Abstract
Tumor cells respond to their microenvironment, which can include hypoxia and malnutrition, and adapt their metabolism to survive and grow. Some oncogenes are associated with cancer metabolism via regulation of the related enzymes or transporters. However, the importance of metabolism and precise metabolic effects of oncogenes in colorectal cancer remain unclear. We found that colorectal cancer cells survived under the condition of glucose depletion, and their resistance to such conditions depended on genomic alterations rather than on KRAS mutation alone. Metabolomic analysis demonstrated that those cells maintained tricarboxylic acid cycle activity and ATP production under such conditions. Furthermore, we identified pivotal roles of GLUD1 and SLC25A13 in nutritional stress. GLUD1 and SLC25A13 were associated with tumor aggressiveness and poorer prognosis of colorectal cancer. In conclusion, GLUD1 and SLC25A13 may serve as new targets in treating refractory colorectal cancer which survive in malnutritional microenvironments.
Similar content being viewed by others
Introduction
Colorectal cancer is the third most frequently diagnosed cancer, fourth leading cause of cancer deaths worldwide, and accounts for 1.3 million new cases and 690,000 deaths annually1. Although effective treatment has led to improvements in survival, colorectal cancer remains a major global health problem2,3. Thus, it is necessary to develop additional novel and efficient treatments.
The physiology of tumor tissues differs from that of normal tissues in many aspects, the majority of which result from differences between the vasculatures of the two tissues4. Poorly formed tumor vasculature leads to a hypoxic microenvironment in which the nutrient levels are low and levels of waste products are high5. Tumor cells respond to such conditions and adapt their metabolism to survive and grow. Cancer cells are well known for having high rates of glucose consumption and lactate production despite bioavailability of sufficient oxygen for complete oxidation of glucose. This phenomenon is termed the Warburg effect6. Since the discovery of the Warburg effect, many researchers have studied the metabolism of cancer cells and tumor tissues. In cancer cells, glutamine, one of the most important nutrients as well as glucose, is reportedly metabolized more abundantly than other non-essential amino acids7. Glutamine metabolism not only provides a source for synthesis of macromolecules, such as lipids, proteins, and nucleotides, but also supports nicotinamide adenine dinucleotide phosphate (NADPH) production and anaplerosis in proliferating tumor cells8. This difference in metabolism between cancer and normal cells is expected to provide opportunities for development of innovative cancer treatments.
Several reports have indicated that tumor cells show changes in metabolism induced by oncogenes. MYC and HIF-1 have been reported to regulate genes associated with glucose metabolism, such as the glucose transporter, GLUT1, hexokinase 2, pyruvate kinase M2 (PKM2), lactate dehydrogenase A, and pyruvate dehydrogenase kinase 19. MYC is also postulated to stimulate glutamine metabolism via regulation of amino acid transporters (e.g., SLC1A5) and glutaminase. Moreover, expressions of the malic enzyme 1 (ME1) gene and malic enzyme 2 gene are inhibited by p5310. KRAS (G12D mutation) has an important role in regulating pancreatic tumor metabolism via stimulation of glucose uptake and activation of the hexosamine biosynthesis and pentose phosphate pathways11. Furthermore, there is a non-canonical pathway of glutamine in pancreatic ductal adenocarcinoma cells that is regulated by the KRAS oncogene12. However, the importance of glutamine metabolism and precise metabolic effects of oncogenes in colorectal cancer cells remain unknown.
The aim of this study is to elucidate metabolic adaptation to nutritional stress and the role of the involved oncogenes in human colorectal cancer. The present study showed that the metabolism of colorectal cancer, distinct from that of pancreatic cancer, depended on genomic alterations, which previously have been uncharacterized and was not restricted to KRAS mutation alone. Colorectal cancer can survive under the condition of glucose depletion while retaining TCA cycle activity. The cells’ survival relies on a delicate balance between energy and reactive oxygen species (ROS) production. Glutamate dehydrogenase 1 (GLUD1) and SLC25A13 have pivotal roles under glucose-deprived conditions and are associated with tumor aggressiveness and colorectal cancer prognosis.
Results
Survival of colorectal cancer cells under condition of glucose depletion
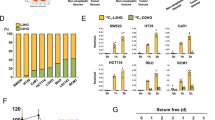
Glucose and glutamine are two of the most abundant nutrients in plasma, and together, they account for most of the carbon and nitrogen metabolism occurring in mammalian cells. Both nutrients are essential for growth of pancreatic ductal adenocarcinoma cells with KRAS mutation12. To assess the role of glucose and glutamine in colorectal cancer cells, a proliferation assay was performed under various media conditions (Fig. 1A and Supplementary Fig. S1A). For the assay, we confirmed that DLD1 and HCT116 cells had a KRAS mutation at codon 13 involving a nucleotide change from GGC to GAC, and that HT29 and CaR1 cells did not have this KRAS mutation (Fig. 1B and Supplementary Fig. S1B). Notably, DLD1, HCT116, and CaR1 cells could survive under the glucose-deprived conditions (Fig. 1C–E and Supplementary Fig. S1C–E). Furthermore, DLD1 cells that had strong resistance to the condition of glucose depletion were able to survive for 14 days (Fig. 1F and G), and the passage of DLD1 cells was possible under that condition. The rate of apoptotic cells under the glucose-deprived conditions was lower in DLD1 cells than in HT29 cells (1.5% vs. 24.7%, respectively) (Fig. 1H). These findings show that colorectal cancer cells can survive under conditions of glucose depletion (glutamine sufficiency), which is profoundly different from pancreatic cancer cells in which both nutrients are indispensable.
Colorectal cancer cell lines survive under glucose depleted conditions.
(A) Cells were cultured in complete medium, which was replaced the following day with glucose- and glutamine-deficient media supplemented with 10% dialyzed fetal bovine serum and glucose (10 mM) or glutamine (2 mM), respectively. At the indicated time points, the cells were fixed in 80% methanol and stained with 0.1% crystal violet. OD determined the relative cell proliferation at 595 nm. Glc: glucose, Gln: glutamine. (B) RNA was extracted from DLD1 and HT29 cells, and the KRAS gene was PCR-amplified and sequenced. (C,D) Relative growth of DLD1 (C) and HT29 (D) under the indicated conditions. The difference in relative growth between Gln and Neither in DLD1 was significant. (E) The representative images are shown for day 3 (scale bar, 200 mm). (F) DLD1 cells survived for 2 weeks under the condition of glucose depletion. (G) The representative images are shown for day 14. (H) DLD1 and HT29 cells were cultured under the indicated conditions for 24 h. Apoptotic cells were analyzed by FACS using annexin V-FITC and propidium iodide. Representative results are shown. Data are presented as the mean ± SD of at least three independent experiments **P < 0.01).
Resistance to glucose-deprived conditions in colorectal cancer depends on genomic alterations not restricted to KRAS mutation alone
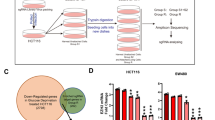
Many reports indicate that the oncogene KRAS has an important role not only in cellular transformation and metabolic reprograming during tumorigenesis but also in chemotherapy for colorectal cancer, considering that mutant KRAS transduces oncogenic signaling and is the most reliable predictor for cancer response to anti-EGFR monoclonal antibodies11,12,13. We were interested in how genomic alterations, including alteration of the KRAS oncogene, in colorectal cancer influence cell proliferation under glucose-deprived conditions. To this end, mouse transformed NIH3T3 cells (Cle-H3) by human colorectal cancer genome with KRAS mutations were used in a proliferation assay14. The results showed that the Cle-H3 cells survived under glucose-deprived conditions, but the parental NIH3T3 did not (Fig. 2B–D), which suggested a role of the colorectal cancer genome in cell survival under glucose-deprived conditions. To assess whether these findings resulted solely from the oncogene KRAS, the proliferation assay using oncogenic KRAS-overexpressing NIH3T3 cells was similarly performed (Fig. 2E). The data showed that both KRAS-overexpressing NIH3T3 cells and control NIH3T3 cells did not survive under glucose-deprived conditions (Fig. 2F and G). Furthermore, knockdown of KRAS in the Cle-H3 and DLD1 cells did not influence their survival under glucose-depleted conditions (Supplementary Fig. S2A–F). These results demonstrated that resistance to the glucose-deprived conditions in colorectal cancer depended on uncharacterized genomic alterations that were not restricted to KRAS mutation alone.
Resistance to glucose-deprived conditions in colorectal cancer depended on whole genomic alterations rather than on KRAS mutation alone.
(A) The scheme shows the transfection methods of whole genomic DNA derived from a human colon cancer cell line (MA) harboring a KRAS mutation at codon 12 involving a nucleotide change from GGT to GAT. Transfection into NIH3T3 cells was performed using the calcium phosphate sedimentation method to establish Cle-H3 cells. (B,C) Relative growth of NIH3T3 (B) and Cle-H3 (C) under the indicated conditions. The difference in relative growth between Gln and Neither in Cle-H3 was significant. (D) Representative images are shown for day 3 (scale bar, 200 mm). (E) Western blot analysis of NIH3T3 cells transfected with control or KRAS overexpression (OE) vector. (F,G) Relative growth of control (F) and KRAS-overexpressing NIH3T3 cells (G) under the indicated conditions. Data are presented as the mean ± SD for at least three independent experiments (**P < 0.01).
Metabolism of colorectal cancer was not restricted to KRAS mutation alone
We hypothesized that KRAS mutation alone may not markedly influence colorectal cancer metabolism. In the metabolomic analysis using control and KRAS knockdown DLD1 cells, which can survive under glucose-deprived conditions, we found that the metabolism of colorectal cancer did not depend on KRAS mutation alone but depended on the medium condition as well (Fig. 3A and B). In addition, k-means clustering and the Calinski criterion showed that k = 3 gave the optimal number of clusters, which indicated low dependence of metabolism of colorectal cancer on KRAS mutation alone (Supplementary Fig. S3A and B). Then, we measured the metabolites in plasma from wild type and KRAS mutation (Cre/LSL-KRASmut) mice. The principal component analysis showed no significant difference in cellular metabolites between two groups (Supplementary Fig. S3C). Considering that a mutation of another oncogene BRAF plays a role in tumor aggressiveness of retractable colorectal cancer in down stream pathway of KRAS, a role of BRAF was studied. We used two kinds of BRAF specific compounds, PLX4032 and PLX4720, in colorectal cancer HT29 cells in culture, and measured whole metabolites by CE-MS analysis. The data indicated that BRAF inhibition resulted in the suppression of metabolites in TCA cycle (Supplementary Fig. S4A–G), confirming that series of our experiments demonstrated the concept that mutant KRAS status does not predict glucose dependence in culture, and that glutamine fuels oxidative phosphorylation in TCA cycle, as observed in a stress condition such as BRAF specific inhibition in colorectal cancer cells. This observation suggests that the survival of colorectal cancer cells depends on a metabolomic mechanism different from those of other cancers, including pancreatic cancer12. On the basis of previous reports that showed that metabolism was important in DNA methylation15, DNA methylation analysis was performed under various conditions (Supplementary Fig. S3D). The data showed that DNA methylation frequency in colorectal cancer was not markedly influenced by depletion of glucose or glutamine.
Metabolomic analysis for identification of metabolic effects of oncogene KRAS.
(A,B) KRAS knockdown DLD1 cells and control cells were cultured at 3 × 106 cells per 10-cm dish in 10 ml complete medium; the medium was replaced the following day with glucose(−) and glutamine(−) medium supplemented with 10% dialyzed fetal bovine serum. Glucose (10 mM) or glutamine (2 mM) was added, respectively. For hypoxia treatment, cells were incubated for 24 h with 1% O2. Principal component analysis (A) and hierarchical cluster analysis (B) performed under the indicated conditions at the indicated time points. The red circles indicate the groups that have similar principal component scores. Data for two independent experiments are presented.
DLD1 cells retain TCA cycle activity under condition of glucose depletion
To study the mechanism of resistance to glucose deprivation, the levels of glycolytic and TCA cycle metabolites were assessed by metabolomic analysis in living cells (DLD1 and HT29) pulse-chased for 24 h under glucose-deprived conditions. Notably, TCA cycle metabolites were maintained under the condition of glucose depletion in DLD1 cells but not in HT29 cells (Fig. 4). Under glucose-deprived conditions, the levels of glycerate 3-phosphate, 2-phosphoglycerate, and phosphoenolpyruvic acid increased in HT29 cells but not in DLD1 cells, which suggested that glucose depletion in HT29 cells induces gluconeogenesis. It is known that nicotinamide adenine dinucleotide consists of an oxidized form (NAD+) and a reduced form (NADH) and has a pivotal role in the electron transport chain in oxidative phosphorylation16; it is produced by glycolysis, the TCA cycle, and β-oxidation of fatty acids. The results showed that, in DLD1 cells, NADH decreased under glucose-deprived conditions compared with glucose- and glutamine-containing conditions, but NAD+ increased (Supplementary Fig. S5A). It is known that NADPH (reduced form) also has an oxidized form (NADP+) and provides reducing equivalents for various reactions related to biosynthesis; the molecule is used for generation of glutathione-scavenging ROS. The data showed that NADPH increased in DLD1 cells under glucose-deprived conditions, whereas NADPH decreased remarkably and NADP+ increased in HT29 cells (Supplementary Fig. S5B). Under glucose-deprived conditions, DLD1 and HT29 cells had few metabolites in the pentose phosphate pathway, which is the main source of nicotinamide adenine dinucleotide phosphate (Supplementary Fig. S5C), although other pathways, including those related to malic enzyme, isocitrate dehydrogenase, and nicotinamide nucleotide transhydrogenase, may be involved in its supply17. We performed the sphere formation assay, which is sensitive for intracellular redox status. Cells exhibiting high levels of ROS are susceptible to cell death and some cancer cells develop the redox pathway, which contributes to the cell survival and sphere formation. The results showed that the sphere formation frequency under the glucose depletion condition was greater in DLD1 cells when compared with HT29 cells, suggesting that cell survival was associated with the redox balance (Supplementary Fig. S5D). Together, these results indicate that DLD1 cells retain TCA cycle activity under the condition of glucose depletion and resist increases in ROS.
DLD1 cells retain TCA cycle activity under condition of glucose depletion.
DLD1 and HT29 cells were cultured at 3 × 106 cells per 10-cm dish in 10 ml complete medium; this medium was replaced the following day with glucose(−) and glutamine(−) medium supplemented with 10% dialyzed fetal bovine serum. Glucose (10 mM) or glutamine (2 mM) was added, respectively. Cells incubated for 24 h were analyzed. Data are presented as mean ± SD for two independent experiments.
Increase in amino acids levels in colorectal cancer cells under condition of glucose depletion
In addition to glycolytic and TCA cycle metabolites, the levels of amino acids in DLD1 and HT29 cells were studied by metabolomic analysis based on a method described in a previous report18 (Fig. 5). Compared with glucose- and glutamine-containing conditions, glucose-deprived conditions led to an increase in most of the amino acids in the DLD1 and HT29 cells. In particular, aspartic acid and asparagine levels increased remarkably in DLD1 cells (25.3 and 13.4 times, respectively). The total level of amino acids increased in both cell lines.
Increase in amino acids levels in colorectal cancer cells under condition of glucose depletion.
DLD1 and HT29 cells were plated at 3 × 106 cells per 10-cm dish in 10 ml complete medium; the medium was replaced the following day with glucose(−) and glutamine(−) media supplemented with 10% dialyzed fetal bovine serum. Glucose (10 mM) or glutamine (2 mM) was added, respectively. Cells incubated for 24 h were analyzed. Data are presented as mean ± SD for two independent experiments.
Reliance of survival of DLD1 cells on the delicate balance between ROS and energy production
To adapt to nutritional stress, cells exposed to starvation have reportedly used autophagy to maintain ATP energy production and macromolecular synthesis19. Indeed, glucose depletion caused a decline in ATP levels in DLD1 and HT29 cells (Supplementary Fig. S6A) and induced autophagy in HT29 cells but not in DLD1 cells, as shown by LC3-II levels (Supplementary Fig. S6B). These findings suggest that the influence of autophagy on the increase in amino acids was limited to DLD1 cells (versus HT29 cells). Asparagine has reportedly been shown to suppress expression of C/EBP homologous protein (CHOP), which induces apoptosis20. In parallel with the effect of asparagine level, glucose depletion for 24 h induced CHOP in HT29 cells but not in DLD1 cells (Supplementary Fig. S6B), which suggested that the resistance to glucose-deprived conditions in colorectal cancer was partially associated with the level of asparagine. Given that the NADPH level in HT29 cells was low under glucose-deprived conditions, we studied ROS levels. The data showed that the level of ROS in HT29 cells under glucose-deprived conditions was 5.3 times higher than the levels under glucose- and glutamine-containing conditions, but it was only 1.7 times higher in DLD1 cells (Supplementary Fig. S6C). We were interested in whether the cell survival in glucose deprivation may depend on oxidative phosphorylation in mitochondria. Metformin reportedly inhibits the respiratory chain complex I associated with ATP generation in mitochondria21. As expected, the addition of metformin in culture under glucose-deprived conditions resulted in inhibition of growth of DLD1 cells in a concentration-dependent manner (Supplementary Fig. S6D). Taken together, the data support the idea that retention of resistance to glucose-deprived conditions may rely on oxidative phosphorylation and ROS level control.
Significant association of GLUD1 with resistance to glucose-deprived conditions in colorectal cancer
Glutaminolysis has a profound influence on the maintenance of redox balance and energy production. Under glucose-deprived conditions, its importance appears to further increase, considering that it may change according to the surrounding environment. To verify this hypothesis, gene microarray analysis was performed (Supplementary Fig. S7A) to identify candidate enzymatic genes related to the metabolism of glutamate and glutamine. GLUD1 expression was greatly increased in DLD1 but not in HT29 cells. The results are consistent with those of a previous report in which GLUD1 was shown to have an important role under the condition of glucose deprivation in glioblastoma cells22. The changes in GLUD1 expression were confirmed by quantitative real-time PCR (Supplementary Fig. S7B). Changes in glutamate dehydrogenase activity related to expression of GLUD1 were also confirmed (Supplementary Fig. S7C). Knockdown of GLUD1 in DLD1 cells under glucose-deprived conditions significantly decreased their growth (Supplementary Fig. S7D and E). Taken together, the present study showed that GLUD1 was important in resistance to glucose-deprived conditions in colorectal cancer.
Combined expression of GLUD1 and SLC25A13 is significantly associated with prognosis in colorectal cancer
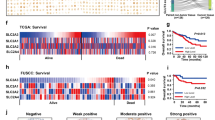
Cancer cells that have resistance to nutritional stress are expected to contribute to worse prognosis. Therefore, the published GSE17536 database was used for screening of genes related to the prognosis of colorectal cancer patients, as described previously23,24 (Supplementary Table S1). SLC25A13 was identified by analyses of the GLUD1 gene and other genes involved in amino acid metabolism in all possible combinations as pairs. SLC25A13 codes for mitochondrial aspartate–glutamate carrier (AGC), which has a central role in the malate–aspartate shuttle25. The reducing equivalents of NADH plus H+ through glycolysis are transferred from the cytosol to the mitochondria for the electron transport chain in oxidative phosphorylation by this shuttle, which is concerned with ROS and ATP production. In the DLD1 cells, the mRNA level of SLC25A13 decreased under glucose-deprived conditions relative to the levels in glucose- and glutamine-containing conditions but increased conversely in HT29 cells (Supplementary Fig. S7F). Then we measured the enzyme activity of TCA related enzymes (succinate dehydrogenase and fumarase). The data showed that DLD1 had higher activity of succinate dehydrogenase, fumarase and malate dehydrogenase than HT29 cells did (Supplementary Fig. S8A–C). These findings suggest that the resistance to glucose-deprived conditions in colorectal cancer may result from inhibition of ROS production via repressed AGC. Subsequently, the relevance of GLUD1 and SLC25A13 expression in colorectal cancer with respect to clinicopathological characteristics and prognosis was evaluated by immunohistochemistry (Fig. 6A). High GLUD1 expression or low SLC25A13 expression was found to be associated with tumor aggressiveness, including depth of tumor invasion, lymph node and distant metastasis, lymphatic and venous invasion, and stage (Supplementary Table S2). Notably, high GLUD1 expression combined with low SLC25A13 expression had a higher association with tumor aggressiveness than did individual expression of each (Table 1). This combination also mirrored the effect on prognosis (Fig. 6B–G). There was no significant association between the individual expression of GLUD1 and SLC25A13. Then we assess the association of GLUD1 and PKM2 protein expression; the PKM2 is a splicing form of pyruvate kinase gene. The experimental data indicated that the GLUD1 expression was independent of PKM2 expression (Supplementary Fig. S8D), supporting the notion that both glucose metabolism and glutamine metabolism are important for the colorectal cancer. Taken together, colorectal cancer cells that can adapt to nutritional stress via regulation of GLUD1 and SLC25A13 contribute to tumor aggressiveness and result in worse prognosis (Fig. 6H).
Combined expression of GLUD1 and SLC25A13 was shown to be significantly associated with prognosis in colorectal cancer.
(A) Negative staining (−), positive staining (+), moderate staining (++), and strong staining (+++) for GLUD1 and SLC25A13 in colorectal cancer (scale bar, 200 mm). (B and C) Kaplan–Meier curves for recurrence-free survival (RFS) (B) and overall survival (OS) (C) according to GLUD1 expression. Differences between the two groups were evaluated by the log-rank test. Ordinate survival rate, abscissa years after surgery. (D,E) Kaplan–Meier curves for RFS (D) and OS (E) according to SLC25A13 expression. (F and G) Kaplan–Meier curves for RFS (F) and OS (G) according to combined expression of GLUD1 and SLC25A13. (H) Schematic overview of GLUD1-mediated Glu metabolism and SLC25A13-mediated ATP production. The citrin (SLC25A13) and aralar (SLC25A12; not shown) are components of aspartate-glutamate carrier (AGC), which functions collaboratively with oxoglutarate carrier (OGC) in the malate aspartate shuttle (MAS), a major intracellular pathway to transfer reducing equivalents. Glu: glutamic acid, Gln: glutamine, OAA: oxaloacetic acid, Asp: aspartic acid, α-KG: α-ketoglutaric acid, Mal: malic acid.
Discussion
We found that the metabolism of colorectal cancer cells that could survive under glucose-deprived conditions was significantly different from that of pancreatic ductal adenocarcinoma cells. In addition, the metabolism depended on genomic alterations that are uncharacterized and not restricted to KRAS mutation alone. In pancreatic ductal adenocarcinoma cells, a non-canonical glutamine pathway mediated by the oncogene KRAS that regulates glutamic-oxaloacetic transaminase 1 (GOT1) and GLUD1 has been described12. In non-small cell lung cancer, KRAS has a profound influence on glutamine metabolism through regulation of ME1 and GOT1, the high expression of which has been shown to correlate with poorer prognosis after radiotherapy26. A human breast carcinoma cell line that expresses an activated KRAS protein has been shown to have high glycolytic flux, low TCA cycle activity, and increased usage of glutamine for anabolic synthesis13. The solitary effects of oncogenic KRAS in metabolism are reportedly significant in various cancers; however, these effects were not observed for colorectal cancer in the present study. It would appear that in different cancers, metabolism varies, and the influence of oncogenes on metabolism is also diverse. Understanding the inherent metabolisms of different cancer types and how they are affected by oncogenes may lead to the development of specific cancer therapies.
In glutaminolysis, glutamine is first converted by glutaminase to glutamate and ammonia and then into α-ketoglutaric acid by either GLUD1 or less prominently by other transaminases. Jin et al. reported that expression of GLUD1 was elevated in breast and lung cancer tissue relative to that in normal tissue27. In addition, the mammalian target of rapamycin complex 1 the activity of which is dysregulated in many cancers, activates GLUD1 via inhibition of SIRT428. Cancer cells enhance glutaminolysis via positive regulation of GLUD1 to respond to increased demand of glutamine, which is used as a carbon source for energy generation, a component of protein and nucleotide, and in production of glutathione and NADPH. On the basis of the results of immunohistochemistry for 104 colorectal cancer specimens, Liu et al. reported that GLUD1 expression was high and associated with poorer prognosis29. Our results showing an association between GLUD1 expression and prognosis in colorectal cancer were consistent with those of Liu et al. Enhanced glutaminolysis by increased expression and activity of GLUD1 under nutritional stress may contribute to these findings. A recent study investigated regulation of redox homeostasis by GLUD1. Fumarate, the level of which is controlled by GLUD1, has been shown to positively regulate a ROS-scavenging enzyme, glutathione peroxidase 127. Our results showed that the increase in ROS levels under glucose-deprived condition was suppressed in DLD1 cells in which expression and activity of GLUD1 increased. Our results support those of the study by Jin et al. (Figs S5C, S6B and S6C).
A mutation in SLC25A13 coding for citrin has been shown to cause citrin deficiency that results in type-II citrullinemia and is characterized by hyperammonemia, steatohepatitis, and neuropsychiatric symptoms25. Although expression of SLC25A13 is known to be high in the liver, the significance of its expression in the colon has not been understood fully. In the present study, SLC25A13 expression was observed in the glandular cells of normal colorectal tissue and colorectal cancer tissue by immunohistochemistry (Fig. 6A). Mutations of SLC25A13 reportedly lead to hepatic overload and increase the risk of hepatocellular carcinoma30. Shohet et al. reported that higher expression of SLC25A13 correlated with a poorer outcome in neuroblastoma31. However, there are only a few studies that have examined the relevance of SLC25A13 to cancer. To the best of our knowledge, this is the first study to identify the importance of SLC25A13 in colorectal cancer. We showed that SLC25A13 was negatively associated with the depth of tumor invasion, extent of lymph node metastasis, distant metastasis, lymphatic invasion, and stage in colorectal cancer. Furthermore, SLC25A13 was profoundly correlated with colorectal cancer prognosis (Fig. 6D–G and Table S2). In addition, colorectal cancer cells were found to adapt to nutritional stress through regulation of SLC25A13. Therefore, SLC25A13 may have an important role in development of colorectal cancer.
In conclusion, we found that the metabolism of colorectal cancer, which is different from that of pancreatic cancer, depended on genomic alterations that are uncharacterized and not restricted to KRAS mutation alone. Colorectal cancer cells that have resistance to glucose depletion retain TCA cycle activity and rely on the delicate balance between ROS production and ATP production. GLUD1 and SLC25A13 have pivotal roles in nutritional stress and are associated with tumor aggressiveness and poorer prognosis of colorectal cancer. These proteins may serve as new targets in the treatment of refractory colorectal cancer.
Materials and Methods
Cell lines, culture and serum
The human colorectal cancer cell line HT29 was obtained from the ATCC (Manassas, VA, USA). Other colorectal cancer cell lines included DLD1, which was obtained from the Cell Resource Center for Biomedical Research Institute of Development, Aging, and Cancer (Tohoku University), and HCT116 and CaR1, which were obtained from the Japan Cancer Research Resources Bank (JCRB, Tokyo, Japan). Mouse embryonic fibroblast cell lines, parental NIH3T3, and transformed NIH3T3 cells by whole genomic DNA derived from a human colon cancer cell line (MA) were purchased from the RIKEN Cell Bank (Tsukuba, Japan). Cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM D6046; Sigma Aldrich, St. Louis, MO, USA) containing 10% fetal bovine serum (FBS), 100 U/ml penicillin, and 100 μg/ml streptomycin (Life Technologies, Carlsbad, CA, USA) at 37 °C in a humidified incubator with 5% CO2. Mice serum were obtained under anesthesia from wild type and KRAS mutation (Cre/LSL-KRASmut) BL6 mice, which were purchased from the Jackson laboratory (Bar Harbor, ME, USA), and maintained under the ethical agreement of animal facility at Osaka University (24–122; approved by chairman professor Kaneda).
Cell proliferation assay
Cells were added to 24-well plates at 10,000–15,000 cells per well in 0.5 ml of medium that was replaced the following day with glucose- and glutamine-deficient medium (DMEM D5030; Sigma Aldrich) supplemented with 10% dialyzed FBS (#26400–044; Invitrogen, Carlsbad, CA). Glucose (Wako Pure Chemical Industries, Osaka, Japan) and glutamine (Nacalai Tesque, Kyoto, Japan) were added. After cells were fixed in 80% methanol and stained with 0.2% crystal violet, the relative cell proliferation was quantified by absorbance at 595 nm.
Apoptosis detection
Apoptotic cells were analyzed using an annexin V [fluorescein isothiocyanate (FITC)-conjugated] apoptosis kit (K101–400; BioVision, Mountain View, CA, USA). In brief, cells growing on 6-cm dishes at 1–2 × 106 cells/dish for 24 h were loaded with 0.5 ml binding buffer, 5 μl annexin V-FITC, and 5 μl propidium iodide. After incubation at room temperature for 5 min in the dark, annexin V-FITC binding and PI staining were analyzed by flow cytometry.
Transfection of vector
Cells overexpressing KRAS were generated using a pCMV6-Entry plasmid containing the KRASG12V sequence with SgfI–MluI restriction sites (OriGene, Rockville, MD, USA) and Fugene6 (Roche Applied Science, Indianapolis, USA). KRAS knockdown cells were generated using the Lenti-vpak Packaging Kit (OriGene). A lentiviral shRNA construct, NM_033360 that targets human KRAS was obtained from the MISSION TRC-Hs1.0 library (Sigma). Control cells were transfected using the same procedure but with an empty control vector. DLD1 cells were selected using 2 μg/ml puromycin to establish stable cell lines.
Western blot analysis
Western blot analysis was performed as previously described32. KRAS antibody (SAB1404011; Sigma Aldrich; 1:250 dilution), ERK (#9102; Cell Signaling Technology, Beverly, MA; 1:1000 dilution), phosphorylated ERK (#9101 S; Cell Signaling Technology; 1:1000 dilution), CHOP (#5554; Cell Signaling Technology; 1:1000 dilution), Bcl-2 (#2870; Cell Signaling Technology; 1:1000 dilution), and ACTB antibody (A2066; Sigma Aldrich; 1:1000 dilution) were used.
Metabolomic analysis
Cells were cultured at 3 × 106 cells per 10-cm dish in 10 ml complete medium; the medium was replaced the following day with glucose(−) and glutamine(−) medium supplemented with 10% dialyzed fetal bovine serum. Glucose (10 mM) or glutamine (2 mM) was added, respectively. For hypoxia treatment, cells were incubated for 24 h with 1% O2. After culture medium was aspirated from the dish, the cells were washed twice using 5% mannitol solution and treated with 800 μl of methanol and 550 μl of Milli-Q water containing internal standards [H3304–1002; Human Metabolome Technologies (HMT), Tsuruoka, Japan]. The metabolite extracts were centrifuged at 2,300 g at 4 °C for 5 min. Next, to remove macromolecules, 800 μl of the upper aqueous layer was filtered by centrifugation using a Millipore 5-kDa cutoff filter at 9,100 g at 4 °C for 120 min, and resuspended in 50 μl of Milli-Q water for metabolomic analysis. Metabolomic analysis was performed using a C-SCOPE package in HMT and capillary electrophoresis–time-of-flight mass spectrometry for cation analysis and capillary electrophoresis–tandem mass spectrometry for anion analysis33,34. To obtain information regarding the peak, including the m/z ratio, migration time (MT), and peak area, the peaks were identified using automatic integration software (MasterHands; Keio University, Tsuruoka, Japan and MassHunter Quantitative Analysis B.04.00; Agilent Technologies, Santa Clara, CA, respectively). Peak areas were normalized against those of the internal standards, and the relative area values were further normalized by sample amounts. Hierarchical cluster analysis and principal component analysis were performed using HMT’s proprietary softwares, PeakStat and SampleStat, respectively.
Immunohistochemical analysis
Immunohistochemical analysis was performed as previously described32. Colorectal cancer tissue samples were obtained from 151 consecutive patients who underwent surgery at the Osaka University Hospital between 2006 and 2009. None of the patients had received chemotherapy or radiotherapy before surgery. All patients provided their written informed consent for use of their clinical samples in this study, which was approved by the Institutional Review Board. GLUD1 antibody (ab166618; Abcam; 1:500 dilution), SLC25A13 antibody (PA5–21991; Thermo Fisher Scientific; 1:500 dilution) were used.
Evaluation of staining
According to a method described in a previous report35, we scored the samples as follows: (A) fraction of positive stained cells: ≤5% scored 0, 6%–25% scored 1, 26%–50% scored 2, 51%–75% scored 3, and ≥75% scored 4; (B) intensity of staining: no staining scored 0, weak staining scored 1, moderate staining scored 2, and strong staining scored 3. The immunoreactivity score (IRS) was obtained by multiplying both parameters. The samples were divided into two groups as follows: low expression (IRS < 6) and high expression (IRS ≥ 6) groups.
Statistical analysis
Data were indicated as the mean ± standard deviation (SD). We performed Student’s t-test and Fisher’s exact test to determine statistically significant differences using JMP Pro 10 software (SAS Institute, Cary, NC, USA). The Kaplan–Meier method was used to assess recurrence-free survival and overall survival. A P-value < 0.05 was considered to indicate statistical significance.
Additional Information
How to cite this article: Miyo, M. et al. Metabolic Adaptation to Nutritional Stress in Human Colorectal Cancer. Sci. Rep. 6, 38415; doi: 10.1038/srep38415 (2016).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Torre, L. A. et al. Global cancer statistics, 2012. CA Cancer J Clin 65, 87–108, doi: 10.3322/caac.21262 (2015).
Nagasaki, T. et al. Feasibility and safety of laparoscopic surgery for metachronous colorectal cancer. Surg Today 45, 434–438, doi: 10.1007/s00595-014-0925-1 (2015).
Stattner, S. et al. Evolution of surgical microwave ablation for the treatment of colorectal cancer liver metastasis: review of the literature and a single centre experience. Surg Today 45, 407–415, doi: 10.1007/s00595-014-0879-3 (2015).
Brown, J. M. & Giaccia, A. J. The unique physiology of solid tumors: opportunities (and problems) for cancer therapy. Cancer Res 58, 1408–1416 (1998).
Dasu, A., Toma-Dasu, I. & Karlsson, M. Theoretical simulation of tumour oxygenation and results from acute and chronic hypoxia. Phys Med Biol 48, 2829–2842 (2003).
Warburg, O. On the origin of cancer cells. Science 123, 309–314 (1956).
Reitzer, L. J., Wice, B. M. & Kennell, D. Evidence that glutamine, not sugar, is the major energy source for cultured HeLa cells. J Biol Chem 254, 2669–2676 (1979).
DeBerardinis, R. J. et al. Beyond aerobic glycolysis: transformed cells can engage in glutamine metabolism that exceeds the requirement for protein and nucleotide synthesis. Proc Natl Acad Sci USA 104, 19345–19350, doi: 10.1073/pnas.0709747104 (2007).
Dang, C. V., Le, A. & Gao, P. MYC-induced cancer cell energy metabolism and therapeutic opportunities. Clin Cancer Res 15, 6479–6483, doi: 10.1158/1078-0432.ccr-09-0889 (2009).
Jiang, P., Du, W., Mancuso, A., Wellen, K. E. & Yang, X. Reciprocal regulation of p53 and malic enzymes modulates metabolism and senescence. Nature 493, 689–693, doi: 10.1038/nature11776 (2013).
Ying, H. et al. Oncogenic Kras maintains pancreatic tumors through regulation of anabolic glucose metabolism. Cell 149, 656–670, doi: 10.1016/j.cell.2012.01.058 (2012).
Son, J. et al. Glutamine supports pancreatic cancer growth through a KRAS-regulated metabolic pathway. Nature 496, 101–105, doi: 10.1038/nature12040 (2013).
Gaglio, D. et al. Oncogenic K-Ras decouples glucose and glutamine metabolism to support cancer cell growth. Mol Syst Biol 7, 523, doi: 10.1038/msb.2011.56 (2011).
Takiguchi, Y., Takahashi, Y., Kuriyama, T. & Miyamoto, T. NIH3T3 transfectant containing human K-ras oncogene shows enhanced metastatic activity after in vivo tumor growth or co-culture with fibroblasts. Clin Exp Metastasis 10, 351–360 (1992).
Ulrey, C. L., Liu, L., Andrews, L. G. & Tollefsbol, T. O. The impact of metabolism on DNA methylation. Hum Mol Genet 14 Spec No 1, R139–147, doi: 10.1093/hmg/ddi100 (2005).
Eng, J., Lynch, R. M. & Balaban, R. S. Nicotinamide adenine dinucleotide fluorescence spectroscopy and imaging of isolated cardiac myocytes. Biophys J 55, 621–630, doi: 10.1016/s0006-3495(89)82859-0 (1989).
Hanukoglu, I. & Rapoport, R. Routes and regulation of NADPH production in steroidogenic mitochondria. Endocr Res 21, 231–241 (1995).
Kami, K. et al. Metabolomic profiling of lung and prostate tumor tissues by capillary electrophoresis time-of-flight mass spectrometry. Metabolomics 9, 444–453, doi: 10.1007/s11306-012-0452-2 (2013).
Wei, Y., Pattingre, S., Sinha, S., Bassik, M. & Levine, B. JNK1-mediated phosphorylation of Bcl-2 regulates starvation-induced autophagy. Mol Cell 30, 678–688, doi: 10.1016/j.molcel.2008.06.001 (2008).
Zhang, J. et al. Asparagine plays a critical role in regulating cellular adaptation to glutamine depletion. Mol Cell 56, 205–218, doi: 10.1016/j.molcel.2014.08.018 (2014).
Marini, C. et al. Direct inhibition of hexokinase activity by metformin at least partially impairs glucose metabolism and tumor growth in experimental breast cancer. Cell Cycle 12, 3490–3499, doi: 10.4161/cc.26461 (2013).
Yang, C. et al. Glioblastoma cells require glutamate dehydrogenase to survive impairments of glucose metabolism or Akt signaling. Cancer Res 69, 7986–7993, doi: 10.1158/0008-5472.can-09-2266 (2009).
Smith, J. J. et al. Experimentally derived metastasis gene expression profile predicts recurrence and death in patients with colon cancer. Gastroenterology 138, 958–968, doi: 10.1053/j.gastro.2009.11.005 (2010).
Koseki, J. et al. Mathematical analysis predicts imbalanced IDH1/2 expression associates with 2-HG-inactivating beta-oxygenation pathway in colorectal cancer. Int J Oncol 46, 1181–1191, doi: 10.3892/ijo.2015.2833 (2015).
Palmieri, F. The mitochondrial transporter family (SLC25): physiological and pathological implications. Pflugers Arch 447, 689–709, doi: 10.1007/s00424-003-1099-7 (2004).
Chakrabarti, G. Mutant KRAS associated malic enzyme 1 expression is a predictive marker for radiation therapy response in non-small cell lung cancer. Radiat Oncol 10, 145, doi: 10.1186/s13014-015-0457-x (2015).
Jin, L. et al. Glutamate dehydrogenase 1 signals through antioxidant glutathione peroxidase 1 to regulate redox homeostasis and tumor growth. Cancer Cell 27, 257–270, doi: 10.1016/j.ccell.2014.12.006 (2015).
Csibi, A. et al. The mTORC1 pathway stimulates glutamine metabolism and cell proliferation by repressing SIRT4. Cell 153, 840–854, doi: 10.1016/j.cell.2013.04.023 (2013).
Liu, G. et al. Glutamate dehydrogenase is a novel prognostic marker and predicts metastases in colorectal cancer patients. J Transl Med 13, 144, doi: 10.1186/s12967-015-0500-6 (2015).
Rahman, N. Realizing the promise of cancer predisposition genes. Nature 505, 302–308, doi: 10.1038/nature12981 (2014).
Shohet, J. M. et al. A genome-wide search for promoters that respond to increased MYCN reveals both new oncogenic and tumor suppressor microRNAs associated with aggressive neuroblastoma. Cancer Res 71, 3841–3851, doi: 10.1158/0008-5472.can-10-4391 (2011).
Miyo, M. et al. Tumour-suppressive function of SIRT4 in human colorectal cancer. Br J Cancer 113, 492–499, doi: 10.1038/bjc.2015.226 (2015).
Ohashi, Y. et al. Depiction of metabolome changes in histidine-starved Escherichia coli by CE-TOFMS. Mol Biosyst 4, 135–147, doi: 10.1039/b714176a (2008).
Ooga, T. et al. Metabolomic anatomy of an animal model revealing homeostatic imbalances in dyslipidaemia. Mol Biosyst 7, 1217–1223, doi: 10.1039/c0mb00141d (2011).
Zhou, J. et al. RhoE is associated with relapse and prognosis of patients with colorectal cancer. Ann Surg Oncol 20, 175–182, doi: 10.1245/s10434-012-2472-6 (2013).
Acknowledgements
Research reported in this paper was supported by Ministry of Education, Culture, Sports, Science, and Technology (22130005, 25112708, 25134711, 24390315 and 26670604), by Ministry of Health, Labour and Welfare (H23–003) and by National Institute of Biomedical Innovation (12–4).
Author information
Authors and Affiliations
Contributions
M.M., M.K., N.N., T.S., K.N., H.M., J.K., K.K. and H.C. performed the experiments. N.H., J.N., T.H., N.G, F.M., T.S, T.M, H.S, Y.D., M.M. and H.I. designed the experiments. All authors contributed to writing the manuscript.
Ethics declarations
Competing interests
Institutional endowments were partly received from Taiho Pharmaceutical Co., Ltd. (http://www.taiho.co.jp/english/), the Evidence Based Medical (EBM) Research Center (http://ebmrce.co.jp/index.html), Chugai Co., Ltd. (http://www.chugai-pharm.co.jp/english/index.html), Yakult Honsha Co., Ltd. (http://www.yakult.co.jp/english/index.html), and Merck Co., Ltd. (http://www.merck.co.jp/en/index.html). Those funders had no role in the main experimental equipment, supply expenses, study design, data collection and analysis, decision to publish, or preparation of the manuscript for this work.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Miyo, M., Konno, M., Nishida, N. et al. Metabolic Adaptation to Nutritional Stress in Human Colorectal Cancer. Sci Rep 6, 38415 (2016). https://doi.org/10.1038/srep38415
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep38415
This article is cited by
-
Metabolic networks in mutant KRAS-driven tumours: tissue specificities and the microenvironment
Nature Reviews Cancer (2021)
-
A shift in glutamine nitrogen metabolism contributes to the malignant progression of cancer
Nature Communications (2020)
-
Changes in lipids composition and metabolism in colorectal cancer: a review
Lipids in Health and Disease (2019)
-
Rewiring urea cycle metabolism in cancer to support anabolism
Nature Reviews Cancer (2018)
-
Colorectal Cancer and Metabolism
Current Colorectal Cancer Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.