Abstract
SIRT1, the NAD+-dependent protein deacetylase, controls cell-cycle progression and apoptosis by suppressing p53 tumour suppressor. Although SIRT1 is known to be phosphorylated by JNK1 upon oxidative stress and subsequently down-regulated, it still remains elusive how SIRT1 stability and activity are controlled. Here, we have unveiled that CHFR functions as an E3 Ub-ligase of SIRT1, responsible for its proteasomal degradation under oxidative stress conditions. CHFR interacts with and destabilizes SIRT1 by ubiquitylation and subsequent proteolysis. Such CHFR-mediated SIRT1 inhibition leads to the increase of p53 acetylation and its target gene transcription. Notably, CHFR facilitates SIRT1 destabilization when SIRT1 is phosphorylated by JNK1 upon oxidative stress, followed by prominent apoptotic cell death. Meanwhile, JNK inhibitor prevents SIRT1 phosphorylation, leading to elevated SIRT1 protein levels even in the presence of H2O2. Taken together, our results indicate that CHFR plays a crucial role in the cellular stress response pathway by controlling the stability and function of SIRT1.
Similar content being viewed by others
Introduction
CHFR (checkpoint protein with FHA and RING finger domains) is a RING-type E3 ubiquitin (Ub)-ligase, which regulates numerous important cellular proteins, i.e., PLK1, Aurora A, HLTF, and HDAC1, to function as a mitotic checkpoint and a tumour suppressor1,2,3,4. Notably, CHFR is able to modulate acetylation levels of histones as well as non-histone proteins like p53 and further induce p21 expression by suppressing the HDAC1 activity4. Histone deacetylases (HDACs) are divided into four classes based on the sequence homology: class I (HDAC1~3 and 8), class II (HDAC4~7 and HDAC9~10), class III (SIRT1~7), and class IV (HDAC11). Class I, II, and IV are considered “classical” HDACs that utilize Zn+ as a cofactor and generally inhibited by trichostatin A (TSA). Meanwhile, class III HDACs, also known as sirtuins, are NAD+-dependent histone deacetylases and homologous to yeast Sir2 (silent information regulator 2)5,6,7.
SIRT1 is the most representative NAD+-dependent deacetylase, which belongs to the class III HDAC family6,7. SIRT1 deacetylates not only histones but also many non-histone proteins including FOXO, Ku70, p300, Rb, E2F1, NF-κB, and p538,9,10. For example, SIRT1 removes an acetyl moiety from p53, resulting in the inhibition of p53-dependent cell cycle arrest and apoptosis11,12,13,14, suggesting that SIRT1 could act against p53. Consequently, through this deacetylation activity for various target substrates, SIRT1 plays a pivotal role in controlling diverse cellular processes, e.g., aging, autophagy, centrosome duplication, energy metabolism, inflammation, and tumorigenesis15,16,17.
Although SIRT1 is known to be regulated by several transcription factors, microRNAs, endogenous regulators such as AROS (active regulator of SIRT1) and DBC1 (deleted in breast cancer 1)18,19,20,21, and post-translational modifications, including SUMOylation22 and deubiquitylation23, the molecular machinery to regulate the expression and the activity of SIRT1 are quite complex and still remains under investigation. It has recently been reported that SIRT1 is phosphorylated by c-Jun N-terminal kinase 1 (JNK1)24,25 and subsequently degraded in a proteasome-dependent manner. Ubiquitylation is a reversible post-translational modification, which plays key roles in determining protein stability and conveying important cellular signals26. Therefore, it is plausible that ubiquitin-proteasome system (UPS) might be directly linked to control SIRT1 stability and function.
In the present study, we demonstrated that SIRT1 is a new target substrate of CHFR E3 Ub-ligase. CHFR binds to and ubiquitylates SIRT1, leading to its proteasomal degradation. CHFR also elevates p53 acetylation by destabilizing SIRT1, resulting in the increase of its transcriptional activity and apoptotic cell death. Especially, SIRT1 is destabilized in the presence of CHFR under oxidative stress, followed by enhanced apoptotic cell death. These results provide evidence that CHFR plays a crucial role in the regulation of SIRT1 stability and function.
Results and Discussion
CHFR interacts with SIRT1
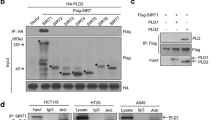
As CHFR is able to interact and suppress the classical HDAC4, we tested the possibility whether CHFR is also able to regulate another subtype of HDAC, the sirtuin family protein, SIRT1. We first performed a co-immunoprecipitation (IP) assay to determine the interaction between CHFR and SIRT1 in HEK293T cells transiently expressing FLAG-CHFR and MYC-SIRT1. When SIRT1 was immunoprecipitated with anti-FLAG M2 affinity resin, CHFR was readily detectable in the IP eluates, indicating that SIRT1 and CHFR interact with each other in vivo (Fig. 1A). It is worthy of note that SIRT1 protein levels were lower when SIRT1 was co-expressed with CHFR compared to when SIRT1 alone was transfected (Fig. 1A, lanes 2 and 3), suggesting that CHFR might be responsible for this SIRT1 destabilization. We also examined whether endogenous CHFR and SIRT1 could associate with each other. Since CHFR is often epigenetically silenced by DNA hypermethylation in most immortalized cancer cells27, we have generated CHFR-stable cells, which are identical except CHFR expression compared to its parental cancer cells, to mimic CHFR activation. SIRT1 was present in the immunoprecipitated eluates with anti-CHFR antibody and similarly, CHFR was present in anti-SIRT1 IP eluates, indicating that CHFR and SIRT1 bind together in HeLa-CHFR cells (Fig. 1B). The interaction between SIRT1 and CHFR was further validated by a GST pull-down assay using GST-SIRT1 purified from Escherichia coli and His-CHFR purified from Sf9 cells. His-CHFR protein was pulled down together with GST-SIRT1, but not with GST alone (Fig. 1C), indicating that the interaction between SIRT1 and CHFR is rather direct. Next, we mapped the region of CHFR required for the interaction with SIRT1. FLAG-tagged CHFR truncated mutants (ΔN, amino acids (a.a.) 143–664; ΔCR, a.a 1–475) were generated (Fig. 1D, upper panel) and tested the degree of interaction with SIRT1 in HEK293T cells in the presence of proteasome inhibitor MG132. SIRT1 was present in the IP eluates from CHFR wild-type (WT) and CHFR-ΔN, but not from CHFR-ΔCR, suggesting that the cysteine rich (CR; a.a 476–664) domain of CHFR is critical for the interaction with SIRT1 (Fig. 1D, lower panel). Taken together, CHFR binds to SIRT1 directly through its CR domain.
SIRT1 interacts with CHFR.
(A) CHFR binds to SIRT1 in vivo. HEK293T cells were transiently transfected as indicated. Cell lysates were immunoprecipitated with anti-FLAG M2 resin and immunoblotted with indicated antibodies. (B) CHFR and SIRT1 are endogenously associated with each other. HeLa-CHFR cell lysates were immunoprecipitated with either anti-CHFR or anti-SIRT1 in combination with preimmune serum or IgG, respectively, and analyzed by immunoblotting with indicated antibodies. Asterisk and arrowhead designate IgG and immunoprecipitated target proteins, SIRT1 or CHFR, respectively. (C) Recombinant GST-SIRT1 and His-CHFR proteins directly interact with each other in vitro. Purified proteins were pulled down with glutathione-sepharose resin and immunoblotted with anti-CHFR or anti-GST antibodies. (D) CR domain of CHFR is required for the interaction with SIRT1. A schematic representation of CHFR with its functional domains is shown in the top panel: FHA, a forkhead-associated; R, a RING finger; and CR, a cysteine-rich domain, respectively. FLAG-CHFR DNA plasmids (WT, wild-type; ΔN, a.a. 143~664; ΔCR, a.a. 1~475, respectively) were transfected into HEK293T cells. At 24 h post-transfection, cells were treated with 2 μM MG132 for 12 h. Cell lysates were immunoprecipitated with anti-FLAG M2 resin and immunoblotted with indicated antibodies.
CHFR ubiquitylates and promotes the proteasomal degradation of SIRT1
Given that SIRT1 is destabilized in the presence of CHFR as shown in Fig. 1A and they bind to each other, it is plausible that CHFR could act as a specific E3 Ub-ligase of SIRT1 to modulate its protein levels. In order to test this possibility, SIRT1 was transfected into HeLa cells, where CHFR is normally not expressed4, together with either mock- or CHFR-expression vector. SIRT1 protein levels were decreased in a CHFR dose-dependent manner and this reduction was blocked by the treatment of MG132 (Fig. 2A), indicating that CHFR leads to the proteasomal degradation of SIRT1. On the contrary, either E3 Ub-ligase-defective CHFR-I306A4 or SIRT1-binding-defective CHFR-ΔCR mutant failed to decrease SIRT1 protein levels compared to CHFR WT (Fig. 2B), suggesting that both an E3 Ub-ligase activity and a substrate binding ability of CHFR are necessary for SIRT1 destabilization.
CHFR negatively regulates SIRT1.
(A) CHFR overexpression results in the decrease of endogenous SIRT1. HeLa cells were transfected with FLAG-CHFR (0, 0.5, or 2 μg) and treated with 2 μM MG132 for 12 h as indicated. (B) Both an E3 Ub-ligase activity and a binding capacity to SIRT1 are necessary for CHFR to down-regulate SIRT1. FLAG-CHFR DNA plasmids encoding WT, I306A (Ile306Ala), or ΔCR mutants were transfected into HeLa cells as indicated. Cell lysates were immunoblotted with anti-FLAG and anti-SIRT1 antibodies. (C) CHFR ubiquitylates SIRT1 in vivo. HEK293T cells were transiently transfected with indicated plasmids and treated with 2 μM MG132 for 12 h. Cell lysates were immunoprecipitated with anti-FLAG M2 resin and probed with anti-HA antibody. (D) SIRT1 is ubiquitylated by CHFR in vitro. Purified SIRT1 is incubated with either CHFR WT or I306A mutant in the presence of E1, UbcH5b, ATP, ubiquitin as indicated. After in vitro ubiquitylation reaction, samples were analysed by immunoblotting with anti-SIRT1 antibody.
We next sought to examine whether CHFR is able to ubiquitylate SIRT1 prior to its proteolytic degradation. HEK293T cells were transiently transfected with expression vectors encoding HA-Ub, FLAG-SIRT1, and MYC-CHFR WT or -I306A mutant, and treated with MG132. SIRT1 was heavily ubiquitylated by CHFR WT, but not by CHFR-I306A mutant (Fig. 2C). In addition, in vitro ubiquitylation assay under defined conditions was performed using purified E1, E2 (UbcH5b), FLAG-SIRT1, and His-CHFR (WT or I306A). CHFR WT efficiently catalyzed poly-ubiquitylation of SIRT1. However, there is no or little ubiquitylation of SIRT1 with CHFR-I306A mutant (Fig. 2D). Taken together, these results suggest that CHFR serves as a specific E3 Ub-ligase for SIRT1 ubiquitylation.
CHFR enhances p53 acetylation and its transcriptional activity
As SIRT1 is able to deacetylate p53 and suppress its transcriptional activity12, and CHFR facilitates SIRT1 degradation, we speculated that CHFR affects p53 functions through the inhibition of the SIRT1 activity. To test this hypothesis, p53 acetylation was monitored in HCT116 cells expressing p53, p300, SIRT1, and CHFR. We have previously reported that CHFR binds to and down-regulates HDAC1, resulting in the increase of p53 acetylation4. To rule out such possibility that CHFR-mediated HDAC1 destabilization influences p53 acetylation, cells were then treated with TSA, a class I/II HDAC inhibitor. As expected, p53 is deacetylated by the SIRT1 introduction (Fig. 3A, lanes 2 and 3). Ectopic expression of CHFR highly elevated the levels of p53 acetylation in accordance with the reduced SIRT1 protein levels (Fig. 3A, lanes 3 and 4). These results indicate that CHFR is able to inhibit SIRT1 function not to deacetylate p53. To further validate the biological consequences of CHFR-induced SIRT1 degradation, we examined the effect of CHFR on the p53 transcriptional activity using p53 response element-containing luciferase genes, i.e., PG13-luc and p21-luc. Consistent with a previous finding, SIRT1 inhibited p53-driven gene expression and this decrease was restored by CHFR co-expression (Fig. 3B,C). These data suggest that CHFR enhances the p53 transcriptional activity by destabilizing SIRT1 (as shown in the bottom panel of Fig. 3B) and inhibiting its deacetylase activity, which are illustrated in Fig. 3D.
CHFR enhances the p53 transcriptional activity by destabilizing SIRT1.
(A) p53 acetylation is restored in the presence of CHFR. MYC-p53, HA-p300, FLAG-SIRT1, and/or FLAG-CHFR were transfected into HCT116 cells and treated with 0.5 μM TSA for 6 h. Cell lysates were immunoblotted with anti-acetyl p53, anti-p53, anti-FLAG, and anti-GAPDH antibodies. (B–D) CHFR increases p53-dependent gene expression by suppressing SIRT1. HCT116 cells were transiently transfected with expression vectors for p53, SIRT1, CHFR, and β-galactosidase along with either (B) PG13- or (C) p21 promoter-driven luciferase reporter gene, and treated with 0.5 μM TSA for 6 h as indicated. Luciferase activity was measured and normalized to β-galactosidase activity. Values were expressed as mean ± SEM of three independent experiments. Corresponding immunoblots are shown below (B). CHFR-mediated p53 transactivation is illustrated in (D).
CHFR is responsible for SIRT1 degradation under oxidative stress conditions
As SIRT1 is known to be down-regulated upon oxidative stress28 and we have shown thus far that SIRT1 is destabilized by CHFR, we investigated whether CHFR is involved in this oxidative stress-induced SIRT1 destabilization. To test this hypothesis, we utilized HeLa-CHFR stable cells to assess endogenous SIRT1 protein levels and found that the treatment of 1 mM H2O2 for 6 h in cells was sufficient to decrease SIRT1 proteins (Supplementary Fig. S1A). The turn-over rate of SIRT1 in the presence of H2O2 was further determined using HeLa-control and HeLa-CHFR stable cells, which are identical except for expressing CHFR. Upon oxidative stress, SIRT1 was quickly destabilized only in HeLa-CHFR cells, indicating that CHFR is responsible for H2O2-induced SIRT1 degradation (Fig. 4A). These reduced SIRT1 protein levels were restored by the co-treatment of MG132 with H2O2, suggesting that SIRT1 is degraded by the ubiquitin-proteasome system upon oxidative stress (Fig. 4B). We have then performed the co-immunoprecipitation assay in both endogenous and transiently transfected conditions to examine whether H2O2 treatment affects the interaction between CHFR and SIRT1. Since SIRT1 is destabilized in the presence of CHFR upon H2O2 treatment, we have utilized either CHFR–stable cells or E3 Ub ligase-defective CHFR-I306A mutant and analysed the binding differences under the H2O2-induced oxidative stress conditions. While the binding degree of CHFR to SIRT1 was slightly increased upon H2O2 treatment, apparently, it did not seem the all-or-none differences in their interaction (Fig. 4C and Supplementary Fig. S1B). Given that SIRT1 is phosphorylated by JNK1 under oxidative stress conditions24,25, we investigated whether JNK signaling is linked to CHFR-mediated SIRT1 turn-over upon oxidative stress. When cells were treated with H2O2, the JNK pathway was activated, which was validated by the induction of phosphorylated JNK and phosphorylated c-Jun, and consequently, SIRT1 was destabilized. On the other hand, the treatment of JNK inhibitor SP600125 in cells together with H2O2 inactivated the JNK pathway and simultaneously blocked SIRT1 destabilization (Fig. 4D). Since the treatment of hydrogen peroxide is more likely to induce acute and instant damage to cells due to its quick removal by cells, we have reiterated the H2O2-driven SIRT1 destabilization under chronic oxidative stress conditions by the glucose oxidase (Gox) enzyme (Fig. 4E). Although glucose oxidase-induced chronic oxidative stress resulted in much stronger and prolonged damage to cells, consistent with the previous results shown in Fig. 4A, SIRT1 protein levels were decreased only in the presence of CHFR upon oxidative stress regardless of the type of oxidative stress triggers. Therefore, these results indicate that phosphorylated SIRT1 by JNK1 under oxidative stress conditions is destabilized by CHFR.
CHFR facilitates SIRT1 degradation under oxidative stress conditions.
(A) CHFR is responsible for SIRT1 degradation upon oxidative stress. HeLa-control and –CHFR stable cells were treated with H2O2 as indicated and immunoblotted with anti-SIRT1, anti-CHFR, and anti-GAPDH antibodies. SIRT1 protein levels were quantitated by ImageJ and normalized to GAPDH levels. (B) SIRT1 is degraded via the ubiquitin-proteasome pathway upon oxidative stress. HeLa-CHFR stable cells were either left untreated or treated with 1 mM hydrogen peroxide for 6 h along with MG132 proteasome inhibitor as indicated. The treatment of MG132 proteasome inhibitor in cells restored SIRT1 protein levels upon oxidative stress. (C) CHFR interacts with SIRT1 more upon oxidative stress. HeLa-CHFR cell lysates treated with either mock- or 1 mM hydrogen peroxide for 6 h were immunoprecipitated with anti-SIRT1 and analyzed by immunoblotting with anti-CHFR and anti-SIRT1 antibodies. Asterisk and arrowhead designate IgG and immunoprecipitated CHFR. (D) HeLa-CHFR stable cells were treated with H2O2 in the presence or absence of SP600125 as indicated. Cell lysates were immunoblotted with anti-SIRT1, anti-CHFR, anti-phospho-JNK, anti-JNK, anti-phospho-c-Jun, and anti-GAPDH antibodies. (E) SIRT1 is destabilized by glucose oxidase. HeLa-control and –CHFR stable cells were treated with glucose oxidase as indicated and immunoblotted with anti-SIRT1, anti-CHFR, and anti-GAPDH antibodies. SIRT1 protein levels were quantitated by ImageJ and normalized to GAPDH levels.
CHFR promotes oxidative stress-induced cell death by destabilizing SIRT1
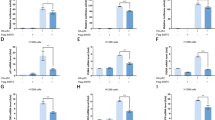
Given that CHFR is able to negatively regulate SIRT1 by ubiquitylation-mediated proteasomal degradation, we aimed to explore the biological outcomes of SIRT1 destabilization by CHFR, especially under oxidative stress conditions. In line with our previous results shown in Fig. 4, SIRT1 protein levels were significantly lower in H2O2-treated HeLa-CHFR cells compared to mock-treated cells. On the contrary, there was not much difference of SIRT1 in between mock- and H2O2-treated HeLa-control cells, indicating that CHFR is the underlying cause of SIRT1 degradation upon oxidative stress (Fig. 5A, top panel). This was further illustrated by the apoptosis assay measuring Annexin V and propidium iodide (PI) fluorescence in H2O2-treated HeLa-CHFR cells. Cell death was increased in HeLa-CHFR cells compared to control cells, and much greatly augmented by the H2O2 treatment (Fig. 5A, bottom panel). Next, we took a closer look at apoptotic events to further delineate how CHFR affects H2O2-driven cell death in HCT116-CHFR stable cells. Stained cells were sub-divided into four quadrants according to Annexin V and PI positivity. As cells were treated with H2O2 for 6 h, viable (Annexin V−/PI−) cells were decreased, while apoptotic (Annexin V+) and necrotic (PI+) cells were increased. Notably, H2O2-treated HCT116-CHFR cells showed the highest early apoptotic (Annexin V+/PI−) cell death among all tested (Fig. 5B). Moreover, Annexin V and PI fluorescence microscopy in HCT116-CHFR cells also revealed that CHFR is responsible for increased cell death upon oxidative stress (Fig. 5C). We have then investigated the effect of chronic oxidative stress-induced by Gox on apoptotic cell death along with SIRT1 protein levels. Cell death was much greatly augmented according to the strength of oxidative stress in CHFR-expressing cells (Fig. 5D,E) and similarly, SIRT1 protein levels were decreased in line with this increased cell death (Supplementary Fig. S1C). This was further validated by the cell viability assay, indicating that chronic oxidative stress by Gox leads to massive cell death (Fig. 5F).
CHFR augments apoptotic cell death upon oxidative stress.
(A) SIRT1 is destabilized in a CHFR- and H2O2-dependent manner. Cells were either left untreated or treated with H2O2 for 6 h, and immunoblotted with indicated antibodies. Annexin V/propidium iodide-based apoptotic cell death assay was performed and percent cell death values from the flow cytometry analysis were expressed as mean ± SEM of three independent experiments. (B–F) Oxidative stress-induced cell death is prominent in CHFR-stable cells. Cell viability was determined by annexin V/propidium iodide double staining following the treatment of either hydrogen peroxide (H2O2) or glucose oxidase (Gox) for 6 h. (B) Flow cytometric dot plots were sub-divided into four quadrants according to Annexin V and PI positivity and a detailed distribution of cell death events was shown. The lower left quadrant (Annexin V−/PI−) represents viable cells. The lower right quadrant (Annexin V+/PI−) was considered as early-stage apoptotic cells, the upper left (Annexin V−/PI+) as necrotic or dead cells, and the upper right quadrant (Annexin V+/PI+) as late apoptotic cells, respectively. (C) Annexin V−and/or PI-stained cells were monitored by the EVOSTM fluorescence cell imaging system. The scale bar represents 200 μm to visualize a wider area. (D) Gox-induced percent total cell death, (E) Gox- and H2O2-induced percent apoptotic cell death, and (F) cell viability examined by WST-1-based CellVia assay were expressed as mean ± SEM of three independent experiments. (G) A proposed model for CHFR to control SIRT1 upon oxidative stress.
Collectively, our data highlight that SIRT1 stability and function were negatively regulated by CHFR-mediated ubiquitylation and subsequent proteolysis. The inhibition of SIRT1 in human cancer cells by CHFR expression leads to elevated acetylation of p53 and simultaneous trans-activation of p53-driven target genes to elicit apoptosis in response to oxidative stress (Fig. 5G).
It is worthy of note that not only SIRT1 but also CHFR were destabilized by H2O2 in CHFR-stable cells. Since CHFR is known to be regulated by its own auto-ubiquitylation activity29, it would be of interest to study whether the E3 Ub-ligase activity of CHFR is controlled by the JNK signaling pathway. As both CHFR and SIRT1 have been implicated in cell cycle control and tumorigenesis, it would also be of particular interest to investigate how and when CHFR and/or SIRT1 respond to diverse cellular stresses during tumour progression. Here, we aimed to add a new line of evidence how CHFR contributes to tumour suppression. Especially, we have shown that CHFR is able to suppress not only HDAC1- class I HDAC4, but also SIRT1- class III HDAC. CHFR is often epigenetically inactivated in various cancer cells27, and reduced CHFR expression in normal cells leads to tumorigenic phenotypes30. Accordingly, such CHFR malfunction may lead to SIRT1 stabilization, which in turn represses p53 and other tumour suppressors to accelerate tumour initiation and metastasis. We have shown thus far that CHFR elevates the p53 activity by destabilizing SIRT1 when CHFR was re-introduced into cancer cells. Since p53 is widely regarded as “the guardian of genome”31,32, it is plausible that CHFR becomes a part of the watchman to keep cells under surveillance. This reinforces the role of CHFR as a tumour suppressor. CHFR also acts as a cell cycle checkpoint33, therefore, CHFR helps to maintain the cellular integrity against harmful stimuli and the threshold for apoptosis and cell senescence.
Methods
Cell culture, Transfection, and Reagents
HCT116, HeLa, and HEK293T cells were cultured in Dulbecco’s modified Eagle’s medium supplemented with 100 U ml−1 penicillin, 100 μg ml−1 streptomycin, and 10% FBS (Gibco) at 37 °C in a humidified 5% CO2 condition. Transient and stable transfections were carried out using either lipofectamine 2000 (Invitrogen) or polyethylenimine (Sigma) according to the manufacturer’s instructions. Following chemical reagents used in the study were obtained from Sigma or otherwise stated: TSA, hydrogen peroxide, glucose oxidase, and SP600125.
Immunoprecipitation and Immunoblotting
For immunoprecipitation, cells were lysed in TNET buffer (20 mM Tris-HCl [pH 7.5], 150 mM NaCl, 0.1 mM EDTA, and 0.2% Triton X-100) and 1x protease inhibitor cocktail (Roche). Cell lysates were incubated with anti-FLAG M2 affinity resin (Sigma) for 2 h at 4 °C. Resins were collected by centrifugation and washed three times with TNET buffer. Bound proteins were eluted, resolved by SDS-PAGE, and immunoblotted with appropriate antibodies. The following antibodies were used: anti-SIRT1, anti-MYC, anti-GST, anti-GAPDH, anti-HA, and anti-p53 (Santa Cruz Biotechnology); anti-acetyl p53 (Millipore); anti-FLAG and anti-β-actin (Sigma); anti-Xpress (Invitrogen); peroxidase-conjugated AffiniPure goat anti-rabbit and anti-mouse IgGs (Jackson ImmunoResearch); anti-CHFR antiserum was raised against a recombinant His-CHFR. Relative protein levels in the immunoblot figures were quantitated by ImageJ and normalized to either β-actin or GAPDH levels. Values are plotted as the mean ± SEM of at least three independent experiments.
GST pull-down assay
GST-SIRT1 was purified from Escherichia coli and His-CHFR was purified from Sf9 insect cells. GST-SIRT1 (1 μg) and His-CHFR (1 μg) were incubated with Glutathione Sepharose 4 Fast Flow (GE Healthcare) for 1 h at 4 °C. After incubation, bound proteins were eluted, resolved by SDS-PAGE, and analysed by immunoblotting with anti-CHFR and anti-GST antibodies.
Ubiquitylation assay
For the in vitro ubiquitylation assay, FLAG-SIRT1 protein (0.3 μg) purified from HEK293T cells was incubated with E1 (0.2 μg), UbcH5b (0.2 μg), Ubiquitin (2 μg), CHFR (1 μg), and ATP-regenerating system (50 mM Tris-HCl [pH 7.5], 5 mM MgCl2, 10 mM creatine phosphate, 5 U ml−1 of phosphocreatine kinase, and 5 mM ATP) at 37 °C for indicated times. For the in vivo ubiquitylation assay, cells were transfected with appropriate expression vectors and treated with 2 μM MG132 (A.G. Scientific) for 12 h. Cell lysates were incubated with anti-FLAG M2 affinity resin (Sigma). After incubation, bound proteins were eluted and analysed by immunoblotting.
Reporter assay
HCT116 cells were transfected with indicated plasmids with β-gal constructs and treated with 0.5 μM TSA for 6 h before harvest. Luciferase activity was measured in a luminometer with a luciferase system (Promega) and normalized to β-galactosidase activity. Values were expressed as mean ± SEM from three independent experiments.
Apoptosis assay
Cells were treated with either hydrogen peroxide or glucose oxidase for indicated times to induce oxidative stress, stained with either Alexa Fluor® 488 Annexin V/Dead Cell Apoptosis Kit (Molecular Probes) or FITC Annexin V Apoptosis Detection Kit (BD Biosciences) according to manufacturers’ instructions, and analysed using the TaLi® image-based cytometer (Invitrogen) or visualized under the EVOSTM cell imaging system (Thermo Fisher Scientific). DAPI was used to counterstain the nuclei.
Cell viability assay
Cells were seeded at a density of 104 cells/well in 100 μL of culture medium in a 96-well plate and treated with either hydrogen peroxide or glucose oxidase for 6 h to induce oxidative stress. Cells were treated with 10 μL of CellVia (water-soluble tetrazolium salt, WST-1; Young In Frontier) and incubated for an additional 2 h at 37 °C. Cell viability was measured using a multiwell microplate reader at a wavelength of 450 nm along with a reference wavelength of 650 nm. The same volume of culture medium plus CellVia reagent in the absence of cells were used as a blank control to subtract the background absorbance. Values were expressed as mean ± SEM from three independent experiments.
Additional Information
How to cite this article: Kim, M. et al. CHFR negatively regulates SIRT1 activity upon oxidative stress. Sci. Rep. 6, 37578; doi: 10.1038/srep37578 (2016).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Kang, D., Chen, J., Wong, J. & Fang, G. The checkpoint protein Chfr is a ligase that ubiquitinates Plk1 and inhibits Cdc2 at the G2 to M transition. The Journal of cell biology 156, 249–259, doi: 10.1083/jcb.200108016 (2002).
Yu, X. et al. Chfr is required for tumor suppression and Aurora A regulation. Nature genetics 37, 401–406, doi: 10.1038/ng1538 (2005).
Kim, J. M. et al. CHFR functions as a ubiquitin ligase for HLTF to regulate its stability and functions. Biochemical and biophysical research communications 395, 515–520, doi: 10.1016/j.bbrc.2010.04.052 (2010).
Oh, Y. M. et al. Chfr is linked to tumour metastasis through the downregulation of HDAC1. Nature cell biology 11, 295–302, doi: 10.1038/ncb1837 (2009).
Guarente, L. Sir2 links chromatin silencing, metabolism, and aging. Genes Dev 14, 1021–1026 (2000).
Tanny, J. C., Dowd, G. J., Huang, J., Hilz, H. & Moazed, D. An enzymatic activity in the yeast Sir2 protein that is essential for gene silencing. Cell 99, 735–745 (1999).
Smith, J. S. et al. A phylogenetically conserved NAD+-dependent protein deacetylase activity in the Sir2 protein family. Proceedings of the National Academy of Sciences of the United States of America 97, 6658–6663 (2000).
Brooks, C. L. & Gu, W. How does SIRT1 affect metabolism, senescence and cancer? Nature reviews. Cancer 9, 123–128, doi: 10.1038/nrc2562 (2009).
Morris, B. J. Seven sirtuins for seven deadly diseases of aging. Free radical biology & medicine 56, 133–171, doi: 10.1016/j.freeradbiomed.2012.10.525 (2013).
Vaquero, A. et al. Human SirT1 interacts with histone H1 and promotes formation of facultative heterochromatin. Molecular cell 16, 93–105, doi: 10.1016/j.molcel.2004.08.031 (2004).
Sykes, S. M. et al. Acetylation of the p53 DNA-binding domain regulates apoptosis induction. Molecular cell 24, 841–851, doi: 10.1016/j.molcel.2006.11.026 (2006).
Luo, J. et al. Negative control of p53 by Sir2alpha promotes cell survival under stress. Cell 107, 137–148 (2001).
Tang, Y., Zhao, W., Chen, Y., Zhao, Y. & Gu, W. Acetylation is indispensable for p53 activation. Cell 133, 612–626, doi: 10.1016/j.cell.2008.03.025 (2008).
Cheng, H. L. et al. Developmental defects and p53 hyperacetylation in Sir2 homolog (SIRT1)-deficient mice. Proceedings of the National Academy of Sciences of the United States of America 100, 10794–10799, doi: 10.1073/pnas.1934713100 (2003).
Wang, Z. & Chen, W. Emerging Roles of SIRT1 in Cancer Drug Resistance. Genes & cancer 4, 82–90, doi: 10.1177/1947601912473826 (2013).
Haigis, M. C. & Sinclair, D. A. Mammalian sirtuins: biological insights and disease relevance. Annual review of pathology 5, 253–295, doi: 10.1146/annurev.pathol.4.110807.092250 (2010).
Ling, H., Peng, L., Seto, E. & Fukasawa, K. Suppression of centrosome duplication and amplification by deacetylases. Cell Cycle 11, 3779–3791, doi: 10.4161/cc.21985 (2012).
Kim, E. J., Kho, J. H., Kang, M. R. & Um, S. J. Active regulator of SIRT1 cooperates with SIRT1 and facilitates suppression of p53 activity. Molecular cell 28, 277–290, doi: 10.1016/j.molcel.2007.08.030 (2007).
Kim, J. E., Chen, J. & Lou, Z. DBC1 is a negative regulator of SIRT1. Nature 451, 583–586, doi: 10.1038/nature06500 (2008).
Wang, C. et al. Interactions between E2F1 and SirT1 regulate apoptotic response to DNA damage. Nature cell biology 8, 1025–1031, doi: 10.1038/ncb1468 (2006).
Yamakuchi, M., Ferlito, M. & Lowenstein, C. J. miR-34a repression of SIRT1 regulates apoptosis. Proceedings of the National Academy of Sciences of the United States of America 105, 13421–13426, doi: 10.1073/pnas.0801613105 (2008).
Yang, Y. et al. SIRT1 sumoylation regulates its deacetylase activity and cellular response to genotoxic stress. Nature cell biology 9, 1253–1262, doi: 10.1038/ncb1645 (2007).
Lin, Z. et al. USP22 antagonizes p53 transcriptional activation by deubiquitinating Sirt1 to suppress cell apoptosis and is required for mouse embryonic development. Molecular cell 46, 484–494, doi: 10.1016/j.molcel.2012.03.024 (2012).
Nasrin, N. et al. JNK1 phosphorylates SIRT1 and promotes its enzymatic activity. PloS one 4, e8414, doi: 10.1371/journal.pone.0008414 (2009).
Gao, Z. et al. Sirtuin 1 (SIRT1) protein degradation in response to persistent c-Jun N-terminal kinase 1 (JNK1) activation contributes to hepatic steatosis in obesity. The Journal of biological chemistry 286, 22227–22234, doi: 10.1074/jbc.M111.228874 (2011).
Hershko, A. & Ciechanover, A. The ubiquitin system. Annual review of biochemistry 67, 425–479, doi: 10.1146/annurev.biochem.67.1.425 (1998).
Toyota, M. et al. Epigenetic inactivation of CHFR in human tumors. Proceedings of the National Academy of Sciences of the United States of America 100, 7818–7823, doi: 10.1073/pnas.1337066100 (2003).
Cao, C. et al. SIRT1 confers protection against UVB- and H2O2-induced cell death via modulation of p53 and JNK in cultured skin keratinocytes. J Cell Mol Med 13, 3632–3643, doi: 10.1111/j.1582-4934.2008.00453.x (2009).
Oh, Y. M., Yoo, S. J. & Seol, J. H. Deubiquitination of Chfr, a checkpoint protein, by USP7/HAUSP regulates its stability and activity. Biochemical and biophysical research communications 357, 615–619, doi: 10.1016/j.bbrc.2007.03.193 (2007).
Privette, L. M., Gonzalez, M. E., Ding, L., Kleer, C. G. & Petty, E. M. Altered expression of the early mitotic checkpoint protein, CHFR, in breast cancers: implications for tumor suppression. Cancer research 67, 6064–6074, doi: 10.1158/0008-5472.CAN-06-4109 (2007).
Lane, D. P. Cancer. p53, guardian of the genome. Nature 358, 15–16, doi: 10.1038/358015a0 (1992).
Efeyan, A. & Serrano, M. p53: guardian of the genome and policeman of the oncogenes. Cell Cycle 6, 1006–1010 (2007).
Scolnick, D. M. & Halazonetis, T. D. Chfr defines a mitotic stress checkpoint that delays entry into metaphase. Nature 406, 430–435, doi: 10.1038/35019108 (2000).
Acknowledgements
This work was supported by Mid-career Researcher Program (NRF-2014R1A2A2A01005258, NRF-2010-0014738 to J.H.S.), Basic Research Laboratory Program (NRF-2014R1A4A1005259 to J.H.S.), and Basic Science Research Program (NRF-2015R1D1A1A01061219 to M.K.) through the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIP and MOE).
Author information
Authors and Affiliations
Contributions
M.K., Y.E.K., J.O.S., and S.J.B. performed the experiments. M.K. and J.H.S. analysed the data and wrote the manuscript. J.H.S. conceived the study and supervised the project.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Kim, M., Kwon, Y., Song, J. et al. CHFR negatively regulates SIRT1 activity upon oxidative stress. Sci Rep 6, 37578 (2016). https://doi.org/10.1038/srep37578
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep37578
This article is cited by
-
CHFR promotes metastasis of human gastric carcinoma by activating AKT and ERK via NRF2- ROS axis
BMC Gastroenterology (2023)
-
SIRT1 ubiquitination is regulated by opposing activities of APC/C-Cdh1 and AROS during stress-induced premature senescence
Experimental & Molecular Medicine (2023)
-
Oxidative stress promotes SIRT1 recruitment to the GADD34/PP1α complex to activate its deacetylase function
Cell Death & Differentiation (2018)
-
Exogenous nicotinamide supplementation and moderate physical exercise can attenuate the aging process in skeletal muscle of rats
Biogerontology (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.