Abstract
Post-transcriptional upregulation is an effective way to increase the expression of transgenes and thus maximize the yields of target chemicals from metabolically engineered organisms. Refractory elements in the 3′ untranslated region (UTR) that increase mRNA half-life might be available. In Saccharomyces cerevisiae, several terminator regions have shown activity in increasing the production of proteins by upstream coding genes; among these terminators the DIT1 terminator has the highest activity. Here, we found in Saccharomyces cerevisiae that two resident trans-acting RNA-binding proteins (Nab6p and Pap1p) enhance the activity of the DIT1 terminator through the cis element GUUCG/U within the 3′-UTR. These two RNA-binding proteins could upregulate a battery of cell-wall–related genes. Mutagenesis of the DIT1 terminator improved its activity by a maximum of 500% of that of the standard PGK1 terminator. Further understanding and improvement of this system will facilitate inexpensive and stable production of complicated organism-derived drugs worldwide.
Similar content being viewed by others
Introduction
Strong expression of the enzyme-encoding genes involved in target metabolic pathways is essential for the efficient production of target chemicals in the cells of metabolically engineered microorganisms, where multiple reactions proceed simultaneously. In Saccharomyces cerevisiae, this goal typically is achieved through upregulation at the transcriptional level, such as by incorporating strong promoters and multiple copies of transgenes1. A successful example of this strategy is the transgenic S. cerevisiae strain that produces the precursor for the antimalarial drug artemisinin2. Specifically, the GAL1/10 promoter—one of the strongest promoters in S. cerevisiae3—was used in combination with the multi-copy 2μ vector to strongly express several genes encoding enzymes involved in the synthesis of the isoprenoid2. However, upregulation at the transcriptional level comes with the burden of mRNA turnover and the accompanying energy consumption. The greater the increase in the number of transgenes to be introduced, the greater the reduction in yield of the target chemical. Therefore, alternative means that upregulate gene expression at the post-transcriptional level are required for the efficient production of complicated organism-derived drugs, such as opioids4.
Terminator regions are located within about 200 bp downstream of protein-coding sequences and have distinct functions, namely transcriptional termination, poly(A) addition, and post-transcriptional regulation via the 3′-UTR5,6. The amount of protein accumulated depends upon mRNA stability, which might in turn depend on secondary structures in the 3′-UTR7. Refractory elements in the 3′ untranslated region (UTR) that increase mRNA half-life might be available. In Saccharomyces cerevisiae, several terminator regions have shown activity in increasing the production of proteins by upstream coding genes8,9,10,11,12. Whereas some cis-elements within the 3′-UTR and its corresponding trans proteins downregulate gene expression via mRNA degradation or translational repression13,14, in vertebrates a few promote the accumulation of protein through translation initiation by cytoplasmic polyadenylation15, stabilization of fragile mRNA16, or translational activation17,18 by formation of mRNA–protein complexes. These elaborate mechanisms, which function transiently during developmental process or stress responses, have yet to be applied in metabolic engineering.
Among the terminator regions in the S. cerevisiae genome, the dityrosine-deficient 1 (DIT1) terminator (DIT1t), which was identified by comprehensive screening9, has the strongest ability to increase transgene expression under various growth conditions and among various S. cerevisiae strains19. The DIT1 terminator also upregulates the expression of upstream transgenes that encode various proteins, including fluorescent proteins and cellulases, under different promoters19,20. Only the activity of DIT1t is enhanced in the stationary phase, whereas those of the other top-five-ranked terminators decrease, suggesting that DIT1t activity is post-transcriptionally upregulated by unknown factors19, which might interact with the DIT1 3′-UTR (Fig. 1A). In this study, we identified these factors to improve the strong DIT1 terminator for protein production and metabolic engineering.
Identification of DIT1t-activating factors.
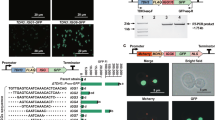
(A) Sequence of the DIT1 3′-UTR. The length is 208 bp. The cis-element GUUCG is indicated in red. Five deleted regions are indicated by lines. One point mutation (m22) is indicated by an asterisk. Two effective deletion mutations (d7 and d21) are indicated in cyan. (B) Genetic interaction between PAP1 and NAB6. The GFP fluorescence intensity of the control NAB6 strain harboring pGP564 was used as the standard. (C) GFP mRNA levels analyzed by RT-PCR. Each total RNA was extracted from the corresponding cells, as denoted in the caption to Fig. 1B. (D) Deletion analysis of the DIT1 terminator to identify Pap1p- and Nab6p-recruiting cis region(s). A series of DIT1t mutants (d7 to d21) with 3-bp deletions in the DIT1t region (Supplementary Table 3) and a 10-bp-deleted d2 mutant were constructed. The GFP fluorescence intensity of the wild-type DIT1t strain harboring pGP564 was used as the standard control (indicated as “WT”). A d2-deleted region is indicated by the line. Four deleted regions (d7, d15, d16, and d21) are indicated. (E) Gain-of-function analyses using mutated PGK1 terminators. Two strains harboring a mutated terminator (Supplementary Fig. 7) were constructed. The GFP fluorescence intensities were measured. (F) Identification of DIT1t-activating cis-element sequences by using point mutagenesis. A series of DIT1t mutants (m1 to m30) with point mutations in the DIT1t region (Supplementary Table 3) were constructed. One point mutation (m22) is indicated by a cyan circle. The GFP fluorescence intensity of the wild-type DIT1t strain harboring pGP564 was used as the standard. Data are means of three or four independent experiments. (G) The protein production system involving the cis element GUUCG in the DIT1 3′-UTR. Both Nab6p and Pap1p are considered to be trans-acting factors in this system. The empty control vector (white), pGP564 with NAB6 insert (gray), pGP564 with PAP1 insert (blue), and pGP564 with PAP1–NAB6 combined insert (green) were separately transformed into the corresponding strains. Data are means ± 1 SD of at least three independent experiments. n.s., not significant; *p < 0.01. See also Supplementary Table 5.
Results and Discussion
Identification of DIT1t-activating genes
To shed light on these unknown DIT1t-activating factors, we introduced the Yeast Genomic Tiling Collection21 into a DIT1t strain that had a genome-integrated green fluorescent protein (GFP) upstream of DIT1t under the control of the TDH3 gene promoter (Supplementary Figs 1–3 and Supplementary Table 1). We identified two DIT1t-activating genes: nucleic acid-binding protein 622 (NAB6), which had one RNA-binding domain and three poly(A) binding domains, and poly(A) RNA polymerase23 (PAP1). Each gene enhanced the fluorescence intensity of GFP upon overexpression in the DIT1t strain (Fig. 1B), but not in the other top-five-ranked strains (Supplementary Fig. 4). A transformant overexpressing both NAB6 and PAP1 in the DIT1t strain exhibited greater GFP fluorescence than did the transformants harboring each gene separately. Deletion of NAB6 (nab6Δ) in the DIT1t strain background decreased the GFP fluorescence to 60% and virtually prevented PAP1 overexpression from increasing GFP fluorescence (Fig. 1B). In contrast, no significant differences in fluorescence intensity between the NAB6 background and the nab6Δ background were observed in the other terminator backgrounds (Supplementary Fig. 5). These results indicated that NAB6 and PAP1 increased DIT1t activity specifically and coordinately. In taking these results together with the physical interaction between Nab6p and Pap1p24, we hypothesized that Nab6p might preferentially bind with the DIT1 3′-UTR of mRNA and recruit Pap1p to sterically inhibit the degradation of the mRNA. Alternatively, RNA polymerase II-dependent gene looping25 between the DIT1 terminator and the cognate promoter via a Nab6p–Pap1p complex might facilitate transcription reinitiation.
mRNA level
To evaluate these two possibilities, we conducted quantitative PCR to investigate the amounts of GFP mRNA containing the DIT1 3′-UTR that were produced by these strains (Fig. 1C). Overexpression of NAB6 increased the amount of GFP mRNA by 10% independently of PAP1, whereas the GFP mRNA level was not affected by the PAP1-overexpressing or nab6Δ background. These results indicated that gene looping likely does not promote transcription of the transgenes harboring DIT1t. These results raise the possibility that Nab6p binds to the DIT1 3′-UTR and thus stabilizes the mRNA to increase protein yield; however, the contribution of Pap1p to the increase strongly suggests a mechanism other than mRNA stabilization.
Identification of cis-element
To genetically identify the cis-elements associated with DIT1t activation, we made six DIT1t mutants with 10-bp deletions (d1 to d6) from the 208-bp sequence of the wild-type DIT1 3′-UTR, as determined by 3′-RACE (Supplementary Fig. 6A). Of the six deletions, only the d2 deletion (i.e., UUAGUUAGUU [+51 to +60 from the stop codon]; Fig. 1D and Supplementary Fig. 6B) prevented both NAB6 and PAP1 overexpression from increasing DIT1t activity. This result suggested that a specific cis-element was present near the d2 deletion. As a result of comprehensive comparative analyses of diverse eukaryotic genomes, the length of the cis-element within the 3′-UTR is considered to be about 5 to 10 bp26. To conduct fine mapping of the cis-element, we made another series of DIT1t mutants with 3-bp deletions between regions d1 and d3 (d7 to d21). Of the 15 deletions, two consecutive ones, d15 (AGT [+57 to +59]) and d16 (TCG [+60 to +62]) prevented the activation of DIT1t by overexpression of both NAB6 and PAP1 (Fig. 1D). These deleted regions overlapped with the d2 deletion, suggesting that AGUUCG was the cis-element involved in DIT1t activation. Gain-of-function analyses using mutated PGK1 terminators with the AGTTCG insertion or substitution showed that the cis-element might increase terminator activity by overexpression of NAB6 (Fig. 1E and Supplementary Fig. 7). Overexpression of PAP1 has not been observed to significantly increase the fluorescence intensity of GFP (Fig. 1E). Saturation mutagenesis with 1-bp mutations around the core candidate sequence was conducted to determine the functional cis-element sequences (Fig. 1F). Among the mutated sequences shown in this figure, substitution of any of the first five nucleotides only slightly reduced DIT1t activation compared with that in the wild type, although substitution of the first T for a G reduced activation more markedly. Substitution of the last five nucleotides markedly reduced the degree of activation, whereas substitution of the last G with a T made DIT1t activation only slightly lower than in the wild type. Taken together, the cis-element GUUCG/U in the 3′-UTR might enhance upstream protein production in a way that is dependent upon Nab6p and Pap1p (Fig. 1G). Pap1p might modulate the length of the poly(A) tail of the mRNA harboring the DIT1 3′-UTR to increase translation. Alternatively, an Nab6p–Pap1p complex directly interacts with the initiation factor complex to increase the frequency of translation initiation or to stabilize polyribosomes to promote translation.
Other target mRNAs
In eukaryotes, RNA-binding-protein–mRNA complexes have been proposed to function as hubs to regulate genetic networks at the posttranscriptional level but remains to be well elucidated26,27. The mRNAs associated with Nab6p have been identified comprehensively and the significance of the interactions ranked whereas the sequence of the Nab6p binding motif has yet to be determined27. DIT1 mRNA was not detected as an Nab6p-binding mRNA in the previous study27; this could have been because DIT1 is expressed strictly at a specific time during sporulation28. We investigated whether the terminators from Nab6-binding mRNA genes were activated by overexpression of NAB6 and PAP1 in the same way as was DIT1t (Table 1). Eleven of the top-18-ranked terminators exhibited Nab6p–Pap1p-dependent activation, and all 11 terminator regions had at least one core cis-element sequence, GUUCG/U, within 200-bp downstream from the stop codon, as was found in DIT1t (Table 1). Nab6p–Pap1p-dependent activation appeared irrelevant to the intensity of terminator activity (Table 1). Six of the 11 terminators were derived from cell wall–related genes (Table 1). DIT1 is also involved in the synthesis of spore walls29, suggesting that Nab6p–Pap1p–mRNA complexes could form part of the networks regulating cell-wall modification. This putative network could also regulate the other cell wall-related genes. These results also support the hypothesis that Nab6p directly associates with the cis-element GUUCG/U within the 3′-UTR.
Effects on cell growth
In batch fermentation, both the rate of production and the overall product yield often are correlated with growth rate. We investigated whether the deletion of NAB6 or overexpression of NAB6 or PAP1 influenced the growth rates of the yeasts in the absence of the expression of other transgenes upstream of DIT1t. At 30 °C, the growth of the nab6Δ mutant was comparable to that of the control strain. In contrast, strains overexpressing either NAB6 or PAP1 showed decreased growth compared with the control strain, and the strain overexpressing both NAB6 and PAP1 exhibited even greater growth retardation than did either of the single protein–overexpressing strains (Fig. 2A). This detrimental effect could have resulted from exacerbation of NAB6 and PAP1 gene expression by the action of the Nab6p–Pap1p complex, followed by disordered composition, or modification, of the cell wall. Overexpression of cell wall–related genes decreases growth rates during the vegetative stage in S. cerevisiae30. Excess Nab6p could non-specifically bind poly(A) within mRNAs via its poly(A) binding domains to indiscriminately affect gene expression. In metabolically engineering yeasts, we must design the elements in such a way that specifically increases the expression of the target enzyme genes. Our result here implied that directly overexpression of NAB6 and/or PAP1 was inappropriate as a means of enhancing the expression of transgenes.
(A) Growth retardation by overexpression of NAB6 and PAP1. (Left) Empty pGP564 vector, pGP564-NAB6, pGP564-PAP1, and pGP564-PAP1-NAB6 were separately transformed into the wild-type W303-1A strain (upper; marked NAB6) or the nab6Δ strain (lower; marked nab6Δ). Two days after inoculation on SD-leucine medium at 30 °C, the colonies were observed. (Right) Optical density (OD) was measured every 30 min from 0 to 24 h after reinoculation at OD660 = 0.1. The averages of three replicate experiments from 6 to 24 h are indicated. The nab6Δ/DIT1t strain (gold) and the NAB6/DIT1t strains harboring pGP564 (white), pGP564-NAB6 (gray), pGP564-NAB6 (blue), or pGP564-PAP1-NAB6 (green) are indicated. (B) Identification of the strongest terminator derived from DIT1t. (a) Activation of strong DIT1t-derived terminators by overexpression of NAB6 and PAP1. The fluorescence intensity of the wild-type DIT1t strain harboring pGP564 was used as the standard. These experiments were conducted independently of those in Fig. 1. (b) Comparison of terminator activities among DIT1t strains and the standard PGK1t strain in various growth phases. The fluorescence intensities of terminator strains with the GFP gene under the control of the TDH3 promoter were measured by flow cytometry. Sampling times were 6, 12, 24, and 30 h after reinoculation at OD660 = 0.1. (c) Characterization of the strongest DIT1t-d22 terminator. (Left) Effects of promoter and reporter gene exchange on 3′-UTR activity. These strains expressed either the GFP gene under the control of ACT1pro (left) or the mKO2 gene under the control of TDH3pro (right), as indicated. (Right) Effect of the host strain on 3′-UTR activity. Two wild-type Saccharomyces cerevisiae strains (A451 and TDO2) were transformed individually. The fluorescence intensities of the terminator strains containing the GFP gene under the control of the TDH3 promoter were measured by flow cytometry. The terminator strains examined were PGK1t (purple), DIT1t (orange). DIT1t-m22 (yellow), DIT1t-d7 (gray), DIT1t-d21 (dark gray), and DIT1t-d22 (pink). Data are means ± 1 SD of at least three independent experiments. n.s., not significant; *p < 0.01; **p < 0.05. See also Supplementary Table 5.
Improvement of the DIT1 terminator
To improve this post-transcriptional expression system without overexpressing NAB6 and PAP1, we modified the sequence of the DIT1 3′-UTR. We found that three mutated DIT1 terminators had 5% to 10% greater activity than the wild-type DIT1t when NAB6 and PAP1 were overexpressed (Fig. 2B (a)). The activity of DIT1t-d22, which was made by accumulation of the d7 and d21 mutations, was greater than those of DIT1t-d7 and DIT1t-d21 and comparable to that of DIT1t-m22. Among five DIT1t-derived terminators, DIT1t-d22 had the greatest activity from the early log phase to the stationary phase in the W303-1a background (Fig. 2B (b)). During the stationary phase, the maximum activity of DIT1t-d22 was about 5 times that of the standard PGK1 terminator and 1.5 times that of the wild-type DIT1 terminator. We examined the activities of both DIT1t and DIT1t-d22 in terms of the effects of promoter and reporter gene exchanges, namely by using the weak ACT1 promoter instead of the strong TDH3 promoter and by using an mKO2 red fluorescent protein gene instead of the GFP gene (Left panel in Fig. 2B (c)). The DIT1t-d22 strain showed greater terminator activity than the DIT1t strain while a significant difference was observed only at log phase in both cases. In another haploid strains (A451) and one diploid strain (TDO2) used in the abovementioned study19 and transfected with constructs harboring DIT1t or DIT1t-d22, we observed that transfection with the DIT1t-d22 strain gave significantly greater terminator activity than did transfection with the wild-type DIT1t strain at almost all phases (Right panel in Fig. 2B (c)).
Conclusions
Like in the case of opioid-producing yeasts with dozens of transgenes4, use of our system might improve the efficiency of production of target chemicals. The molecular mechanism by which Nab6p–Pap1p–3′-UTR complexes increase protein production by transgenes remains to be determined. Two mutated PGK1 terminators were associated with an increase in upstream protein production in a manner dependent on Nab6p but not on Pap1p (Fig. 1E). These results suggested that cis-elements other than GUUCG/U are needed for Pap1p to interact with Nab6p or the 3′-UTR to enhance protein production. Further investigation is needed to determine all of the components of this mechanism and their functions. With such knowledge, we would be able to improve this regulatory system to increase protein production by transgenes and apply it to other organisms by using synthetic biological methods. The use of these methods likely will contribute to the development of metabolically-engineered organisms for producing a wide variety of beneficial chemicals.
Materials and Methods
Host strain and media
Escherichia coli JM109 was used as the host cell for DNA manipulation. Luria–Bertani (LB) medium supplemented with 100 mg/L ampicillin or 30 mg/L kanamycin was used for E. coli cell culture to select the transformants. Saccharomyces cerevisiae strain BY4741 (MATa URA3Δ0 LEU2Δ0 HIS3Δ1 MET15Δ0) was used for screening and cloning. Strains expressing codon-optimized GFP under the control of the TDH3 promoter and either DIT1t (DIT1t strain), the RPL15BA terminator (RPL15BAt strain), the RPL41B terminator (RPL41B strain), the RPL3 terminator (RPL3t strain), the IDP1 terminator (IDP1t strain), or the PGK1 terminator (PGK1t strain) were constructed previously19. Saccharomyces cerevisiae strain W303-1A (MATa leu2-3, 112 trp1-1 can1-100 ura3-1 ade2-1 his3-11, 15) was also used as a host strain for genetic analyses. Three S. cerevisiae strains were used as wild-type strains to examine the activity of DITt-d22: they were A451 (MATα can1 leu2 trp1 ura3 aro7); TDO2 (MATa/α trp1/trp1 ura3/ura3) made from OC-2T31 with the use of 5-fluoroorotic acid; and YPH499 (MATa ura3-52 lys2-801 ade2-101 trp1-Δ63 his3-Δ200 leu2-Δ1). Yeast cells were grown in synthetic complete medium, which contained 0.67% yeast nitrogen base without amino acids (Difco, Detroit, MI), 0.082% complete supplement mixture (CSM), CSM-Leucine (ForMedium, Norfolk, UK), and adenine (40 mg/L), supplemented with 2% glucose (Synthetic Dextrose minimal medium; SD) and with 2% agar as needed.
Plasmid construction
Except in the case of the pGP564-PAP1-NAB6 vector, all plasmid construction was performed by using the gap-repair cloning method32 and the BY4741 strain. For each gene, the region from the predicted promoter region to the terminator region was cloned into the pGP564 vector (#YSC5034, Thermo Fisher Scientific Inc., MA). The pGP564 vector was digested with BamHI and XhoI restriction enzymes at 37 °C to prepare linear DNA for gap-repair cloning. The genes used in this study are as follows, and the locations of the regions amplified are given relative to the start codon of the gene (=position 1). NAB6 (−391 to +3952) and YML119W (−1000 to +1720) were amplified from 12B3 vector. PAP1 (−509 to +2149) and ECM9 (−746 to +1312) were amplified from 10A10 vector. CCC1 (−487 to +1540) was amplified from 11E10 vector. MSC3 (−500 to +2674) was amplified from the BY4704 strain chromosome. Plasmids pGP564-NAB6, -YML119W, -PAP1, -ECM9, -CCC1, and -MSC3 were extracted from the transformants and then used to transform the JM109 strain, as described above. The pGP564 vector containing both the PAP1 and NAB6 gene sequences (pGP564-PAP1-NAB6) was constructed by using an In-Fusion HD cloning kit (Takara Bio Inc.). After the digestion of pGP564-NAB6 vector with SpeI restriction enzyme, the PAP1 PCR product was reacted with the digested vector to produce the pGP564-PAP1-NAB6 vector.
Screening for DIT1t-activating factors by using the Yeast Genomic Tiling Collection
A simple scheme of the screening strategy is given in Supplementary Fig. 1. The Yeast Genomic Tiling Collection, which was constructed previously21, was purchased from Thermo Fisher Scientific (catalog no. YSC4613). This collection is stored in 16 wells of a 96-well microplate. For each well, we mixed the Escherichia coli Yeast Genomic Tiling Collection strains together and cultured them at 37 °C overnight in 100 mL of LB medium containing 30 μg/mL kanamycin. The DNA of each of the 16 plasmid pools was then isolated by using the alkali-sodium dodecyl sulfate (SDS) method without the addition of RNase and used to transform the DIT1t strain. Yeast transformation was performed as described previously33. Briefly, yeast cells were grown in yeast extract peptone dextrose (YPD) medium overnight, then diluted 10-fold with YPD and incubated for 5 h. Cells were precipitated and washed with deionized water, and then mixed with transformation solution (120 μL of 60% PEG3350, 5 μL of 4 M lithium acetate, 10 μL of 1 M DTT, and 10 μL of 10 mg/mL denatured salmon sperm DNA) and the plasmid DNA. The mixture was incubated at 42 °C for 40 min and then spread on SD-leucine medium. The resultant pool of transformants was transiently collected and stored at −80 °C until further use. The pool of transformants was spread on SD-leucine medium, and individual transformants were randomly picked into 96-deep well plates and incubated at 30 °C at 1800 rpm in an MBR-022UP Bio Shaker Incubator (Taitec, Saitama, Japan) overnight. A 10-μL aliquot of culture was reinoculated into 500 μL of SD-leucine and grown at 30 °C for 6 h. The GFP fluorescence intensity (FI) and side-scatter intensity (SS) of the cells were measured by using a Cell Lab Quanta SC MPL flow cytometer (Beckman-Coulter, CA), and a histogram of the FI/SS ratio in each cell was fitted with IgorPro 6.1 software (Wavemetrics, Lake Oswego, OR), as described previously34.
Construction of nab6Δ strain
NAB6 gene deletion in the W303-1a strain (nab6Δ) was performed as follows. First, the KanMX selectable marker region was amplified from the genome of the faa4::KANMX4/faa4::KANMX4 strain in the diploid non-essential homozygous deletion strain collection (Research Genetics, Huntsville, AL). We then conducted nested PCR with NAB6-nested-f (5′-CCTTGTTTCAGGTTACGTGAAAAGCATCCAGAGAAGAT-3′) and NAB6-nested-c (5′-AAGAAATCGGAGAAAAAAAGGGAGAAACACCTTGGCAG-3′), followed by NAB6–150(40)-KANMX4-334 (5′-AAAGCATCCAGAGAAGATATCCCAAAGACTGAAGAGGTGTcccagaataccctccttgacag-3′) and NAB6+3500c(40)-KANMX4+1023c (5′-GGAGAAACACCTTGGCAGGCCGCCTTAGGCTTCCTTGGGCgaatcgacagcagtatagcgacca-3′). The PCR product was transformed into the DIT1t strain, and the resultant transformants were plated on YPD medium containing 100 μg/mL G418 to select for the presence of the KanMX marker.
Analysis of genetic interaction between NAB6 and PAP1
Four plasmids (pGP564, pGP564-NAB6, pGP564-PAP1, and pGP564-NAB6-PAP1) were introduced into the DIT1t strain in the W303-1a NAB6 background. Two plasmids (pGP564 and pGP564-PAP1) were introduced into the DIT1t strain in the nab6Δ background. As described above, the intensities of GFP fluorescence of these six strains were investigated 6 h after reinoculation.
Quantitative real-time PCR analysis
mRNA levels were measured as described in our previous paper8. Briefly, the DIT1t strain was cultured for 6, 12, or 24 h, and then total RNA was extracted by using a High Pure RNA Isolation Kit (Roche, Basel, Switzerland). cDNA was synthesized from a 1-μL aliquot of total RNA by using a High Capacity cDNA Reverse Transcription Kit (Life Technologies, Carlsbad, CA). Real-time PCR analysis was performed with SYBR Green PCR Master Mix and an ABI PRISM 7000 sequence detection system (Life Technologies). To quantify the amount of cDNA, the following primers were used: PAP1-1 (5′-TGGAGTGGTCTTGTTGAAAGTAAGG-3′), PAP1-2 (5′-AAGGGTTTGGTGAAAGGATGTG-3′), NAB6-1 (5′-TGGGTTCAGACATAGGCAATAGAAC-3′), NAB6-2 (5′-TCCTCCACGCACGACATTA-3′), GFP-1 (5′-CCAATTGGTGATGGTCCAGTCT-3′), GFP-2 (5′-CGGTGACGAACTCCAACAAAA-3′), TUB1-1 (5′-CCAACTGGTTTCAAGATCGGTA-3′), and TUB1-2 (5′-TCCACAGTGGCCAATTGTGA-3′). The amount of TUB1 mRNA was used as an internal control.
Identification of the sequence of the DIT1 3′-UTR
Rapid amplification of cDNA ends PCR (RACE-PCR) was used to identify the complete sequence of the DIT1 3′-UTR (3′-Full RACE Core set; Takara Bio Inc., Shiga, Japan). One microgram of total RNA isolated from the DIT1t strain was reverse-transcribed by using the oligo(dT) adaptor primer, and this first-strand cDNA was amplified with PrimeSTAR HS DNA polymerase (Takara Bio Inc.) by using an optimized GFP-specific primer (5′-CCAGACAACCATTATTTG-3′) and an anchor primer. Amplified 3′-RACE products were cloned into the pCR4Blunt-TOPO vector (Invitrogen); more than 20 independent clones were sequenced.
Identification of the sequence of the DIT1t-activating cis-element
A series of DNA fragments containing 21 deletion-mutated terminators (d1 to d21) and 30 point-mutated terminators (m1 to m30) were synthesized (Genscript, Inc., Piscataway, NJ). These mutated DIT1 terminators are listed in Supplementary Table 2. Strains expressing codon-optimized GFP under the control of the TDH3 promoter and each DIT1t (DIT1t strains) were constructed as described above. Three plasmids (pGP564, pGP564-NAB6, or pGP564-PAP1) were introduced into the deletion-mutated DIT1t strains. Four plasmids (pGP564, pGP564-NAB6, pGP564-PAP1, or pGP564-NAB6-PAP1) were introduced into the point-mutated DIT1t strains. As described above, the intensities of GFP fluorescence of these 183 strains were investigated 6 h after reinoculation.
Gain-of-function analyses using the mutated PGK1 terminators
DIT1t-inserted PGK1t was designed as follows: the 17-bp DIT1t sequence containing a cis-element between +46 and +62 from the stop codon was inserted into the PGK1 terminator between +28 and +29 from the stop codon (Supplementary Table 2 and Supplementary Fig. 7). The DIT1t-substituted PGK1t was designed as follows: the 11-bp PGK1t sequence between +17 and +29 from the stop codon was substituted for the 14-bp DIT1t sequence (Supplementary Table 2 and Supplementary Fig. 7). Four plasmids (pGP564, pGP564-NAB6, pGP564-PAP1, or pGP564-NAB6-PAP1) were introduced into the three PGK1t (PGK1t, DIT1t-inserted PGK1t, and DIT1t-substituted PGK1t) strains. As described above, the intensities of GFP fluorescence of these 12 strains were investigated 6 h after reinoculation.
Assay of the terminator activities of Nab6p-binding mRNA genes
Eighteen terminator regions of Nab6p-binding mRNA genes were amplified by PCR as described previously8. All cloning procedures were performed with an In-Fusion Advantage PCR Cloning Kit (Clontech, Mountain View, CA). Each PCR primer set was designed as 5′-AATTGTATAACTGAGGTACC-3′ plus a gene-specific forward primer, and 5′-TAATGTCGTTGGATCC-3′ plus a gene-specific reverse primer. Each cloned terminator region was inserted into the vector and identified by sequencing analysis. DNA sequences of both the gene-specific primers and the amplified regions are listed in Supplementary Table 3. Each terminator strain was constructed, and the activity of the terminator of each strain was measured as described above (Supplementary Fig. 8).
Assay of activities of mutation-accumulated DIT1 terminators
A DNA fragment containing DIT1t-d22 was synthesized (Genscript, Inc., Piscataway, NJ). These DNA sequences are described in Supplementary Table 3. Four plasmids (pGP564, pGP564-NAB6, pGP564-PAP1, or pGP564-NAB6-PAP1) were introduced into the DIT1t-d22 strain. As described above, the intensities of GFP fluorescence were investigated 6 h after reinoculation.
Additional Information
How to cite this article: Ito, Y. et al. Enhancement of protein production via the strong DIT1 terminator and two RNA-binding proteins in Saccharomyces cerevisiae. Sci. Rep. 6, 36997; doi: 10.1038/srep36997 (2016).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Da Silva, N. A. & Srikrishnan, S. Introduction and expression of genes for metabolic engineering applications in Saccharomyces cerevisiae. FEMS Yeast Res. 12, 197–214 (2012).
Paddon, C. J. et al. High-level semi-synthetic production of the potent antimalarial artemisinin. Nature 496, 528–532 (2013).
Lohr, D., Venkov, P. & Zlatanova, J. Transcriptional regulation in the yeast GAL gene family: a complex genetic network. FASEB J. 9, 777–787 (1995).
Galanie, S., Thodey, K., Trenchard, I. J., Filsinger Interrante, M. & Smolke, C. D. Complete biosynthesis of opioids in yeast. Science 349, 1095–1100 (2015).
van Helden, J., del Olmo, M. & Perez-Ortin, J. E. Statistical analysis of yeast genomic downstream sequences reveals putative polyadenylation signals. Nucleic Acids Res. 28, 1000–1010 (2000).
Kuersten, S. & Goodwin, E. The power of the 3′ UTR: translational control and development. Nat. Rev. Genet. 4, 626–637 (2003).
Babiskin, A. H. & Smolke, C. D. A synthetic library of RNA control modules for predictable tuning of gene expression in yeast. Mol. Syst. Biol. 7, 471 (2011).
Yamanishi, M., Katahira, S. & Matsuyama, T. TPS1 terminator increases mRNA and protein yield in a Saccharomyces cerevisiae expression system. Biosci. Biotechnol. Biochem. 75, 2234–2236 (2011).
Yamanishi, M. et al. A Genome-Wide Activity Assessment of Terminator Regions in Saccharomyces cerevisiae Provides a “Terminatome” Toolbox. ACS Synth. Biol. 2, 337–347 (2013).
Curran, K. A., Karim, A. S., Gupta, A. & Alper, H. S. Use of expression-enhancing terminators in Saccharomyces cerevisiae to increase mRNA half-life and improve gene expression control for metabolic engineering applications. Metab. Eng. 19, 88–97 (2013).
Curran, K. A. et al. Short Synthetic Terminators for Improved Heterologous Gene Expression in Yeast. ACS Synth. Biol. 4, 824–832 (2015).
Ito, Y., Yamanishi, M., Ikeuchi, A., Imamura, C. & Matsuyama, T. Combinatorial Screening for Transgenic Yeasts with High Cellulase Activities in Combination with a Tunable Expression System. PloS one 10, e0144870 (2015).
Parker, R. RNA degradation in Saccharomyces cerevisae. Genetics 191, 671–702 (2012).
Miller, M. A. & Olivas, W. M. Roles of Puf proteins in mRNA degradation and translation. Wiley Interdiscip. Rev. RNA 2, 471–492 (2011).
Barnard, D. C., Ryan, K., Manley, J. L. & Richter, J. D. Symplekin and xGLD-2 are required for CPEB-mediated cytoplasmic polyadenylation. Cell 119, 641–651 (2004).
Beckel-Mitchener, A. C., Miera, A., Keller, R. & Perrone-Bizzozero, N. I. Poly(A) tail length-dependent stabilization of GAP-43 mRNA by the RNA-binding protein HuD. J. Biol. Chem. 277, 27996–28002 (2002).
Uniacke, J. et al. An oxygen-regulated switch in the protein synthesis machinery. Nature 486, 126–129 (2012).
Fukao, A. et al. MicroRNAs trigger dissociation of eIF4AI and eIF4AII from target mRNAs in humans. Mol. Cell 56, 79–89 (2014).
Ito, Y. et al. Characterization of five terminator regions that increase the protein yield of a transgene in Saccharomyces cerevisiae. J. Biotechnol. 168, 486–492 (2013).
Ito, Y., Yamanishi, M., Ikeuchi, A. & Matsuyama, T. A highly tunable system for the simultaneous expression of multiple enzymes in Saccharomyces cerevisiae. ACS Synth. Biol. 4, 12–16 (2015).
Jones, G. M. et al. A systematic library for comprehensive overexpression screens in Saccharomyces cerevisiae. Nat. Methods 5, 239–241 (2008).
Samanta, M. P. & Liang, S. Predicting protein functions from redundancies in large-scale protein interaction networks. Proc. Natl. Acad. Sci. USA 100, 12579–12583 (2003).
Lingner, J., Kellermann, J. & Keller, W. Cloning and expression of the essential gene for poly(A) polymerase from S. cerevisiae. Nature 354, 496–498 (1991).
Ezeokonkwo, C., Ghazy, M. A., Zhelkovsky, A., Yeh, P. C. & Moore, C. Novel interactions at the essential N-terminus of poly(A) polymerase that could regulate poly(A) addition in Saccharomyces cerevisiae. FEBS Lett. 586, 1173–1178 (2012).
Ansari, A. & Hampsey, M. A role for the CPF 3′-end processing machinery in RNAP II-dependent gene looping. Genes Dev. 19, 2969–2978 (2005).
Ray, D. et al. A compendium of RNA-binding motifs for decoding gene regulation. Nature 499, 172–177 (2013).
Hogan, D. J., Riordan, D. P., Gerber, A. P., Herschlag, D. & Brown, P. O. Diverse RNA-binding proteins interact with functionally related sets of RNAs, suggesting an extensive regulatory system. PLoS Biol. 6, e255 (2008).
Friesen, H., Hepworth, S. & Segall, J. An Ssn6-Tup1-dependent negative regulatory element controls sporulation-specific expression of DIT1 and DIT2 in Saccharomyces cerevisiae. Mol. Cell. Biol. 17, 123–134 (1997).
Briza, P., Eckerstorfer, M. & Breitenbach, M. The sporulation-specific enzymes encoded by the DIT1 and DIT2 genes catalyze a two-step reaction leading to a soluble LL-dityrosine-containing precursor of the yeast spore wall. Proc. Natl. Acad. Sci. USA 91, 4524–4528 (1994).
Yoshikawa, K. et al. Comprehensive phenotypic analysis of single-gene deletion and overexpression strains of Saccharomyces cerevisiae. Yeast 28, 349–361 (2011).
Saitoh, S., Mieno, Y., Nagashima, T., Kumagai, C. & Kitamoto, K. Breeding of a new type of baker’s yeast by δ-integration for overproduction of glucoamylase using a homothallic yeast. J. Ferment. Bioeng. 81, 98–103 (1996).
Ma, H., Kunes, S., Schatz, P. J. & Botstein, D. Plasmid construction by homologous recombination in yeast. Gene 58, 201–216 (1987).
Yamanishi, M. & Matsuyama, T. A modified Cre-lox genetic switch to dynamically control metabolic flow in Saccharomyces cerevisiae. ACS Synth. Biol. 1, 172–180 (2012).
Matsuyama, T., Yamanishi, M. & Takahashi, H. Improvement of galactose induction system in Saccharomyces cerevisiae. J. Biosci. Bioeng. 111, 175–177 (2011).
Acknowledgements
We thank Dr. Hisao Moriya for his helpful discussions. We would like to thank Prof. Rinji Akada, Dr. Hisashi Hoshida, Sayo Takei, Mieko Horikiri, Mami Kanemitsu and Takako Nakamura for their technical assistance. Financial support from the Ministry of Education, Science, Sports, and Culture of Japan Grant-in-Aid for Exploratory Research (JP25660071) (TM) and Scientific Research (C) (JP15K07378) (TM) is acknowledged.
Author information
Authors and Affiliations
Contributions
Y.I., T.K., S.I., K.I., M.F.-S. and T.M. conceived and designed the experiments. Y.I., M.Y. and T.K. identified the trans-acting factors. S.K. and T.M. determined the cis elements. S.I., K.I., M.F.-S. and T.M. co-wrote the paper. All of the authors discussed the data and commented on the manuscript.
Ethics declarations
Competing interests
The authors have the following interests. This work was mainly supported by TOYOTA Central Research and Development Laboratories Incorporation, and partially supported by JSPS. Some authors including Y.I., M.Y., S.K., and T.M., were employed by TOYOTA CENTRAL R&D LABS. INC. at the time of the study. There are two patents (JP-A-2015-136303 and JP-A-2016-3052), but no products in development or marketed products to declare. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript. The other authors including T.K., S.I., K.I., and M.F.-S. declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Ito, Y., Kitagawa, T., Yamanishi, M. et al. Enhancement of protein production via the strong DIT1 terminator and two RNA-binding proteins in Saccharomyces cerevisiae. Sci Rep 6, 36997 (2016). https://doi.org/10.1038/srep36997
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep36997
This article is cited by
-
Yeasts of the Blastobotrys genus are promising platform for lipid-based fuels and oleochemicals production
Applied Microbiology and Biotechnology (2021)
-
Improving the functionality of surface-engineered yeast cells by altering the cell wall morphology of the host strain
Applied Microbiology and Biotechnology (2021)
-
Transcriptional control of gene expression in Pichia pastoris by manipulation of terminators
Applied Microbiology and Biotechnology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.