Abstract
Carbaryl (1-naphthyl N-methylcarbamate) is a most widely used carbamate pesticide in the agriculture field. Soil isolate, Pseudomonas sp. strain C5pp mineralizes carbaryl via 1-naphthol, salicylate and gentisate, however the genetic organization and evolutionary events of acquisition and assembly of pathway have not yet been studied. The draft genome analysis of strain C5pp reveals that the carbaryl catabolic genes are organized into three putative operons, ‘upper’, ‘middle’ and ‘lower’. The sequence and functional analysis led to identification of new genes encoding: i) hitherto unidentified 1-naphthol 2-hydroxylase, sharing a common ancestry with 2,4-dichlorophenol monooxygenase; ii) carbaryl hydrolase, a member of a new family of esterase; and iii) 1,2-dihydroxy naphthalene dioxygenase, uncharacterized type-II extradiol dioxygenase. The ‘upper’ pathway genes were present as a part of a integron while the ‘middle’ and ‘lower’ pathway genes were present as two distinct class-I composite transposons. These findings suggest the role of horizontal gene transfer event(s) in the acquisition and evolution of the carbaryl degradation pathway in strain C5pp. The study presents an example of assembly of degradation pathway for carbaryl.
Similar content being viewed by others
Introduction
The advent of industrialization has led to release of anthropogenic chemicals into the environment imposing a selective pressure on microbes. This pressure is countered by displaying the high propensity for rapid evolution of novel metabolic pathways as well as its spread through facile horizontal transfer of catabolic genes within the microbial population1,2,3,4,5,6,7. Carbaryl (1-naphthyl N-methylcarbamate), a carbamate class of compound, is the third most widely used broad-spectrum insecticide in the agriculture field since 1960. The insecticidal activity is due to the ester bond which inhibits acetylcholine esterase competitively8. Microorganisms have been reported to utilize carbaryl9,10,11,12,13,14,15,16, and the complete pathway has been demonstrated at the functional level in soil isolates, Pseudomonas sp. strains C4, C5 and C615. Carbaryl is metabolized to the central carbon cycle intermediates via 1-naphthol, 1,2-dihydroxynaphthalene, salicylate and gentisate15. Based on the metabolic studies, the pathway has been hypothesized to be organized into ‘upper’ (carbaryl to salicylate), ‘middle’ (salicylate to gentisate) and ‘lower’ (gentisate to TCA cycle intermediate) segments17. The detailed genetic organization and evolutionary origin of the carbaryl degradation pathway have not yet been reported. The enzyme, 1-naphthol 2-hydroxylase (1NH) responsible for the conversion of 1-naphthol to 1,2-dihydroxynaphthalene in the upper pathway, has been purified and characterized at kinetic level from various carbaryl degrading Pseudomonas sp.18,19,20, however the gene encoding 1NH has not been reported so far.
Microorganisms have been exposed to carbaryl since 1960 s. Pseudomonas sp. strain C5pp draft genome analysis provides a unique opportunity to explore the probable ancestral origin of various genes and evolutionary events responsible for the evolution of carbaryl metabolic pathway. In the present study, we report the genetic organization of degradation genes and functional identification of carbaryl hydrolase (CH), 1-naphthol 2-hydroxylase (1NH) and 1,2-dihydroxynaphthalene dioxygenase (12DHNDO) involved in carbaryl metabolism in Pseudomonas sp. strain C5pp (formerly known as strain C5). Based on the analysis of Supercontig-A, we hypothesize that the degradation property must have been acquired through horizontal gene transfer (HGT) which has further evolved to degrade carbaryl as the carbon source more efficiently.
Results and Discussion
Construction and functional screening of genomic DNA library
The genomic DNA library of Pseudomonas sp. strain C5pp was constructed in CopyControl fosmid pCC2FOS with a phage titer of 3 × 106 CFU.ml−1. The G1 clone, with the ability to degrade gentisate as the carbon source, was digested with BamHI, partially sequenced and annotated (Table S1).
E. coli EPI300 cells harboring fosmid clone G1-DNA (here onward referred as G1 cells) showed a good growth (O.D540 nm = 0.8 in 24 h) in M9 medium containing gentisate as the sole carbon source. The G1-DNA showed amplification of genes viz. salicylaldehyde dehydrogenase (SalDH, 0.5 kb), salicylate 5-hydroxylase (S5H) β-subunit (0.5 kb) and gentisate dioxygenase (GDO, 0.25 kb) by PCR using primers reported earlier17, however G1 cells failed to grow on carbaryl or salicylate. The cell-free extract (CFE) prepared from G1 cells grown on M9-glucose showed significantly lower activity of GDO (1 nmole.min−1.mg−1) as compared to M9-gentisate (276 nmole.min−1.mg−1) suggesting the inducible nature of GDO. The CFE prepared from G1 (Fig. 1A) as well as strain C5pp (Fig. 1B) cells in the presence of gentisate (100 μM) showed the formation of maleylpyruvate (~47 μM) by the action of GDO. Addition of GSH led to isomerization of maleylpyruvate to fumarylpyruvate by maleylpyruvate isomerase (MPI) and further conversion of fumarylpyruvate to fumarate and pyruvate by fumarylpyruvate hydrolase (FPH, Fig. 2A). The carbon-source dependent enzyme activity and induction studies suggest that the G1-DNA harbors essential ‘lower’ pathway, gentisate metabolic genes along with regulatory elements, thus enabling E. coli to use gentisate as the sole source of carbon and energy. Inability of G1 cells to utilize carbaryl is due to lack carbaryl hydrolase (CH) and 1,2-dihydroxy naphantlene dioxygenase (12DHNDO, Fig. 2B). Further cells failed to utilize salicylate, this is probably due to inability of mcbH regulatory element to express or function in E. coli.
Time-dependent spectral changes observed in the cell-free extracts prepared from cells of (A) E. coli harboring G1 fosmid DNA (G1 cells) grown on gentisate (0.1%) and (B) Pseudomonas sp. strain C5pp grown on carbaryl (0.1%) as the carbon source. The enzyme reactions (Phosphate buffer, 50 mM, pH 7.5; gentisate 100 μM and 100 μg of total protein) were scanned from 240–400 nm at every 5 min interval (solid lines). After 20 or 15 min of reaction, glutathione (1 mM) was added in the reaction mixture and spectral changes were recorded at interval of 1 min (dashed lines). The solid lines in both panel represents gentisate dioxygenase (GDO) mediated conversion of gentisate to maleylpyruvate (increase in absorbance at 330 nm, ε340nm = 13,000 M−1.cm−1). The dashed lines represent the spectral scans observed after addition of glutathione. The observed shift in the absorption peak at 340 nm was due to conversion of maleylpyruvate to fumarylpyruvate by GSH-dependent maleylpyruvate isomerase (MPI). Further, the observed decrease in the absorbance at 340 nm was due to hydrolysis of fumarylpyruvate to fumarate and pyruvate by fumarate-pyruvate hydrolase (FPH). Based on the metabolic studies, the gentisate metabolic pathway in strain C5pp and G1 cells is proposed in panel.
The carbaryl degradation pathway in Pseudomonas sp. strain C5pp.
The metabolic steps involved in the degradation of carbaryl are depicted in panel (A). The arrangement of genes on the Supercontig-A involved in the carbaryl metabolism is represented in panel (B). The arrow indicates the direction of gene transcription, numbers indicate the gene length in bp and the number in parenthesis indicate the intergenic distances in bp. mge indicates the genes encoding various mobile genetic elements. The details of genes are given in Table 1. The ‘upper’ pathway gens are marked with pink, ‘middle’ pathway with mango yellow and ‘lower’ pathway with green color. The probable functional regulators are marked in blue color. Regulator genes probably not related to the carbaryl metabolism are marked in grey color. ‘RTase’ and ‘endase’ indicates reverse transcriptase and endonuclease, respectively. Supercontig-A consist of contigs from the draft genome sequence of strain C5pp in the order 83-68-92-62-76-61-76-47. The G1 DNA is mapped on Supercontig-A from 17,600 bp to 60,826 bp and is marked by filled arrow head.
Genome analysis and validation of genes involved in the carbaryl degradation
The draft genome features and the statistics of Pseudomonas sp. strain C5pp (JWLN00000000.1 ref. 21) are summarized in Fig. S1. The sequences obtained from six sub-clones of G1-fosmid DNA (Table S1), when used as a query against C5pp draft genome, retrieved contig 47 (32.72 kb, ‘lower’ pathway), 62 (13.65 kb, ‘upper’ pathway), 61 (14.04 kb, ‘middle’ pathway) and 76 (2.64 kb). Gaps present between these contigs were filled by primer walking and gap filling PCR reactions (Table S2). The assembled contig (76334 bp) here onward is referred as Supercontig-A (KU522233). The genomic region covered in G1-fosmid (~40 kb) is depicted in Fig. 2B and annotation details are summarized in Table S3. Genes involved in the carbaryl degradation and their homology/identity and arrangement are summarized in Table 1 and Fig. 2.
The activities of CH, 1NH, 12DHNDO, SalDH and GDO from strain C5pp has been demonstrated earlier15,22. NCBI annotation and BLASTp analysis of Supercontig-A identifies following genes with putative functions: i) mcbD, 2-hydroxychromene 2-carboxyl isomerase; mcbE, trans-o-hydroxybenzilidinepyruvate aldolase-hydratase; and mcbF, SalDH involved in the ‘upper’ pathway; ii) mcbIJKL, salicylate 5-hydroxylase involved in the ‘middle’ pathway; and iii) mcbO, GDO; mcbP, FPH; mcbQ, MPI involved in the ‘lower’ pathway of carbaryl metabolism (Fig. 2, Table 1). Beside these genes, it was also found to harbor mcbA, mcbB and mcbC which were analyzed by sequence and function based approaches to ascertain their role in the carbaryl degradation.
mcbA encodes carbaryl hydrolase
The partially purified CH from strain C5pp was found to be a monomeric protein with mol. wt. of ~80 kDa (Fig. S2). BLASTp analysis of McbA (769 a.a.) annotated as hypothetical protein (Table 1) displayed 24% identity with CehA from Rhizobium sp. AC100 23. The predicted molecular mass of McbA (83 kDa, Expasy), was found to be similar to that observed for partially purified CH from strain C5pp (Fig. S2) hence, this hypothetical protein was predicted to be a putative CH. The mcbA was cloned into pET-28a(+) and expressed into E. coli (Fig. S3A). The CFE (soluble fraction) prepared from IPTG-induced E. coli harboring pET28a-CH showed a CH activity of 28 nmole.min−1.mg−1 as compared to uninduced cells (11 nmole.min−1.mg−1). Time-dependent spectral scan showed the decrease in the absorbance at 280 nm with concomitant increase in the absorbance at 322 nm due to formation of 1-naphthol (Fig. 3). The observed spectral changes were similar to those reported from wild type strain C5pp15. In HPLC analysis, besides carbaryl peak (RT 4.4 min), the reaction mixture gave an additional peak (RT 5.2 min) corresponding to the standard 1-naphthol (Fig. S3B–E). Carbaryl possesses an amide as well as ester linkage, hence hydrolysis can be catalyzed either by esterase or amidase type of enzyme to yield 1-naphthol. The CH from Rhizobium sp. strain AC10023 and Arthrobacter sp. strain RC10024 was reported to act as esterase and amidase, respectively. To determine the mode of hydrolysis (either as esterase or amidase type), the activity was tested on 1-naphthylacetate23,24 or 1-naphthalene acetamide. Recombinant CH from the soluble fraction was found to be active on carbaryl (16 nmole.min−1.mg−1, 100%), 1-naphthylacetate (5.7 nmole.min−1.mg−1, 36%) but not on 1-naphthalene acetamide. Similar results were obtained for the partially purified CH from strain C5pp (data not shown), suggesting that the enzyme from strain C5pp is an esterase type CH.
Functional analysis of carbaryl hydrolase (rCH).
Time-dependent spectral changes observed in the CFE prepared from E. coli BL21(DE3) cells induced with IPTG harboring pET28-CH. The enzyme reaction was scanned from 200-340 nm every 3 min interval for 10 cycles with carbaryl (400 μM) as the substrate. The decrease in the absorbance at 280 nm (down arrow) and increase in the absorbance peak at 322 nm (up arrow) indicated the conversion of carbaryl to 1-naphthol, respectively. The crossover point at 290 nm indicates the isobestic point for the conversion of the carbaryl into 1-naphthol. The lysate from E. coli carrying vector alone failed to show any increase in the absorbance at 322 nm over the course of 30 min of incubation indicating absence of CH activity in E. coli.
Significant divergence at the functional (amidase or esterase) as well as at the sequence level has been observed for CH. The phylogenetic analysis revealed the clustering of CH from strain C5pp with esterase type of enzymes. The amino acid sequences of CH from strain C5pp and Rhizobium sp. AC100 were compared with functionally characterized esterase belonging to fifteen different families. The functionally characterized CHs clustered together and showed unique conserved motif {W-X-S-[AGST]-D-X-H-[ILV]-H-[AIL]-X(3)-[APST]} suggesting a new family of esterases (Fig. S4 and Fig. S5).
mcbB encodes 1,2-dihydroxynaphthalene dioxygenase
The BLASTp analysis of McbB (275 a.a., ~30 kDa) retrieved homologs which belong to type-II extradiol dioxygenase (EDO). Phenanthrene and naphthalene degrading Burkholderia sp. was reported to harbor phnC gene (828 bp, 275 a.a.) which encodes type-II EDO with activity on 1,2-dihydroxynaphthalene25. So far, the reported 12DHNDOs are 302 a.a. long, moderately conserved and found to be a member of type-I EDO catalyzing the ring-fission of 1,2-dihydroxynaphthalene to 2-hydroxychromene 2-carboxylate. Thus it was speculated that the protein McbB, encoded by mcbB in strain C5pp might catalyze the ring-fission of 1,2-dihydroxynaphthalene. The sequence alignment of the putative McbB with 12DHNDO from Burkholeria sp. RP007 showed 68% identity and three conserved motifs- DHY, DHG and GXSH (Fig. S6A) essential for the ring-cleavage activity26. The phylogenetic analysis reveals that McbB clustered with type-II EDO of Burkholderia and Ralstonia (Fig. 4). So far, there are no reports on 12DHNDO from Pseudomonas sp. which belongs to type-II EDO.
The phylogenetic analysis of 1,2-dihydroxynaphthalene dioxygenase (12DHNDO) from strain C5pp with representative members of three types of EDO
. The EDO from strain C5pp clusters with type II EDO which includes functionally characterized PhnC from Burkholderia. The numbers in parentheses indicates the protein accession id. Enzyme abbreviations: 12DHNDO, 1,2-dihydroxynaphthalene dioxygenase; BphC1, 2,3-dihydroxybiphenyl 1,2 dioxygenase; HPCD, homoprotocatechuate 2,3-dioxygenase; C23DO, catechol 2,3-dioxygenase; PcpA, 2,6-dichlorohydroquinone 1,2-dioxygenase; QueD, quercetin 2,3-dioxygenase; GDO, gentisate 1,2-dioxygenase; P23DO, protocatechuate 2,3-dioxygenase; MhpB, 2,3-dihydroxyphenylpropionate 1,2-dioxygenase; LigAB, protocatechuate 4,5-dioxygenase.
To validate that mcbB encodes 12DHNDO, the gene was cloned into pET-28a(+) and over-expressed in E. coli (Fig. S6B). The CFE of IPTG-induced E. coli cells harboring pET28a-12DHNDO construct showed 12DHNDO activity (95 nmole.min−1.mg−1). Partially purified enzyme (Fig. S6B) showed the specific activity of 1.9 μmole.min−1.mg−1 with 1,2-dihydroxy naphthalene as the substrate. The observed activity for the partially purified r12DHNDO appears to be low as compared to purified 12DHNDO from Pseudomonas putida (75 μmole.min−1.mg−1, type-I EDO27) and r12DHNDO form Burkholderia sp RP007 (564 μmole.min−1.mg−1, type-II EDO25). Though NCBI server annotated mcbB as protocatechuate 3,4-dioxygenase, the partially purified enzyme failed to show activity on protocatechuate, suggesting that mcbB codes for 12DHNDO.
mcbC encodes 1-naphthol 2-hydroxylase
1NH belongs to class-A monooxygenase which consist of single component external flavoproteins requiring NAD(P)H as electron donor18,19,20,28. However, the gene encoding 1NH has not been reported till date. N-terminal (MLXNIFLKDE) and partial peptide (FTLLTGIGGEGWIR) sequences of 1-NH purified from strain C5pp showed identity (100%) with amino acid sequence derived from mcbC (1773 bp, 590 a.a.) suggesting the gene probably encodes 1NH (Fig. 2). The RAST analysis29 predicted McbC of strain C5pp to be a 2,4-dichlorophenol monooxygenase (24DCPM, Table 1). The mcbC gene was cloned into pET-28a(+) and over-expressed in E. coli (Fig. S7A). The CFE of IPTG-induced E. coli cells harboring pET28a-1NH construct showed 1.78 μmole.min−1.mg−1 1NH activity. The recombinant 1NH (r1NH) was purified to homogeneity (Fig. S7B, specific activity of 8.2 μmole.min−1.mg−1; fold purification 3.5 and yield 33%) by Ni-NTA followed by Sephacryl S-200HR gel filtration chromatography (homodimeric protein, native mol. wt. of ~145 kDa and subunit mol. wt. of ~66 kDa; Fig. 5A). The MALDI-TOF/TOF analysis of r1NH and 1NH purified from strain C5pp confirmed that both proteins are identical (Fig. S7C). The observed molecular, kinetic properties and substrate specificities (Fig. 5B) are similar to those reported for wild type 1NH from strain C5pp20. Phylogenetic study showed 1NH to be related to monooxygenases acting on phenol and its derivatives (Fig. 5C). r1NH showed activity on 2,4-dichlorophenol (47%) as compared to activity on 1-naphthol (100%), but failed to hydroxylate phenol (Fig. 5B). Results suggests the identification of a new gene mcbC encoding 1NH in Pseudomonas sp. strain C5pp. The promiscuity of 1NH on unrelated substrate like 2,4-dichlorophenol and the good amino acid sequence identity with 24DCPM (55%) suggests a probable common ancestral origin for enzymes 1NH and 24DCPM. Upon acquiring by strain C5pp, the gene has probably evolved to code for efficient 1NH, thus allowing cells to mineralize carbaryl more rapidly at higher concentrations.
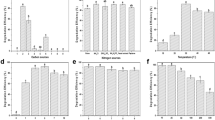
Characterization of 1-naphthol 2-hydroxylase (1NH).
(A) Gel filtration (Sephacryl S-200 HR) elution profile for recombinant 1NH. The native mol. wt. was determined to be ~145 kDa. Gel filtration column was calibrated with β-amylase (200 kDa), alcohol dehydrogenase (150 kDa), BSA (66 kDa), carbonic anhydrase (29 kDa), and cytochrome c (12.4 kDa). Inset represents the SDS-PAGE profile of purified 1NH (sub unit mol wt. ~66 kDa). The image has been cropped to depict one of the fractions of Sephacryl S-200HR. The complete gel image can be seen in the Supplementary information (Figure S7B). (B) Substrate specificity and kinetic constants observed for recombinant 1NH. (C) Phylogenetic analysis of 1NH from C5pp with members of PHBH family of monooxygenase. The numbers in parentheses indicates the protein accession id. Enzyme abbreviations are : 24DCPM, 2,4-dichlorophenol monooxygenase; PH, phenol hydroxylase; HbpA, 3-hydroxybiphenyl monooxygenase; PM, phenol monoxygenase; ChqA, chlorobenzoquinol monooxygenase; OhpB 3-(2-hydroxyphenyl) propionic acid monooxygenase; 1NH, 1-naphthol 2-hydroxylase; MhqA, methylbenzoquinol monooxygenase; AKH, aklavinone 11-hydroxylase; RdmE, aklavinone 12-hydroxylase; PHBH, p-hydroxybenzoate hydroxylase; 4ABH, 4-aminobenzoate hydroxylase; PnpH, p-nitrophenol hydroxylase; 3HBH, 3-hydroxybenzoate hydroxylase; 26DHPH, 2,6-dihydroxyphenol hydroxylase; SALH, salicylate 1-hydroxylase. Na; no activity, -; not determined.
Phylogenetic analysis of ‘upper’ and ‘middle’ pathway genes
The carbaryl degradation pathway follows metabolic steps similar to naphthalene pathway from 1,2-dihydroxynaphthalene onwards. Based on this observation, we speculate that the genes for carbaryl degradation pathway in strain C5pp must have been acquired and evolved from these more prevalent systems in the presence of positive selection pressure. One of the non-parametric indicators is the systematic phylogenetic studies to aid in understanding the evolutionary relatedness. The analysis of McbD, McbE and McbF indicates that their putative functions as hydroxychromene 2-carboxylate isomerase, trans-o-hydroxybenzylidenepyruvate hydratase-aldolase, and SalDH, respectively (Fig. S8A–C). Phylogeny analysis of McbIJKL suggested the close relatedness to functionally characterized S5H from Ralstonia sp. U230 (Fig. S8D–G).
Analysis of putative regulators involved in the carbaryl metabolism
Supercontig-A harboring carbaryl degradation genes was found to contain five genes (mcbG, mcbH, mcbN, mcbR and mcbS) encoding putative regulators. The phylogenetic analysis of McbG, McbH, McbN, McbR and McbS reveals that they are branched into five distinct clusters of LysR/TetR regulators (Fig. S9A).
The amino acid sequence of the McbG (290 a.a., putative regulator of ‘upper’ pathway segment) was similar to that of LysR-type transcriptional regulators. The closest homolog of McbG was found to be LysR-type PhnS (58% identity at a.a. level) regulator involved in phenanthrene metabolism in Burkholderia sp. strain RP00725. PhnS was found to be a part of polycistronic mRNA with direction of transcription in-sync with the downstream genes involved in phenanthrene metabolism. The distance between mcbG and its upstream metabolic genes (mcbF encoding SalDH) was found to be 1146 bp in contrast to 52 bp observed for phnS and its downstream genes. The analysis of the intervening sequence between mcbG and their respective upstream genes indicated the presence of fragmented transposases, suggesting that the regulator is acquired by HGT and probably involved in the regulation of ‘upper’ pathway enzymes.
Gene mcbH (903 bp, 300 a.a.), a part of the ‘middle’ pathway segment (salicylate degradation) encodes a McbH and showed 67–69% identity to a group of NagR/DntR/NahR type LysR transcriptional regulators from Pseudomonas sp. and Burkholderia sp. Members of this family are reported to recognize salicylate as the specific effector molecule to induce the degradation of aromatics31.
In the vicinity of genes involved in the utilization of gentisate (lower pathway), three genes encoding putative regulator proteins viz. McbN (951 bp, 316 a.a., LysR type), McbR (927 bp, 308 a.a., LysR type) and McbS (636 bp, 211 a.a., TetR type) were identified. Among these three regulators, McbN was found to share 79% identity with HbzR (LysR type regulator) from Pseudomonas alcaligenes NCIMB 9867 which was reported to be involved in the gentisate metabolism32. McbR and S showed relatedness with LysR family transcription regulator from Burkholderia sp. lig30 (68%) and TetR family transcription regulator from Brenneria sp. EniD312 (64%) with unknown functions.
The ClustalW alignment supports the possibility that McbG, McbH and McbN might recognize same co-inducers as their orthologs (Fig. S9B). Though the co-inducer for PhnS is not known, NahR and HbzR have been shown to recognize salicylate and gentisate, respectively. Overall, the observed gene organization with respective regulator into three distinct ‘upper’, ‘middle’ and ‘lower’ pathways for the carbaryl degradation in strain C5pp corroborates well with biochemical studies reported earlier17.
Evolution of carbaryl degradation pathway
The G+C profile viewer33 (http://tubic.tju.edu.cn/GC-Profile/) revealed a skewing in the G+C content of Supercontig-A (Fig. 6A). The G+C content in the region 10629–36324 bp, which harbored ‘upper’ (carbaryl to salicylic acid) and ‘middle’ (salicylate to gentisate) pathway genes, was significantly lower (~54%) than that of strain C5pp (62.65%). The G+C content from 36325–76334 bp, which harbors genes involved in the gentisate metabolism, was observed to be ~60%. The remarkable difference in the G+C content suggests probably a different ancestral origin for genes involved in the ‘upper’ and ‘middle’ pathway. RAST analysis identified 42 transposases in C5pp draft genome, of which 17 (40%) were present in Supercontig-A (Table 1), indicating this region to be hotspot for genome alterations. The upstream region of ‘upper’ pathway genes was found to harbor class-I integron with features like presence of transposase, 25 bp left-end repeat (IRi), attI site, 5′ and 3′ conserved segment (CS) (Fig. 6B). It also harbors additional 25 bp direct repeat (92% homology to IRi), aminoglycoside nucleotidyl transferase (Smr, streptomycin resistance, ANT3class), dihydropteroate synthase (folic acid biosynthesis) and N-acetyltransferase (GCN5 family) genes. The invert repeat (IRt) at the right-end of integron was absent. Downstream, a transposase (tnpA) of IS6100 family flanked by 25 bp invert repeat was found to be present as seen for Tn6217 followed by genes for the ‘upper’ pathway. Interestingly, the regulator McbG proposed to be involved in the transcription of upper pathway genes was flanked by two truncated transposases (Fig. 6B). The presence of transposon fragments suggest that they have been lost by decay-linked recombination events and might be the remnant of previously functional integrative conjugative element.
Analysis of carbaryl degradation cluster.
(A) G+C skew plot of 76333 bp region proposed to be involved in the carbaryl degradation. Thin line indicates G+C content as calculated by G+C viewer (http://tubic.tju.edu.cn/GC-Profile/). Thick line indicates G+C content variation as calculated manually with 500 bp window. Horizontal dashed lines indicate maximum (max), minimum (min) and average (avg) G+C content. The genetic organization of carbaryl degrading genes is depicted at the top. The area shaded in grey represents the G+C content of upper, middle and lower pathway involved in the carbaryl degradation. Genetic organization of genes and mobile genetic elements involved in (B) ‘upper’ pathway segment genes, filled yellow oval indicates attI, filled red box indicate 25 bp IRi and black rectangular boxes indicate invert repeats IRL and IRR, Smr - streptomycin resistance; DHP, dihydropterate synthase; NAT, N-acetyl transferase; IRt site of the integron is missing. (C) ‘middle’ pathway segment genes, Red box depicts IS element containing transpsosases flanked by IRs highlighted in black boxes. Blue arrowhead indicates group II intron and (D) ‘lower’ pathway segment genes; Red box depicts IS element containing transpsosases flanked by IRs highlighted in black boxes; double lined box represents recombinase; MetA, a conserved protein; FAR, fusaric acid resistance; S/T-Pase, serine/threonine protein phosphatase; In-Pase, inositol phosphatase; HP, hypothetical protein; regulators are marked in blue arrow heads. The orientation of arrow heads indicate the direction of transcription. Genes coding for ‘upper’ pathway enzymes are marked with pink, ‘middle’ pathway with mango yellow and ‘lower’ pathway with green color.
Strain C5pp carries a catabolic transposon ‘TnC5ppSal’, which harbors mcbIJKL (‘middle’ pathway) and exhibits class-I composite transposon features (IS21 family insertion element repeats, Fig. 6C). The presence of IS elements in inverted orientation probably imparts stability to TnC5ppSal in the genome. The salicylate gene arrangment (mcbIJKL) is similar to Ralstonia sp. strain U234. However, the region containing transpsoase at either ends showed ~80% identity to Pseudomonas resinovorans. This indicates that though there is a synteny between strain C5pp and U2, the transposon for salicylate in strain C5pp has probably a different ancestral origin. Similarly, the ‘lower’ pathway was found to be a part of class-I composite transposon flanked by non-identical IS elements (Fig. 6D). The transposon, is bordered by IS element similar to ISPa1635 and IS481, respectively. Overall, these observations indicate the acquisition of pathway as result of multiple transposition events.
In conclusion, the draft genome sequence of Pseudomonas sp. strain C5pp along with concerted approach involving bioinformatics, molecular biology and biochemical analysis led to decipher the carbaryl degradation pathway at genetic level into three distinct putative operons. Genome analysis further reveals that the degradation capability must have been acquired through HGT events. The present study helps to understand the molecular mechanisms involved in the construction and evolution of metabolic pathway for carbaryl. In this perspective, the information becomes particularly important giving avenue to engineer a strain for effective degradation of array of aromatic compounds.
Materials and Methods
Bacterial culture and Genomic DNA isolation
Soil isolate Pseudomonas sp. strain C5pp (earlier referred as Pseudomonas sp. strain C5) utilizes carbaryl as the sole source of carbon and energy was used in this study15. Strain C5pp was grown on MSM supplemented with glucose (0.25%) or appropriate aromatics (carbaryl/gentisate, 0.1%) as described previously15. E. coli EPI300 was grown on Luria-Bertani (LB) broth35 or M9-medium36 containing glucose or aromatics (carbaryl/salicylate/gentisate, 0.1%) supplemented with leucine (100 μg.ml−1), thiamine (10 μg.ml−1) and arabinose (0.001%).
The genomic DNA was isolated from strain C5pp grown on carbaryl (0.1%) using UltraClean® Microbial DNA isolation kit (MoBio Laboratories, USA).
Construction and functional screening of Pseudomonas sp. strain C5pp genomic fosmid library
The genomic DNA isolated from strain C5pp was end-repaired using T4 DNA polymerase and T4 polynucleotide kinase (NEB, USA). The fragments of ~40 kb size were eluted from low melting agarose gel using β-agarse (NEB, USA), ligated to the fosmid vector (pCC2FOS) and packaged as per manufacturer’s instructions (Epicentre, USA). The packaged fosmids were transfected into E. coli EPI300 and appropriate dilutions were plated onto LB plates containing chloramphenicol (12.5 μg.ml−1). The genomic DNA library of Pseudomonas sp. strain C5pp was constructed in CopyControl fosmid pCC2FOS with a phage titer of 3 × 106 CFU.ml−1 and a pool of ~9600 colonies was used for screening.
The library pool was plated onto M9 medium supplemented with leucine (100 μg.ml−1), thiamine (10 μg.ml−1), arabinose (0.001%), aromatic compound (carbaryl/salicylate/gentisate, 0.1%) and chloramphenicol (12.5 μg.ml−1). Twenty eight colonies were observed on M9 plates containing gentisate at the end of 8th day indicating the ability to utilize gentisate as the carbon source, whereas plates containing carbaryl or salicylate did not show any colonies.
Five colonies were picked up randomly from M9-gentisate plates and analyzed further. Fosmid DNA was isolated by Fosmid Max DNA purification kit (FMAX 046, USA) and restriction digested with BamHI (~12, 8, 5, 2.8, 2.5 and 1.8 kb) or NotI (20, 11, 5, 4.8, 2.7 and 2.3 kb). One of the randomly chosen clone designated as G1 was digested with BamHI and all six fragments were sub-cloned into pUC19 and partially sequenced using universal M13 forward and reverse primers (M13F, 5′-GTTTTCCCAGTCACGAC-3′ and M13R, 5′-CAGGAAACAGCTATGAC-3′, Promega, USA) after cloning it into pUC19 at BamHI site.
Preparation of cell-free extracts and enzyme assays
Randomly selected single gentisate positive colony of E. coli EPI300 was grown for 24 h on M9 medium (150 ml) supplemented with: a) gentisate (0.1%), chloramphenicol (12.5 μg ml−1); and b) glucose (0.25%), chloramphenicol (12.5 μg.ml−1). Cells were harvested by centrifugation, suspended in ice-cold phosphate buffer (50 mM, pH 7.5) and disrupted by sonication at 4 °C (four cycles with 4 min interval; each cycle: 15 pulses, output 11 W, Ultrasonic processor GE130). Cell homogenate was centrifuged at 40,000×g for 30 min. The clear supernatant obtained was referred as CFE and used as the enzyme source. Protein was estimated by Bradford method using bovine serum albumin as the standard37. CH, 1NH, and GDO were monitored spectrophotometrically (Perkin Elmer; model Lambda 35) as described15,22. 12DHNDO was assayed as described38 after reactivation. Specific activities are reported as nmole.min−1.mg−1 of protein. To identify the reaction product of CH, the reaction mixture was subjected to HPLC (Agilent 1200 series) using RP-C18 column (4.6 × 250 mm, particle size 5 μM, Eclipse plus C-18, Agilent) using solvent system methanol:water (60:40 v/v, flow rate 1 ml.min−1). The detection was with diode array detector at 280 and 322 nm for carbaryl and 1-naphthol, respectively.
Identification of genes involved in carbaryl degradation
To predict the genes for carbaryl degradation pathway, the partial sequences obtained from six sub-clones of G1 were blast analysed against the draft genome of strain C5pp, which retrieved contigs 47, 61, 62 and 76. These contigs were further subjected to ORF finder (http://www.ncbi.nlm.nih.gov/projects/gorf/). The predicted genes were analysed further using the function and sequence based approach. Contigs were assembled using gap-filling reactions using primers as mentioned in Table S2.
Phylogenetic analysis
The phylogenetic trees were inferred for amino acid sequences of similar enzymes/proteins obtained from NCBI BLAST using Neighbor-Joining (NJ) algorithm by using MEGA6 software. The protein sequences were aligned using ClustalW program. The robustness of the tree topology was assessed by bootstrap analysis of 500 replicons using Jones-Taylor-Thornton (JTT) model39.
Cloning and over-expression of carbaryl hydrolase, 1-naphthol 2-hydroxylase and 1,2-dihydroxynaphthalene dioxygenase
The gene predicted for CH, 1NH and 12DHNDO were PCR amplified from genomic DNA of strain C5pp using primers: CHF- 5′-CTAGCTAGCATGGCGGTCACGGCAAATTATTTGC-3′ (under line represents restriction site for NheI) and CHR- 5′-CCGCTCGAGTCAATGCGCGGCAAGCCGG-3′ (XhoI); 1NHF- 5′-CCGGAATTCCATATGCTGAAAAATATTT-3′ (EcoRI and NdeI) and 1NHR- 5′-CCGGAATTCTTAAAGACAGAGAATTGC-3′ (EcoRI); and 12DHNDOF- 5′-CATGCCATGGCTCAGATTGTAGCTGG-3′ (NcoI) and 12DHNDOR- 5′CCGCTCGAGGCTAGGCTTCATCTCCATATACCC-3′ (XhoI), respectively. The gel purified PCR products (2.31, 1.77, and 0.825 kb for CH, 1NH and 12DHNDO, respectively) were digested with appropriate restriction enzymes and cloned into pET-28a(+) (Novagen). The expressed protein contains His6-tag at N-terminus. pET28-CH, pET28–1NH and pET28-12DHNDO were transformed into E. coli BL21 (DE3). A single colony obtained was grown on LB medium (500 ml) containing kanamycin (30 μg.ml−1) at 37 °C till OD600 nm = 0.8. The culture was chilled on ice for an hour and induced by addition of IPTG (100 μM) for 16 h at 18 °C. Cells were harvested, re-suspended (1:10 g wt./vol.) in potassium-phosphate buffer (50 mM, pH7.5) with glycerol (5%) [Buffer-A] and CFE was prepared. CFE from E. coli cells as well as E. coli carrying vector alone (absence of gene of interest) were used as control.
Purification and characterization of recombinant 12DHNDO and 1NH
The recombinant 12DHNDO and 1NH (r1NH) were purified using Ni-NTA matrix (Qiagen, USA, See Supporting Information). r1NH was characterized for substrate specificity and kinetic constants as described earlier (Trivedi et al., 2014). r1NH and 1NH purified from strain C5pp were subjected to MALDI-TOF/TOF-MS/MS analysis as described40.
Additional Information
How to cite this article: Trivedi, V. D. et al. Insights into functional and evolutionary analysis of carbaryl metabolic pathway from Pseudomonas sp. strain C5pp. Sci. Rep. 6, 38430; doi: 10.1038/srep38430 (2016).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Copley, S. D. Evolution of a metabolic pathway for degradation of a toxic xenobiotic, the patchwork approach. Trends Biochem Sci 25, 261–265 (2000).
Copley, S. D. Evolution of efficient pathways for degradation of anthropogenic chemicals. Nat Chem Biol 5, 559–566 (2009).
Copley, S. D. et al. The whole genome sequence of Sphingobium chlorophenolicum L-1, insights into the evolution of the pentachlorophenol degradation pathway. Genome Biol Evol 4, 184–198 (2012).
Fenner, K., Canonica, S., Wackett, L. P. & Elsner, M. Evaluating pesticide degradation in the environment, blind spots and emerging opportunities. Science 341, 752–758 (2013).
Hacker, J. & Carniel, E. Ecological fitness, genomic islands and bacterial pathogenicity. A Darwinian view of the evolution of microbes. EMBO Rep 2, 376–381 (2001).
Maeda, K. et al. Complete nucleotide sequence of carbazole/dioxin-degrading plasmid pCAR1 in Pseudomonas resinovorans strain CA10 indicates its mosaicity and the presence of large catabolic transposon Tn4676. J Mol Biol 326, 21–33 (2003).
Top, E. M. & Springael, D. The role of mobile genetic elements in bacterial adaptation to xenobiotic organic compounds. Curr Opin Biotechnol 14, 262–269 (2003).
Smulders, C. J., Bueters, T. J., Van Kleef, R. G. & Vijverberg, H. P. Selective effects of carbamate pesticides on rat neuronal nicotinic acetylcholine receptors and rat brain acetylcholinesterase. Toxicol Appl Pharmacol 193, 139–146 (2003).
Sud, R. K., Sud, A. K. & Gupta, K. G. Degradation of sevin (1-naphthyl-N-methyl carbamate by Achromobacter sp.). Arch Mikrobiol 87, 353–358 (1972).
Larkin, M. J. & Day, M. J. The metabolism of carbaryl by three bacterial isolates, Pseudomonas spp. (NCIB 12042 & 12043) and Rhodococcus sp. (NCIB 12038) from garden soil. J Appl Bacteriol 60, 233–242 (1986).
Chapalamadugu, S. & Chaudhry, G. R. Hydrolysis of carbaryl by a Pseudomonas sp. and construction of a microbial consortium that completely metabolizes carbaryl. Appl Environ Microbiol 57, 744–750 (1991).
Hayatsu, M. & Nagata, T. Purification and characterization of carbaryl hydrolase from Blastobacter sp. strain M501. Appl Environ Microbiol 59, 2121–2125 (1993).
Hayatsu, M., Mizutani, A., Hashimoto, M., Sato, K. & Hayano, K. Purification and characterization of carbaryl hydrolase from Arthrobacter sp. RC100. FEMS Microbiol Lett 201, 99–103 (2001).
Doddamani, H. P. & Ninnekar, H. Z. Biodegradation of carbaryl by a Micrococcus species. Curr Microbiol 43, 69–73 (2001).
Swetha, V. P. & Phale, P. S. Metabolism of carbaryl via 1,2-dihydroxynaphthalene by soil isolates Pseudomonas sp. strains C4, C5, and C6. Appl Environ Microbiol 71, 5951–5956 (2005).
Seo, J. S., Keum, Y. S. & Li, Q. X. Metabolomic and proteomic insights into carbaryl catabolism by Burkholderia sp. C3 and degradation of ten N-methylcarbamates. Biodegradation 24, 795–811 (2013).
Singh, R., Trivedi, V. D. & Phale, P. S. Metabolic regulation and chromosomal localization of carbaryl degradation pathway in Pseudomonas sp. strains C4, C5 and C6. Arch Microbiol 195, 521–535 (2013).
Swetha, V. P., Basu, A. & Phale, P. S. Purification and characterization of 1-naphthol-2-hydroxylase from carbaryl-degrading Pseudomonas strain C4. J Bacteriol 189, 2660–2666 (2007).
Sah, S. & Phale, P. S. 1-naphthol 2-hydroxylase from Pseudomonas sp. strain C6, purification, characterization and chemical modification studies. Biodegradation 22, 517–526 (2011).
Trivedi, V. D., Majhi, P. & Phale, P. S. Kinetic and spectroscopic characterization of 1-naphthol 2-hydroxylase from Pseudomonas sp. strain C5. Appl Biochem Biotechnol 172, 3964–3977 (2014).
Trivedi, V. D., Jangir, P. K., Sharma, R. & Phale, P. S. Draft genome sequence of carbaryl degrading soil isolate Pseudomonas sp. strain C5pp. Genome Announc. 9, 4(3) pii, e00526-16. doi: 10.1128/genomeA.00526-16 (2016).
Singh, R., Trivedi, V. D. & Phale, P. S. Purification and characterization of NAD+ -dependent salicylaldehyde dehydrogenase from carbaryl-degrading Pseudomonas sp. strain C6. Appl Biochem Biotechnol 172, 806–819 (2014).
Hashimoto, M., Fukui, M., Hayano, K. & Hayatsu, M. Nucleotide sequence and genetic structure of a novel carbaryl hydrolase gene (cehA) from Rhizobium sp. strain AC100. Appl Environ Microbiol 68, 1220–1227 (2002).
Hashimoto, M. et al. Cloning and nucleotide sequence of carbaryl hydrolase gene (cahA) from Arthrobacter sp. RC100. J Biosci Bioeng 101, 410–414 (2006).
Laurie, A. D. & Lloyd-Jones, G. The phn genes of Burkholderia sp. strain RP007 constitute a divergent gene cluster for polycyclic aromatic hydrocarbon catabolism. J Bacteriol 181, 531–540 (1999).
Eltis, L. D. & Bolin, J. T. Evolutionary relationships among extradiol dioxygenases. J Bacteriol 178, 5930–5937 (1996).
Patel, T. R. & Barnsley, E. A. Naphthalene metabolism by pseudomonads, purification and properties of 1,2-dihydroxynaphthalene oxygenase. J Bacteriol 143, 668–673 (1980).
van Berkel, W. J., Kamerbeek, N. M. & Fraaije, M. W. Flavoprotein monooxygenases, a diverse class of oxidative biocatalysts. J Biotechnol 124, 670–689 (2006).
Overbeek, R. et al. The subsystems approach to genome annotation and its use in the project to annotate 1000 genomes. Nucleic Acids Res 33, 5691–5702 (2005).
Zhou, N. Y., Al-Dulayymi, J., Baird, M. S. & Williams, P. A. Salicylate 5-hydroxylase from Ralstonia sp. strain U2, a monooxygenase with close relationships to and shared electron transport proteins with naphthalene dioxygenase. J Bacteriol 184, 1547–1555 (2002).
Lonneborg, R., Smirnova, I., Dian, C., Leonard, G. A. & Brzezinski, P. In vivo and in vitro investigation of transcriptional regulation by DntR. J Mol Biol 372, 571–582 (2007).
Yeo, C. C., Tan, C. L., Gao, X., Zhao, B. & Poh, C. L. Characterization of hbzE-encoded gentisate 1,2-dioxygenase from Pseudomonas alcaligenes NCIMB 9867. Res Microbiol 158, 608–616 (2007).
Gao, F. & Zhang, C. T. GC-Profile, a web-based tool for visualizing and analyzing the variation of GC content in genomic sequences. Nucleic Acids Res 34, W686–W691 (2006).
Fuenmayor, S. L., Wild, M., Boyes, A. L. & Williams, P. A. A gene cluster encoding steps in conversion of naphthalene to gentisate in Pseudomonas sp. strain U2. J Bacteriol 180, 2522–2530 (1998).
Sezonov, G., Joseleau-Petit, D. & D’Ari, R. Escherichia coli physiology in Luria-Bertani broth. J Bacteriol 189, 8746–8749 (2007).
Nicolaou, S. A., Gaida, S. M. & Papoutsakis, E. T. Coexisting/Coexpressing Genomic Libraries (CoGeL) identify interactions among distantly located genetic loci for developing complex microbial phenotypes. Nucleic Acids Res 39, e152 (2011).
Bradford, M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72, 248–254 (1976).
Kuhm, A. E., Stolz, A., Ngai, K. L. & Knackmuss, H. J. Purification and characterization of a 1,2-dihydroxynaphthalene dioxygenase from a bacterium that degrades naphthalenesulfonic acids. J Bacteriol 173, 3795–3802 (1991).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6, Molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30, 2725–2729 (2013).
Basu, B. & Apte, S. K. Gamma radiation-induced proteome of Deinococcus radiodurans primarily targets DNA repair and oxidative stress alleviation. Mol Cell Proteomics 11, M111 (2012).
Acknowledgements
VDT acknowledges CSIR, Govt. of India for senior research fellowship. We thank Dr. Basu, BARC, Mumbai for MALDI-TOF/TOF analysis. PP acknowledges research grant from Department of Biotechnology, Govt. of India, RS is thankful to Director, CSIR-IGIB, New Delhi for support and encouragement.
Author information
Authors and Affiliations
Contributions
Vikas D.T.: Genome and HGT analysis, cloning, over-expression and analysis of CH, 1NH and 12DHNDO, fosmid library construction and functional analysis of G1 clone, experimental work from strain C5pp Pramod J.: Fosmid library construction, genome sequencing and analysis Sharma R. and Phale P.: Concepts, design of experiments All authors analyzed the data critically and reviewed the manuscript before submission.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Trivedi, V., Jangir, P., Sharma, R. et al. Insights into functional and evolutionary analysis of carbaryl metabolic pathway from Pseudomonas sp. strain C5pp. Sci Rep 6, 38430 (2016). https://doi.org/10.1038/srep38430
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep38430
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.