our central hypothesis in this paper is that, these miRNAs in such superfamilies may interact directly or indirectly, by forming a core miRNA-miRNA co-regulatory network and thereby acting as a signature component for prognosis, prediction, and early diagnosis of any disease including cancer.
Several computational efforts have been implemented to study and discover the disease-miRNA interaction networks based on functional enrichment analysis14, social network analysis methods15, similarity-based methods16, and diffusion-based methods17. Some studies have integrated genomic and phenotype data sets to infer novel miRNA-disease associations18. A miRNA regulatory network was also constructed by integrating multidimensional high-throughput data and was used to identify the cancer-associated miRNAs19. Similarly, co-regulating miRNA clusters and prioritized candidate miRNAs across multiple types of diseases have been predicted. Using co-regulating functional modules, a miRNA-miRNA synergistic network was constructed to study the aspect of homophily among miRNAs associated with the same disease and subsequently disease-specific miRNAs were detected based on their network topological features. In this study, a miRNA-miRNA co-regulation network was constructed by selecting common miRNAs across various data sets related to the same disease, pairing them based on their sharing of common targets, and subsequently performing a GO enrichment analysis of their predicted targets. These miRNAs were qualified as co-regulating if they shared a significant amount of GO enrichment analyses of predicted targets20. Disease-specific miRNAs were also identified using the miRNA target-dysregulated network built on the assumption that causative miRNAs show abnormal regulation of their target genes21. Similarly, disease-specific miRNAs were also identified by integrating phenotype associations of diseases which had matching miRNA and mRNA expression profiles22. Network theoretic algorithms such as the biclique-based method23, biclustering technique24 and maximum weighted matching25 among others have been deployed to discover and predict the patterns of miRNA regulation. Graph theoretical methods and network inference models have also been applied to analyze complex regulatory interactions and reconstruct the causative gene regulatory network and other biological networks26,27,28,29.
In this work, we have used the miRNA expression data sets available at the PhenomiR30 database to predict miRNA-miRNA core/signature interactions across several cancers using a combination of (i) six state-of-the-art network inference algorithms, (ii) a wisdom of crowds31 based consensus approach32 to generate disease-specific miRNA interaction networks with higher accuracy, and (iii) a simplified graph intersection analysis to identify the miRNA-miRNA core interactions across multiple diseases belonging to a particular disease class.
Methods
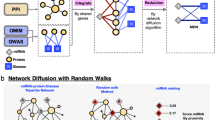
The methodology adopted in this paper is comprised of i) translating the miRNA-disease expression scores from the PhenomiR database into a miRNA expression matrix (Fig. 1, Step 1); ii) deploying six network inference algorithms on the expression matrix and deriving the miRNA-miRNA interaction scores from each algorithm (Fig. 1, Step 2); iii) performing a consensus-based approach, i.e. estimating an average score for every miRNA-miRNA interaction across its six predicted scores (Fig. 1, Step 3); iv) validating the resultant interactions using precision-recall analysis with a hypothetical true network generated using the PubMed IDs from PhenomiR; v) analyzing the miRNA-miRNA interaction networks for every disease and detection of the conserved miRNA-miRNA interactions across various groups of cancers and finally vi) validating the conserved miRNA-miRNA interactions in the identified group of cancers via manual literature search.
Step 1 consists of translating the PhenomiR data set into three miRNA expression matrices (a, b and c) based on three approaches. In Step 2, each of these matrices are subjected to six network inference algorithms which produce the interaction scores across the different MiDj nodes. In Step 3, the six individual MiDj − MxDy interaction scores are averaged into a final score designating its confidence.
Data preparation and modeling
The data from the PhenomiR database is freely available and was used in this study. PhenomiR 2.0 was downloaded for the purposes of this study. PhenomiR 2.0 is a comprehensive data set containing 535 database entries across 345 articles recording miRNA expressions in diseases30. As shown in Fig. 2, the data from PhenomiR was converted into a disease-specific miRNA expression matrix (shown in Fig. 3). The miRNAs whose fold-change values were not available in PhenomiR 2.0 data set were discarded from the study; this also includes some misformatted lines of data that were excluded from further processing as they were also missing the fold-change values. Here, the core idea is to consider a pair of miRNA and disease as a single miRNA-disease (MD) node, as seen in Fig. 3; note that, for ease of reference, we consider an MiDj pair as an MD node which conceptually designates a disease-specific miRNA. The same miRNA participating in multiple diseases will have different expression profiles in each of them and hence the disease specific miRNA terminology, i.e., MD, signifies a miRNA’s expression profile in a particular disease. Thus, every unique miRNA-disease pair constitutes a unique MD type node. In this disease-specific miRNA expression matrix (Fig. 3-b), each row represents a study/experiment and each column represents an MD’s expression score in that study. The resultant expression matrix herein, has 4,343 unique nodes/columns (i.e., unique MDs in the network) for 267 samples (i.e., rows).
In the PhenomiR data set, some MDs have two fold-change values indicating minimum and maximum expression scores while other MDs only report a minimum fold-change expression score (for e.g., see Fig. 1, Step 1, PhenomiR data set, row 2). To assess these scenarios, we devised three different methodologies (described in the next section), generated separate expression matrices based on each methodology and performed the subsequent analysis on each of them.
Average scoring
Under Average scoring method, for the MDs having both minimum and maximum fold-change values per sample, their average was taken and considered as the final expression value in the expression matrix. As shown in Fig. 1, Step-1, the entry M1-D1 has two expression values - 2.3 and 2.9, i.e., minimum fold-change and maximum fold-change respectively, which were averaged to 2.6 in the Expression Matrix 1 (see Fig. 1, Step 1-a). For MDs with only their minimum expression values reported, this single value was also considered to be its average expression value.
Retaining maximum and minimum expression values
The Average scoring method can lead to a potential loss of information as the individual maximum and minimum expression values (when available) were not retained. Hence we designed the following two methods to generate the expression matrix.
- 1
Max-Min scoring
Under Max-Min scoring method, for the MDs having minimum and maximum fold-change expression values, (instead of taking their average) both these data points were considered as separate entries; thus, the same MD was considered twice in the expression matrix with the duplicate entry designating a new experiment. As displayed in Fig. 1, Step 1, the first row entry, M1-D1 in Study-1 has two expression values; these values were individually considered as separate data points and included in the expression matrix accordingly along with their co-expressing miRNAs’ expression values, providing us with Expression Matrix 2 (see Fig. 1, Step 1-b).
- 2
Computing Missing Max. scoring
Under Computing Missing Max scoring method, for the MDs which did not have a maximum fold-change expression value, we took an average of its maximum fold-change values across all its other samples and substituted this average score as it’s maximum fold-change expression value. As shown in Fig. 1, Step 1, the entry M2-D1 on 2nd row does not have a maximum fold-change value. However, M2-D1 combination has maximum fold-change expression values of 6.7 and 3.1 from sample #5 and #6, respectively. Herein, we took an average of these two values, i.e. 6.4 and substituted it for the original missing value for M2-D1 in the 2nd row. This method overcomes the limitation posed due the non-availability of the expression value by giving its closest approximation, based on the particular MD’s expression pattern across the sample spectrum. After applying this method, the Average Scoring method was performed on this matrix to obtain Expression Matrix 3 (see Fig. 1, Step 1-c).
After the three expression matrices were derived, a reverse engineering methodology32 was adopted to reconstruct the MD-MD regulatory network from these expression matrices (Fig. 3, Network Inference), by applying six widely used network inference algorithms along with a consensus-based ranking algorithm, which is explained in the next section.
Network inference algorithms
Each expression matrix has 4,343 nodes and therefore, there are potentially 4,343 × 4,343 (i.e. 18,861,649) MD-MD interactions in the network. Six different network inference algorithms were applied on the miRNA expression matrix, which gave prediction scores for every MD-MD interaction. We used the mutual information-based algorithm, Context Likelihood of Relatedness (CLR)33, Maximum Relevance Minimum Redundancy Backward (MRNETB)34, Basic Correlation methods (Pearson and Spearman), Distance Correlation (DC)35, and regression-based Gene Network Inference with Ensemble of Trees (GENIE3)36 algorithms for network inference. The details of the algorithms are given in Supplementary File S1. Note that, the Basic Correlation methods resulted in two different network inference algorithms based on the type of correlations implemented, i.e., one each for Pearson and Spearman correlations.
Consensus based network inference approach
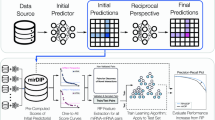
Each of the six individual network inference algorithms produced a ranked list of prediction scores for every MD-MD interaction (see Fig. 1, Step-2). Thereafter, we used the wisdom of crowds31 approach, which proposes that the aggregation of information from the community yields better results than the individual few. In this study, the consensus based approach aggregates the collective information (i.e. prediction scores) from the six individual network inference algorithms and computes a more accurate final score for MD-MD interactions. This rank is computed by taking an average of the predicted ranks of each interaction derived from the corresponding network inference algorithms. Figure 4 displays the workflow of this approach. This approach was earlier implemented to infer gene-regulatory networks and yielded highest accuracy compared to each of the individual network inference algorithms32.
This consensus based network inference approach is executed in the Average Rank32 algorithm which essentially computes the average score of a particular MD-MD interaction by taking the mean of its six predicted ranks. The ranking methodology used in this algorithm is based on the Borda count method. This method is used in elections during which voters rank candidates as per their preferences. The winning candidate is the one with the best average rank. Here, all the interactions are first ranked in descending order of their predicted scores (as seen in the column Rank in Table 1). Describing briefly, the Borda count method allocates points to each rank. The highest ranked interaction (meaning, 1) get the maximum Borda points (number of interactions - 1) and the lowest ranked interaction has 0 Borda points as demonstrated in the column Borda points in Table 1. In order to derive the final rank between 0 and 1, these points are thereafter normalized to derive a relative Borda rank. Thus, each rank has been translated to its new relative Borda rank. Note that, the Borda count ranking method is among the many other methods to perform averaging of the ranks in the consensus methodology.
The six network inference algorithms generate six different ranks for each interaction and the consensus algorithm next computes an average Borda rank for the interaction. Tables 1 and 2 display a scenario of ranking four MD-MD interactions I1, I2, I3 and I4 via a consensus-based approach as executed in AverageRank algorithm. Table 1 displays the ranked list of predictions for these interactions by all the six network inference algorithms based on their prediction scores. For example, in Table 1, Algorithm 1 ranks MD-MD interactions in this order — I4, I2, I1 and I3 based on their prediction scores. The individual ranks for miRNA-miRNA interaction I4 are 1, 3, 4, 2, 2 and 3 by the six algorithms respectively (noted with *), and their relative respective Borda ranks are 1, 0.333, 0, 0.666. 0.666 and 0.333. The final rank of interaction I4 is the average of all the Borda ranks, i.e., 0.49, as demonstrated in Table 2 (noted with *). Similarly the final ranks of every other interaction is computed using the following formula,

where, K is the number of algorithms (six, in our case). These results are displayed in Table 2.
An example of the final result listing of our MD-MD interactions is shown in Table 3 (also see Fig. 1 Step-3).
In these results, we noted all the different possibilities of interactions that can occur considering the miRNA-disease pair, i.e. MD as a node. There are essentially four types of interactions that can exist in this network. These are explained in Table 4. Among these types, type 1 is a self-loop and not applicable for our purposes. For application purposes of our methodology, we focused on analyzing the set of interactions belonging to type 3 which is further elaborated in the next section. Interactions of type 2 and type 4 will be studied in the future to analyze the relationship between diseases sharing a common miRNA (type 2) and the proximity between dissimilar miRNAs and dissimilar diseases (type 4) having high probabilities of interaction.
Disease-specific miRNA network construction
In this section, the results of the type 3 interactions were selected for disease-specific analysis. There were 66 unique diseases in the final predicted list of interactions from the Average Rank algorithm; this list of diseases are provided in the Supplementary File S2. Under a specific disease Dx, all the miRNA-miRNA edges, i.e. M1Dx − M2Dx edges were collected into a single Dx disease network; thereby giving us the disease-specific miRNA-miRNA interaction network (DMIN) (Fig. 5, Step 1). DMIN is a network G = (V, E), where V = {M1Dx, M2Dx, … MnDx} (i.e., set of miRNAs under disease name Dx) and E is the ordered set of edges, where edge e = {Mi, Mj}. We performed a similar network construction for every cancer-related disease, Dx. To pursue a more definitive and cancer-specific analysis, only cancer-related diseases were chosen and grouped into classes based on their tissue/organ specificity. We created four major classes: i) gastrointestinal cancers (esophageal, gastroesophageal, gastrointestinal, gastric, and colorectal cancer), ii) endocrine cancers (hepatocellular, pancreatic, and thyroid carcinoma follicular, and thyroid carcinoma papillary), iii) leukemia/blood cancers (hematological tumors, acute myeloid leukemia, chronic lymphatic leukemia, and acute myelogenous leukemia), and iv) nerve cancers (neuroblastoma, medulloblastoma, and glioblastoma).
Under a particular disease class, all the corresponding DMINs were combined into a single network (Fig. 5, Step 2). Using graph intersection analysis, we mined the miRNA-miRNA interaction networks of all the cancers within the specific class to identify a conserved (signature/core) miRNA-miRNA interaction component. This identified miRNA-miRNA interaction component was present in all the diseases of that particular class. These findings are reported in the pan-cancer miRNA signatures section and the results are discussed in the Discussion section.
Results
Validation of interactions
After executing the Consensus based network inference approach on three input miRNA expression matrices derived from the three approaches mentioned in the Data preparation and modeling section (Average scoring, Retaining Max-Min and Computing Missing Max.), we obtained three sets of predicted miRNA-miRNA interactions. Each predicted interaction was validated by querying for PubMed IDs in the PhenomiR database which cited and reported the occurrence of miRNAs’ association with the specific disease in a single PubMed ID. For e.g., for each predicted interaction, i.e. MaDx to MbDx, if a PubMed ID cited the occurrence of the association between the miRNAs (Ma, Mb) and the disease (Dx), the interaction was termed as true/validated (1); else the predicted interaction was termed as unknown/unverified (0). Based on this, labels were generated for every interaction in the resultant set forming the true network. We performed a precision-recall analysis to ascertain the accuracy of the consensus-based network inference method. The precision-recall values were calculated using the formula:

where tp, fp, and fn are true-positives, false-positives and false-negatives respectively.
Figure 6 displays the results of the precision-recall analysis and the ROC curve for all the three approaches used. As demonstrated in the figure, the Average scoring method fared better than the other two methods; in fact the Computing Missing Max. method also performed well for low recall but gradually degraded for higher recall values. Based on this precision-recall curve, our proposed methodology displays a high precision (for up to a 30% recall) demonstrating its effectiveness in providing high confidence to the results. The ROC curve shows that both the Average scoring and Computing Missing Max. methods are comparable in predicting the true positives when compared to the number of false positives seen alongside.
The figure demonstrates that the Average scoring (blue curve) method fared better than Retaining Max-Min (green curve) and Computing Missing Max. (red curve) methods. The inset image shows that the precision of Average scoring method slightly outperformed the Computing Missing Max. and was the best overall performer.
Note that our true network generation method has some obvious limitations. While a true edge constituting the association of the two miRNAs with the same disease in the same PubMed ID is still acceptable (specifically because these edges were manually curated), the unverified edges may simply mean that a study has not yet been reported associating the miRNAs to the same disease. Hence, a high precision performance should be the best judge of our methodology whereas the recall curve can be somewhat circumstantial.
Pan-cancer miRNA signatures
After the Validation of interactions, in order to confidently detect miRNA signatures in the specified disease classes, only the top 10% interactions with the highest confidence scores were used in the construction of DMIN (Fig. 5, Step 1) and the subsequent graph intersection approach (Fig. 5, Step 2). Hence, all the considered miRNA-miRNA interactions had a confidence score of 0.9 and above. As reported in Fig. 7, under gastrointestinal cancers, we detected a signature component of three miRNAs (hsa-mir-30a, hsa-mir-181a-1, and hsa-mir-29c). For endocrine cancers, the signature component consisted of hsa-mir-221, hsa-mir-222, hsa-mir-155, hsa-mir-224, hsa-mir-181a-1, and hsa-mir-181b-1. For leukemia cancers, the signature component consisted of hsa-mir-29b-1, hsa-mir-106a, hsa-mir-20a, hsa-mir-126, and hsa-mir-130a. We observed two different signatures for nerve cancers. For subsequent validation of these cancer-specific signature set of miRNAs, we manually mined PubMed articles which corroborate our results, as reported in Fig. 7. We queried both the PhenomiR database and the PubMed Central database for these reported PubMed IDs; the results from these two sources are shown in different colors in Fig. 7. We also observed that, while hsa-mir-30 is common in gastrointestinal and nerve cancers; hsa-mir-181 is shared by gastrointestinal, endocrine and nerve cancers. The miRNA signature component of the category leukemia is found to possess a distinct group of miRNAs (Fig. 7). The role and involvement of these miRNAs in their associated diseases are further elaborated in the Discussion section.
The individual steps involved in the manual search process from PubMed Central are shown in Fig. 8. To summarize, we first searched PubMed Central with the list of core miRNAs and each disease for which they form a signature component. We next manually checked the ‘search’ results to confirm the associations (i.e., the pruning step for PMIDs). If not enough results were retrieved from this search, we entered each miRNA, disease pair individually for all the miRNAs forming the signature component in that disease; each of these results were then manually pruned and collated to give us the set of PMIDs corresponding to the core miRNAs for that disease. This process was repeated for all the other diseases of a particular disease class.
miRsig - an online tool
In order to aid researchers to identify disease-specific miRNA-miRNA interaction networks across several diseases, we developed the miRsig tool, available at http://bnet.egr.vcu.edu/miRsig. miRsig allows the user to visualize the miRNA-miRNA interaction network for each disease recorded in PhenomiR and also across multiple diseases. The results are based on the consensus-based network inference approach. miRsig also allows users to search for a common/core miRNA-miRNA interaction component in a user-specified selection of diseases (see Fig. 9). Users can create their own class/category of cancers by selecting more diseases, as shown in Fig. 9. The edges in the interaction have confidence scores as weights, from 0 (minimum) to 1 (maximum). Hence, the tool also allows the user to view only the higher/lower/specific confidence interactions by changing the Maximum and Minimum confidence score ranges. Currently, the total number of edges across the entire miRNA-miRNA interaction networks are more than 18 million. Hence, to avoid cluttering of the result set and to allow clear visibility and comprehension of the network, the Minimum score is set to 0.5, if not specified by the user. Users can also view and analyze the topological properties of miRNA clusters interacting in each or a set of diseases. The signature/core miRNA-miRNA interactions among esophageal, gastroesophageal, gastrointestinal, gastric, and colorectal cancers, as predicted and visualized is shown in Fig. 9. This network component consisting of three miRNAs (has-mir-30a, has-mir-181a-1, and has-mir-29c) is the signature component for all the aforementioned five cancers, and can be validated using simple literature search on PubMed Central database as demonstrated in Fig. 8.Users can also download the miRNA-interaction network in the format of an edge-list in a CSV file. This edge-list can be imported in various network analysis tools such as, NodeXL, Cytoscape, etc. for further study and analysis of the interaction network.
miRsig tool has been developed using MySQL as the back-end database and HTML, PHP, JavaScript, AJAX for front-end design. The interactive network visualization has been implemented using data visualization library, D3.js37.
Discussion
miRNA-mRNA interactions have been substantially documented38 and is a prime area of ongoing research. Similarly, miRNA- miRNA interactions through mutual co-expression39, via transcription factor40, and miRNA-disease associations6 have also been reported. However, miRNA-miRNA interactions towards identification of a core miRNA-miRNA module that could potentially be a signature component for a particular disease have not been studied enough. Many studies have used computational approaches to study this aspect. A miRNA-miRNA co-regulation network in lung cancer was identified using a progressive data refining approach20. Similarly, miRNA expression profiling along with a genome-wide SNP approach was used to create a miRNA-miRNA synergistic network to study coronary artery disease41. miRNA-miRNA interactions were also identified in esophageal cancer using K-clique analysis on a bipartite network consisting of miRNAs and subpathways42. Additionally, miRNA-target interactions were integrated with miRNA and mRNA expressions to deduce miRNA-miRNA interactions in prostate cancer43. A network topological approach was also undertaken to identify disease miRNAs by constructing a miRNA-miRNA synergistic network consisting of co-regulating functional modules44.
In this work, we adopted a strategy that takes a miRNA expression profile and uses six different network inference algorithms (CLR33, MRNETB34, Basic Correlation (Pearson and Spearman), DC35, GENIE336), each varying in their inference strategies, integrated with a consensus approach and graph intersection to identify the conserved miRNA-miRNA interaction signature across a group of diseases (cancers, in this case). The identified signatures were validated via manual literature search and were found to be associated within the classes of the selected cancers, demonstrating the efficacy of the method. Under validation, we retrieved the PMIDs reporting the associations from the PhenomiR database and also performed a manual literature search in the PubMed Central database to separately corroborate our results, as displayed in Fig. 7.
Our results show that, the expression profile of hsa-mir-30a, hsa-mir-181a-1, and hsa-mir-29c could be a signature for gastrointestinal cancers that comprises of esophageal, gastroesophageal, gastrointestinal, gastric, and colorectal cancers (Fig. 7). These miRNAs are already reported to be associated with these cancers45,46,47,48. miRNAs (hsa-mir-30a, hsa-mir-29c, hsa-mir-181a-1) displayed the same trend of expression in a study of esophageal adenocarcinoma (EAC) and Barrett’s esophagus (BE) and were differentially up-regulated in both the disease tissues. hsa-mir-181a and hsa-mir-29c showed higher expression levels in EAC to that of BE with high grade dysplasia48. Studies have also reported hsa-mir-181a, hsa-mir-30a and hsa-mir-29c being overexpressed in esophagela carcinoma (EC) and hsa-mir-29c to be underexpressed in EC49,50 and therefore, this group of miRNAs may be considered for developing a pan-diagnostic tool for the aforementioned cancers.
We identified that hsa-mir-221, hsa-mir-222, hsa-mir-155, hsa-mir-224, hsa-mir-181a-1, and hsa-mir-181b-1 make the signature for endocrine cancers (hepatocellular, pancreatic, and thyroid cancers) (Fig. 7). Reports suggest that these miRNAs are predominantly associated with this group of cancers51,52,53,54. In another study analyzing molecular signatures for aggressive pancreatic cancer, all the miRNAs (hsa-mir-221, hsa-mir-222, hsa-mir-155, hsa-mir-224, hsa-mir-181a-1, and hsa-mir-181b-1) were significantly altered due to chronic exposure to conventional anti-cancer drugs55. A large-scale meta-analysis investigating candidate miRNA biomarkers for pancreatic ductal adenocarcinoma (PDAC) across eleven miRNA expression profiling studies, reported all the miRNAs to be up-regulated and having a consistent direction of change. miRNAs hsa-mir-221, hsa-mir-222, hsa-mir-155 were reported to be upregulated together in at least five of these studies with a consistent direction. Among them, miRNAs hsa-mir-221, hsa-mir-155 were identified as part of a meta-signature and biomarkers for PDAC56. Studies also report all these miRNAs to be associated with lung cancer57. Thus this set of miRNAs may be used/tested as a diagnostic tool for all the endocrine cancers considered here.
Seven miRNAs (hsa-mir-29b-1, hsa-mir-146a, hsa-mir-20a, hsa-mir-126, hsa-mir-99a, hsa-mir-199b and hsa-mir-130a) that are well documented for their association with various kinds of leukemia54,58,59,60,61,62,63 are found to form the signature component of leukemia from our analysis (Fig. 7). miRNAs (hsa-mir-29b-1, hsa-mir-20a, hsa-mir-126, hsa-mir-146a, hsa-mir-199b) were differentially expressed in a blood stem cell study in which the blood stem cells were treated with plerixafor and granulocyte colony-stimulating factor. The miRNAs were recorded to be expressed in this treated cell study analyzing acute lymphocytic leukemia conditions64. miRNAs (hsa-mir-126, hsa-mir-130a, hsa-mir-99a, hsa-mir-146a, hsa-mir-199b) have also been reported to express together in a myeloid cell study exploring transcription factor binding site motifs65. Therefore, this signature group of miRNAs can be potentially used as a screening or diagnostic tool for a range of different types of leukemia.
In case of neurone cancers (neuroblastoma, medulloblastoma, and glioblastoma) we detected two signatures: i) hsa-mir-323, hsa-mir-129-1, hsa-mir-137, hsa-mir-330, hsa-mir-149, hsa-mir-107, hsa-mir-30c-1, hsa-mir-181b-1 and ii) hsa-mir-30b, hsa-mir-331, hsa-mir-150, hsa-let-7a-1 (Fig. 7). Regarding the first signature network component, hsa-mir-137, hsa-mir-330, hsa-mir-149, hsa-mir-107, hsa-mir-181b were among the miRNAs whose experimentally validated targets (such as CTBP1, CDC42, CDK6, E2F1, VEGFA, AKT1, KAT2B) affect the pathways which play a crucial role in glioblastoma biology. Deregulations of hsa-mir-137, hsa-mir-330 and hsa-mir-149 lead to effects in the glioma de novo pathway, VEGF signaling pathway and Notch signaling pathway66. Among the miRNAs reported in the second signature component, hsa-mir-330 and hsa-mir-30b are among the top ten miRNAs having least coefficient of variation in the expression of benign kidney tumor and hsa-mir-150 is differentially expressed in metastatic clear cell renal cell carcinoma67.
Comparing our results with other similar works has been challenging, primarily because there are not many studies that have reported direct miRNA-miRNA co-regulations across these disease classes. Similar studies13,20,68 have used different disease and miRNA data sets which makes a one-to-one comparison challenging. In some previous works, miRNA-miRNA regulatory associations have been deduced based on the semantic similarities between the associated diseases69 and based on the analysis of shared transcription factors, common targets, KEGG pathway analysis and corroboration from literature20. However, none of these methods allow for a network-level miRNA-miRNA analysis for a variety of diseases and hence cannot be used for comparison purposes to the predicted interaction networks in this paper.
Online analysis and visualization of results is an aid to the research community. Along these lines, several network analysis and visualization tools have been developed, such as VisANT for integrative online visual analysis of biological networks and pathways70, miRegulome for miRNA regulome visualization and analysis12 and miRNet for functional analysis of miRNAs within a high-performance network visual analytics system71 among others. However, no tool is available so far which can perform an online visualization and analysis of signature miRNAs across multiple diseases. The miRsig tool developed here bridges this gap and provides an intuitive analysis and visualization of core/signature miRNA-miRNA interaction components for several diseases.
Conclusion
In this work, we have developed a novel consensus-based network analysis pipeline to identify disease-specific miRNA-miRNA interactions by combining the expression profiles of various miRNAs in specific diseases. This method can effectively identify the signature/core miRNA-miRNA interactions for a group of diseases; here tested on cancer. These signature miRNAs may have potential use for diagnostic, prognostic, or therapeutic applications for a group of related diseases such as cancers. The predicted miRNA-miRNA signature patterns were extensively validated by the PMIDs reported in the PhenomiR database as well as an independent manual literature search from PubMed Central. miRsig thus provides a powerful prediction and visualization tool for the identification of core/signature miRNA-miRNA interactions amongst a number of diseases. Our future work includes investigating the (i) miRNAsameDiseasedifferent category of interactions to study the dynamics of similar miRNAs across multiple diseases and also (ii) the miRNAdifferentDiseasedifferent category of interactions to understand the evolution of diseases based on the underlying miRNA expression patterns. As miRNAs may potentially serve as biomarkers for a wide variety of diseases, our proposed pipeline may motivate the study of several interesting questions both for particular diseases or across multiple diseases.
Additional Information
How to cite this article: Nalluri, J. J. et al. miRsig: a consensus-based network inference methodology to identify pan-cancer miRNA-miRNA interaction signatures. Sci. Rep. 7, 39684; doi: 10.1038/srep39684 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Filipowicz, W., Bhattacharyya, S. N. & Sonenberg, N. Mechanisms of post-transcriptional regulation by micrornas: are the answers in sight? Nature Reviews Genetics 9, 102–114 (2008).
Place, R. F., Li, L.-C., Pookot, D., Noonan, E. J. & Dahiya, R. Microrna-373 induces expression of genes with complementary promoter sequences. Proceedings of the National Academy of Sciences 105, 1608–1613 (2008).
Saraiya, A. A., Li, W. & Wang, C. C. Correction: Transition of a microrna from repressing to activating translation depending on the extent of base pairing with the target. PloS one 8 (2013).
Tüfekci, K. U., Meuwissen, R. L. J. & Genç, Ş. The role of micrornas in biological processes. miRNomics: MicroRNA Biology and Computational Analysis 15–31 (2014).
Blenkiron, C. & Miska, E. A. mirnas in cancer: approaches, aetiology, diagnostics and therapy. Human molecular genetics 16, R106–R113 (2007).
Ardekani, M. A. & Moslemi Naeini, M. The role of micrornas in human diseases. Avicenna journal of medical biotechnology 2, 161–180 (2011).
Ye, S. et al. Bioinformatics method to predict two regulation mechanism: Tf-mirna-mrna and lncrna-mirna-mrna in pancreatic cancer. Cell biochemistry and biophysics 70, 1849–1858 (2014).
Barh, D., Malhotra, R., Ravi, B. & Sindhurani, P. Microrna let-7: an emerging next-generation cancer therapeutic. Current Oncology 17, 70 (2010).
Tang, R. et al. Mouse mirna-709 directly regulates mirna-15a/16-1 biogenesis at the posttranscriptional level in the nucleus: evidence for a microrna hierarchy system. Cell research 22, 504–515 (2012).
Shi, B., Zhu, M., Liu, S. & Zhang, M. Highly ordered architecture of microrna cluster. BioMed research international 2013 (2013).
Lu, M. et al. An analysis of human microrna and disease associations. PloS one 3, e3420 (2008).
Barh, D. et al. miregulome: a knowledge-base of mirna regulomics and analysis. Scientific reports 5 (2015).
Hamilton, M. P. et al. Identification of a pan-cancer oncogenic microrna superfamily anchored by a central core seed motif. Nature communications 4 (2013).
Yuan, D. et al. Enrichment analysis identifies functional microrna-disease associations in humans. PloS one 10, e0136285 (2015).
Zou, Q. et al. Prediction of microrna-disease associations based on social network analysis methods. BioMed research international 2015, 810514 (2015).
Chen, H. & Zhang, Z. Similarity-based methods for potential human microrna-disease association prediction. BMC medical genomics 6, 12 (2013).
Liao, B., Ding, S., Chen, H., Li, Z. & Cai, L. Identifying human microrna-disease associations by a new diffusion-based method. Journal of bioinformatics and computational biology 13, 1550014 (2015).
Shi, H. et al. Integration of multiple genomic and phenotype data to infer novel mirna-disease associations. PloS one 11, e0148521 (2016).
Yang, J.-H. & Qu, L.-H. Discovery of microrna regulatory networks by integrating multidimensional high-throughput data. In MicroRNA Cancer Regulation 251–266 (Springer, 2013).
Song, R., Catchpoole, D. R., Kennedy, P. J. & Li, J. Identification of lung cancer mirna-mirna co-regulation networks through a progressive data refining approach. Journal of Theoretical Biology (2015).
Xu, J. et al. Prioritizing candidate disease mirnas by topological features in the mirna target-dysregulated network: Case study of prostate cancer. Molecular cancer therapeutics 10, 1857–1866 (2011).
Xu, C. et al. Prioritizing candidate disease mirnas by integrating phenotype associations of multiple diseases with matched mirna and mrna expression profiles. Mol. BioSyst. 10, 2800–2809 (2014).
Yoon, S. & De Micheli, G. Prediction of regulatory modules comprising micrornas and target genes. Bioinformatics 21, ii93–ii100 (2005).
Bandyopadhyay, S., Mitra, R., Maulik, U. & Zhang, M. Q. Development of the human cancer microrna network. Silence 1, 1 (2010).
Nalluri, J. J. et al. Dismira: Prioritization of disease candidates in mirna-disease associations based on maximum weighted matching inference model and motif-based analysis. BMC Genomics 16, S12 (2015).
Pavlopoulos, G. A. et al. Using graph theory to analyze biological networks. BioData mining 4, 1 (2011).
Cho, H., Berger, B. & Peng, J. Reconstructing causal biological networks through active learning. PloS one 11, e0150611 (2016).
Nagarajan, N. & Kingsford, C. Giraf: robust, computational identification of influenza reassortments via graph mining. Nucleic acids research gkq1232 (2010).
Pati, A., Vasquez-Robinet, C., Heath, L. S., Grene, R. & Murali, T. Xcisclique: analysis of regulatory bicliques. BMC bioinformatics 7, 1 (2006).
Ruepp, A. et al. Phenomir: a knowledgebase for microrna expression in diseases and biological processes. Genome biology 11, R6 (2010).
Surowiecki, J. The wisdom of crowds (Anchor, 2005).
Marbach, D. et al. Wisdom of crowds for robust gene network inference. Nature methods 9, 796–804 (2012).
Faith, J. J. et al. Large-scale mapping and validation of escherichia coli transcriptional regulation from a compendium of expression profiles. PLoS biology 5, e8 (2007).
Meyer, P., Marbach, D., Roy, S. & Kellis, M. Information-theoretic inference of gene networks using backward elimination. In Biocomp 700–705 (2010).
Guo, X., Zhang, Y., Hu, W., Tan, H. & Wang, X. Inferring nonlinear gene regulatory networks from gene expression data based on distance correlation. PloS one 9, e87446 (2014).
Irrthum, A., Wehenkel, L., Geurts, P. et al. Inferring regulatory networks from expression data using tree-based methods. PloS one 5, e12776 (2010).
Bostock, M. Data-driven documents. http:https://d3js.org/. Accessed: 2016-04-18.
Afonso-Grunz, F. & Müller, S. Principles of mirna-mrna interactions: Beyond sequence complementarity. Cellular and Molecular Life Sciences 72, 3127–3141 (2015).
Guo, L., Sun, B., Wu, Q., Yang, S. & Chen, F. mirna-mirna interaction implicates for potential mutual regulatory pattern. Gene 511, 187–194 (2012).
Arora, S., Rana, R., Chhabra, A., Jaiswal, A. & Rani, V. mirna-transcription factor interactions: a combinatorial regulation of gene expression. Molecular genetics and genomics 288, 77–87 (2013).
Hua, L., Xia, H., Zhou, P., Li, D. & Li, L. Combination of microrna expression profiling with genome-wide snp genotyping to construct a coronary artery disease-related mirna-mirna synergistic network. Bioscience trends 8, 297–307 (2014).
Wu, B. et al. Dissection of mirna-mirna interaction in esophageal squamous cell carcinoma. PloS one 8, e73191 (2013).
Alshalalfa, M. Microrna response elements-mediated mirna-mirna interactions in prostate cancer. Advances in bioinformatics 2012 (2012).
Xu, J. et al. Mirna-mirna synergistic network: construction via co-regulating functional modules and disease mirna topological features. Nucleic acids research 39, 825–836 (2011).
Monzo, M. et al. Overlapping expression of micrornas in human embryonic colon and colorectal cancer. Cell research 18, 823–833 (2008).
Ueda, T. et al. Relation between microrna expression and progression and prognosis of gastric cancer: a microrna expression analysis. The lancet oncology 11, 136–146 (2010).
Zhang, Y. et al. Profiling of 95 micrornas in pancreatic cancer cell lines and surgical specimens by real-time pcr analysis. World journal of surgery 33, 698–709 (2009).
Yang, H. et al. Microrna expression signatures in barrett’s esophagus and esophageal adenocarcinoma. Clinical Cancer Research 15, 5744–5752 (2009).
Guo, Y. et al. Distinctive microrna profiles relating to patient survival in esophageal squamous cell carcinoma. Cancer research 68, 26–33 (2008).
Zhou, S.-L. & Wang, L.-D. Circulating micrornas: novel biomarkers for esophageal cancer. World J Gastroenterol 16, 2348–2354 (2010).
Jiang, J. et al. Association of microrna expression in hepatocellular carcinomas with hepatitis infection, cirrhosis, and patient survival. Clinical Cancer Research 14, 419–427 (2008).
Roldo, C. et al. Microrna expression abnormalities in pancreatic endocrine and acinar tumors are associated with distinctive pathologic features and clinical behavior. Journal of Clinical Oncology 24, 4677–4684 (2006).
Nikiforova, M. N., Tseng, G. C., Steward, D., Diorio, D. & Nikiforov, Y. E. Microrna expression profiling of thyroid tumors: biological significance and diagnostic utility. The Journal of Clinical Endocrinology & Metabolism 93, 1600–1608 (2008).
Jiang, J., Lee, E. J., Gusev, Y. & Schmittgen, T. D. Real-time expression profiling of microrna precursors in human cancer cell lines. Nucleic acids research 33, 5394–5403 (2005).
Ali, S., Almhanna, K., Chen, W., Philip, P. A. & Sarkar, F. H. Differentially expressed mirnas in the plasma may provide a molecular signature for aggressive pancreatic cancer. Am J Transl Res 3, 28–47 (2010).
Ma, M.-Z. et al. Candidate microrna biomarkers of pancreatic ductal adenocarcinoma: meta-analysis, experimental validation and clinical significance. Journal of Experimental & Clinical Cancer Research 32, 1 (2013).
Leidinger, P., Keller, A. & Meese, E. Micrornas-important molecules in lung cancer research. Frontiers in genetics 2, 104 (2012).
Zanette, D. et al. mirna expression profiles in chronic lymphocytic and acute lymphocytic leukemia. Brazilian Journal of Medical and Biological Research 40, 1435–1440 (2007).
Garzon, R. et al. Microrna signatures associated with cytogenetics and prognosis in acute myeloid leukemia. Blood 111, 3183–3189 (2008).
Garzon, R. et al. Distinctive microrna signature of acute myeloid leukemia bearing cytoplasmic mutated nucleophosmin. Proceedings of the National Academy of Sciences 105, 3945–3950 (2008).
Colquhoun, J. A. S. With the Kurram Field Force, 1878-79 (WH Allen & Company, 1881).
Contreras, J. R. et al. Microrna-146a modulates b-cell oncogenesis by regulating egr1. Oncotarget 6, 11023 (2015).
Favreau, A. J., McGlauflin, R. E., Duarte, C. W. & Sathyanarayana, P. mir-199b, a novel tumor suppressor mirna in acute myeloid leukemia with prognostic implications. Experimental hematology & oncology 5, 1 (2016).
Donahue, R. E. et al. Plerixafor (amd3100) and granulocyte colony-stimulating factor (g-csf) mobilize different cd34+ cell populations based on global gene and microrna expression signatures. Blood 114, 2530–2541 (2009).
Jansen, B. J. et al. Microrna genes preferentially expressed in dendritic cells contain sites for conserved transcription factor binding motifs in their promoters. BMC genomics 12, 1 (2011).
Singh, S. K., Vartanian, A., Burrell, K. & Zadeh, G. A microrna link to glioblastoma heterogeneity. Cancers 4, 846–872 (2012).
Wu, X. et al. Identification of a 4-microrna signature for clear cell renal cell carcinoma metastasis and prognosis. PloS one 7, e35661 (2012).
Bandyopadhyay, S. & Bhattacharyya, M. Analyzing mirna co-expression networks to explore tf-mirna regulation. BMC bioinformatics 10, 163 (2009).
Wang, D., Wang, J., Lu, M., Song, F. & Cui, Q. Inferring the human microrna functional similarity and functional network based on microrna-associated diseases. Bioinformatics 26, 1644–1650 (2010).
Hu, Z. et al. Visant 4.0: Integrative network platform to connect genes, drugs, diseases and therapies. Nucleic acids research 41, W225–W231 (2013).
Lab, X. miRNet, network-based visual analysis of miRNAs, targets and functions. http://www.mirnet.ca/ (2015). [Online; Last accessed 4-April-2016].
Acknowledgements
We thank Daniel Marbach, Ph.D. for promptly responding to our queries regarding the network inference tools hosted on the GenePattern website.
Author information
Authors and Affiliations
Contributions
J.J.N. and P.G. conceived, conceptualized and implemented the computational methods. D.B. and V.A. provided biological insights and validated miRsig tool. J.J.N., D.B. and P.G. wrote the manuscript. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Nalluri, J., Barh, D., Azevedo, V. et al. miRsig: a consensus-based network inference methodology to identify pan-cancer miRNA-miRNA interaction signatures. Sci Rep 7, 39684 (2017). https://doi.org/10.1038/srep39684
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep39684
This article is cited by
-
Neuron enriched extracellular vesicles’ MicroRNA expression profiles as a marker of early life alcohol consumption
Translational Psychiatry (2024)
-
Network pharmacology-based strategy to investigate the effect and mechanism of α-solanine against glioma
BMC Complementary Medicine and Therapies (2023)
-
Non-coding RNAs in human health and disease: potential function as biomarkers and therapeutic targets
Functional & Integrative Genomics (2023)
-
Determining causal miRNAs and their signaling cascade in diseases using an influence diffusion model
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.