Abstract
Recent studies have shown that circular RNAs (circRNAs) are abundant, widely expressed in mammals, and can display cell-type specific expression. However, how production of circRNAs is regulated and their precise biological function remains largely unknown. To study how circRNAs might be regulated during colorectal cancer progression, we used three isogenic colon cancer cell lines that differ only in KRAS mutation status. Cellular RNAs from the parental DLD-1 cells that contain both wild-type and G13D mutant KRAS alleles and isogenically-matched derivative cell lines, DKO-1 (mutant KRAS allele only) and DKs-8 (wild-type KRAS allele only) were analyzed using RNA-Seq. We developed a bioinformatics pipeline to identify and evaluate circRNA candidates from RNA-Seq data. Hundreds of high-quality circRNA candidates were identified in each cell line. Remarkably, circRNAs were significantly down-regulated at a global level in DLD-1 and DKO-1 cells compared to DKs-8 cells, indicating a widespread effect of mutant KRAS on circRNA abundance. This finding was confirmed in two independent colon cancer cell lines HCT116 (KRAS mutant) and HKe3 (KRAS WT). In all three cell lines, circRNAs were also found in secreted extracellular-vesicles, and circRNAs were more abundant in exosomes than cells. Our results suggest that circRNAs may serve as promising cancer biomarkers.
Similar content being viewed by others
Introduction
Circular RNAs (circRNAs) were first reported more than 30 years ago1,2,3,4, but had long been perceived as occasional RNA splicing errors until recent genome-wide analyses powered by next generation sequencing (NGS) technologies have shown these are bona fide RNA species. Studies during the past several years have identified a large number of exonic and intronic circRNAs across the eukaryotic lineage, including human, mouse, zebrafish, worms, fungi, and plants5,6,7,8. Based on the assumption that the abundance of circRNAs is much lower than that of linear RNAs, early studies typically use RNase R, a magnesium-dependent 3′ to 5′ exoribonuclease, to deplete linear RNAs before sequencing9. However, recent work showed that the abundance of circRNAs is similar to or higher than that of linear transcripts for about one in eight human genes10, which can be partially explained by higher cellular stability and longer half-life of circRNAs compared to linear mRNAs11. The observed high abundance of circRNAs suggests that RNase R treatment is likely to be unnecessary in NGS-based analysis of circRNAs, consistent with the identification of 7112 circRNA candidates from non-poly(A)-selected libraries generated by the ENCODE project12,13. It is now clear that circRNAs are evolutionarily conserved, exhibit cell-specific expression patterns, and are regulated independent of their linear transcripts10,14,15. For example, circRNAs are enriched in brain and accumulate to the highest levels in the aging central nervous system16,17. Recent studies also showed that circRNAs can be transferred to human exosomes18, where they are enriched and stable19. These findings suggest that circRNAs are prevalent, abundant, and potentially functional.
Knowledge about the general sequence features, biogenesis, and putative functions of circRNAs, especially exonic circRNAs, has gradually accumulated11. Because both circRNAs and linear RNAs are spliced from pre-mRNAs, the competition between circularization and linear splicing may play a role in the regulation of gene expression20. Moreover, introns between exons may be retained when exons are circularized21. Circularization of exonic circRNAs typically involves the canonical GU-AG splice site pairs22 and can contain one or multiple exons. On average, single-exon circRNAs form with exons that are three times longer than non-circularized exons10. Exon circularization is promoted by pairing of reverse complementary sequences within introns bracketing circRNAs; reverse complimentary sequences are primarily Alu repeats23,24,25. Two possible mechanisms for the formation of exonic circRNAs have been proposed, and both involve the canonical spliceosome11. Two circRNAs in mammals have been shown to function as miRNA sponges5, but significant enrichment of miRNA binding sites was not found for the majority of circRNA candidates12,13.
Although other non-coding RNAs have been shown to play critical roles in cancer, the association between circRNAs and cancer is largely unknown26,27,28. In this study, we performed deep RNA-Seq analysis of rRNA-depleted total RNA libraries to characterize circRNA expression in three isogenically-matched human colon cancer cell lines that differ only in the mutation status of the KRAS oncogene. The parental DLD-1 cells contain both wild-type and G13D mutant KRAS alleles, whereas the isogenically-matched derivative cell lines DKO-1 and DKs-8 contain only a mutant KRAS and a wild-type KRAS allele, respectively. KRAS mutations occur in approximately 34–45% of colon cancers29,30 and have been associated with a wide range of tumor-promoting effects31. We developed an integrated bioinformatics pipeline to identify, confirm and annotate circRNAs based on RNA-Seq data. Using the pipeline, we studied both cellular and exosomal circRNAs in the three cell lines, with confirmation of altered circRNAs in a second set of isogenically matched cell lines. To our knowledge, this is the first report describing the impact of a well-established oncogene on the abundance of circRNAs.
Results
Bioinformatics pipeline
Exonic circRNAs largely result from back-spliced exons, in which splice junctions are formed by an upstream 5′ splice acceptor and a downstream 3′ splice donor. Back-splice reads mapping to such junctions are the most important indicator for circRNAs that can be gleaned from RNA-Seq data5,11,16,23,32,33. Similar to the existing pipeline used by Memczak et al.5, our pipeline (Fig. 1A) uses the presence of back-splice reads to identify exonic circRNA candidates. However, multiple mapping positions are allowed when mapping anchors in our pipeline. Find-circ only reports a random mapping position and may therefore miss some circRNAs (false negatives). Moreover, because one read may be considered as a back-splicing candidate at one position or a linear gapped mapping at another position, find-circ may also introduce false positives. Thus, allowing multiple mapping position in our pipeline may help reduce both false positives and false negatives. Briefly, one paired-end read was used as two single-end reads for mapping to the genome. Mappable reads were discarded because back-splice reads cannot be mapped to the genome directly. The 5′ and 3′ termini of unmapped reads were then extracted as anchors, which were aligned to the genome independently with multiple mapping allowed. Because multiple mapping is allowed, all possible pairs of anchor alignments were evaluated. If any of these pairs correspond to a normal linear gapped mapping, the read was discarded. For the remaining reads, all the possible extensions that could be extended to reconstruct the original read with a maximum of two mismatches were further considered. Then we will search the GU/AC splice sites for each extension. If any extensions with the GU/AC splice sites, the read was considered as with GU/AC splice sites. Extended alignments flanked by GU/AG splice sites were used to define a back-splice read.
Bioinformatics pipeline and analysis of cell circRNAs.
(A) Computational pipeline. (B–D) Pearson Correlation analysis between cell replicates for DKs-8, DLD-1, and DKO-1 colorectal cell lines, respectively. (E) Distribution of percentages of back-splice mates with corresponding mates that can be mapped (PCMM) for each sample.
Contamination from other biological sources may affect both the identification and quantification of circRNAs. To check possible contaminations from bacteria and viruses, we built a database with all bacterial and viral sequences and blasted all back-splicing mates against the database. For cellular RNAs, 99.6% to 99.8% of the mates had no hits to the database and none of them had a hit with two or less mismatches. For exosomal RNAs, 91.8% to 99.4% of the mates had no hits to the database and only a few had a hit with two or less mismatches (Table S1). Next, all back-splicing mates were mapped to the bovine genome34 both linearly and using the back-splicing detection algorithm. The linear mapping percentages were close to 0 for all samples, and no more than 2.2% of the back-splicing mates could be back-splicing-mapped to the bovine genome (Table S2). These results show that the vast majority of the identified circular RNAs are not from bacterial and viral contamination and the potential contamination from the bovine sources is very limited. We discarded all back-splicing reads that can be mapped to bacterial, viral, or bovine genomes from downstream analysis to avoid any influence from possible contamination.
Sequence fragments supported by two or more remaining back-splice reads were considered as circRNA candidates, and those supported by ten or more back-splice reads were considered as high quality candidates. Finally, circRNA candidates with sequence fragment lengths between 100 and 1000,000 bp were reported by the pipeline.
Identification of circRNA candidates in colorectal cancer cells
We prepared cellular RNA libraries from the three isogenic-KRAS CRC cell lines, each with two biological replicates. RNase R treatment was not applied during library construction. Sequencing was performed at high depth, with ~100 million reads per sample. Applying the above-described pipeline to cellular RNAs from the three cell lines identified thousands of circRNA candidates (Table S3) and hundreds of high quality circRNA candidates in each biological replicate (Table 1). Among the 1620 high quality candidates detected in our study, 1395 (86.1%) were found in the circBase database35.
To assess the reproducibility of the data, we generated scatter plots comparing the back-splice read counts of individual circRNAs from replicates of the three cell lines (Fig. 1B–D). As shown, the vast majority of all candidates were supported by consistent identification of back-splice reads in all replicates. Person’s correlations between replicates were 0.99, 0.93, and 0.94 for DKs-8, DLD-1, and DKO-1, respectively. These scatter plots show circRNA candidates with higher read counts are closer to the diagonal, suggesting that reproducibility tended to be higher for circRNA candidates with high back-splice read counts. Therefore, our downstream analyses focused only on high quality candidates with at least ten back-splice reads.
To further evaluate the reliability of the identified circRNA candidates, we leveraged the paired end information. As shown in Figure S1, if one mate of a paired end read mapped to a back-splice junction (Mate a), the corresponding mate could be mapped to the candidate circRNA sequence either within the circle (Mate b’) or crossing the back-splice junction (Mate b). For each high quality candidate, we calculated the Percentage of back-splice mates with Corresponding Mates that can be mapped to the candidate circRNA sequence (PCMM), i.e. the percentage of properly paired back-splice mates. As shown in Fig. 1E, the median percentages ranged from 88.2% to 90.0% across the six samples, suggesting high reliability of these circRNA candidates. We also tested RNase R resistance of circRNAs in the DKO-1 and DKs-8 cell lines with the top four most abundant circRNAs. As shown in Figure S2, these circRNAs were enriched by RNase R (R+) treatment compared to mock treated controls (R−). Thus circRNAs are resisted to RNase R treatment. Taken together, these results suggest that a large number of circRNAs can be reliably and reproducibly identified and quantified in the three cell lines.
Down-regulation of circRNAs in KRAS mutant cells
To test whether the expression levels of circRNAs are regulated by KRAS, we compared the levels of circular RNA candidates between the mutant and wild-type KRAS cell lines. circRNAs were globally down-regulated in the mutant KRAS DKO-1 (Fig. 2A) and DLD-1 (Fig. 2B) cell lines compared to the wild-type KRAS DKs-8 cell line. Specifically, 443 and 305 circRNAs were significantly down-regulated in DKO-1 and DLD-1 cells, respectively (False Discovery Rate [FDR] < 0.01 and Fold Change [FC] >2). In contrast, only 5 and 13 circRNAs were significantly up-regulated in DKO-1 and DLD-1 cells, respectively. Among the top ten most abundant circRNAs in distinct genes, seven were significantly down-regulated in DKO-1 cells and five of them were also significantly down-regulated in DLD-1 cells (Table 2). These results suggest that circRNAs are down-regulated in KRAS mutant cells at a global level.
Differential expression analysis for cellular circRNAs.
(A,B) circRNA differential expression analysis between mutant and wild-type KRAS cells. (C,D) Differential expression results for circRNAs and their host genes. Histograms of each gene and corresponding circRNAs log2FCs are shown above the X and Y-axes, respectively. (E) qRT-PCR results for seven selected circRNAs between mutant and wild-type cells. (F) qRT-PCR results for host genes of selected circRNAs. (G) qRT-PCR results for seven selected circRNAs between HCT116 and HKe3 cells. (H) qRT-PCR results for host genes between HCT116 and HKe3 cells. (Two-tailed, paired t-test was used for the analysis. *denote p values ≤ 0.1 and **≤0.05).
We next sought to determine whether circRNA down-regulation was due to down-regulation of corresponding host genes. Figure 2C and D provide a direct comparison of the differential expression results for circRNAs and their host genes between each of the two mutant cell lines and the wild-type cell line. While the log-fold changes of the host genes exhibited a symmetrical distribution around 0, the log-fold changes of circRNAs were negatively shifted toward decreased abundance in mutant KRAS cell lines. The correlations between log-fold changes of circRNAs and host genes were 0.19 and 0.16, respectively, for the two comparisons. Using the most abundant circRNA candidate circRNA chr4:187627717-187630999 as an example, we found that this circRNA was down regulated by 6.6- and 5.3-fold in DLD-1 and DKO-1 cells, respectively compared to DKs-8 cells. In contrast, the host gene FAT1 was only down-regulated by 1.7- and 1.8-fold, respectively. These data suggest that circRNAs can be regulated independently of their corresponding host genes.
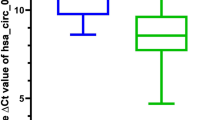
To validate our findings, we performed qRT-PCR analysis for seven out of the ten most abundant circRNA candidates. As shown in Fig. 2E, all were confirmed by qRT-PCR and six out of the seven circRNA candidates were significantly down-regulated in at least one mutant cell line compared with the wild-type cell line (two-tailed, paired t-tests was used for the analysis, where *are p values ≤ 0.1 and **≤0.05). As a comparison, Fig. 2F shows different trends for the host genes of these circRNAs. These results further confirm our finding that circRNAs are down-regulated in mutant KRAS cells and that the regulation of circRNAs can occur independent of their host genes.
To further strengthen our conclusion, we performed additional experiments using another pair of isogenically-matched human colon cancer cell lines, HCT116 and HKe3. Derived from a completely different cancer, HCT116 harbors mutant G13D KRAS while its clonal derivative HKe3 contains wild-type KRAS36. Consistent with our previous results, all seven circRNAs assayed were down regulated in the HCT116 cells compared to the HKe3 cell line as shown in Fig. 2G. Among them, circFAT1 was significantly down-regulated in the mutant KRAS cell line (HCT116). Furthermore, the host genes for these candidates were not significantly differentially expressed between HCT116 and HKe3 cell lines (Fig. 2H). These results support our finding that circRNAs are down-regulated in mutant KRAS cells and that the regulation of circRNAs can occur independently of their host genes.
circRNAs in exosomes
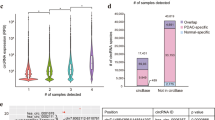
Several recent reports have identified extracellular circRNAs18,19. To test whether circRNAs could be detected in the exosomes of colon cancer cell lines, we performed RNA-Seq analysis for exosomal RNAs from the three cell lines, each with three biological replicates. High quality circRNA candidates were identified in all three cell lines (Table S4). However, the number of high quality candidates varied among the replicates. Because the variation between DKs-8 exosomal replicates was relatively low, we focused our downstream analyses on data from DKs-8 derived exosomes. High quality exosomal circRNA candidates identified in this cell line were well supported by paired end information. Specifically, the median percentages of properly paired back-splice mates were 90.0%, 91.3%, and 91.7% for the three replicates, respectively (Fig. 3A). Table 3 shows the ten most abundant exosomal circRNA candidates in distinct genes in DKs-8 cells. Interestingly, seven of these circRNAs were also the top ten most abundant circRNAs candidates in DKs-8 cells (Table 2).
Bioinformatics analysis of circRNAs in exosomes.
(A) Distribution of PCMM values for three DKs-8 exosomes replicates. (B) RT-PCR results for five selected circRNAs between mutant and wild-type cell lines in exosomes. (C) RT-PCR results for host genes of confirmed circRNAs in exosome. (Two-tailed, paired t-test was used for the analysis. *Denote p values ≤ 0.1 and **≤0.05).
To validate the RNA-Seq results, qRT-PCR analysis was also performed on these consistently present and abundant circRNA candidates in exosomes. As shown in Fig. 3B, five of these circRNAs were confirmed as present in exosomes and three of them were differentially expressed in at least one set of mutant cell line derived exosomes compared with the wild-type cell line exosomes (two-tailed, paired t-tests was used for the analysis, where *are p values ≤ 0.1 and **≤0.05). Among them, circFAT1 was significantly down regulated in DKO-1 as compared to DKs-8 exosomes (Fig. 3B); this circRNA followed the same trend in cells (Fig. 2E). Meanwhile, circRTN4 was significantly up regulated in DLD-1 exosomes (Fig. 3B), while it was significantly down regulated in DLD-1 cells (Fig. 2E). The mRNA expression levels of these circRNA host genes were also tested by qRT-PCR and the results shown in Fig. 3C. The mRNA expression levels of both FAT1 and RTN4 were up regulated in exosomes from mutant KRAS cells. Therefore the shift in the relative circRNA levels was not the same as that for their linear mRNA host genes when comparing mutant and wild-type KRAS derived exosomes. These results suggested that there is a complex exosomal trafficking mechanism for circular RNAs. This is interesting given the increased abundance of RNA-binding proteins present in wild-type KRAS as compared to mutant KRAS cell-derived exosomes37. Results from the proteomic analysis of these exosomes may explain both the relative differences in circRNA and linear RNA content in DKs-8 as compared to DKO-1 and DLD-1 exosomes, as well as the relatively consistent levels of such RNAs in DKs-8 exosomes, given that such RNAs might be trafficked by these specifically exosomally-localized DKs-8 enriched RNA-binding proteins.
Relative abundance of circular and linear transcripts
Because RNAse R treatment was not applied during the RNA library construction in this study, the resulting RNA-Seq data allowed us to directly compare the abundance of circRNAs and their linear host RNAs. Similar to previous studies15,17, we used the ratio between Expression level of exons With Circular RNAs and Expression level of exons with No Circular RNAs (EWC/ENC) to quantify the relative abundance of these two types of transcripts.
For cellular RNAs, the median EWC/ENC ratios ranged from 1.57 to 1.84 across the three cell lines (Fig. 4A). Similar analysis was performed on exosomal RNAs, where the median EWC/ENC ratios were much higher and ranged from 2.56 to 4.26 (Fig. 4B). Figure 4C and D show the read coverage depth plots for the most abundant circRNA circFAT1 (chr4:187627717-187630999) in DKs-8 cells and exosomes, respectively. The exon corresponding to circFAT1 (red) had a much higher read depth compared with other exons (blue). The EWC/ENC ratios were 3.5 and 3.0 for the two cell replicates, respectively, and 7.1, 7.6, and 7.7 for the three exosome replicates, respectively. These results are consistent with recent reports that circRNAs are more abundant than their host linear RNAs15,17 and provide additional evidence that circRNAs are likely to be more stable than their linear transcripts10,11. In addition, our results suggest that cirRNAs are enriched in exosomes, which is consistent with a recent publication19.
Relative expression levels of circRNAs compared to linear transcripts.
(A,B) Distribution of EWC/ENC values for cellular and exosomal RNAs, respectively. DKO-1.exo.1 was excluded because only 2 high quality candidates were identified in this replicate (Table S4). (C,D) Expression levels of exons with and without circRNA with the FAT1 gene in DKs-8 cells and exosomes, respectively.
Discussion
In this work, we determined the circRNA expression profiles in both cells and exosomes from three CRC cell lines that differ only in KRAS mutation status. Hundreds of high quality circRNA candidates were identified in cellular RNAs and we discovered that they could be transferred to exosomes. circRNAs tended to be more abundant in exosomes. Importantly, we showed that circRNA abundance was down-regulated at a global level in mutant KRAS cell lines, suggesting a potential involvement of circRNAs in oncogenesis.
There are complex regulatory mechanisms for both circRNA and host gene expression. Although circRNAs were down-regulated in both DLD-1 and HCT166 based cell lines, it is difficult to conclude that the circRNAs are directly regulated by KRAS. One possibility is that down-regulation of circRNAs in KRAS mutant cells is caused by their increased exporting to exosomes. However, as shown in Fig. 4B, the EWC/ENC median values were 2.77, 4.15, 3.38, 2.96 and 2.56 for KRAS mutant exosomes and were 4.26, 3.43 and 3.25 for KRAS WT exosomes (Fig. 4B). The median values in KRAS mutant exosomes were comparable to that in KRAS WT exosomes. Moreover, Fig. 3B shows that two of three significantly regulated circRNAs between KRAS mutant and WT exosomes were down-regulated in KRAS mutant (circFAT1 and circARHGAP5). These data suggest that circRNAs are not enriched in exosomes of the KRAS mutant cells. We also examined the expression levels of the RNA-editing enzymes ADAR and the RNA-binding protein QKI, which were reported as circRNA regulators25,28. The ADAR was decreased in the KRAS mutant cells, which may lead to an increase of circRNAs. QKI was down-regulated in KRAS mutant cells, which may lead to down-regulation of circRNAs. More broadly, we studied the expression levels of all RNA-binding proteins from RBPDB38, following the approach taken by Conn et al.28. Six were found to be differentially expressed (FDR < 0.01 and absolute log2FC > 1) in KRAS mutant cell lines compared with wild-type cell lines (ELAVL2, RBMS3, BICC1, MSI1, RBM44, and LARP6). Three of these were up regulated and the others were down regulated. The most up-regulated gene, ELAVL2, can function as an alternative pre-mRNA splicing regulator in mammalian neurons39,40. The most down-regulated gene, MSI1, is also an important post-transcriptional regulator41,42. These genes may serve as candidate circRNA regulators. However, our previous work shows that the correlation of mRNA and protein expression level is low for RNA-binding proteins43. Further investigation will be needed to precisely define how circRNAs are regulated.
Methods
Cell Culture
Cells were cultured in DMEM supplemented with 10% bovine growth serum until 80% confluent. To collect exosomes, cells were then washed 3 times with PBS and cultured for 24 hr in serum-free medium. The medium was collected and replaced with ionomycin-containing media for 1 hr, after which ionomycin-containing media was collected and pooled with the previously collected serum-free medium.
Exosome isolation
Exosomes were isolated from conditioned medium of DKO-1, Dks-8, and DLD-1 cells, with slight modification44. Pooled media as describted above was centrifuged for 10 min at 300 × g to remove cellular debris, and the resulting supernatant was then filtered through a 0.22-um polyethersulfone filter (Nalgene, Rochester, NY) to reduce microparticle contamination. The filtrate was concentrated ~300-fold with a 100,000 molecular-weight cutoff centrifugal concentrator (Millipore). The concentrate was then subjected to high-speed centrifugation at 150,000 × g for 2 hr. The resulting exosome-enriched pellet was resuspended in PBS containing 25 mM HEPES (pH 7.2) and washed by centrifuging again at 150,000 × g for 3 hr. The wash steps were repeated a minimum of 3 times until no trace of phenol-red was detected. The resulting pellet was resuspended in PBS containing 25 mM HEPES (pH 7.2) and protein concentrations were determined with a MicroBCA kit (Pierce). The number of exosomes per ug of protein was determined by nanoparticle tracking analysis (NanoSight, Wiltshire, UK) and the results can be found in a recent publication from us in which the same exosome preparations were used (Figure S1A in that paper)37. Analysis was performed on three independent preparations of exosomes.
RNA purification
Total RNA from exosomes and cells was isolated using TRIzol (Life Technologies). In the case of exosomal RNA isolation TRIzol was incubated with 100 ul or less of concentrated exosomes for an extended 15 min incubation prior to chloroform extraction. RNA pellets were resuspended in 60 μl of RNase-free water and were then re-purified using the miRNeasy kit (QIAGEN). Final RNAs were eluted with two rounds of 30 ul water extraction.
mRNA library preparation and sequencing
Total RNA containing both long RNA as well as miRNA fractions was extracted from exosomes or cell lines using Trizol followed by miRNeasy Kit purification. Final elution was in 60 μl RNase free sterile distilled water. The concentration and integrity of the extracted total RNA was estimated by Qubit® 2.0 Fluorometer (Invitrogen, Carlsbad, California), and Agilent 2100 Bioanalyzer (Applied Biosystems, Carlsbad, CA), respectively. RNA samples with a RIN value of at least 7.0 or higher were used for further processing.
Approximately 500 ng of total RNA was required for proceeding to downstream RNA-seq applications. Briefly, a Ribo-zero Magnetic Gold rRNA removal kit (Epicenter, IIlumina Inc.) was used to remove ribosomal RNA from the total RNA. Next, first strand synthesis was performed using NEBNext RNA first strand synthesis module (New England BioLabs Inc., Ipswich, MA, USA). Immediately, directional second strand synthesis was performed using NEBNExt Ultra Directional second strand synthesis kit. Following this, cDNAs were used for standard library preparation protocol using NEBNext® DNA Library Prep Master Mix Set for Illumina® with slight modifications. Briefly, end-repair was performed followed by polyA addition and custom adapter ligation. Post-ligated materials were individually barcoded with unique in-house genomics service lab (GSL) primers. Library quality was assessed by Qubit 2.0 Fluorometer, and the library concentration was estimated by utilizing a DNA 1000 chip on an Agilent 2100 Bioanalyzer. Accurate quantification for sequencing applications was determined using the qPCR-based KAPA Biosystems Library Quantification kit (Kapa Biosystems, Inc., Woburn, MA). Each library was diluted to a final concentration of 12.5 nM and pooled equimolar prior to clustering. Paired-End (PE) sequencing was performed on all samples. Raw reads were de-multiplexed using a bcl2fastq conversion software v1.8.3 (Illumina, Inc.) with default settings.
circRNA identification
Reads with length 100 bp were mapped to the UCSC hg19 human genome (with mitochondrial sequences) by Bowtie 2 with up to 2 mismatches (version 2.2.3)45. Paired 3′ and 5′ end anchors with length 20 bp were extracted for each unmapped read. Anchor pairs were mapped to the above genome with no mismatches and up to 40 mapping positions using Bowtie 2. Refseq gene annotations from UCSC were used to annotate circRNA candidates46. Custom PERL scripts were used to implement the pipeline (Fig. 1A).
Contamination analysis
We built a database with all bacterial and viral sequences from the NCBI nt database47 (ftp://ftp.ncbi.nlm.nih.gov/genomes/refseq/bacteria/assembly_summary.txt and ftp://ftp.ncbi.nih.gov/genomes/Viruses/). All back-splicing mates were blasted48 (ncbi-blast-2.3.0+) against the database with default parameters. Next, back-splicing mates were linearly mapped to the bovine genome34 by Tophat2 (version 2.0.12) with up to 2 mismatches49. Moreover, these mates were mapped to the bovine genome using the back-splicing detection algorithm described above.
Differential expression analysis
To count reads mapped normally to genes, paired end reads were mapped to the hg19 human genome using Tophat2 with up to 2 mismatches. Htseq-count (version 0.6.0) with default parameters was used to count reads mapped to genes with the refSeq annotation50. It is worth noting that host gene expression was quantified using reads from both linear and circRNAs because existing tools cannot separate linear and circRNA counts based on RNA-Seq data from total RNAs. Accordingly, our results may have underestimated the difference between linear and circRNAs levels. The correlation between the regulation of linear and circRNAs would be even lower if we were able to separate the read counts. The EdgeR R package (version 3.6.8) was used for differential expression analysis51. This package uses the Trimmed Mean of M-values (TMM) normalization method to remove systematic technical effects that occur in the data to minimize the impact of technical bias on differential expression analysis results52. Moreover, the empirical Bayes method used in the package enables gene-specific variation estimates even when the number of replicate samples is very small. This method has been demonstrated in experiments with only two replicates53, and thus is particularly appropriate for our study. For differential expression analysis of circRNAs, back-splicing read counts of circRNAs were added to the bottom of gene count list as new genes for the normalization purpose. The cutoffs for log2 fold change (log2FC) and FDR were |log2FC| > 1 and FDR < 0.01.
Evaluate circRNA candidates by paired end information
To evaluate circRNA candidates by paired end sequencing information, corresponding mates of paired end reads were initially extracted for back-splice mapped mates. Then, fragments from the 5′ ends of linear transcripts of circRNAs with length 100 nt were copied to the 3′ end of these linear transcripts. These mates were then mapped to the modified linear sequences using Tophat2 with up to 2 mismatches. PCMM values were calculated as the number of reads both mates are mappable/the number of reads with back-splice mapped mate.
Compare expression levels between circRNAs and linear transcripts
To compare the relative expression levels between exons with and without circRNA candidates, DEPTH tool from Samtools package (version 0.1.19–44428 cd) was used to report read depths for genes with circRNA candidates54. The mean value of read depths from an exon was used as the read depth of the exon. EWC/ENC value was calculated as the mean depth of exons with circRNAs/without circRNAs.
RT-PCR
To validate circRNA species, 0.5 ug of total RNA was reverse transcribed in a 30 μl reaction using AccuScript Hi-Fi RT kit with random hexamers according to manufactures protocol (#200820, Agilent Technologies). The resultant cDNA was diluted 4-fold in RNase- and DNase-free water and approximately 14 ng was used as template for each qPCR reaction. qPCR was performed in technical triplicates for each amplicon using SsoAdvanced Universal SYBR® Green Supermix (Bio-Rad). qPCR reactions were conducted on a Bio-Rad CFX384 instrument and relative expression levels were obtained using cycle threshold (Ct) values obtained by instrument software. All Ct values ≥31 were considered as background and discarded from further analysis. Triplicate C(t) values were averaged and normalized to U6 snRNA. Fold-changes were calculated using the ΔΔC(t) method, where: Δ = C(t)circRNA - C(t)U6 snRNA, and ΔΔC(t) = ΔC(t)DKO or DLD − ΔC(t)DKs, and FC = 2^ΔΔC(t). Analysis was performed on three independent cell and exosomal samples. Forward (F) and reverse (R) primers used in qPCR analysis were designed against head-to-tail junctions of putative circRNA products as follows: FAT1- (F) ACGCCAGAGCCATCTCTAAT, (R) GCAATGGGGAGACATTTGGC; HIPK3- (F) ATGGCCTCACAAGTCTTGGT, (R) TGGCCGACCCAAAGTCTATT; ARHGAP5- (F) TGATCTTGAAGATGTTTCTGCACAG, (R) CATCTAACTCCTGGTCAGAAGTG; MAN1A2- (F) TTCGAGCTGATCATGAGAAGG, (R) GCAAGTAGGCCTCCAATAAA; RHOBTB3- (F) TAAAGGCTGAAGCGTCACATTAT, (R) CTCGATTACATTTGAAACATCCCCA; RTN4- (F) CAACTAAGAAGAGGCGCCTG, (R) AGACTGGAGTGGTGTTTGGT; SMARCA5- (F) GGCTTGTGGATCAGAATCTGAACA, (R) TCTCTATAGTCTTCTCCTTCGAAGT. All primer sequences are 5′ to 3′ (Table S5). Table S6 includes primer details and sequence information for the linear RNA species. The primer sequences were blasted against the NCBI human genomic + transcript database to ensure specific amplification of the intended targets. Moreover, the melt curves showed that each primer set only had one specific peak, suggesting that the amplicon was specific and no other secondary targets were being amplified.
Additional Information
How to cite this article: Dou, Y. et al. Circular RNAs are down-regulated in KRAS mutant colon cancer cells and can be transferred to exosomes. Sci. Rep. 6, 37982; doi: 10.1038/srep37982 (2016).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Hsu, M. T. & Coca-Prados, M. Electron microscopic evidence for the circular form of RNA in the cytoplasm of eukaryotic cells. Nature 280, 339–340 (1979).
Nigro, J. M. et al. Scrambled exons. Cell 64, 607–613 (1991).
Cocquerelle, C., Daubersies, P., Majerus, M. A., Kerckaert, J. P. & Bailleul, B. Splicing with inverted order of exons occurs proximal to large introns. The EMBO journal 11, 1095–1098 (1992).
Saad, F. A. et al. A 3′ consensus splice mutation in the human dystrophin gene detected by a screening for intra-exonic deletions. Human molecular genetics 1, 345–346 (1992).
Memczak, S. et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 495, 333–338, doi: 10.1038/nature11928 (2013).
Wang, P. L. et al. Circular RNA Is Expressed across the Eukaryotic Tree of Life. Plos One 9, doi: 10.1371/journal.pone, 0090859 (2014).
Zhang, Z. et al. Discovery of Replicating Circular RNAs by RNA-Seq and Computational Algorithms. PLoS pathogens 10, e1004553, doi: 10.1371/journal.ppat.1004553 (2014).
Zhang, Y. et al. Circular intronic long noncoding RNAs. Molecular cell 51, 792–806, doi: 10.1016/j.molcel.2013.08.017 (2013).
Cheng, Z. F. & Deutscher, M. P. An important role for RNase R in mRNA decay. Molecular cell 17, 313–318, doi: 10.1016/j.molcel.2004.11.048 (2005).
Jeck, W. R. et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. Rna 19, 141–157, doi: 10.1261/rna.035667.112 (2013).
Jeck, W. R. & Sharpless, N. E. Detecting and characterizing circular RNAs. Nature biotechnology 32, 453–461, doi: 10.1038/nbt.2890 (2014).
Guo, J. U., Agarwal, V., Guo, H. & Bartel, D. P. Expanded identification and characterization of mammalian circular RNAs. Genome biology 15, 409, doi: 10.1186/s13059-014-0409-z (2014).
Consortium, E. P. The ENCODE (ENCyclopedia Of DNA Elements) Project. Science 306, 636–640, doi: 10.1126/science.1105136 (2004).
Salzman, J., Gawad, C., Wang, P. L., Lacayo, N. & Brown, P. O. Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. Plos One 7, e30733, doi: 10.1371/journal.pone.0030733 (2012).
Salzman, J., Chen, R. E., Olsen, M. N., Wang, P. L. & Brown, P. O. Cell-type specific features of circular RNA expression. PLoS genetics 9, e1003777, doi: 10.1371/journal.pgen.1003777 (2013).
Westholm, J. O. et al. Genome-wide Analysis of Drosophila Circular RNAs Reveals Their Structural and Sequence Properties and Age-Dependent Neural Accumulation. Cell reports 9, 1966–1980, doi: 10.1016/j.celrep.2014.10.062 (2014).
Rybak-Wolf, A. et al. Circular RNAs in the Mammalian Brain Are Highly Abundant, Conserved, and Dynamically Expressed. Molecular cell 58, 870–885, doi: 10.1016/j.molcel.2015.03.027 (2015).
Bahn, J. H. et al. The Landscape of MicroRNA, Piwi-Interacting RNA, and Circular RNA in Human Saliva. Clinical chemistry 61, 221–230, doi: 10.1373/clinchem.2014.230433 (2015).
Li, Y. et al. Circular RNA is enriched and stable in exosomes: a promising biomarker for cancer diagnosis. Cell Research, doi: 10.1038/cr.2015.82 (2015).
Ashwal-Fluss, R. et al. circRNA biogenesis competes with pre-mRNA splicing. Molecular cell 56, 55–66, doi: 10.1016/j.molcel.2014.08.019 (2014).
Li, Z. et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nature structural & molecular biology, doi: 10.1038/nsmb.2959 (2015).
Black, D. L. Mechanisms of alternative pre-messenger RNA splicing. Annual review of biochemistry 72, 291–336, doi: 10.1146/annurev.biochem.72.121801.161720 (2003).
Zhang, X. O. et al. Complementary sequence-mediated exon circularization. Cell 159, 134–147, doi: 10.1016/j.cell.2014.09.001 (2014).
Liang, D. & Wilusz, J. E. Short intronic repeat sequences facilitate circular RNA production. Genes & development 28, 2233–2247, doi: 10.1101/gad.251926.114 (2014).
Ivanov, A. et al. Analysis of Intron Sequences Reveals Hallmarks of Circular RNA Biogenesis in Animals. Cell reports, doi: 10.1016/j.celrep.2014.12.019 (2014).
Hansen, T. B., Kjems, J. & Damgaard, C. K. Circular RNA and miR-7 in cancer. Cancer research 73, 5609–5612, doi: 10.1158/0008-5472.CAN-13-1568 (2013).
Martens-Uzunova, E. S. et al. Long noncoding RNA in prostate, bladder, and kidney cancer. European urology 65, 1140–1151, doi: 10.1016/j.eururo.2013.12.003 (2014).
Conn, S. J. et al. The RNA Binding Protein Quaking Regulates Formation of circRNAs. Cell 160, 1125–1134, doi: 10.1016/j.cell.2015.02.014 (2015).
Vogelstein, B. et al. Genetic alterations during colorectal-tumor development. N Engl J Med 319, 525–532, doi: 10.1056/NEJM198809013190901 (1988).
Wong, R. & Cunningham, D. Using Predictive Biomarkers to Select Patients With Advanced Colorectal Cancer for Treatment With Epidermal Growth Factor Receptor Antibodies. J Clin Oncol 26, 5668–5670 (2008).
Velho, S. & Haigis, K. M. Regulation of homeostasis and oncogenesis in the intestinal epithelium by Ras. Experimental cell research 317, 2732–2739, doi: 10.1016/j.yexcr.2011.06.002 (2011).
Hansen, T. B. et al. Natural RNA circles function as efficient microRNA sponges. Nature 495, 384–388, doi: 10.1038/Nature11993 (2013).
Bachmayr-Heyda, A. et al. Correlation of circular RNA abundance with proliferation - exemplified with colorectal and ovarian cancer, idiopathic lung fibrosis, and normal human tissues. Scientific reports 5, 8057, doi: 10.1038/srep08057 (2015).
Elsik, C. G. et al. Bovine Genome Database: new tools for gleaning function from the Bos taurus genome. Nucleic Acids Res 44, D834–839, doi: 10.1093/nar/gkv1077 (2016).
Glazar, P., Papavasileiou, P. & Rajewsky, N. circBase: a database for circular RNAs. Rna 20, 1666–1670, doi: 10.1261/rna.043687.113 (2014).
Shirasawa, S., Furuse, M., Yokoyama, N. & Sasazuki, T. Altered Growth of Human Colon Cancer Cell-Lines Disrupted at Activated Ki-Ras. Science 260, 85–88, doi: 10.1126/science.8465203 (1993).
Demory Beckler, M. et al. Proteomic analysis of exosomes from mutant KRAS colon cancer cells identifies intercellular transfer of mutant KRAS. Mol Cell Proteomics 12, 343–355, doi: 10.1074/mcp.M112.022806 (2013).
Cook, K. B., Kazan, H., Zuberi, K., Morris, Q. & Hughes, T. R. RBPDB: a database of RNA-binding specificities. Nucleic Acids Res 39, D301–D308, doi: 10.1093/nar/gkq1069 (2011).
Ince-Dunn, G. et al. Neuronal Elav-like (Hu) proteins regulate RNA splicing and abundance to control glutamate levels and neuronal excitability. Neuron 75, 1067–1080, doi: 10.1016/j.neuron.2012.07.009 (2012).
Izquierdo, J. M. Hu antigen R (HuR) functions as an alternative pre-mRNA splicing regulator of Fas apoptosis-promoting receptor on exon definition. J Biol Chem 283, 19077–19084, doi: 10.1074/jbc.M800017200 (2008).
Katz, Y. et al. Musashi proteins are post-transcriptional regulators of the epithelial-luminal cell state. Elife 3, e03915, doi: 10.7554/eLife.03915 (2014).
Ratti, A. et al. Post-transcriptional regulation of neuro-oncological ventral antigen 1 by the neuronal RNA-binding proteins ELAV. J Biol Chem 283, 7531–7541, doi: 10.1074/jbc.M706082200 (2008).
Zhang, B. et al. Proteogenomic characterization of human colon and rectal cancer. Nature 513, 382–387, doi: 10.1038/nature13438 (2014).
Higginbotham, J. N. et al. Amphiregulin exosomes increase cancer cell invasion. Current biology: CB 21, 779–786, doi: 10.1016/j.cub.2011.03.043 (2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nature methods 9, 357–359, doi: 10.1038/nmeth.1923 (2012).
Kent, W. J. et al. The human genome browser at UCSC. Genome Res 12, 996–1006, doi: 10.1101/Gr.229102 (2002).
Pruitt, K. D., Tatusova, T. & Maglott, D. R. NCBI reference sequences (RefSeq): a curated non-redundant sequence database of genomes, transcripts and proteins. Nucleic Acids Res 35, D61–65, doi: 10.1093/nar/gkl842 (2007).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25, 3389–3402 (1997).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome biology 14, R36, doi: 10.1186/gb-2013-14-4-r36 (2013).
Anders, S., Pyl, P. T. & Huber, W. HTSeq-a Python framework to work with high-throughput sequencing data. Bioinformatics, doi: 10.1093/bioinformatics/btu638 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140, doi: 10.1093/bioinformatics/btp616 (2010).
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome biology 11, R25, doi: 10.1186/gb-2010-11-3-r25 (2010).
Chen, Y., Aaron, T. L. & Gordon, K. S. In Statistical Analysis of Next Generation Sequencing Data 51–74 (Springer International Publishing, 2014).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079, doi: 10.1093/bioinformatics/btp352 (2009).
Acknowledgements
This work was supported by a grant from the NIH, U19 CA179514.
Author information
Authors and Affiliations
Contributions
Y.D. and B.Z. designed the study. Y.D. performed all computational, bioinformatics, and statistical analyses. D.J.C., J.L.F., J.N.H., D.K.J., A.M.W., R.J.C. and J.G.P. developed the experimental protocol and prepared the samples. N.P. and S.L. performed the RNA-Seq experiments. D.J.C. performed all RT-PCR analysis. Y.D., J.G.P., J.L.F. and B.Z. wrote the manuscript with comments and final approval from all authors.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Dou, Y., Cha, D., Franklin, J. et al. Circular RNAs are down-regulated in KRAS mutant colon cancer cells and can be transferred to exosomes. Sci Rep 6, 37982 (2016). https://doi.org/10.1038/srep37982
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep37982
This article is cited by
-
Tumor-derived small extracellular vesicles in cancer invasion and metastasis: molecular mechanisms, and clinical significance
Molecular Cancer (2024)
-
The effects of exercise on epigenetic modifications: focus on DNA methylation, histone modifications and non-coding RNAs
Human Cell (2024)
-
The emerging landscape of exosomal CircRNAs in solid cancers and hematological malignancies
Biomarker Research (2022)
-
Emerging role of non-coding RNAs in the regulation of KRAS
Cancer Cell International (2022)
-
Roles and clinical application of exosomal circRNAs in the diagnosis and treatment of malignant tumors
Journal of Translational Medicine (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.