Abstract
Our study combined 16S rRNA-pyrosequencing and whole genome sequencing to analyze the fecal metagenomes of the divergently selected lean (LL) and fat (FL) line chickens. Significant structural differences existed in both the phylogenic and functional metagenomes between the two chicken lines. At phylum level, the FL group had significantly less Bacteroidetes. At genus level, fourteen genera of different relative abundance were identified, with some known short-chain fatty acid producers (including Subdoligranulum, Butyricicoccus, Eubacterium, Bacteroides, Blautia) and a potentially pathogenic genus (Enterococcus). Redundancy analysis identified 190 key responsive operational taxonomic units (OTUs) that accounted for the structural differences between the phylogenic metagenome of the two groups. Four Cluster of Orthologous Group (COG) categories (Amino acid transport and metabolism, E; Nucleotide transport and metabolism, F; Coenzyme transport and metabolism, H; and Lipid transport and metabolism, I) were overrepresented in LL samples. Fifteen differential metabolic pathways (Biosynthesis of amino acids, Pyruvate metabolism, Nitrotoluene degradation, Lipopolysaccharide biosynthesis, Peptidoglycan biosynthesis, Pantothenate and CoA biosynthesis, Glycosaminoglycan degradation, Thiamine metabolism, Phosphotransferase system, Two-component system, Bacterial secretion system, Flagellar assembly, Bacterial chemotaxis, Ribosome, Sulfur relay system) were identified. Our data highlighted interesting variations between the gut metagenomes of these two chicken lines.
Similar content being viewed by others
Introduction
Host genetic background is a major determinant factor that controls the phenotype. The gut microbiota is recognized as an important environmental factor that interacts with its host through metabolic exchange and contributes to host energy absorption1. The present work investigated the host gut metagenome of a divergently selected chicken model based on fatness traits. Strong reasons have supported us to perform this work in broiler chicken.
Firstly, poultry are important protein sources in human diet and hence they are of enormous economic value; and domesticated chickens are the most common domestic animals in the world. The formation of close symbiotic relationship between the host-gut microbiota is known to be crucial in sustaining host health. Accumulating evidence shows that gut dysbiosis is linked to a variety of diseases, especially metabolic-associated syndromes like obesity2, diabetes3, and rheumatoid arthritis4. One vital function of the gut microbiota is to provide microbial-based metabolic pathways that are otherwise not indigenously encoded within the host genome; and in turn the gut microbiota participates in regulating the host nutritional metabolism, including nutrient assimilation and energy capture. Moreover, they act in protecting the host from pathogens, detoxification, and immune development5. Taking into account the crucial health-related roles, without any doubt the gut microbiota can be regarded as a potential target for improving the general health, growth performance and productivity of broiler chickens, which have always been of intense interest to breeders.
Secondly, numerous studies have concluded that the gut microbiota is linked with host obesity and energy metabolism1,6,7, potentially via the connection between the gut microbes and body fat absorption and disposition8,9. For instance, the phylum, Firmicutes, is more abundant in obese than lean individuals, and vice versa for Bacteroidetes. Moreover, the relative abundance of Bacteroidetes increases, accompanied with a decrease in Firmicutes, after a weight loss program for obese individuals1. By transferring the gut microbiota from obese or lean mice to germ-free mice, it was shown that a high Firmicutes to Bacteroidetes gut microbial ratio increased body fat accumulation10. Apart from bacteria, the dominant human gut archaeon, Methanobrevibacter smithii, affects host calorie harvest and adiposity through the digestion of dietary polysaccharides11. In contrast, the reduction of some normal gut bacteria along with the enrichment of certain opportunistic pathogens in the gut microbiota may raise the risk of weight gain. The opportunistic pathogen, Enterobacter cloacae B29, isolated from obese human’s gut is able to induce obesity and insulin resistance in germ-free mice12.
However, most available data in this aspect are based on rodents and human models10,13,14, which may not be completely suited in the case of chicken because of its unique anatomy and physiological functioning. Chickens swallow food and store it in the crop before digestion and absorption taking part in the gizzard, small intestine and cecum. Only a relatively short time of 2.5 hours is required for the food to pass through the upper intestine, contrasting to that of 12–20 hours in the ceca5. The prolonged retention of digesta in chicken ceca, where the largest quantity and diversity of microbes are hosted along the chicken gastrointestinal tract15, may indicate the essential nutritional function of the locality, including nutrient absorption, digestion of non-start polysaccharides (often present in high proportion in feed), recycling of nitrogen via uric acid dissimilation, and provision of B vitamins and essential amino acids5,16.
Additionally, most currently published studies only described the structure and function of the chicken gut microbiota, and the spatial and temporal changes upon specific stimulations15,17,18,19,20. Our study compared and contrasted the gut metagenome chickens selectively bred based on fatness traits. Selective breeding is a common agricultural method that allows preferential selection of desirable genetic traits. This work has taken the advantage of the availability of two broiler lines21, the fat (FL) and lean (LL) chicken, which have been selectively bred for 15 generations (at the time of this experiment) from the parental Arbor Acres broilers based on the plasma very low density lipoprotein (VLDL) concentration and abdominal fat percentage (AFP). The LL chickens are characterized by a high efficiency in extracting and incorporating food source energy into lean meat, while the FL chickens have a great tendency of abdominal fat deposition. Polymorphisms of several gene loci may be responsible for the fatness traits in the divergent chicken lines, including the insulin-like growth factor binding protein 222, adipocyte fatty acid-binding protein23, acetyl-CoA carboxylase α24, and the PC1/PCSK1 region of the Z chromosome25. The divergent chicken lines also displayed variations in the preadipocyte microRNA expression profile26 and specific genes (namely SLC9A3, GNAL, SPOCK3, ANXA10, HELIOS, MYLK, CCDC14, SPAG9, SOX5, VSNL1, SMC6, GEN1, MSGN1 and ZPAX) within the copy number variation regions27.
We hypothesized that, apart from the host genetics, variation of the fatness trait may also link with the composition of the gut microbial metagenome. Thus, in this study, we compared the faecal microbial metagenome of these two chicken lines by integrating the 16S rRNA-pyrosequencing with whole genome sequencing (WGS). We aimed to investigate whether the process of lean and fat chicken selection simultaneous led to colonization of different spectra of gut microbiota. Data generated from this study have provided interesting insights into the role of environmental factors, e.g. the gut microbiota, in relation to the host phenotype in a genetically-predisposed model.
Materials and Methods
Ethics statement
All animal work was conducted according to the guidelines for the care and use of experimental animals established by the Ministry of Science and Technology of the People’s Republic of China (Approval number: 2006–398) and approved by the Laboratory Animal Management Committee of Northeast Agricultural University and the Ethical Committee of the Inner Mongolia Agricultural University.
Animals and sample collection
Two chicken lines, FL and LL, were provided by the Northeast Agricultural University. These two chicken lines were both originated from Arbor Acres broiler, which underwent 15 generations of selection since 1996 based on the VLDL and AFP at 7 weeks of age. Briefly, the VLDL and AFP for all the first generation male chickens were measured at 7 weeks of age. The VLDL and AFP of the next generation were measured and compared to that of the previous one, and broilers of lower or higher average VLDL and AFP were chosen for subsequent breeding, as described previously28. In this way, the chicken lines were divergently selected for 15 generations. At 15th generations, the AFP of the FL was on average 4.57 higher than that of the LL (Supplementary Fig. S1 and Supplementary Table S1). Each chicken was raised in an individual cage in order to prevent contamination from uncontrolled particle intake and feathers29. All birds had free access to water. Birds and environmental controls were checked twice daily by trained staff. The temperature was maintained at 16–18 °C, and the humidity was maintained at 50–60%. At different life stages, the birds were fed with different diets, according to the Arbor Acres Plus parent stock breeder management guide and nutrition specifications (http://en.aviagen.com/). At the time of sampling (bird age between week 37 to 40), the birds were feed-restricted on a standard diet containing 14.2% crude protein and metabolic energy of 2745 kcal/kg. The feed nutritive content and digestible amino acid supplementation at the time of sampling are provided in Table 1.
Fecal samples of each chicken (15 lean and 14 fat line chickens) were collected by laying clean papers on the cage floor, and the droppings were then transferred to 5 ml tube by 1000 μL pipette. All of the 29 samples were used for bacterial 16S rRNA genes V1-V3 region pyrosequencing and for whole-genome shotgun (WGS) sequencing.
DNA extraction
Genomic DNA was extracted from each fecal sample using Qiagen DNA Stool Mini Kit (Qiagen, Hilden, Germany) following the instructions of the manufacturer. The quality of extracted DNA was assessed by 0.8% agarose gel electrophoresis and spectrophotometry (optical density at 260 nm/280 nm). All extracted DNA samples were stored at −20 °C for further analysis.
PCR Amplification and 16S rRNA pyrosequencing
The V1-V3 region of the 16S ribosomal RNA (rRNA) genes in all the samples was amplified by PCR for barcoded pyrosequencing. For PCR amplification, 10 ng of extracted DNA was amplified in a 20 μL reaction buffer containing 4 μL 5× FastPfu Buffer, 2 μL 2.5 mM dNTPs, 0.4 μL FastPfu polymerase, 0.8 μL of each 5 μM primer (TransGen Biotech) and double deionized water. The primers used to amplify the bacterial V1-V3 region of the 16S rRNA gene were 27 F (5′-AGAGTTTGATCCTGGCTCAG-3′) and 533 R (5′-TTACCGCGGCTGCTGGCAC-3′). The PCR program was as follows: 94 °C for 5 min; 30 cycles of 94 °C for 45 s, 55 °C for 40 s, and 72 °C for 1 min; and a final extension of 72 °C for 7 min). The primers used for amplifying archaeal 16S rRNA gene were Arch344F (5′-ACGGGGYGCAGCAGGCGCGA-3′) and Arch915 (5′-GTGCTCCCCCGCCAATTCCT-3′). The PCR program was as follows: 95 °C for 2 min; 30 cycles of 95 °C for 30 s, 55 °C for 30 s, and 72 °C for 30 s with a final extension of 72 °C for 5 min.
The quality of the PCR products was ensured using Agilent 2100 Bioanalyzer (Agilent Technologies, Palo Alto, Calif.) in accordance with the manufacturer’s instructions. The PCR products were pooled in equimolar ratios with a final concentration of 100 nmol/L for pyrosequencing (Roche GS FLX), performed by Shanghai Majorbio Bio-pharm Technology Co., Ltd.
Bioinformatics processing of 16S rRNA pyrosequencing data
The extraction of high-quality sequences was performed with the QIIME package (Quantitative Insights Into Microbial Ecology) (v1.7). Raw sequences were selected based on sequence length, quality, primer and tag, according to the following criteria: the length of the raw sequences was more than 300 bp; the variable region was more than 300 bp in length and lying between the two primers; the read sequence had a perfect match with the barcode; the high-quality score (>20) for the proportion of bases was larger than 93% in the raw read.
PyNAST and UCLUST were applied to align sequences under 100% clustering of sequence identity in order to obtain unique representative sequences. Operational taxonomic units (OTUs) were classified under the threshold of 97% identity by using UCLUST. FastTree was applied to construct the de novo phylogenetic tree. Ribosomal Database Project (RDP) Classifier (Release 11.4) was applied to assign taxonomic to each OTU representative sequence. Owing to the relatively short length of the V1-V3 region of 16S rRNA genes, BLASTN from the NCBI website was applied when RDP failed to classify an OTU taxonomically. Differently defined thresholds were set to assign the reference sequence into the matching taxonomic group, which were 95%, 92%, 91%, 85%, and 75% for genus, family, order, class, and phylum, respectively30.
Whole-genome shotgun (WGS) sequencing and quality control
All samples were sequenced with the Illumina HiSeq2000 instrument. Libraries were prepared with a fragment length of approximately 300 bp. Paired-end reads were generated with 100 bp in the forward and reverse directions. The length of each read was trimmed with Sickle. Reads that aligned to the chicken genome were also removed. This set of high-quality reads was then used for further analysis. The sample taxonomic profile was also determined directly from the metagenome dataset using two online software, i.e. Parallel-META31 and MetaPhlAn32, following the instructions of the two software developers.
Illumina short reads de novo assembly, gene prediction and construction of the non-redundant gene catalogue
The microbial Illumina reads were assembled into contigs using IDBA-UD33 with default parameters. Genes were predicted on the contigs with MetaGeneMark34. A non-redundant gene catalogue was constructed with CD-HIT35 using a sequence identity cut-off of 0.95, with a minimum coverage cut-off of 0.9 for the shorter sequences.
Functional annotation
Putative amino acid sequences, translated from the gene catalogue, were aligned against the proteins/domains in the Cluster of Orthologous Group (COG) and Kyoto Encyclopedia of Genes and Genomes (KEGG) databases (submit online) using BLASTP (e-value ≤ 1e-5 with a bit score higher than 60). Each protein was assigned to the KEGG orthologue group (KO) or COG by the highest scoring annotated hit.
Computation of relative gene abundance
To assess the relative gene abundance, reads were aligned against the gene catalogue with Bowtie236 using parameters: -p 12 -x nt -1 R1.fastq -2 R2.fastq -S R.sam. Then, to normalize the sequencing coverage, the relative abundance instead of the raw read count was used to quantify the gut microbial genes. The calculation process was as follows:
Step 1: Calculation of the copy number of each gene:

Step 2: Calculation of the relative abundance of gene i:

ai: the relative abundance of gene i.bi: the copy number of gene i from sample N.Li: the length of gene i.xi: the number of mapped reads.
The relative gene abundances and other calculation were completed using Python scripts.
Statistical analyses
Statistical analyses were performed mainly using R packages (http://www.r-project.org/), together with Python37, Canoco for Windows 4.5 (Microcomputer Power, NY, USA), and PAST38. Rarefaction analysis, Shannon diversity index and Simpson’s diversity index were used to estimate the richness and diversity of OTUs. Principal coordinate and principal component analyses (PCoA and PCA) were used to assess the gut microbiota and metagenome structure of different samples, respectively. Redundancy analysis (RDA) was applied to identify bacterial groups which significantly contributed to the structural difference. The relative abundances of differential taxonomic groups were visualized by heatmap in R using the “pheatmap” package. Differences in the relative abundances of taxonomic groups and at gene level between samples were evaluated with Mann-Whitney test. False discovery rate (FDR) values were estimated using the Benjamini–Yekutieli method to control for multiple testing39. P-values less than 0.05 were considered statistically significant. Differential genes were also tested with DESeq240. Significantly differentiating KEGG modules were identified according to the final reporter score calculated from the aggregated Z-score of individual KOs with the cut-off level of ≥1.6 (90% confidence according to normal distribution)41. Whether the significantly differentiating modules were enriched in the FL or LL chickens was further determined by comparing the number of individual KOs that was enriched in the specific chicken line.
Nucleotide sequence accession numbers
Datasets generated by the shotgun and 16S rRNA amplicon sequencing approaches reported in this study have been deposited in the GenBank database (accession number: SRP083441) and MG-RAST project (ID: 19742), respectively.
Results
Sequence abundance and diversity of 16S rRNA gene
A total of 263,158 of v1-v3 region of 16S rRNA raw sequence reads were generated from 29 chickens (15 LL and 14 FL), with an average of 9074 sequence read for each sample. 46,415 sequence reads were delimited through PyNAST alignment and 100% sequence identity clustering for further analysis. 7312 OTUs were identified at 97% sequence similarity level with high threshold identity and with an average of 788 OTUs for each sample. The lowest level of taxon abundance for each OTU was determined by combining homologous sequence alignment method and clustering which based on information extracted from RDP and Greengenes (Release 13.5) databases. Results showed that 19.6% of sequences were unable to be assigned to the genus level. The Shannon diversity curves for all samples plateaued (Supplementary Fig. S2), even though the individual rarefaction curves did not reach saturation phase. This suggested that the increasing of sequencing depth would possibly identify more new phylotypes, but the current analysis had already captured the majority of the microbial diversity, respectively.
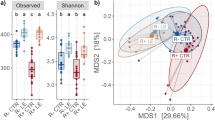
The Shannon and Simpson indices were applied to evaluate the diversity of gut microbiota, while the chao1 and observed species indices were indicators for species abundance. No significant difference was found between lean and fat chickens for all four indices, as assessed by Mann-Whitney test (Supplementary Table S2), and p-values for Shannon, Simpson, chao1 and observed species were 0.15, 0.18, 0.39 and 0.47.
Core gut microbiota detected in all samples based on 16S rRNA amplicon sequencing
In this study, the ‘core gut microbiota’ in chicken was defined as the taxonomical units (genus and OTUs) that were detected in all samples. Nine core genera, including Clostridium (23.44%), Bacteroides (18.78%), Lactobacillus (8.77%), Ruminococcus (3.97%), Hallella (1.61%), Subdoligranulum (1.07%), Faecalibacterium (1.04%), Roseburia (0.98%), and Eubacterium (0.31%), were identified (Supplementary Table S3).
Comparison of gut bacterial composition between LL and FL at phylum and genus levels based on 16S rRNA amplicon sequencing
Nineteen phyla were identified within the complete dataset. The 4 most dominant phyla (Firmicutes, Bacteroidetes, Proteobacteria and Actinobacteria) accounted for 99.11% of the total sequence, contributing to 53.44%, 41.09%, 3.22% and 1.33% for LL and 71.36%, 23.40%, 3.43% and 0.98% for FL, respectively. The proportion of Bacteroidetes, but not the other 3 dominant phyla, were significantly different between the LL and FL samples (LL>FL, 41.09% versus 23.40%, p = 0.034). Minor phyla, including Thermotogae, Verrucomicrobia, Cyanobacteria, Acidobacteria, Fibrobacteres, Chloroflexi, Gemmatimonadetes, Synergistetes, Fusobacteria, TM7, Deinococcus-Thermus, Tenericutes, Deferribacteres, Lentisphaerae, Spirochaetes, together contributed to 0.1% of the total sequence.
The dominant gut microbiota genera mainly belonged to the phyla, Firmicutes and Bacteroidetes (Fig. 1). However, considerable difference was noted between the fecal microbiota of the two chicken lines. In the gut microbiota of lean chickens, the relatively abundant genera (>1%) of Firmicutes included Clostridium (19.06%), Ruminococcus (5.50%), Lactobacillus (3.09%), Streptococcus (2.39%), Oscillibacter (1.52%), Subdoligranulum (1.30%), Faecalibacterium (1.18%), Enterococcus (1.14%), Roseburia (1.03%), whereas the Bacteroidetes was comprised with a wider diversity of relatively abundant genera, namely Bacteroides (23.45%), Hallella (2.36%), Parabacteroides (1.79%), Alistipes (1.23%), Paraprevotella (1.21%) (Fig. 1A). For FL chickens, the dominant and subdominant genera of over 1% relative abundance included members of the Firmicutes (Clostridium, Lactobacillus, Enterococcus, Turicibacter, Ruminococcus, Oscillibacter, Streptococcus, Megamonas) of 28.14%, 14.86%, 4.26%, 3.30%, 2.32%, 1.68%, 1.40%, and 1.02%, respectively) and Bacteroidetes (Bacteroides and Parabacteroides of 13.77% and 1.13%, respectively) (Fig. 1B).
Moreover, the proportions of 5 out of the 9 identified core genera were significantly higher in LL as compared to FL, including Ruminococcus, Subdoligranulum, Eubacterium, Bacteroides, Hallella (p values ranged from 0.001 to 0.047). Other genera which were also significantly enriched in LL included Alistipes, Butyricicoccus, Hespellia, Anaerosporobacter, Odoribacter, Prevotella, Collinsella and Blautia (p values ranged from 0.003 to 0.05). In contrast, the proportions of Enterococcus and Corynebacterium were lower in LL (p = 0.022, 0.045, respectively) (Supplementary Table S4).
Multivariate analysis of the gut microbiota structure of LL and FL chickens based on 16S rRNA amplicon sequencing
In order to compare the overall difference of the gut microbiota of LL and FL chickens, PCoA was performed. The PCoA score plots based on weighted and unweighted UniFrac are shown in Fig. 2. Although sample overlapping occurred between the two samples groups, clustering tendencies were observed, suggesting a moderate difference existed in the gut microbiota structure between the LL and FL chickens. Results from the multivariate analysis of variance (MANOVA) revealed the significant difference of gut microbiota between LL and FL (p < 0.05). Moreover, clustering analysis based on the weighted (p = 0.0024), but not unweighted (p = 0.7767), UniFrac distances revealed significant difference between the two chicken lines (Supplementary Fig. S3), indicating that the difference lied on the more prevalent lineages rather than the rare taxonomical groups.
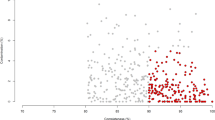
To further identify the specific bacterial groups principally accounting for the difference observed between the LL and FL gut microbiota, RDA was carried out. In this case, the LL and FL chickens were the “nominal environmental variables”, and the relative abundances of all OTUs (at 97% similarity) were the response variables. Monte Carlo Permutation Test showed that the constrained ordination model by lean or obese was statistically significant (p = 0.002) and the first canonical axis was able to explain 9.8% of the variability of response variables. Moreover, a total of 190 OTUs were identified as key variables which had good correlation with sample scores on the RDA canonical axis. Among them, 109 and 81 OTUs were enriched in the gut microbiota of LL and FL chickens, respectively (Fig. 3). The heatmap constructed by the relative abundance of these 190 OTUs is shown in Fig. 4.
Biplot of redundancy analysis (RDA) on the basis of OTU relative abundance at 98% similarity level.
‘ ’ and ‘
’ and ‘ ’ represent the constrained explanatory variables, fat (F) and lean (L) lines. Blue arrows represent response variables with the first ordination axis explaining for at least 9.8% of the variability. A p-value of 0.002 was generated from Monte Carlo Permutation Test.
’ represent the constrained explanatory variables, fat (F) and lean (L) lines. Blue arrows represent response variables with the first ordination axis explaining for at least 9.8% of the variability. A p-value of 0.002 was generated from Monte Carlo Permutation Test.
As assessed by Mann-Whitney test, 49 (18 Bacteroidaceae, 6 Ruminococcaceae, 6 Prevotellaceae, 5 Lachnospiraceae, 3 Clostridiaceae, 11 from other or unidentified families) out of the 190 key OTUs were significantly more abundant in lean chickens (p = 0.048–0.002), whereas the relative abundance of 47 OTUs (14 Clostridiaceae, 5 Lactobacillaceae, 4 Ruminococcaceae, 3 Micrococcaceae, 3 Erysipelotrichaceae, 18 from other or unidentified families) were significantly enriched in obese chickens (p = 0.046–0.002) (Supplementary Table S5).
Comparison of the gut archaeal composition between LL and FL
Archaeal members were also identified in the gut microbiota of both LL and FL chickens (Table 2). All archaeal sequences could be assigned to the phylum Euryarchaeota. At the genus level, most sequences belonged to Methanobrevibacter and Methanocorpusculum, and a few of the sequences correspond to Methanosarcina. Each of the 3 genera occupied less than 0.1% of the total sequences. No significant difference was found in the archeal composition between the two sample groups.
To investigate the archaea in the gut of both LL and FL, two steps UCLUST method combining with multivariate statistics were applied to conduct OTUs. By PCoA, the first two principal coordinates for weighted UniFrac were 86.09% and 5.78%, and 35.57% and 37.30% for unweighted UniFrac. Furthemore, the two groups overlapped each other; thus, no obvious difference in the gut archaea profile existed between the two groups (p > 0.05) (Supplementary Fig. S4).
Coverage of WGS sequencing
To profile the gut microbial metagenome of the chickens, WGS sequencing was performed on all 29 samples. However, one of them failed the quality test (data not shown); thus, no data was included in the metagenome analysis from this sample. The 28 samples yielded a total of 234.4 Gb of pair-end reads (an average of 52,485,882 high-quality reads and 239,690 genes per sample) that were of high-quality and were free of chicken DNA and adaptor contaminants (Supplementary Table S6).
Structures and functions of the chicken gut microbial metagenome
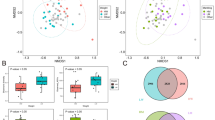
The structure and function of the LL and FL gut microbial metagenome were analyzed based on the COG and KEGG functions. The relative abundances of all the COG functional groups in the LL and FL chickens are represented by box plots (Fig. 5). Four of the functional categories were significantly different between the two chicken lines (all with relative abundance of lean > fat line), which were Amino acid transport and metabolism (E) (p = 0.0350), Nucleotide transport and metabolism (F) (p = 0.0042), Coenzyme transport and metabolism (H) (p = 0.0186), and Lipid transport and metabolism (I) (p = 0.0079), respectively. To visualize the functional difference between the two sample groups, PCA analysis was performed based on all detected KO (Fig. 6A) (with PC1 and PC2 representing 57.01% and 18.03% of the total variance). Although only a weak clustering pattern was observed on the score plot with mild overlap of symbols representing the two groups, further analysis by MANOVA revealed that they were significantly different (p = 0.0012) (Fig. 6B).
Comparison of the structure of lean and fat chicken gut metagenomes.
(A) Differential abundance of COG functional categories of the two chicken lines. COG category codes are as follows: A, RNA processing and modification; B, Chromatin structure and dynamics; C, Energy production and conversion; D, Cell cycle control, cell division, chromosome partitioning; E, Amino acid transport and metabolism; F, Nucleotide transport and metabolism; G, Carbohydrate transport and metabolism; H, Coenzyme transport and metabolism; I, Lipid transport and metabolism; J, Translation, ribosomal structure and biogenesis; K, Transcription; L, Replication, recombination and repair; M, Cell wall/membrane/envelope biogenesis; N, Cell motility; O, Posttranslational modification, protein turnover, chaperones; P, Inorganic ion transport and metabolism; Q, Secondary metabolites biosynthesis, transport and catabolism; R, General function prediction only; S, Function unknown; T, Signal transduction mechanisms; U, Intracellular trafficking, secretion, and vesicular transport; V, Defense mechanisms; W, Extracellular structures; Y, Nuclear structure; Z, Cytoskeleton. P-values of <0.01 and <0.05 are represented by ‘**’ and ‘*’, respectively.
Differential chicken gut microbial KEGG genes, modules and pathways
A total of 6739 KEGG genes could be assigned from the whole dataset, among which 320 genes (179 and 141, representing 2.66% and 2.09%, overrepresented in lean and fat line chickens with p-values ranging from 0.0003 to 0.0491 and 0.0010 to 0.0499, respectively) were differentially enriched, as detected by pairwise Mann-Whitney test (Supplementary Table S7).
To understand the biological meaning of such gene abundance differences, all the detected genes were further assigned to KEGG module and pathway levels. The detected genes could be assigned to 579 different KEGG modules. Forty (6.91%) of the modules had a final reporter score of over 1.6 (cutoff limit for significantly differentiating pathway). By comparing the number of individual KOs that was enriched in the specific chicken line, 31 (5.35%) and 8 (1.38%) modules were found to be more abundant in the lean and fat line chickens, respectively, while 1 (0.12%) module displayed similar level of activity between the two chicken lines (Supplementary Table S8). Table 3 includes only the modules with a high (≥ 80%) or a low (≤ 20%) proportion of KO that had higher relative gene abundance in lean line.
Similarly, based on the same level of final reporter score cutoff, a total of 15 differential pathways could be identified; most of them were relating to the category of Metabolism (namely Biosynthesis of amino acids, Pyruvate metabolism, Nitrotoluene degradation, Lipopolysaccharide biosynthesis, Peptidoglycan biosynthesis, Pantothenate and CoA biosynthesis, Glycosaminoglycan degradation, Thiamine metabolism), followed by Environmental Information Processing (namely Phosphotransferase system (PTS), Two-component system, Bacterial secretion system), Cellular Processes associated with Cell motility (Flagellar assembly, Bacterial chemotaxis), and Genetic Information Processing (Ribosome, Sulfur relay system) (Supplementary Table S9). To verify the profiles of differential pathways and modules, a second method, DESeq2, was used to compare the functional metagenomes of the two sample groups. Comparing to the results generated by Mann-Whitney test, highly similar results were obtained at both KEGG pathway (Supplementary Table S10) and module levels (Supplementary Table S11).
Discussion
Our study analyzed the gut microbiome of the divergently selected lean and fat broiler chicken lines. We asked whether different spectra of gut microbiota will be retained in these two chicken lines; and if so, whether the two spectra of gut microbiota potentially carry genomes that confer different metabolic capacity.
To answer our first question, we comparatively analyzed the fecal microbiota profiles of the two chicken lines mainly based on the 16S rRNA amplicon sequencing. Our MANOVA analysis revealed a significant difference between the LL and FL chicken fecal microbiota structure. At phylum level, significantly more Bacteroidetes (p = 0.0343) was found in the LL samples. Bacteroidetes is known to be associated with fat accumulation in chickens42 and less of these bacteria are present in obese human individuals1,43.
At genus level, some of the 14 identified differential genera are known short-chain fatty acid (SCFA)-producers. The butyrate-producers (Subdoligranulum, Butyricicoccus, Eubacterium)44,45, propionate-producers (Bacteroides)46 and acetate-producers (Blautia)47,48 were diminished in the FL samples. Subdoligranulum and Butyricicoccus can stimulate the growth of intestinal epithelial cells and thus reduce the invasion and colonization of Salmonella in veterinary medicine49. Eubacterium is a possible butyrate-producer in animal guts44. Low grade inflammation is involved actively in obesity development; and butyrate is anti-inflammatory and protects the intestinal barrier function50,51. Thus, fewer gut butyrate-producers in the FL group may contribute to its fat accumulation.
Meanwhile, Enterococcus sequences were approximately 4-fold more abundant in the FL samples. Enterococcus is reported to be associated with colorectal cancer52,53; and Enterococcus faecalis can damage eukaryotic cellular DNA in colonic epithelial cells by producing extracellular superoxides and hydroperoxides54,55. The local cellular damages and reactive oxygen species caused by these bacteria may alter intestinal permeability, lead to cell death, and hence act as triggers for inflammation and obesity.
No significant difference existed in the relative abundance of Lactobacillus between LL and FL samples, but an apparent variation existed (3.09% ± 0.92% versus 14.86% ± 6.82% in LL and FL samples, respectively). Moreover, ten fatness traits correlated key responsive OTUs identified by RDA belonged to Lactobacillus. In all cases, more were present in the FL samples, with OTUs 7838, 9029, and 8615 exhibiting significant differences (p-values = 0.003–0.014; 16.64-, 12.65-, and 4.31-fold enrichment in FL samples). Probiotics have been used in agricultural practices to increase growth rate, improve digestion, nutrient absorption and nutrient digestion. The probiotics-driven weight gain effect is strain-specific. The administration of Lactobacillus fermentum, Lactobacillus ingluviei, Lactobacillus agilis and Lactobacillus salvarius was associated with weight gain in chickens56,57,58,59, while consuming some other Lactobacillus species resulted in weight loss in rodents or humans60,61,62. The mechanism of Lactobacillus in host weight modification is complex and yet to be elucidated.
Generally, more Methanobrevibacter was found in the FL group, though the difference was not statistically significant. Methanogenic archaea, in particular Methanobrevibacter, participate in regulating gut metabolism by removing excessive bowel hydrogen. This subsequently improves the efficiency of microbial fermentation and enhances host energy capture. In a gnotobiotic mouse model, Methanobrevibacter results in weight gain in the presence of Bacteroides thetaiotaomicron12. Methanobrevibacter improve acetate and butyrate production meanwhile eliminate hydrogen and formate63, which are vital carbon sources for colon epithelium cells. This syntrophism between the gut bacteria and archaea may raise energy extraction when indigestible polysaccharide diets are given. The higher Methanobrevibacter abundance in the FL group may contribute to its obese phenotype.
Apart from describing the sample microbiota using a more traditional approach based on 16S rRNA amplicon sequencing, we also extracted the taxonomic profiles directly from the metagenome dataset by using two publicly accessible online tools, Parallel-META and MetaPhlAn (Supplementary Figs S5 and S6). We found some discrepancies between the phylogenic metagenome profiles as determined by the three methods; such discrepancies are possibly caused by the differences in the algorithm used in taxonomic assignment in each case64, as well as bias between the metagenomic- and 16S rRNA-based sequencing approaches65,66,67. The current study detected a broader range of bacterial genera with the 16S rRNA amplicon sequencing approach (246 versus 88 and 90 different bacterial genera with Parallel-META and MetaPhlAn, respectively) (data not shown), although many of these genera comprised only a minute proportion of the entire bacterial microbiota community.
To answer our second question, we analyzed the functional fecal metagenomes of the two chicken lines. Even though symbols representing the two groups mildly overlapped in the PCA plot based on KEGG orthology distribution (Fig. 6A), further MANOVA analysis confirmed that structural difference existed between the two KO-annotated functional metagenomes. Sequences encoding for the Amino acid transport and metabolism (E), Nucleotide transport and metabolism (F), Coenzyme transport and metabolism (H), and Lipid transport and metabolism (I) were overrepresented in the COG annotation of LL dataset.
Annotation of KEGG KO revealed a total of 15 differential pathways. Many of these KEGG pathways potentially relate to obesity, adiposity and energy balance regulation. For example, Lipopolysaccharide biosynthesis and Flagellar assembly are indirectly involved in host adiposity. Obesity is considered to be connected with chronic low-grade inflammation. Obesity causes noticeable structural alterations at the gut lining, resulting in elevated intestinal permeability that favors the gut microbiota-derived lipopolysaccharide translocation to the bloodstream. The endotoxemia may activate toll-like receptor 4 to proinflammatory status68. Flagellin is the major flagellar structural protein, which is specifically detected by the host toll-like receptor 5. Mice deficient of toll-like receptor become hyperphagic and develop obesity; and the transfer of gut microbiota from these mice to wild type germ-free mice confer metabolic syndrome features to the recipients, suggesting that the Flagellar assembly plays a role in obesity development69. On the other hand, peptidoglycan exerts an anti-inflammatory effect via the peptidoglycan recognition protein 370, which may counter the low-grade inflammation in obesity status. Some other pathways associate with nutrient absorption (including Biosynthesis of amino acids, Pyruvate metabolism and Phosphotransferase system (PTS)) and bacterial colonization and proliferation (including Two-component system, Bacterial chemotaxis, Bacterial secretion system). These pathways are required for substrate sensing and foraging, as well as carbohydrate substrate transporting and utilizing; thus, they are crucial for energy extraction and provision.
Only few reports have discussed the roles of vitamin metabolism in obesity development. The administration of thiamine prevents from obesity and metabolic disorders in Otsuka Long-Evans Tokushima Fatty rats that resemble human metabolic syndrome and obesity71, while pantothenic acid derivatives were found to exert hypolipidemic effect to hypothalamic obesity mice induced by aurothioglucose, potentially by the mechanisms of insulin resistance reduction and lipolysis in serum and adipose tissue72. The functions of other differential pathways in obesity development are less clear (e.g. Ribosome, Sulfur relay system, Nitrotoluene degradation, Glycosaminoglycan degradation).
Microbiome-derived SCFAs are considered as key players in regulating host obesity. Generally, butyrate and propionate are antiobesogenic, while acetate is obesogenic73. At KEGG module level, of particular interest is the higher abundance of lysine and isoleucine biosynthesis pathways of the LL group. Some gut commensal bacteria (including members of Eubacterium) are able to produce butyrate from lysine, although no gut microbe is known to contain the complete pathway74. Several amino acids, including both lysine and isoleucine, can serve as precursors for gut butyrogenesis75. Thus, we speculate that the gut microbiota of LL chicken may produce butyrate using amino acids as substrates.
At KEGG module level, two methanogenesis (M00357 and M00567) and the Pyridoxal biosynthesis (M00124) modules were enriched in the FL group, which may contribute to fat accumulation. The higher activity of methanogenesis is in line with the higher relative abundances of methanogenic archaea in the FL samples. A recent study found that the active form of vitamin B6 (pyridoxal 5-phosphate) is linked with adipogenesis using a comparative metabolomic approach76.
On the other hand, the ascorbate biosynthesis module (M00129) is enriched in the LL sample. Ascorbic acid supplementation may suppress adiposity and insulin resistance gene expression in high-fat diet induced obese rats77. Such effect may due to the intrinsic ascorbate anti-oxidative activity that ameliorates free radical-induced oxidative stress and thus lessens inflammation.
To sum up, we highlighted the overall structural differences between the fecal phylogenic and functional metagenomes of the LL and FL chickens. Between the two groups, significant differences were found in the relative abundances of some energy metabolite-related bacteria (especially the SCFA-producers) and potential pathogens (e.g. Enterococcus), and in numerous biochemical pathways relating to obesity, adiposity and energy balance. Our experimental design does not allow us to draw a concrete conclusion on whether the microbiota residing in the FL or LL chicken gut significantly or at least partially contributed to obesity or, alternatively, the modified metabolism driven by the host fatness traits selection resulted in modulated intestinal tract environment and thus the shift of microbiota composition to adapt to host obesity. Nevertheless, this study has provided a deeper insight into the possible contribution of gut microbiota in modulating obesity.
Additional Information
How to cite this article: Hou, Q. et al. Differential fecal microbiota are retained in broiler chicken lines divergently selected for fatness traits. Sci. Rep. 6, 37376; doi: 10.1038/srep37376 (2016).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Ley, R. E., Turnbaugh, P. J., Klein, S. & Gordon, J. I. Microbial ecology: human gut microbes associated with obesity. Nature 444, 1022–1023 (2006).
Bäckhed, F. et al. The gut microbiota as an environmental factor that regulates fat storage. Proc. Natl. Acad. Sci. USA 101, 15718–15723 (2004).
Larsen, N. et al. Gut microbiota in human adults with type 2 diabetes differs from non-diabetic adults. PloS One 5, e9085 (2010).
Toivanen, P. Normal intestinal microbiota in the aetiopathogenesis of rheumatoid arthritis. Ann. Rheum. Dis. 62, 807–811 (2003).
Stanley, D., Hughes, R. J. & Moore, R. J. Microbiota of the chicken gastrointestinal tract: influence on health, productivity and disease. Appl. Microbiol. Biotechnol. 98, 4301–4310 (2014).
Ley, R. E. et al. Obesity alters gut microbial ecology. Proc. Natl. Acad. Sci. USA 102, 11070–11075 (2005).
Palau-Rodriguez, M. et al. Metabolomic insights into the intricate gut microbial–host interaction in the development of obesity and type 2 diabetes. Front. Microbiol. 6 (2015).
Zhang, C. et al. Interactions between gut microbiota, host genetics and diet relevant to development of metabolic syndromes in mice. ISME J. 4, 232–241 (2010).
Zhang, C. et al. Structural resilience of the gut microbiota in adult mice under high-fat dietary perturbations. ISME J. 6, 1848–1857 (2012).
Turnbaugh, P. J. et al. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444, 1027–1031 (2006).
Samuel, B. S. et al. Genomic and metabolic adaptations of Methanobrevibacter smithii to the human gut. Proc. Natl. Acad. Sci. USA 104, 10643–10648 (2007).
Fei, N. & Zhao, L. An opportunistic pathogen isolated from the gut of an obese human causes obesity in germfree mice. ISME J. 7, 880–884 (2013).
Devaraj, S., Hemarajata, P. & Versalovic, J. The Human Gut Microbiome and Body Metabolism: Implications for Obesity and Diabetes. Clin. Chem. 59, 617–628 (2013).
Osto, M. & Lutz, T. A. Translational value of animal models of obesity—Focus on dogs and cats. Eur. J. Pharmacol. 759, 240–252 (2015).
Sergeant, M. J. et al. Extensive Microbial and Functional Diversity within the Chicken Cecal Microbiome. PLoS ONE 9, e91941 (2014).
Yeoman, C. J. et al. The microbiome of the chicken gastrointestinal tract. Anim. Health Res. Rev. 13, 89–99 (2012).
Choi, J. H., Kim, G. B. & Cha, C. J. Spatial heterogeneity and stability of bacterial community in the gastrointestinal tracts of broiler chickens. Poult. Sci. 93, 1942–1950 (2014).
Oakley, B. B. & Kogut, M. H. Spatial and Temporal Changes in the Broiler Chicken Cecal and Fecal Microbiomes and Correlations of Bacterial Taxa with Cytokine Gene Expression. Front. Vet. Sci. 3 (2016).
Singh, K. M. et al. Taxonomic and gene-centric metagenomics of the fecal microbiome of low and high feed conversion ratio (FCR) broilers. J. Appl. Genet. 55, 145–154 (2014).
Stanley, D., Geier, M. S., Chen, H., Hughes, R. J. & Moore, R. J. Comparison of fecal and cecal microbiotas reveals qualitative similarities but quantitative differences. BMC Microbiol. 15 (2015).
Wang, Q. et al. Polymorphism of Heart Fatty Acid-Binding Protein Gene Associatied with Fatness Traits in the Chicken. Anim. Biotechnol. 18, 91–99 (2007).
Leng, L., Wang, S., Li, Z., Wang, Q. & Li, H. A polymorphism in the 3′-flanking region of insulin-like growth factor binding protein 2 gene associated with abdominal fat in chickens. Poult. Sci. 88, 938–942 (2009).
Shi, H., Wang, Q., Zhang, Q., Leng, L. & Li, H. Tissue expression characterization of chicken adipocyte fatty acid-binding protein and its expression difference between fat and lean birds in abdominal fat tissue. Poult. Sci. 89, 197–202 (2010).
Tian, J. et al. A Single Nucleotide Polymorphism of Chicken Acetyl-CoA Carboxylase A Gene Associated with Fatness Traits. Anim. Biotechnol. 21, 42–50 (2009).
Zhang, H. et al. Selection Signature Analysis Implicates the PC1/PCSK1 Region for Chicken Abdominal Fat Content. PLoS ONE 7, e40736 (2012).
Wang, W. et al. Expression profiling of preadipocyte microRNAs by deep sequencing on chicken lines divergently selected for abdominal fatness. PloS One 10, e0117843 (2015).
Zhang, H. et al. Detection of genome-wide copy number variations in two chicken lines divergently selected for abdominal fat content. BMC Genomics 15, 517 (2014).
Guo, L. et al. Comparison of adipose tissue cellularity in chicken lines divergently selected for fatness. Poult. Sci. 90, 2024–2034 (2011).
Meyer, B., Bessei, W., Vahjen, W., Zentek, J. & Harlander-Matauschek, A. Dietary inclusion of feathers affects intestinal microbiota and microbial metabolites in growing Leghorn-type chickens1. Poult. Sci. 91, 1506–1513 (2012).
Trivedi, P., Duan, Y. & Wang, N. Huanglongbing, a systemic disease, restructures the bacterial community associated with citrus roots. Appl. Environ. Microbiol. 76, 3427–3436 (2010).
Su, X., Pan, W., Song, B., Xu, J. & Ning, K. Parallel-META 2.0: enhanced metagenomic data analysis with functional annotation, high performance computing and advanced visualization. PloS One 9, e89323 (2014).
Segata, N. et al. Metagenomic microbial community profiling using unique clade-specific marker genes. Nat. Methods 9, 811–814 (2012).
Peng, Y., Leung, H. C. M., Yiu, S. M. & Chin, F. Y. L. IDBA-UD: a de novo assembler for single-cell and metagenomic sequencing data with highly uneven depth. Bioinforma. Oxf. Engl. 28, 1420–1428 (2012).
Zhu, W., Lomsadze, A. & Borodovsky, M. Ab initio gene identification in metagenomic sequences. Nucleic Acids Res. 38, e132 (2010).
Li, W. & Godzik, A. Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinforma. Oxf. Engl. 22, 1658–1659 (2006).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Sanner, M. F. Python: a programming language for software integration and development. J. Mol. Graph. Model. 17, 57–61 (1999).
Hammer, Ø., D. A. T. Harper & P. D. Ryan . PAST: Paleontological statistics software package for education and data analysis. Palaeontol. Electron. 4, 9 (2001).
Benjamini, Y. & Yekutieli, D. The control of the false discovery rate in multiple testing under dependency. Ann. Stat. 29, 1165–1188 (2001).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15 (2014).
Feng, Q. et al. Gut microbiome development along the colorectal adenoma–carcinoma sequence. Nat. Commun. 6, 6528 (2015).
Torok, V. A., Allison, G. E., Percy, N. J., Ophel-Keller, K. & Hughes, R. J. Influence of Antimicrobial Feed Additives on Broiler Commensal Posthatch Gut Microbiota Development and Performance. Appl. Environ. Microbiol. 77, 3380–3390 (2011).
De Filippo, C. et al. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proc. Natl. Acad. Sci. 107, 14691–14696 (2010).
Louis, P. & Flint, H. J. Diversity, metabolism and microbial ecology of butyrate-producing bacteria from the human large intestine. FEMS Microbiol. Lett. 294, 1–8 (2009).
Eeckhaut, V. et al. Butyrate production in phylogenetically diverse Firmicutes isolated from the chicken caecum: Butyrate-producing bacteria from the chicken caecum. Microb. Biotechnol. 4, 503–512 (2011).
Wall, R. et al. Contrasting effects of Bifidobacterium breve NCIMB 702258 and Bifidobacterium breve DPC 6330 on the composition of murine brain fatty acids and gut microbiota. Am. J. Clin. Nutr. 95, 1278–1287 (2012).
Turroni, S. et al. Enterocyte-Associated Microbiome of the Hadza Hunter-Gatherers. Front. Microbiol. 7 (2016).
Kettle, H., Louis, P., Holtrop, G., Duncan, S. H. & Flint, H. J. Modelling the emergent dynamics and major metabolites of the human colonic microbiota: Emergent microbial dynamics in the colon. Environ. Microbiol. 17, 1615–1630 (2015).
Eeckhaut, V. et al. Butyricicoccus pullicaecorum gen. nov., sp. nov., an anaerobic, butyrate-producing bacterium isolated from the caecal content of a broiler chicken. Int. J. Syst. Evol. Microbiol. 58, 2799–2802 (2008).
Roelofsen, H., Priebe, M. G. & Vonk, R. J. The interaction of short-chain fatty acids with adipose tissue: relevance for prevention of type 2 diabetes. Benef. Microbes 1, 433–437 (2010).
Brahe, L. K., Astrup, A. & Larsen, L. H. Is butyrate the link between diet, intestinal microbiota and obesity-related metabolic diseases? Obes. Rev. Off. J. Int. Assoc. Study Obes. 14, 950–959 (2013).
Balamurugan, R., Rajendiran, E., George, S., Samuel, G. V. & Ramakrishna, B. S. Real-time polymerase chain reaction quantification of specific butyrate-producing bacteria, Desulfovibrio and Enterococcus faecalis in the feces of patients with colorectal cancer. J. Gastroenterol. Hepatol. 23, 1298–1303 (2008).
Huycke, M. M., Abrams, V. & Moore, D. R. Enterococcus faecalis produces extracellular superoxide and hydrogen peroxide that damages colonic epithelial cell DNA. Carcinogenesis 23, 529–536 (2002).
Huycke, M. M. & Moore, D. R. In vivo production of hydroxyl radical by Enterococcus faecalis colonizing the intestinal tract using aromatic hydroxylation. Free Radic. Biol. Med. 33, 818–826 (2002).
Jones, B. V., Begley, M., Hill, C., Gahan, C. G. M. & Marchesi, J. R. Functional and comparative metagenomic analysis of bile salt hydrolase activity in the human gut microbiome. Proc. Natl. Acad. Sci. 105, 13580–13585 (2008).
Capcarova, M., Weiss, J., Hrncar, C., Kolesarova, A. & Pal, G. Effect of Lactobacillus fermentum and Enterococcus faecium strains on internal milieu, antioxidant status and body weight of broiler chickens. J. Anim. Physiol. Anim. Nutr. 94, e215–e224 (2010).
Khan, M., Raoult, D., Richet, H., Lepidi, H. & La Scola, B. Growth-promoting effects of single-dose intragastrically administered probiotics in chickens. Br. Poult. Sci. 48, 732–735 (2007).
Angelakis, E. & Raoult, D. The increase of Lactobacillus species in the gut flora of newborn broiler chicks and ducks is associated with weight gain. PloS One 5, e10463 (2010).
Lan, P. T. N., Binh, L. T. & Benno, Y. Impact of two probiotic Lactobacillus strains feeding on fecal lactobacilli and weight gains in chicken. J. Gen. Appl. Microbiol. 49, 29–36 (2003).
Takemura, N., Okubo, T. & Sonoyama, K. Lactobacillus plantarum strain No. 14 reduces adipocyte size in mice fed high-fat diet. Exp. Biol. Med. Maywood NJ 235, 849–856 (2010).
Karlsson, C. L. J. et al. Effects on weight gain and gut microbiota in rats given bacterial supplements and a high-energy-dense diet from fetal life through to 6 months of age. Br. J. Nutr. 106, 887–895 (2011).
Sanchez, M. et al. Effect of Lactobacillus rhamnosus CGMCC1.3724 supplementation on weight loss and maintenance in obese men and women. Br. J. Nutr. 111, 1507–1519 (2014).
Eckburg, P. B. Diversity of the Human Intestinal Microbial Flora. Science 308, 1635–1638 (2005).
Peabody, M. A., Van Rossum, T., Lo, R. & Brinkman, F. S. L. Evaluation of shotgun metagenomics sequence classification methods using in silico and in vitro simulated communities. BMC Bioinformatics 16 (2015).
Filippidou, S. et al. Under-detection of endospore-forming Firmicutes in metagenomic data. Comput. Struct. Biotechnol. J. 13, 299–306 (2015).
Poretsky, R., Rodriguez-R, L. M., Luo, C., Tsementzi, D. & Konstantinidis, K. T. Strengths and Limitations of 16S rRNA Gene Amplicon Sequencing in Revealing Temporal Microbial Community Dynamics. PLoS ONE 9, e93827 (2014).
Brooks, J. P. et al. The truth about metagenomics: quantifying and counteracting bias in 16S rRNA studies. BMC Microbiol. 15, 66 (2015).
Neves, A. L., Coelho, J., Couto, L., Leite-Moreira, A. & Roncon-Albuquerque, R. Metabolic endotoxemia: a molecular link between obesity and cardiovascular risk. J. Mol. Endocrinol. 51, R51–R64 (2013).
Vijay-Kumar, M. et al. Metabolic syndrome and altered gut microbiota in mice lacking Toll-like receptor 5. Science 328, 228–231 (2010).
Zenhom, M. et al. Peptidoglycan recognition protein 3 (PglyRP3) has an anti-inflammatory role in intestinal epithelial cells. Immunobiology 217, 412–419 (2012).
Tanaka, T. et al. Thiamine Prevents Obesity and Obesity-Associated Metabolic Disorders in OLETF Rats. J. Nutr. Sci. Vitaminol. (Tokyo) 56, 335–346 (2010).
Naruta, E. & Buko, V. Hypolipidemic effect of pantothenic acid derivatives in mice with hypothalamic obesity induced by aurothioglucose. Exp. Toxicol. Pathol. Off. J. Ges. Für Toxikol. Pathol. 53, 393–398 (2001).
Chakraborti, C. K. New-found link between microbiota and obesity. World J. Gastrointest. Pathophysiol. 6, 110 (2015).
Bui, T. P. N. et al. Production of butyrate from lysine and the Amadori product fructoselysine by a human gut commensal. Nat. Commun. 6, 10062 (2015).
Neis, E., Dejong, C. & Rensen, S. The Role of Microbial Amino Acid Metabolism in Host Metabolism. Nutrients 7, 2930–2946 (2015).
Moreno-Navarrete, J. M. et al. Metabolomics uncovers the role of adipose tissue PDXK in adipogenesis and systemic insulin sensitivity. Diabetologia 59, 822–832 (2016).
Campión, J., Milagro, F. I., Fernández, D. & Martínez, J. A. Diferential gene expression and adiposity reduction induced by ascorbic acid supplementation in a cafeteria model of obesity. J. Physiol. Biochem. 62, 71–80 (2006).
Acknowledgements
This work is partly funded by National Natural Science Foundation of China (Project number: 31460389).
Author information
Authors and Affiliations
Contributions
L.Y.K., H.L. and H.Z. designed the study and wrote the manuscript. L.W., W.H., Y.W. and L.L. designed and performed the experiments. Q.H., Y.Z., Z.G. and J.Z. analyzed the data. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Hou, Q., Kwok, LY., Zheng, Y. et al. Differential fecal microbiota are retained in broiler chicken lines divergently selected for fatness traits. Sci Rep 6, 37376 (2016). https://doi.org/10.1038/srep37376
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep37376
This article is cited by
-
Isolation and characterisation of novel Methanocorpusculum species indicates the genus is ancestrally host-associated
BMC Biology (2023)
-
Chicken cecal microbiota reduces abdominal fat deposition by regulating fat metabolism
npj Biofilms and Microbiomes (2023)
-
Systematic profiling of the chicken gut microbiome reveals dietary supplementation with antibiotics alters expression of multiple microbial pathways with minimal impact on community structure
Microbiome (2022)
-
Effect of host breeds on gut microbiome and serum metabolome in meat rabbits
BMC Veterinary Research (2021)
-
Transcriptome landscapes of differentially expressed genes related to fat deposits in Nandan-Yao chicken
Functional & Integrative Genomics (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.