Abstract
Compartment boundaries prevent cell populations of different lineage from intermingling. In many cases, compartment boundaries are associated with morphological folds. However, in the Drosophila wing imaginal disc, fold formation at the anterior/posterior (A/P) compartment boundary is suppressed, probably as a prerequisite for the formation of a flat wing surface. Fold suppression depends on optomotor-blind (omb). Omb mutant animals develop a deep apical fold at the A/P boundary of the larval wing disc and an A/P cleft in the adult wing. A/P fold formation is controlled by different signaling pathways. Jun N-terminal kinase (JNK) and Yorkie (Yki) signaling are activated in cells along the fold and are necessary for the A/P fold to develop. While JNK promotes cell shape changes and cell death, Yki target genes are required to antagonize apoptosis, explaining why both pathways need to be active for the formation of a stable fold.
Similar content being viewed by others
Introduction
Epithelial folds contribute to morphogenetic movements and the separation of different cell groups, thus shaping the animal body1. For example, invagination of the Drosophila mesoderm is initiated by ventral fold formation2,3. Segmental and parasegmental grooves transiently appear in the trunk region of the Drosophila embryonic epidermis separating fields of cells of different fate4,5. A common but not the only mechanism of epithelial fold formation involves apical cell constriction and the acquisition of a bottle-like cell morphology6. By this mechanism tubes can be formed from epithelial sheets as in the development of tracheal and salivary gland primordia7. Folds also arise in postembryonic epithelia. In the Drosophila eye disc, differentiation depends on the progression of the morphogenetic furrow, a Hedgehog-dependent apical indentation of the eye field8,9. Apical and basal folds can also form at the borders which separate cell groups of different fate in the other imaginal discs of Drosophila larvae10,11,12. Folds can arise in the epithelia of all metazoa. In sea urchins, bottle cells have been shown to be required for invagination of the ectoderm13. In vertebrates, classical examples include the formation of the neural tube in chick14 and of the blastopore lip in amphibians15,16. Neuroectodermal grooves are also found during brain development in mouse and zebrafish17,18.
As outlined above, folds occur in many aspects of Drosophila development, making this species an excellent model organism to study mechanisms of fold formation. Drosophila has provided information on the molecular underpinnings of the required cell shape changes in various developmental paradigms. The best studied system is gastrulation which was investigated at the levels of genetics, cell biology and biophysics and which, therefore, can serve as a benchmark for studies in other systems19,20,21,22. In Drosophila gastrulation, the secreted protein Folded gastrulation (Fog) is central to inducing apical constriction of the invaginating cells23,24 but is dispensable in several other epithelial folding processes. In the Drosophila embryonic ectoderm, the formation of segmental grooves was shown to be controlled by engrailed (en) expression in boundary cells as well as by Hedgehog (Hh) and Wingless (Wg) signaling4. These segment polarity genes are not involved in specifying the position of nearby parasegmental grooves; Wg signaling is required, however, in a non-instructive, permissive role5. Formation and progression of the morphogenetic furrow in the larval eye disc is controlled by Hh and Decapentaplegic (Dpp). Induction of the cell adhesion molecule Cad86C by Hh and Dpp may be one of the mechanisms that effects cell shape changes in this tissue25. The way folds form, thus, can differ with regard to molecular and biomechanical requirements even within one epithelium. For example, in Drosophila gastrulation, the ventral furrow forms by apical constriction whereas the dorsal folds arise by a basal shift of the adherens junctions26.
In the larval wing disc pouch (the future wing blade), the normal, graded expression of Dpp and Wg does not instruct folding but rather is involved in maintaining the appropriate position-specific cell shape27. In the columnar main epithelium, loss of Dpp signaling causes extrusion of cells correlated with loss of the apical microtubule web28,29. Similarly, loss of Dpp targets Optomotor-blind (Omb) or Spalt lead to retraction of cells toward basal membrane30,31. Dpp signaling cell-autonomously promotes and maintains the elongated columnar shape of wing disc cells by regulating Rho1 and the regulatory light chain of non-muscle myosin II32. Wg signaling cell-autonomously promotes and maintains the columnar shape of wing disc cells through maintaining Vestigial (Vg) expression33. The Wg gradient, centered on the D/V boundary, instructs similarly shaped gradients of DE-cadherin concentration and apical cell circumference (high and constricted, respectively, close to the D/V boundary)34. The loss of Adenomatous polyposis coli (APC), a negative regulator of Wg signaling, leads to apical constriction and invagination independent of its effect on the DE-cadherin level. Wg, too, acts by activation of Rho1 and Myosin II35.
The folds which separate parasegments in the Drosophila embryonic ectoderm separate fields of cells that are related by lineage (compartments)36. But even in the absence of lineage restriction, groups of epithelial cells differing in gene expression and fated to develop into different structures tend to be separated by a fold. For instance, in the wing imaginal disc, which gives rise to adult notum, hinge, and wing blade, several folds orthogonal to the proximo-distal axis separate gene expression domains without being lineage restriction boundaries37,38. The most distal of these, the blade/hinge fold develops under control of the Omb-related T-box transcription factors Dorsocross (Doc)39. The proximal notum/hinge fold requires the complementary expression of Omb in hinge and Iroquois complex (Iro-C) in notum40. In contrast, the A/P compartment boundary is not associated with a fold and remains morphologically inconspicuous throughout development41, even though it derives from the corresponding infolded parasegmental boundary in the embryonic ectoderm42. It is conceivable that fold formation was selected against because of the structural requirement for the adult wing as a flight appendage. Indeed, fold formation is actively suppressed by a genetic program. Omb which is expressed in most of the pouch43 is required to maintain the normal epithelial structure at the A/P boundary. Reduction of Omb level in the pouch causes an apical morphogenetic defect at the A/P boundary due to contraction of cells along their apical-basal axis44. We here investigate the mechanisms of boundary fold formation elicited by Omb loss. We found that A/P fold formation is dependent on activation of JNK signaling induced by loss of omb. Loss of omb also induced Yki activity which promoted cell survival and attenuated the pro-apoptotic activity of JNK. Our results reveal a network of signaling pathways induced by loss of omb that controls cell shape and ensures cell survival of folded cells at the A/P boundary.
Results and Discussion
We and others have shown before that Omb prevents aberrant apical fold formation at the A/P boundary44,45. When Omb is directly or indirectly repressed in the P compartment, the A/P boundary of the wing develops a deep apical fold in the larval wing disc and a cleft in the adult wing44. We here use this model system to investigate the mechanisms of boundary fold formation under three genetic manupulations, ptc-Gal4 UAS-tkv, en-Gal4 UAS-omb-RNAi, and nub-Gal4 UAS-omb-RNAi (Fig. S1).
JNK is ectopically activated to initiate A/P fold formation
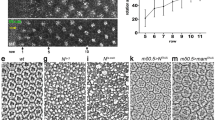
To investigate potential effectors downstream of Omb, we monitored the expression or activity of candidate targets. JNK signaling is an important pathway in the regulation of wing disc morphogenesis46,47. We monitored JNK pathway activity by monitoring transcription of the JNK target gene puckered (puc)48. In both ptc>tkv and en>omb-RNAi wing discs, puc was activated along the A/P fold (Fig. 1A,B). puc was initially activated in cells adjacent to the fold in early-mid L3, then its expression extended further into the A and P compartments (Fig. S2). These data indicate that JNK signaling is activated in the process of A/P fold formation.
JNK signaling is necessary for A/P fold formation.
(A,B) puc-lacZ was ectopically activated at the A/P fold. (C) Repression of JNK signaling by co-expressing a dominant negative form of JNK (bskDN) suppressed the A/P fold formation. Anti-Omb staining (green) demonstrates the efficiency of the posterior knock-down. Activation of JNK signaling by hepCA for a short duration induced extended folding (D) and Mmp1 expression (E) in the dpp-Gal4 domain. (F,F’) Focused induction of Mmp1 expression symmetrically on both sides of the A/P fold in en>omb-RNAi wing disc. (G) Co-expressing Mmp1-RNAi suppressed A/P fold formation. Arrowheads point at the position of the A/P boundary.
We next asked whether JNK activation is required for A/P fold generation. Co-expressing omb-RNAi and a dominant negative form of JNK, bskDN 49, was sufficient to suppress A/P fold formation (Fig. 1C).
This suggests that activation of JNK signaling is required in this process. To test for sufficiency of JNK activation for A/P fold formation, we activated JNK by expressing hepCA (encoding a constitutively active form of JNKK50) for a short duration controlled by dpp-Gal4 and tub-Gal80ts (continuously activation of hepCA induced severe apoptosis thereby disturbing observation of cell morphology). Under these conditions folds occurred throughout the dpp-Gal4 expression domain (Fig. 1D). The broad anterior activation of JNK signaling did not lead, however, to a discrete A/P fold.
The matrix metalloproteinase 1 (Mmp1) is induced by ectopic activation of JNK during morphological reorganization of epithelia51,52,53,54,55. When the JNK pathway was activated for 24 h in the dpp-Gal4 expression domain, Mmp1 was broadly induced anterior to the A/P boundary, similar to the plexus of epithelial folds observed under these conditions (Fig. 1E). However, in en>omb-RNAi wing discs, Mmp1 accumulated in a discrete stripe of A/P fold cells on both sides of the fold (Fig. 1F and F’). Co-expression of omb-RNAi and Mmp1-RNAi with nub-Gal4 rescued the A/P fold with full penetrance (Fig. 1G). Uniform reduction of omb expression on both sides of the A/P boundary, like posterior omb reduction, leads to A/P fold formation (Fig. S1E and F).
However, expression of Mmp1 with dpp-Gal4 along the A/P boundary did not generate a fold (Fig. S3). These data suggests that either additional gene expression changes, induced by the loss of omb, are necessary for A/P fold formation or that Mmp1 must be induced on both sides of the A/P boundary.
Yki-Diap1 signaling is activated parallel to the JNK pathway
It has been reported that Dpp and Wg repress JNK signaling to maintain survival of wing pouch cells. Loss of either Dpp or Wg signaling activates JNK-dependent apoptosis in the wing pouch50. However, omb knock-down did not induce apparent apoptosis at the A/P boundary44, although JNK signaling was activated (Fig. 1A and B). We assume that the apoptosis pathway is repressed in this case. Yki signaling can be induced by the JNK pathway56. Yki targets such as Death-associated inhibitor of apoptosis 1 (Diap1) and the microRNA bantam (ban) can repress apoptosis57,58,59. We analyzed transcription of the Yki target expanded (ex60 and observed that ex-lacZ was up-regulated at the A/P fold generated by en>omb-RNAi (Fig. 2B), suggesting an activation of Yki signaling during A/P fold formation. This was confirmed in nub>omb-RNAi wing discs (Fig. 2C). Upregulation at the A/P fold was also observed for Diap1 (Fig. 2E). The Yki target ban is suppressed by Omb in the medial wing discs61. Consistently, ban was up-regulated in the medial wing disc of nub>omb-RNAi larvae, with the strongest enhancement along the A/P boundary (Fig. 2G). Therefore, during A/P fold generation, ban and Diap1 were both activated and could suppress potential apoptosis induced by changes in cell shape and JNK activation.
Activation of Yki target genes in omb hypomorphic wing discs.
(A) Control experiment to show the normal ex-lacZ expression. (B,C) ex-lacZ was upregulated along the A/P fold in discs in which omb was knocked down in the posterior compartment (B) or in the entire pouch (C). (D) Control experiment to show the normal Diap1-lacZ expression. (E) Diap1-lacZ was upregulated along the A/P fold. (F) Control experiment to show the normal ban-lacZ expression. (G) ban-lacZ was ectopically activated at the A/P fold.
In order to determine whether the Yki targets were induced as a consequence of JNK signaling, we co-expressed bskDN and omb-RNAi in the nub-Gal4 domain. When JNK pathway and A/P fold were suppressed, ex-lacZ expression was still activated at the A/P boundary (Fig. S4A). This also held for Diap1 and ban expression (Fig. S4B and C). This suggests that in A/P fold formation Yki can be activated even when the JNK pathway is blocked. Yki activation, thus, appears to occur parallel to JNK signaling and is not sufficient for fold formation.
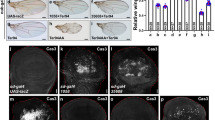
To test whether the suppression of cell death by Yki signaling is required for omb-loss induced A/P fold formation, yki-RNAi was co-expressed with omb-RNAi in the nub-Gal4 domain. As shown in Fig. 3A-A”, co-expressing yki-RNAi was sufficient to suppress the formation of A/P fold. Severe cell death occurred in this double knock-down. When p35 was co-expressed with omb-RNAi and yki-RNAi in the nub-Gal4 domain to inhibit apoptosis, cell death was effectively suppressed (Fig. 3B and B’) and the A/P fold appeared again (Fig. 3B and C). These data indicate that Yki signaling is required for A/P fold by ensuring cell survival.
Suppression of apoptosis is required for fold formation.
(A-A”) Co-expressing omb-RNAi and yki-RNAi by nub-Gal4 suppressed the A/P fold formation but induced severe apoptosis. (B,B’) Whenapoptosis was suppressed by co-expressing p35 an A/P fold was generated. (C) The fold generated in nub>[omb-RNAi + yki-RNAi + p35] wing discs extended along the A/P boundary (anterior compartment marked by Cubitus interruptus (Ci) expression, green).
Generally, abnormal activation of the JNK pathway induces apoptosis. For instance, expression of activated tumor genes or mutation in tumor suppressor genes lead to JNK-induced cell invasion and apoptosis55,62,63,64. However, in omb-knocked-down wing discs, the activation of JNK pathway did not cause cell death along the A/P fold44. We suggest that cell death is suppressed by the simultaneous induction of a cell survival pathway. Yki has an important role in promoting cell survival by driving the expression of downstream genes such as Diap1 and ban65. We found these genes upregulated along the A/P fold (Fig. 2). This suggests that Yki antagonizes apoptosis along the fold. Previous studies identified JNK as a promoter of Yki activity in the wing disc56,63,66. But this regulatory relationship is not absolute62,63,67. We found that co-expression of omb-RNAi and bskDN had no effect on ex, Diap1, and ban expression (Fig. S4). This indicates that, at the A/B boundary, Yki is activated parallel to JNK signaling.
Methods
Drosophila stocks
The transgenes used were as follows: en-Gal4, nub-Gal468, dpp-Gal4, ptc-Gal4, UAS-tkv, UAS-CD8-GFP, UAS-GFP, UAS-omb-RNAi44, tubP-Gal80ts69, UAS-MMP170, UAS-MMP1-RNAi55, UAS-p35, UAS-hepCA, UAS-bskDN, UAS-yki-RNAi (TsingHua Fly Center). Enhancer trap lines were hh-lacZ71, puc-lacZ, ex-lacZ, diap1-lacZ60, and ban-lacZ72. Stocks, if not mentioned otherwise, were obtained from the Bloomington stock center.
Larvae were raised at 25 °C. For efficient expression of RNAi transgenes, larvae were raised at 29 °C. Larvae containing Gal80ts-Gal4 combinations were raised at 18 °C and then were shifted to 29 °C that allows GAL4 to function and activate transcription of UAS controlled transgenes.
Immunohistochemistry
Dissected wing imaginal discs were fixed and stained with antibodies according to the standard procedures. The primary antibodies used were: rabbit anti-Omb, 1:1000; mouse and rabbit anti-beta-galactosidase, 1:2000 (Promega); rabbit anti-caspase3, 1:200 (Santa Cruz), rat anti-2A1 (Ci), 1:200 (Developmental Studies Hybridoma Bank, DSHB); and mouse anti-Mmp1, 1:200 (DSHB). Secondary antibodies used were goat anti-rabbit DyLight 488, goat anti-rat DyLight 488, goat anti-rabbit DyLight 549, goat anti-mouse DyLight 488, goat anti-rabbit Cy3, and goat anti-rabbit Cy5, were diluted 1:200 (Agrisera). Actin was visualized with Rhodamine- phalloidin, 1:1000 (Cytoskeleton). Images were collected using a Leica TCS SP2 AOBS confocal microscope.
Additional Information
How to cite this article: Liu, S. et al. Fold formation at the compartment boundary of Drosophila wing requires Yki signaling to suppress JNK dependent apoptosis. Sci. Rep. 6, 38003; doi: 10.1038/srep38003 (2016).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Pilot, F. & Lecuit, T. Compartmentalized morphogenesis in epithelia: from cell to tissue shape. Dev. Dyn 232, 685–694 (2005).
Oda, H. & Tsukita, S. Real-time imaging of cell-cell adherens junctions reveals that Drosophila mesoderm invagination begins with two phases of apical constriction of cells. J. Cell Sci 114, 493–501 (2001).
Kam, Z., Minden, J. S., Agard, D. A., Sedat, J. W. & Leptin, M. Drosophila gastrulation: analysis of cell shape changes in living embryos by three-dimensional fluorescence microscopy. Development 112, 365–370 (1991).
Larsen, C. W., Hirst, E., Alexandre, C. & Vincent, J. P. Segment boundary formation in Drosophila embryos. Development 130, 5625–5635 (2003).
Larsen, C., Bardet, P. L., Vincent, J. P. & Alexandre, C. Specification and positioning of parasegment grooves in Drosophila. Dev. Biol 321, 310–318 (2008).
Sawyer, J. M. et al. Apical constriction: a cell shape change that can drive morphogenesis. Dev. Biol 341, 5–19 (2010).
Maruyama, R. & Andrew, D. J. Drosophila as a model for epithelial tube formation. Dev. Dyn 241, 119–135 (2012).
Corrigall, D., Walther, R. F., Rodriguez, L., Fichelson, P. & Pichaud, F. Hedgehog signaling is a principal inducer of Myosin-II-driven cell ingression in Drosophila epithelia. Dev. Cell 13, 730–742 (2007).
Wolff, T. & Ready, D. F. The beginning of pattern formation in the Drosophila compound eye: the morphogenetic furrow and the second mitotic wave. Development 113, 841–850 (1991).
Dominguez, M. & Casares, F. Organ specification-growth control connection: new in-sights from the Drosophila eye-antennal disc. Dev. Dyn 232, 673–684 (2005).
Greenberg, L. & Hatini, V. Essential roles for lines in mediating leg and antennal proximodistal patterning and generating a stable Notch signaling interface at segment borders. Dev. Biol 330, 93–104 (2009).
Zirin, J. D. & Mann, R. S. Nubbin and Teashirt mark barriers to clonal growth along the proximal-distal axis of the Drosophila wing. Dev. Biol 304, 745–758 (2007).
Kimberly, E. L. & Hardin, J. Bottle cells are required for the initiation of primary invagination in the sea urchin embryo. Dev. Biol 204, 235–250 (1998).
Schoenwolf, G. C. & Franks, M. V. Quantitative analyses of changes in cell shapes during bending of the avian neural plate. Dev. Biol 105, 257–272 (1984).
Hardin, J. & Keller, R. The behaviour and function of bottle cells during gastrulation of Xenopus laevis. Development 103, 211–230 (1988).
Lee, J. Y. & Harland, R. M. Endocytosis is required for efficient apical constriction during Xenopus gastrulation. Curr. Biol 20, 253–258 (2010).
Inoue, T. et al. Role of cadherins in maintaining the compartment boundary between the cortex and striatum during development. Development 128, 561–569 (2001).
Langenberg, T. & Brand, M. Lineage restriction maintains a stable organizer cell population at the zebrafish midbrain-hindbrain boundary. Development 132, 3209–3216 (2005).
Dawes-Hoang, R. E. et al. Folded gastrulation, cell shape change and the control of myosin localization. Development 132, 4165–4178 (2005).
Fox, D. T. & Peifer, M. Abelson kinase (Abl) and RhoGEF2 regulate actin organization during cell constriction in Drosophila. Development 134, 567–578 (2007).
Kolsch, V., Seher, T., Fernandez-Ballester, G. J., Serrano, L. & Leptin, M. Control of Drosophila gastrulation by apical localization of adherens junctions and RhoGEF2. Science 315, 384–386 (2007).
Polyakov, O. et al. Passive mechanical forces control cell-shape change during Drosophila ventral furrow formation. Biophys. J 107, 998–1010 (2014).
Costa, M., Wilson, E. T. & Wieschaus, E. A putative cell signal encoded by the folded gastrulation gene coordinates cell shape changes during Drosophila gastrulation. Cell 76, 1075–1089 (1994).
Fuse, N., Yu, F. & Hirose, S. Gprk2 adjusts Fog signaling to organize cell movements in Drosophila gastrulation. Development 140, 4246–4255 (2013).
Schlichting, K. & Dahmann, C. Hedgehog and Dpp signaling induce cadherin Cad86C expression in the morphogenetic furrow during Drosophila eye development. Mech. Dev 125, 712–728 (2008).
Wang, Y. C., Khan, Z., Kaschube, M. & Wieschaus, E. F. Differential positioning of adherens junctions is associated with initiation of epithelial folding. Nature 484, 390–393 (2012).
McClure, K. D. & Schubiger, G. Developmental analysis and squamous morphogenesis of the peripodial epithelium in Drosophila imaginal discs. Development 132, 5033–5042 (2005).
Gibson, M. C. & Perrimon, N. Extrusion and death of DPP/BMP-compromised epithelial cells in the developing Drosophila wing. Science 307, 1785–1789 (2005).
Shen, J. & Dahmann, C. Extrusion of cells with inappropriate Dpp signaling from Drosophila wing disc epithelia. Science 307, 1789–1790 (2005).
Shen, J., Dahmann, C. & Pflugfelder, G. O. Spatial discontinuity of optomotor-blind expression in the Drosophila wing imaginal disc disrupts epithelial architecture and promotes cell sorting. BMC Dev. Biol 10, 23 (2010).
Tang, W., Wang, D. & Shen, J. Asymmetric distribution of Spalt in Drosophila wing squamous and columnar epithelia ensures correct cell morphogenesis. Sci. Rep 6, 30236 (2016).
Widmann, T. J. & Dahmann, C. Dpp signaling promotes the cuboidal-to-columnar shape transition of Drosophila wing disc epithelia by regulating Rho1. J. Cell Sci 122, 1362–1373 (2009a).
Widmann, T. J. & Dahmann, C. Wingless signaling and the control of cell shape in Drosophila wing imaginal discs. Dev. Biol 334, 161–173 (2009b).
Jaiswal, M., Agrawal, N. & Sinha, P. Fat and Wingless signaling oppositely regulate epithelial cell-cell adhesion and distal wing development in Drosophila. Development 133, 925–935 (2006).
Zimmerman, S. G., Thorpe, L. M., Medrano, V. R., Mallozzi, C. A. & McCartney, B. M. Apical constriction and invagination downstream of the canonical Wnt signaling pathway require Rho1 and Myosin II. Dev. Biol 340, 54–66 (2010).
Blair, S. S. Lineage compartments in Drosophila. Curr. Biol 13, R548–551 (2003).
Villa-Cuesta, E., Gonzalez-Perez, E. & Modolell, J. Apposition of iroquois expressing and non-expressing cells leads to cell sorting and fold formation in the Drosophila imaginal wing disc. BMC Dev. Biol 7, 106 (2007).
Villa-Cuesta, E. & Modolell, J. Mutual repression between msh and Iro-C is an essential component of the boundary between body wall and wing in Drosophila. Development 132, 4087–4096 (2005).
Sui, L., Pflugfelder, G. O. & Shen, J. The Dorsocross T-box transcription factors promote tissue morphogenesis in the Drosophila wing imaginal disc. Development 139, 2773–2782 (2012).
Wang, D., Li, L., Lu, J., Liu, S. & Shen, J. Complementary expression of optomotor-blind and the Iroquois complex promotes fold formation to separate wing notum and hinge territories. Dev. Biol 416, 225–234 (2016).
Brower, D. L., Smith, R. J. & Wilcox, M. Cell shapes on the surface of the Drosophila wing imaginal disc. J. Embryol. Exp. Morphol 67, 137–151 (1982).
Vincent, J. P. Compartment boundaries: where, why and how? Int. J. Dev. Biol 42, 311–315 (1998).
Grimm, S. & Pflugfelder, G. O. Control of the gene optomotor-blind in Drosophila wing development by decapentaplegic and wingless. Science 271, 1601–1604 (1996).
Shen, J., Dorner, C., Bahlo, A. & Pflugfelder, G. O. optomotor-blind suppresses instability at the A/P compartment boundary of the Drosophila wing. Mech. Dev 125, 233–246 (2008).
Umemori, M., Takemura, M., Maeda, K., Ohba, K. & Adachi-Yamada, T. Drosophila T-box transcription factor Optomotor-blind prevents pathological folding and local overgrowth in wing epithelium through confining Hh signal. Dev. Biol 308, 68–81 (2007).
Agnes, F., Suzanne, M. & Noselli, S. The Drosophila JNK pathway controls the morphogenesis of imaginal discs during metamorphosis. Development 126, 5453–5462 (1999).
Tripura, C., Chandrika, N. P., Susmitha, V. N., Noselli, S. & Shashidhara, L. S. Regulation and activity of JNK signaling in the wing disc peripodial membrane during adult morphogenesis in Drosophila. Int. J. Dev. Biol 55, 583–590 (2011).
Martin-Blanco, E. et al. Puckered encodes a phosphatase that mediates a feedback loop regulating JNK activity during dorsal closure in Drosophila. Genes Dev 12, 557–570 (1998).
Weber, U., Paricio, N. & Mlodzik, M. Jun mediates Frizzled-induced R3/R4 cell fate distinction and planar polarity determination in the Drosophila eye. Development 127, 3619–3629 (2000).
Adachi-Yamada, T., Fujimura-Kamada, K., Nishida, Y. & Matsumoto, K. Distortion of proximodistal information causes JNK-dependent apoptosis in Drosophila wing. Nature 400, 166–169 (1999).
Jezowska, B. et al. A dual function of Drosophila capping protein on DE-cadherin maintains epithelial integrity and prevents JNK-mediated apoptosis. Dev. Biol 360, 143–159 (2011).
Leong, G. R., Goulding, K. R., Amin, N., Richardson, H. E. & Brumby, A. M. Scribble mutants promote aPKC and JNK-dependent epithelial neoplasia independently of Crumbs. BMC Biol 7, 62 (2009).
Srivastava, A., Pastor-Pareja, J. C., Igaki, T., Pagliarini, R. & Xu, T. Basement membrane remodeling is essential for Drosophila disc eversion and tumor invasion. Proc. Natl. Acad. Sci. USA 104, 2721–2726 (2007).
Stevens, L. J. & Page-McCaw, A. A secreted MMP is required for reepithelialization during wound healing. Mol. Biol. Cell 23, 1068–1079 (2012).
Uhlirova, M. & Bohmann, D. JNK- and Fos-regulated Mmp1 expression cooperates with Ras to induce invasive tumors in Drosophila. EMBO J 25, 5294–5304 (2006).
Sun, G. & Irvine, K. D. Regulation of Hippo signaling by Jun kinase signaling during compensatory cell proliferation and regeneration, and in neoplastic tumors. Dev. Biol 350, 139–151 (2011).
Huang, J., Wu, S., Barrera, J., Matthews, K. & Pan, D. The Hippo signaling pathway coordinately regulates cell proliferation and apoptosis by inactivating Yorkie, the Drosophila Homolog of YAP. Cell 122, 421–434 (2005).
Nolo, R., Morrison, C. M., Tao, C., Zhang, X. & Halder, G. The bantam microRNA is a target of the hippo tumor-suppressor pathway. Curr. Biol 16, 1895–1904 (2006).
Thompson, B. J. & Cohen, S. M. The Hippo pathway regulates the bantam microRNA to control cell proliferation and apoptosis in Drosophila. Cell 126, 767–774 (2006).
Graves, H. K., Woodfield, S. E., Yang, C. C., Halder, G. & Bergmann, A. Notch signaling activates Yorkie non-cell autonomously in Drosophila. PLoS One 7, e37615 (2012).
Zhang, X., Luo, D., Pflugfelder, G. O. & Shen, J. Dpp signaling inhibits proliferation in the Drosophila wing by Omb-dependent regional control of bantam. Development 140, 2917–2922 (2013).
Chen, C. L., Schroeder, M. C., Kango-Singh, M., Tao, C. & Halder, G. Tumor suppression by cell competition through regulation of the Hippo pathway. Proc. Natl. Acad. Sci. USA 109, 484–489 (2012).
Enomoto, M. & Igaki, T. Src controls tumorigenesis via JNK-dependent regulation of the Hippo pathway in Drosophila. EMBO Rep 14, 65–72 (2013).
Igaki, T., Pagliarini, R. A. & Xu, T. Loss of cell polarity drives tumor growth and invasion through JNK activation in Drosophila. Curr. Biol 16, 1139–1146 (2006).
Zhao, B., Tumaneng, K. & Guan, K. L. The Hippo pathway in organ size control, tissue regeneration and stem cell self-renewal. Nat. Cell. Biol 13, 877–883 (2011).
Willsey, H. R. et al. Localized JNK signaling regulates organ size during development. Elife 5 (2016).
Doggett, K., Grusche, F. A., Richardson, H. E. & Brumby, A. M. Loss of the Drosophila cell polarity regulator Scribbled promotes epithelial tissue overgrowth and cooperation with oncogenic Ras-Raf through impaired Hippo pathway signaling. BMC Dev. Biol 11, 57 (2011).
Calleja, M., Moreno, E., Pelaz, S. & Morata, G. Visualization of gene expression in living adult Drosophila. Science 274, 252–255 (1996).
McGuire, S. E., Le, P. T., Osborn, A. J., Matsumoto, K. & Davis, R. L. Spatiotemporal rescue of memory dysfunction in Drosophila. Science 302, 1765–1768 (2003).
Page-McCaw, A., Serano, J., Sante, J. M. & Rubin, G. M. Drosophila matrix metalloproteinases are required for tissue remodeling, but not embryonic development. Dev. Cell 4, 95–106 (2003).
Lee, J. J., von Kessler, D. P., Parks, S. & Beachy, P. A. Secretion and localized transcription suggest a role in positional signaling for products of the segmentation gene hedgehog. Cell 71, 33–50 (1992).
Oh, H. & Irvine, K. D. Cooperative regulation of growth by Yorkie and Mad through bantam. Dev. Cell 20, 109–122 (2011).
Acknowledgements
We thank the Bloomington stock center and TsingHua Fly Center for fly stocks. This research was supported by the National Natural Science Foundation of China [NSFC31372255] and the 973 Program [2013CB127603].
Author information
Authors and Affiliations
Contributions
J.S. developed the concept and designed the experiments. S.L. performed the experiments. D.W., G.P. and J.S. analysed the data and wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Liu, S., Sun, J., Wang, D. et al. Fold formation at the compartment boundary of Drosophila wing requires Yki signaling to suppress JNK dependent apoptosis. Sci Rep 6, 38003 (2016). https://doi.org/10.1038/srep38003
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep38003
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.