Abstract
The inverse Ising problem and its generalizations to Potts and continuous spin models have recently attracted much attention thanks to their successful applications in the statistical modeling of biological data. In the standard setting, the parameters of an Ising model (couplings and fields) are inferred using a sample of equilibrium configurations drawn from the Boltzmann distribution. However, in the context of biological applications, quantitative information for a limited number of microscopic spins configurations has recently become available. In this paper, we extend the usual setting of the inverse Ising model by developing an integrative approach combining the equilibrium sample with (possibly noisy) measurements of the energy performed for a number of arbitrary configurations. Using simulated data, we show that our integrative approach outperforms standard inference based only on the equilibrium sample or the energy measurements, including error correction of noisy energy measurements. As a biological proof-of-concept application, we show that mutational fitness landscapes in proteins can be better described when combining evolutionary sequence data with complementary structural information about mutant sequences.
Similar content being viewed by others
Introduction
High-dimensional data characterizing the collective behavior of complex systems are increasingly available across disciplines. A global statistical description is needed to unveil the organizing principles ruling such systems and to extract information from raw data. Statistical physics provides a powerful framework to do so. A paradigmatic example is represented by the Ising model and its generalizations to Potts and continuous spin variables, which have recently become popular for extracting information from large-scale biological datasets. Successful examples are as different as multiple-sequence alignments of evolutionary related proteins1,2,3, gene-expression profiles4, spiking patterns of neural networks5,6, or the collective behavior of bird flocks7. This widespread use is motivated by the observation that the least constrained (i.e. maximum-entropy8) statistical model reproducing empirical single-variable and pairwise frequencies observed in a list of equilibrium configurations is given by a Boltzmann distribution:

with s = (s1,..., sN) being a configuration of N binary variables or ‘spins’. Inferring the couplings J = {Jij}1≤i<j≤N and fields h = {hi}1≤i≤N in the Hamiltonian  from data, known as the inverse Ising problem, is computationally hard for large systems (N ≫ 1). It involves the calculation of the partition function
from data, known as the inverse Ising problem, is computationally hard for large systems (N ≫ 1). It involves the calculation of the partition function  as a sum over an exponential number of configurations. The need to develop efficient approximate approaches has recently triggered important work within the statistical-physics community, cf. e.g. refs 9, 10, 11, 12, 13, 14, 15, 16, 17.
as a sum over an exponential number of configurations. The need to develop efficient approximate approaches has recently triggered important work within the statistical-physics community, cf. e.g. refs 9, 10, 11, 12, 13, 14, 15, 16, 17.
Despite the broad interest in inverse problems, the methodological setting has remained rather limited: all of this literature, including the biological cases mentioned in the beginning, seeks to estimate model parameters starting from a set of configurations s, which are considered to be at equilibrium and independently drawn from P(s). Real data, however, may be quite different. In biological systems, “microscopic spins configurations” (e.g. amino-acid sequences) are increasingly accessible to experimental techniques, and quantitative information for a limited number of particular configurations (e.g. three-dimensional structures, measured activities or thermodynamic stabilities for selected proteins) is frequently available. It seems reasonable to actually integrate such information into the inverse Ising problem instead of ignoring it. In this work, we use two different types of data (cf. Fig. 1):
-
As in the standard inverse Ising problem, part of the data comes as a sample of equilibrium configurations assumed to be drawn from the Boltzmann distribution to be inferred.
-
The second data source is a collection of arbitrary configurations together with noisy measurements of their energy.
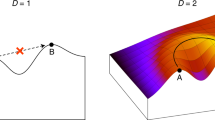
Schematic representation of the inference framework: a sample of equilibrium configurations (dataset Deq) and noisy energy measurements for another set of configurations (dataset DE) are integrated within a Bayesian approach to infer the model  .
.
The dashed lines represent the underlying true landscape, which has to be inferred, the red line the inferred landscape.
These data sets are limited in size and accuracy. Therefore an optimized integration of both data types is expected to improve the overall performance as compared to the individual use of one single data set.
The inspiration to develop this new integrative framework for the inverse Ising problem is taken from protein fitness landscapes in biology, which provide a quantitative mapping from any amino-acid sequence s = (s1,..., sN) to a fitness ϕ(s) measuring the ability of the corresponding protein to perform its biological function. Fitness landscapes are of outstanding importance in evolutionary and medical biology, but it appears impossible to deduce a protein’s fitness from its sequence only. Experimental or computational approaches exploiting other data are urgently needed.
Information about fitness landscapes can be found in the amino-acid statistics observed in natural protein sequences, which are related to the protein of interest. In fact they represent diverse but functional configurations sampled by evolution. It has been recently proposed that their statistical variability can be captured by Potts models (generalization of the Ising model to 21-state amino-acid variables). Indeed, statistical models inferred from large collections of natural sequences have recently led to good predictions of experimentally measured effects18,19,20,21: in a number of systems, the fitness cost Δϕ(s) ≡ ϕ(s) − ϕ(sref) of mutating any amino acid in a reference protein sref strongly correlates with the corresponding energy changes in the inferred statistical model,

suggesting that the Hamiltonian of the inferred models is strictly related to the underlying mutational landscapes.
While evolutionary diverged sequences can be regarded as a global sample of the fitness landscape, further information can be obtained from direct measurements on particular ‘microstates’ of the system, i.e. individual protein sequences. Recent advances in experimental technology allow for conducting large-scale mutagenesis studies: in a typical experiment, a reference protein of interest is chosen, and a large number (103–105) of mutant proteins (having sequences differing by one or few amino acids from the reference) are synthesized and then characterized in terms of fitness. This provides a systematic local measurement of the fitness landscape22,23,24. Regression analysis may be used to globally model mutational landscapes25. A second-order parameterization of ϕ arises naturally in this context, when considering an expansion of effects in terms of independent additive effects and pairwise ‘epistatic’ interactions between sites26,

However, the number of accessible mutant sequences remains small compared to the number of terms in this sum, and mutagenesis data alone are not sufficient to faithfully model fitness landscapes27.
In situations where no single dataset is sufficient for accurate inference, integrative methods accounting for complementary data sources will improve the accuracy of computational predictions. In this paper, (i) we define a generalized inference framework based on the availability of an equilibrium sample and of complementary quantitative information; (ii) we propose a Bayesian integrative approach to improve over the limited accuracy obtainable using standard inverse problems; (iii) we demonstrate the practical applicability of our method in the context of predicting mutational effects in proteins, a problem of outstanding bio-medical importance for questions related to genetic disease and antibiotic drug resistance.
Results
An integrated modeling
The inference setting
Inspired by this discussion, we consider two different datasets originating from a true model  . The first one, Deq = {s1,..., sM}, is a collection of M equilibrium configurations independently drawn from the Boltzmann distribution P0(s). For simplicity, we consider binary variables si ∈ {0, 1}, corresponding to a “lattice-gas” representation of the Ising model in Eq. (1). This implies that energies are measured with respect to the reference configuration sref = (0,..., 0). The standard approach to the inverse Ising problem uses only this type of data to infer parameters of
. The first one, Deq = {s1,..., sM}, is a collection of M equilibrium configurations independently drawn from the Boltzmann distribution P0(s). For simplicity, we consider binary variables si ∈ {0, 1}, corresponding to a “lattice-gas” representation of the Ising model in Eq. (1). This implies that energies are measured with respect to the reference configuration sref = (0,..., 0). The standard approach to the inverse Ising problem uses only this type of data to infer parameters of  : couplings and fields in Eq. (1) are fitted so that the inferred model reproduces the empirical single and pairwise frequencies,
: couplings and fields in Eq. (1) are fitted so that the inferred model reproduces the empirical single and pairwise frequencies,

The second dataset provides a complementary source of information, which shall be modeled as noisy measurements of the energies of a set of P arbitrary (i.e. not necessarily equilibrium) configurations σa. These data are collected in the dataset DE = {(Ea, σa)a=1,...,P}, with

The noise ξa models measurement errors or uncertainties in mapping measured quantities to energies of the Ising model. For simplicity, we consider ξa to be white Gaussian noise with zero mean and variance Δ2: 〈ξaξb〉 = δa,bΔ2.
As schematically represented in Fig. 1, datasets Deq and DE constitute different sources of information about the energy landscape defined by Hamiltonian  . Observables in Eq. (4) are empirical averages computed from equilibrium configurations in Deq, providing global information about the energy landscape. On the contrary, configurations in DE are arbitrarily given, and a (noisy) measurement of their energies provides local information on particular points in the landscape.
. Observables in Eq. (4) are empirical averages computed from equilibrium configurations in Deq, providing global information about the energy landscape. On the contrary, configurations in DE are arbitrarily given, and a (noisy) measurement of their energies provides local information on particular points in the landscape.
A maximum-likelihood approach
To infer the integrated model, we consider a joint description of the probabilities of the two data types for given parameters J and h of tentative Hamiltonian  . The probability of observing the sampled configurations in Deq equals the product of the Boltzmann probability of each configuration,
. The probability of observing the sampled configurations in Deq equals the product of the Boltzmann probability of each configuration,

To derive an analogous expression for the second dataset, we integrate over the Gaussian distribution of the noise  obtaining a Gaussian probability of the energies (remember configurations in DE are arbitrarily given):
obtaining a Gaussian probability of the energies (remember configurations in DE are arbitrarily given):

The combination of these expressions provides the joint log-likelihood for the model parameters given the data:

Maximizing the above likelihood with respect to parameters {hi}1≤i≤N and {Jij}1≤i<j≤N leads to the following self-consistency equations:

with  ,
,  being single and pairwise averages in the model Eq. (1). We have introduced the parameter
being single and pairwise averages in the model Eq. (1). We have introduced the parameter  : in practical applications, the error Δ may not be known, and the parameter 0 ≤ λ < 1 allows to weigh data sources differently. For λ = 0 (i.e. large noise), the standard inverse Ising problem is recovered: optimal parameters are such that the model exactly reproduces magnetizations and correlations of the sample. For λ > 0, the second dataset containing quantitative data is taken into account: whenever energies computed from the Hamiltonian
: in practical applications, the error Δ may not be known, and the parameter 0 ≤ λ < 1 allows to weigh data sources differently. For λ = 0 (i.e. large noise), the standard inverse Ising problem is recovered: optimal parameters are such that the model exactly reproduces magnetizations and correlations of the sample. For λ > 0, the second dataset containing quantitative data is taken into account: whenever energies computed from the Hamiltonian  do not match the measured ones, the model statistics deviates from the sample statistics. Both log-likelihood terms in (8) are concave, and thus their sum: Eq. (9) has a unique solution.
do not match the measured ones, the model statistics deviates from the sample statistics. Both log-likelihood terms in (8) are concave, and thus their sum: Eq. (9) has a unique solution.
Noiseless measurements
The case of noiseless energy measurements in Eq. (5) (i.e. λ → 1) has to be treated separately. First, energies have to be perfectly fitted by the model, by solving the following linear problem:

where we have introduced a N(N + 1)/2 dimensional vectorial representation  of the model parameters, and
of the model parameters, and  contains the exactly measured energies. The matrix
contains the exactly measured energies. The matrix

specifies which parameters contribute to the energies of configurations in the second dataset. If K = N(N + 1)/2 − rank(X) > 0, the parameters cannot be uniquely determined from the measurements: The sample Deq can be used to remove the resulting degeneracy. To do so, we parametrize the set of solutions of Eq. (10) as follows:

where  is any particular solution of the non homogeneous Eq. (10), and
is any particular solution of the non homogeneous Eq. (10), and  a basis of observables spanning the null space of the associated homogeneous problem
a basis of observables spanning the null space of the associated homogeneous problem  . The free parameters
. The free parameters  can be fixed by maximizing their likelihood given sample Deq,
can be fixed by maximizing their likelihood given sample Deq,

with  . The maximization provides conditions for the observables (k = 1,..., K),
. The maximization provides conditions for the observables (k = 1,..., K),

Equation (14) shows that the αk have to be fixed such that empirical averages  equal model averages
equal model averages  . Any possible sparsity of the matrix of measured configurations X (entries are 0 or 1 by definition) can be exploited to find a sparse representation of the
. Any possible sparsity of the matrix of measured configurations X (entries are 0 or 1 by definition) can be exploited to find a sparse representation of the  . In the protein example discussed above, mutagenesis experiments typically quantify all possible single-residue mutations of a reference sequence (denoted (0,..., 0) without loss of generality). In this case, the pairwise quantities sisj with 1 < i < j < N can be chosen as the basis
. In the protein example discussed above, mutagenesis experiments typically quantify all possible single-residue mutations of a reference sequence (denoted (0,..., 0) without loss of generality). In this case, the pairwise quantities sisj with 1 < i < j < N can be chosen as the basis  of the null space. A particular solution of the non-homogeneous system (10) is given by the paramagnetic Hamiltonian
of the null space. A particular solution of the non-homogeneous system (10) is given by the paramagnetic Hamiltonian  , with Ei being the energy shift due to spin flip
, with Ei being the energy shift due to spin flip  .
.
Artificial data
We first evaluate our method on artificial data (Materials and Methods). Random couplings J0 and fields h0 are chosen for a system of N = 32 spins. Dataset Deq is created by Markov chain Monte Carlo (MCMC) sampling, resulting in M = 100 equilibrium configurations. To mimic a protein ‘mutagenesis’ experiment, one of these configurations is chosen at random as the reference sequence, and the energies of all N configurations differing by a single spin flip from the reference (thereafter referred to as single mutants) are calculated, resulting in dataset DE (after adding noise of standard deviation Δ0). Datasets DE and Deq will subsequently called “local” and “global” data respectively.
Equations (9) are solved using steepest ascent, updating parameters J and h in direction of the gradient of the joint log-likelihood (8). Since the noise Δ0 may not be known in practical applications, we solve the equations for several values of λ ∈ [0, 1], weighing data sources differently. We expect the optimal inference to take place at a value λ that maximizes the likelihood in Eq. (8), i.e.  . For λ = 0, this procedure is equivalent to the classical Boltzmann machine28, but for λ > 0, the term corresponding to the quantitative essay constrains energies of sequences in DE to stay close to the measurements. As explained above, the case λ = 1 has to be treated separately; a similar gradient ascent method is used. Since exact calculations of gradients are computationally hard, the mean-field approximation is used (Materials and Methods).
. For λ = 0, this procedure is equivalent to the classical Boltzmann machine28, but for λ > 0, the term corresponding to the quantitative essay constrains energies of sequences in DE to stay close to the measurements. As explained above, the case λ = 1 has to be treated separately; a similar gradient ascent method is used. Since exact calculations of gradients are computationally hard, the mean-field approximation is used (Materials and Methods).
To evaluate the accuracy of the inference, most of the existing literature on the inverse Ising modeling simply compares the inferred parameters with the true ones. However, a low error in the estimation of each inferred parameter does not guarantee that the inferred distribution matches the true one. On the contrary, in the case of a high susceptibility of the statistics with respect to parameter variations, or if the estimation of parameters is biased, the distributions of the inferred and true models could be very different even for small errors on individual parameters. For this reason, we introduce two novel evaluation procedures. First, to estimate the accuracy of the model on a local region of the configuration space, we test its ability to reproduce energies of configurations in DE. Then, we estimate the global similarity of true and inferred distributions using a measure from information theory.
Error correction of local data
We first test the ability of our approach to predict the true single-mutant energies, when noisy measurements are presented in DE, i.e. to correct the measurement noise using the equilibrium sample Deq. For every λ, J and h are inferred and used to compute predicted energies of the N configurations in DE. The linear correlation between such predicted energies (measured with the inferred Hamiltonian) and the true energies (measured with the true Hamiltonian) is plotted as a function of λ in Fig. 2.
Integration of noisy measurements of energies of single mutants with M equilibrium configurations.
The linear correlation between predicted and true single mutants energies is shown in dependence of integration parameter λ, for four different values of the noise Δ0 = 0, 0.43, 0.92, 1.39 added to the energies in the second data set. The integration strength  , which would be naturally used in case of an a priori known noise level, is located close to the optimal inference, cf. the black circles for λ0 = 1, 0.84, 0.54, 0.34. The insert shows the value of the integration strength λ reaching maximal correlation, as a function of the theoretical value
, which would be naturally used in case of an a priori known noise level, is located close to the optimal inference, cf. the black circles for λ0 = 1, 0.84, 0.54, 0.34. The insert shows the value of the integration strength λ reaching maximal correlation, as a function of the theoretical value  , for 200 independent realisations of the input data at different noise levels. Points are found to be closely distributed around the diagonal (red line).
, for 200 independent realisations of the input data at different noise levels. Points are found to be closely distributed around the diagonal (red line).
In the very low noise regime,  , the top curve in Fig. 2 reaches its peak at
, the top curve in Fig. 2 reaches its peak at  , which is expected as local data is then sufficient to accurately “predict” energies from single mutants. On the contrary, in the high noise regime, the maximum is located close to λ = 0, pointing to the fact that local data is of little use in this case. Between those two extremes, an optimal integration strength can be found, yielding a better prediction of energies in DE as for any of the datasets taken individually. It is interesting to notice that even for highly noisy data, integrating the two sources of information with the right weight λ results in an improved modeling.
, which is expected as local data is then sufficient to accurately “predict” energies from single mutants. On the contrary, in the high noise regime, the maximum is located close to λ = 0, pointing to the fact that local data is of little use in this case. Between those two extremes, an optimal integration strength can be found, yielding a better prediction of energies in DE as for any of the datasets taken individually. It is interesting to notice that even for highly noisy data, integrating the two sources of information with the right weight λ results in an improved modeling.
The insert of Fig. 2 shows the integration strength λ at which the best correlation is reached, against the corresponding theoretical value  for different realizations. On average, optimal integration is reached close to the theoretical case of equation (8). This result highlights the possibility of using this integrative approach to correct measurement errors in the energies of single mutants. If a dataset such as Deq provides global information about the energy landscape, and the measurement noise Δ0 can be estimated, an appropriate integration can then be used to infer more accurately the energies of the single mutants.
for different realizations. On average, optimal integration is reached close to the theoretical case of equation (8). This result highlights the possibility of using this integrative approach to correct measurement errors in the energies of single mutants. If a dataset such as Deq provides global information about the energy landscape, and the measurement noise Δ0 can be estimated, an appropriate integration can then be used to infer more accurately the energies of the single mutants.
Global evaluation of the inferred Ising model
To assess the ability of our integrative procedure to provide a globally accurate model, we use the Kullback-Leibler divergence DKL(P0||P) between the true model  and the inferred
and the inferred  (Materials and Methods). The symmetric expression
(Materials and Methods). The symmetric expression

simplifies to the average difference between true and inferred energies. It can be consistently estimated using MCMC samples D (resp. D0) drawn from P (resp. P0), without the need to calculate the partition functions. Σ(P0, P) has an intuitive interpretation in terms of distinguishability of models: It represents the log-odds ratio between the probability to observe samples D and D0 in their respective generating models, and the corresponding probability with models  and
and  swapped:
swapped:

where MK is the number of sampled configurations in  and D0.
and D0.
The inferred model undoubtedly benefits from the integration, as a minimal divergence between the generating and the inferred probability distributions is found for an intermediate value of λ, outperforming both datasets taken individually (Fig. 3). It has to be noted that even in the noiseless case Δ0 = 0, the minimum in Σ(P0, P) obtained at λ = 1 depends crucially on the availability of the equilibrium sample Deq. The local data DE are not sufficient to fix uniquely all model parameters, and the degeneracy in parametrization is resolved using Deq as explained at the end of Sec. 0.
Symmetric Kullback-Leibler divergence ∑ between true and integrated pairwise models versus strength of integration λ, estimated from MK = 3 · 106 MCMC samples.
Different curves correspond to different noise levels added to single mutants energies used for integration, so that datasets DE are the same as in Fig. 2. Insert - Comparison with an independent model using only fields h, with the same methodology.
As a comparison, the same analysis is done using an independent modeling that uses only fields h, and no couplings. The inset of Fig. 3 clearly shows that the pairwise modeling outperforms the independent one. Even the limit λ → 1, where Deq becomes irrelevant in the independent model, the performance of the integrative pairwise scheme is not attained.
Biological data
To demonstrate the practical utility of our integrative framework, we apply it to the challenging problem of predicting the effect of amino-acid mutations in proteins. To do so, we use three types of data: (i) Multiple-sequence alignments (MSA) of homologous proteins containing large collections of sequences with shared evolutionary ancestry and conserved structure and function; they are obtained using HMMer29 using profile models from the Pfam database30. Due to their considerable sequence divergence (typical Hamming distance ~0.8N), they provide a global sampling of the underlying fitness landscape. (ii) Computational predictions of the impact of all single amino-acid mutations on a protein’s structural stability31 are used to locally characterize the fitness landscape around a given protein. The noise term ξa represents the limited accuracy of this predictor, and the uncertainty in using structural stability as a proxy of protein fitness. (iii) Mutagenesis experiments have been used before to simultaneously quantify the fitness effects of thousands of mutants22,23. While datasets (i) and (ii) play the role of Deq and DE in inference, dataset (iii) is used to assess the quality of our predictions (ideally one would use the most informative datasets (i) and (iii) to have maximally accurate predictions, but no complementary dataset to test predictions would be available in that case).
To apply the inference scheme to such protein data, three modifications with respect to simulated data are needed. First, the relevant description in this case is a 21-state Potts Model (Supporting Information), since each variable si, i = 1,..., N, can now assume 21 states (20 amino acids, one alignment gap)32. Second, since measured fitnesses and model energies are found in a monotonous non-linear relation, we have used the robust mapping introduced in ref. 18 (reviewed in the Supporting Information). Third, since correlations observed in MSA are typically too strong for the MF approximation to accurately estimate marginals, we relied on Markov Chain Monte Carlo (Materials and Methods), which has recently been shown to outperform other methods in accuracy of inference for protein-sequence data33,34.
We have tested our approach for predicting the effect of single amino-acid mutations in two different proteins: the β-lactamase TEM-1, a bacterial enzyme providing antibiotic resistance, and the PSD-95 signaling domain belonging to the PDZ family. In both systems computational predictions can be tested against recent high-throughput experiments quantifying the in-vivo functionality of thousands of protein variants22,23. Figure 4 shows the Pearson correlations between inferred energies and measured fitnesses as a function of the weight λ: Maximal accuracy is achieved at finite values of λ when both sources of information are combined, significantly increasing the predictive power of the models inferred considering the statistics of homologs only (λ = 0). When repeating the integrated modeling with a paramagnetic model where all sites are treated independently,  (only single-site frequencies are fitted in this case) the predictive power drops as compared to the Potts model, cf. the red lines in Fig. 4.
(only single-site frequencies are fitted in this case) the predictive power drops as compared to the Potts model, cf. the red lines in Fig. 4.
Linear correlation between experimental fitness effects and predictions from integrated models, at different values of λ, for 742 single mutations in the beta-lactamase TEM-1 (left panel) and for 1426 single mutations in the PSD-95 PDZ domain (right panel).
Error bars represent statistical errors estimated via jack-knife estimation.
Conclusion
In this paper, we have introduced an integrative Bayesian framework for the inverse Ising problem. In difference to the standard setting, which uses only a global sample of independent equilibrium configurations to reconstruct the Hamiltonian of an Ising model, we also consider a local quantification of the energy function around a reference configuration. Using simulated data, we show that the integrated approach outperforms inference based on each single dataset alone. The gain over the standard setting of the inverse Ising problem is particularly large when the equilibrium sample is too small to allow for accurate inference.
This undersampled situation is particularly important in the context of biological data. The prediction of mutational effects in proteins is of enormous importance in various bio-medical applications, as it could help understanding complex and multifactorial genetic diseases, the onset and the proliferation of cancer, or the evolution of antibiotic drug resistance. However, the sequence samples provided by genomic databases, like the multiple-sequence alignments of homologous proteins considered here, are typically of limited size, including even in the most favorable situations rarely more than 103–105 alignable sequences. Fortunately, such sequence data are increasingly complemented by quantitative mutagenesis experiments, which use experimental high-throughput approaches to quantify the effect of thousands of mutants. While it might be tempting to use these data directly to measure mutational landscapes from experiments, it has to be noted that current experimental techniques miss at least 2–3 orders of magnitude in the number of measurable mutants to actually reconstruct the mutational landscape.
In such situations, where no single dataset is sufficient for accurate inference, integrative methods like the one proposed here will be of major benefit.
Methods
Data
Artificial data
For a system of N = 32 binary spins, couplings J0 and fields h0 are chosen from a Gaussian distribution with zero mean, and standard deviation  for J and 0.2 for h (analogous results are obtained for other parameter choices, as long as these correspond to a paramagnetic phase). Dataset Deq is created by Markov chain Monte Carlo (MCMC) simulation, resulting in M = 100 equilibrium configurations. A large number (~105) of MCMC steps are done between each of those configurations to ensure that they are independent. One of these configurations is chosen at random as the reference sequence (“wild-type”), and the energies of all N configurations differing by a single spin flip from the reference are computed (“single mutants”). Gaussian noise of variance
for J and 0.2 for h (analogous results are obtained for other parameter choices, as long as these correspond to a paramagnetic phase). Dataset Deq is created by Markov chain Monte Carlo (MCMC) simulation, resulting in M = 100 equilibrium configurations. A large number (~105) of MCMC steps are done between each of those configurations to ensure that they are independent. One of these configurations is chosen at random as the reference sequence (“wild-type”), and the energies of all N configurations differing by a single spin flip from the reference are computed (“single mutants”). Gaussian noise of variance  can be added to these energies, resulting in dataset DE.
can be added to these energies, resulting in dataset DE.
Biological data
Detailed information about the analysis of biological data is provided in the Supporting Information.
Details of the inference
For artificial data, Eq. (9) are solved using steepest ascent, updating parameters J and h in direction of the gradient. To ensure convergence, we have added an additional  -regularization to the joint likelihood:
-regularization to the joint likelihood:  . A gradient ascent method has been analogously used for the case λ = 1. To estimate the gradient, it is necessary to compute single and pair-wise probabilities pi(J, h) and pij(J, h). Their exact calculation requires summation over all possible configurations of N spins, which is intractable even for systems of moderate size N, so we relied on the following approximation schemes.
. A gradient ascent method has been analogously used for the case λ = 1. To estimate the gradient, it is necessary to compute single and pair-wise probabilities pi(J, h) and pij(J, h). Their exact calculation requires summation over all possible configurations of N spins, which is intractable even for systems of moderate size N, so we relied on the following approximation schemes.
Mean-field inference
In the analysis of artificial data we relied on the mean-field approximation (MF) leading to closed equations for pi and pij:

The main advantage of the MF approximation is its computational efficiency: The first term is solved by an iterative procedure, the second requires the inversion of the couplings matrix J. However, the approximation is only valid and accurate at “high temperatures”, i.e. small couplings35. This condition is verified in the case of the artificial data described above.
MCMC inference
Correlations observed in MSA of protein sequences are typically too strong for the MF approximation to accurately estimate marginals of the model. Therefore we use MCMC sampling of MMC = 104 independent equilibrium configurations to estimate marginals at each iteration of the previously described learning protocol.
Global evaluation of the inferred Ising model
The Kullback-Leibler divergence 
 is a measure of the difference between probability distributions P and Q. It is zero for P ≡ Q, and otherwise positive. In the case of Boltzmann distributions
is a measure of the difference between probability distributions P and Q. It is zero for P ≡ Q, and otherwise positive. In the case of Boltzmann distributions  and
and  , its expression simplifies to
, its expression simplifies to

Evaluating this expression requires the exponential computation of the partition function of both models  and
and  . To overcome this difficulty, we use the symmetrized expression in Eq. (15), which only involves the average of macroscopic observables.
. To overcome this difficulty, we use the symmetrized expression in Eq. (15), which only involves the average of macroscopic observables.
The symmetrized Kullback-Leibler divergence is computed by obtaining MK = 128000 equilibrium configurations from both P and Q, using them to estimate the averages in Eq. (15).
Additional Information
How to cite this article: Barrat-Charlaix, P. et al. Improving landscape inference by integrating heterogeneous data in the inverse Ising problem. Sci. Rep. 6, 37812; doi: 10.1038/srep37812 (2016).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Weigt, M., White, R. A., Szurmant, H., Hoch, J. A. & Hwa, T. Identification of direct residue contacts in protein–protein interaction by message passing. Proceedings of the National Academy of Sciences 106, 67–72 (2009).
Mora, T., Walczak, A. M., Bialek, W. & Callan, C. G. Maximum entropy models for antibody diversity. Proceedings of the National Academy of Sciences 107, 5405–5410 (2010).
Ferguson, A. L. et al. Translating hiv sequences into quantitative fitness landscapes predicts viral vulnerabilities for rational immunogen design. Immunity 38, 606–617 (2013).
Lezon, T. R., Banavar, J. R., Cieplak, M., Maritan, A. & Fedoroff, N. V. Using the principle of entropy maximization to infer genetic interaction networks from gene expression patterns. Proceedings of the National Academy of Sciences 103, 19033–19038 (2006).
Schneidman, E., Berry, M. J., Segev, R. & Bialek, W. Weak pairwise correlations imply strongly correlated network states in a neural population. Nature 440, 1007–1012 (2006).
Cocco, S., Leibler, S. & Monasson, R. Neuronal couplings between retinal ganglion cells inferred by efficient inverse statistical physics methods. Proceedings of the National Academy of Sciences 106, 14058–14062 (2009).
Bialek, W. et al. Statistical mechanics for natural flocks of birds. Proceedings of the National Academy of Sciences 109, 4786–4791 (2012).
Jaynes, E. T. Information theory and statistical mechanics. Physical Review 106, 620 (1957).
Roudi, Y., Tyrcha, J. & Hertz, J. Ising model for neural data: model quality and approximate methods for extracting functional connectivity. Physical Review E 79, 051915 (2009).
Sessak, V. & Monasson, R. Small-correlation expansions for the inverse ising problem. Journal of Physics A: Mathematical and Theoretical 42, 055001 (2009).
Mézard, M. & Mora, T. Constraint satisfaction problems and neural networks: A statistical physics perspective. Journal of Physiology-Paris 103, 107–113 (2009).
Cocco, S., Monasson, R. & Sessak, V. High-dimensional inference with the generalized hopfield model: Principal component analysis and corrections. Physical Review E 83, 051123 (2011).
Cocco, S. & Monasson, R. Adaptive cluster expansion for inferring boltzmann machines with noisy data. Physical Review Letters 106, 090601 (2011).
Nguyen, H. C. & Berg, J. Mean-field theory for the inverse ising problem at low temperatures. Physical Review Letters 109, 050602 (2012).
Aurell, E. & Ekeberg, M. Inverse ising inference using all the data. Physical Review Letters 108, 090201 (2012).
Nguyen, H. C. & Berg, J. Bethe–peierls approximation and the inverse ising problem. Journal of Statistical Mechanics: Theory and Experiment 2012, P03004 (2012).
Decelle, A. & Ricci-Tersenghi, F. Pseudolikelihood decimation algorithm improving the inference of the interaction network in a general class of ising models. Physical Review Letters 112, 070603 (2014).
Figliuzzi, M., Jacquier, H., Schug, A., Tenaillon, O. & Weigt, M. Coevolutionary inference of mutational landscape and the context dependence of mutations in beta-lactamase tem-1. Molecular Biology and Evolution (2016).
Asti, L., Uguzzoni, G., Marcatili, P. & Pagnani, A. Maximum-entropy models of sequenced immune repertoires predict antigen-antibody affinity. PLoS Comput Biol 12, e1004870 (2016).
Morcos, F., Schafer, N. P., Cheng, R. R., Onuchic, J. N. & Wolynes, P. G. Coevolutionary information, protein folding landscapes, and the thermodynamics of natural selection. Proceedings of the National Academy of Sciences 111, 12408–12413 (2014).
Mann, J. K. et al. The fitness landscape of hiv-1 gag: Advanced modeling approaches and validation of model predictions by in vitro testing. PLoS Comput Biol 10, e1003776 (2014).
McLaughlin, R. N. Jr., Poelwijk, F. J., Raman, A., Gosal, W. S. & Ranganathan, R. The spatial architecture of protein function and adaptation. Nature 491, 138–142 (2012).
Jacquier, H. et al. Capturing the mutational landscape of the beta-lactamase tem-1. Proceedings of the National Academy of Sciences 110, 13067–13072 (2013).
Melamed, D., Young, D. L., Gamble, C. E., Miller, C. R. & Fields, S. Deep mutational scanning of an rrm domain of the saccharomyces cerevisiae poly (a)-binding protein. RNA 19, 1537–1551 (2013).
Hinkley, T. et al. A systems analysis of mutational effects in hiv-1 protease and reverse transcriptase. Nature genetics 43, 487–489 (2011).
de Visser, J. A. G. & Krug, J. Empirical fitness landscapes and the predictability of evolution. Nature Reviews Genetics 15, 480–490 (2014).
Otwinowski, J. & Plotkin, J. B. Inferring fitness landscapes by regression produces biased estimates of epistasis. Proceedings of the National Academy of Sciences 111, E2301–E2309 (2014).
Ackley, D. H., Hinton, G. E. & Sejnowski, T. J. A learning algorithm for boltzmann machines. Cognitive Science 9, 147–169 (1985).
Mistry, J., Finn, R. D., Eddy, S. R., Bateman, A. & Punta, M. Challenges in homology search: Hmmer3 and convergent evolution of coiled-coil regions. Nucleic Acids Research 41, e121 (2013).
Finn, R. D. et al. Pfam: the protein families database. Nucleic Acids Research 42, D222–D230 (2014).
Dehouck, Y., Kwasigroch, J. M., Gilis, D. & Rooman, M. Popmusic 2.1: a web server for the estimation of protein stability changes upon mutation and sequence optimality. BMC Bioinformatics 12, 151 (2011).
Morcos, F. et al. Direct-coupling analysis of residue coevolution captures native contacts across many protein families. Proceedings of the National Academy of Sciences 108, E1293–E1301 (2011).
Sutto, L., Marsili, S., Valencia, A. & Gervasio, F. L. From residue coevolution to protein conformational ensembles and functional dynamics. Proceedings of the National Academy of Sciences 112, 13567–13572 (2015).
Haldane, A., Flynn, W. F., He, P., Vijayan, R. S. K. & Levy, R. M. Structural Propensities of Kinase Family Proteins from a Potts Model of Residue Co-Variation. Protein Science 25, 1378–1384 (2016).
Plefka, T. Convergence condition of the tap equation for the infinite-ranged ising spin glass model. Journal of Physics A: Mathematical and general 15, 1971 (1982).
Acknowledgements
MW acknowledges funding by the ANR project COEVSTAT (ANR-13-BS04- 0012-01). This work undertaken partially in the framework of CALSIMLAB, supported by the grant ANR-11-LABX-0037-01 as part of the “Investissements d’ Avenir” program (ANR-11-IDEX-0004-02).
Author information
Authors and Affiliations
Contributions
M.F. and M.W. designed research; P.B.C., M.F. and M.W. performed research; P.B.C. and M.F. analyzed data; and P.B.C., M.F. and M.W. wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Barrat-Charlaix, P., Figliuzzi, M. & Weigt, M. Improving landscape inference by integrating heterogeneous data in the inverse Ising problem. Sci Rep 6, 37812 (2016). https://doi.org/10.1038/srep37812
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep37812
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.