Abstract
Nuclear receptor PPARγ has been proven to affect metabolism in multiple tissues, and has received considerable attention for its involvement in colon cancer and inflammatory disease. However, its role in intestinal metabolism has been largely ignored. To investigate this potential aspect of PPARγ function, we submitted intestinal epithelium-specific PPARγ knockout mice (iePPARγKO) to a two-week period of 25% caloric restriction (CR), following which iePPARγKO mice retained more fat than their wild type littermates. In attempting to explain this discrepancy, we analysed the liver, skeletal muscle, intestinal lipid trafficking, and the microbiome, none of which appeared to contribute to the adiposity phenotype. Interestingly, under conditions of CR, iePPARγKO mice failed to activate their sympathetic nervous system (SNS) and increase CR-specific locomotor activity. These KO mice also manifested a defective control of their body temperature, which was overly reduced. Furthermore, the white adipose tissue of iePPARγKO CR mice showed lower levels of both hormone-sensitive lipase, and its phosphorylated form. This would result from impaired SNS signalling and possibly cause reduced lipolysis. We conclude that intestinal epithelium PPARγ plays an essential role in increasing SNS activity under CR conditions, thereby contributing to energy mobilization during metabolically stressful episodes.
Similar content being viewed by others
Introduction
Caloric restriction (CR) increases life span and improves the health of diverse species under laboratory conditions1,2. Lifelong CR can extend lifespans by up to 50% in rodents, with lesser effects should CR commence later in life3. CR both delays and slows-down the progression of age-related diseases, including neurodegenerative, autoimmune and cardiovascular disease, type 2 diabetes, and cancer4. However, the molecular mechanisms that underlie these beneficial effects remain unknown.
Peroxisome proliferator activated receptor gamma (PPARγ) is a member of the PPAR nuclear receptor subfamily that is primarily known for its insulin sensitizing properties and role as a master regulator of adipogenesis5. PPARγ also modulates a wide range of processes including cellular proliferation, differentiation, glucose and lipid metabolism, and inflammation5,6,7,8,9. PPARγ is highly expressed in adipocytes and in the gastrointestinal (GI) tract, and is expressed at lower levels in the pancreas, liver, and kidney. In the GI tract, PPARγ is present at a relatively high level in the duodenum, and at lower levels throughout the small intestine, reaching its peak expression in the proximal colon, before decreasing again in the distal colon10,11,12. PPARγ expression and activity are induced in the gut by multiple nutrient species, most notably fatty acids and their metabolites, but also by glutamine, curcumin, capsaicin, ginsenosides, vitamin E, and selenium, all of which have anti-inflammatory properties13. Importantly, bacterial metabolites and bacterial by-products such as butyrate14,15, H2O216, and LPS17, also activate PPARγ.
To date, the role of PPARγ in the gastrointestinal tract has, in the main, been restricted to the study of inflammation and cancer, which has led to the identification of PPARγ as a promising therapeutic target in colon cancer. PPARγ inhibits the development of colorectal cancer by decreasing cellular proliferation18,19,20, increasing cell differentiation18,21, inducing apoptosis18,19,22,23,24, and by inhibiting angiogenesis25. PPARγ agonists also mitigate the symptoms of inflammatory bowel disease, reduce inflammation, and are effective in multiple models of ulcerative colitis26,27,28,29,30,31,32,33,34,35, as well as in Crohn’s disease36. Moreover, PPARγ is responsible for the selective killing of bacteria associated with IBD37. Thus, there is a dynamic balance that involves reciprocal interactions between PPARγ and gut microbiota, with PPARγ both activated by bacteria, and capable of regulating the composition of that intestinal microbiota.
The essential contribution of PPARγ to metabolic regulation in multiple organs suggests potential roles in the intestine other than those already described in inflammation and cancer. We therefore challenged intestinal epithelium-specific PPARγ knockout (iePPARγKO) mice with several metabolic stimuli. Here, for the first time, we provide evidence for a connection between CR and intestinal PPARγ. Under conditions of CR, intestinal epithelial PPARγ increases sympathetic nervous system (SNS) activity and promotes the use of stored fat.
Results
Intestinal PPARγ regulates body adiposity in mice subjected to CR
An intestinal epithelium-specific PPARγ knockout mouse was generated by crossing floxed Pparγ (PPARγfl/fl)38 mice with mice expressing the Cre recombinase transgene under the control of the villin promoter (VillinCre+)39. The offspring, PPARγVillinCre+ mice, carry a targeted disruption of Pparγ in their intestinal epithelium; these animals, denoted iePPARγKO mice, were used in parallel with littermate controls (PPARγVillinCre-), or wild-type (WT) mice with the same genetic background. Gene deletion was specific to the gastrointestinal tract (see Supplementary Fig. S1), with successful disruption of Pparγ resulting in a loss of PPARγ protein expression (Supplementary Fig. S1). A very low level of Pparγ deletion was also apparent in the kidney (Supplementary Fig. S1), agreeing with the expression of villin in the epithelial cells of proximal tubules39,40,41. Despite this, we could not detect decreased Pparγ mRNA levels in total RNA samples derived from the kidney (Supplementary Fig. S1). No obvious phenotypes were evident in the iePPARγKO mice during ad libitum feeding, i.e. there were neither differences in body weight nor the size of internal organs when comparing iePPARγKO and WT mice.
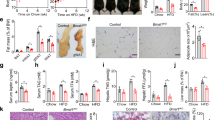
Mice were then exposed to CR, which entailed a 25% reduction in daily food intake for two weeks. As a result of CR, both the iePPARγKO and WT mice lost close to 20% of their body weight (Fig. 1a). Despite having a similar body weight after CR, the remaining body fat content of iePPARγKO CR mice was significantly higher than that of WT CR mice (Fig. 1b). This finding was confirmed by an increased percentage of lean mass identified in WT animals compared to iePPARγKO mice (Fig. 1c), that was not seen in animals fed ad libitum. EchoMRI results were confirmed by the weights of white adipose tissue (WAT) pads. Epididymal and subcutaneous dorsal WAT pads were significantly heavier in iePPARγKO CR mice than in WT CR mice (Fig. 1d, expressed as relative (% body weight), and absolute values (gram), Supplementary Fig. S2). Histological examination of epididymal adipocytes showed generally smaller adipocytes in CR versus ad libitum mice (Fig. 1e–h). Furthermore, adipocytes from iePPARγKO CR mice were bigger than those from WT CR mice, suggesting reduced lipid release during CR (Fig. 1g,h). However, the adipocytes of iePPARγKO CR mice were smaller than their counterparts fed ad libitum, indicating that CR-triggered lipid release still took place in iePPARγKO CR adipocytes, but to a significantly lesser degree than in WT CR adipocytes. This difference in adiposity was reflected by higher plasma leptin levels (Fig. 1i) in CR iePPARγKO mice vs. CR WT mice. We concluded that iePPARγKO mice retain more body fat than WT mice during CR-stimulated energy mobilization.
iePPARγKO mice lose less fat than WT mice when under caloric restriction (CR), or fed a no-sucrose diet (NSD).
WT and iePPARγKO mice were submitted to CR and their body mass measured before and after the challenge (a). Percentage body fat (b) and lean mass (c) were assayed using EchoMRI, with the pad weights of white adipose tissue (WAT) recorded (n = 10–14 mice) (d). Adipocyte size was determined by histological analyses for mice fed ad libitum (e) and after CR (g). The average cell surface area of 50 cells per histologic section (n = 4–5 sections) was determined for mice fed ad libitum (f) and after CR (h). The plasma concentration of leptin was measured for mice fed ad libitum and after CR (i); n = 11–14 mice. Body weight (j) and the percentage of body fat (k) were measured in mice fed ad libitum, in mice after 2 weeks of CR, and in mice after 2 weeks of resumption of normal feeding (n = 8–10 mice). Histological sections from the WAT of mice on a no-sucrose diet (NSD) were analysed (l), and the average adipocyte surface area measured (m). WAT pad weights were measured in mice on a NSD, and a high fat diet (HFD), and are represented as a percentage of body weight (n = 6–16 mice) (n). PPARγ protein levels were assayed by western blot (o), using tissue derived from intestinal epithelium scrapings of ad libitum and CR mice. The signals were quantified, with statistical significance verified using the Student’s t-test; data are presented as means with STD, p = 0.3. All data besides panel o are shown as mean values ± SEM error bar. For panels (f,h,m,j,k,o), the Student’s t test was performed, p < 0.05. For the remainder of the graphs, one-way ANOVA with a Bonferroni post-hoc test was applied. The following symbols #,##,*,***, correspond to statistically significant differences for the following data sets: WT vs. WT CR; KO vs. KO CR; WT CR vs. KO CR; and WT NSD vs. KO NSD, respectively.
As a control, we verified that there was no unscheduled deletion of Pparγ in epididymal, subcutaneous abdominal, as well as subcutaneous dorsal adipose tissues (Supplementary Fig. S2). Theoretically, gene deletion could occur should the villin promoter be induced under CR stress.
Next we assayed whether the difference in adipose tissue mass between iePPARγKO and WT mice was preserved after resumption of a normal dietary regimen for two additional weeks. During this period, the animals regained weight, to levels comparable to those recorded prior to CR (Fig. 1j), with percent body fat also comparable to pre CR levels in both genotypes (Fig. 1k). Hence, the difference in body fat content in CR conditions was reversible for both iePPARγKO and WT mice.
Expression of iePPARγKO adiposity phenotype under diverse, prolonged, metabolic stresses
We then tested whether the adiposity phenotype of iePPARγKO mice could be triggered by other situations involving metabolic stress. Mice were given an ad libitum no-sucrose diet (NSD; sucrose free diet), or a high-fat diet (HFD) for 6 months. The NSD resulted in an increased mass of subcutaneous and epididymal fat, with enhanced adipocyte size in iePPARγKO mice compared to WT mice (Fig. 1l–n, Supplementary Fig. S2). However, total body mass and food intake were similar in both iePPARγKO and WT mice (Supplementary Fig. S2). Both genotypes increased their fat mass to a comparable degree on the HFD (Fig. 1n, Supplementary Fig. S2). Consequently, we concluded that the iePPARγKO adiposity phenotype only develops in situations of reduced energy availability.
Metabolic gene expression was unaltered in the WAT of iePPARγKO CR mice
Next we compared changes in gene expression in the epididymal WAT of iePPARγKO CR mice vs. WT CR mice. As CR stimulates lipolysis, and the release of energy stored in adipocytes, we measured the expression of genes associated with lipid metabolism. Acetyl-CoA carboxylase (Acc) expression increased under CR compared to ad libitum conditions. However, there were no significant differences in the expression of genes related to metabolism, the uncoupling of oxidative phosphorylation, or autophagy, when comparing WAT tissue derived from iePPARγKO CR vs. WT CR mice (Supplementary Fig. S3).
Liver and muscle are not significantly affected by CR in iePPARγKO mice
We then analysed two other metabolic organs, the liver and skeletal muscle, for any differences between iePPARγKO vs. WT mice under conditions of CR. We found no differences in terms of either liver weight or glycogen content (Supplementary Fig. S3). The expression of major genes associated with lipid metabolism, lipid transport, glycogen synthesis (Gys2), and gluconeogenesis were all comparable except for Pparα and fructose-bisphosphatase 1 (Fbp1), which were upregulated in iePPARγKO CR vs. WT CR mice (Supplementary Fig. S3).
During prolonged fasting, muscles are used as an energy supply, leading to sarcopenia. We analysed muscle mass and found that after the 2-week CR diet, the weight of the soleus, gastrocnemius, and tibialis anterior muscles did not differ between iePPARγKO and WT mice (Supplementary Fig. S3). None of the tested genes involved in muscle fibres, autophagy, and metabolism, showed differential expression in iePPARγKO CR vs. WT CR mice in the soleus and tibialis anterior muscles (Supplementary Fig. S3). Thus, the liver and muscles are not significantly affected in CR iePPARγKO mice. We concluded that CR selectively affects adipose tissue in iePPARγKO mice.
The iePPARγKO adiposity phenotype is neither the result of altered intestinal lipid uptake nor lipid transport
Since the phenotype observed in iePPARγKO mice is caused by the deletion of Pparγ in the intestinal epithelium, we then investigated CR-induced changes in the gut. We observed that PPARγ protein levels were similar between ad libitum and CR conditions (Fig. 1o). We then assayed PPARγ transcriptional activity by measuring the expression of several PPARγ target genes (Pparα, Cd36, Fiaf, Scd1). While there were no differences between WT and iePPARγKO mice under ad libitum feeding conditions, there was a higher expression under CR conditions in WT animals compared to iePPARγKO (Fig. 2a). This result suggests that in the intestine, PPARγ is activated by CR. We also found that the expression of the intestinal hormone genes Cck, Gip, Xenin, Tac1, and Glp1, were up-regulated in iePPARγKO CR vs. WT CR mice. Consistently, plasma GLP-1 was significantly increased, while there was only a trend for increased PYY and GIP in the iePPARγKO CR compared to WT CR mice (Fig. 2b). In contrast to hormones, the expression of several genes connected with lipid metabolism were upregulated in WT mice in response to CR, but were unaffected in the iePPARγKO epithelium (Acsl3, Acot4, and Vldlr; Fig. 2a), again demonstrating PPARγ activation under conditions of CR. This suggested a possible differential intestinal lipid uptake in iePPARγKO CR vs. WT CR mice. However, a lipid oral load test revealed no differences in the plasma lipid concentration between iePPARγKO CR and WT CR mice at any time point after oil gavage (Fig. 2c). Furthermore, a direct bomb bomb calorimetry measurement, which assays leftover energy in mouse faeces, showed no differences between the two groups of mice under CR conditions (Fig. 2d). In addition, the quantity of faecal matter generated by iePPARγKO CR and WT CR mice was comparable (Supplementary Fig. S4).
Intestinal lipid uptake and microbiota do not contribute to the iePPARγKO caloric restriction (CR) adipose tissue phenotype.
The relative mRNA expression levels of metabolism-associated genes and intestinal hormones were assayed by RT-qPCR in the intestinal epithelium of WT and iePPARγKO mice fed ad libitum or under CR (n = 10–12 mice) (a). Plasma concentrations of GIP, PYY and GLP-1 were measured for WT and iePPARγKO mice fed ad libitum and after CR (b); n = 7–9. WT CR mice and iePPARγKO CR mice were gavaged with oil, and their blood triglyceride (TG) concentrations measured at the indicated time points (n = 8 mice) (c). The energy content of faeces from WT CR and iePPARγKO CR mice was measured using direct calorimetry (n = 9 mice) (d). The relative mRNA expression levels of inflammatory factors and antibacterial and antiviral peptides were quantified in the intestinal epithelium of WT and iePPARγKO CR mice (n = 10–12 mice) (e). Murine faecal microbiota composition and metabolites were analysed by sequencing (f) and by NMR (g), and the two data sets jointly analysed (h). The statistical difference in plasma TG and faecal energy load was assessed by the Student’s t-test. Gene expression data were analysed using one-way ANOVA fallowed by Bonferroni post-hoc test. Symbols #, # #, and *, correspond to statistically significant differences between WT and WT CR data, KO and KO CR data, and WT CR and KO CR data, respectively. Error bars depict the standard error.
To further investigate whether nutrient uptake was altered in iePPARγKO CR mice, we assayed the levels of various plasma metabolites (see Supplementary Table S1). Triglycerides, cholesterol, and glucose levels differed between the ad libitum fed, CR, and NSD groups. However, these levels were comparable in iePPARγKO and WT mice under all experimental conditions (ad libitum, CR, and NSD). Similarly, we found no genotype-specific differences in plasma lipoprotein triglyceride content, suggesting that there is no difference between WT and iePPARγKO mice in terms of either hepatic lipoprotein loading or lipid distribution to peripheral organs. Furthermore, levels of plasma lipase activity were unaltered in iePPARγKO mice feeding ad libitum vs. CR conditions (Supplementary Fig. S4), suggesting no impact on lipid uptake from plasma by the peripheral organs. We concluded that there is no change in either lipid uptake in the GI tract, or its redistribution to peripheral organs in iePPARγKO mice, and that the iePPARγKO CR adiposity phenotype does not result from the increased uptake of dietary fat.
The microbiome does not impact the iePPARγKO CR adiposity phenotype
As the expression of inflammatory and antibacterial genes are regulated in the GI tract in response to CR (K. Duszka, in preparation), we next tested whether their expression could also be regulated in the intestines of iePPARγKO mice under CR conditions. We found no difference in the expression of inflammatory factor genes for WT vs. iePPARγKO mice. However, compared to WT mice, the iePPARγKO mouse intestine showed a much weaker down-regulation of antibacterial and antiviral peptide gene expression (Reg3β, Reg3γ, IRF7, NOS2, Oas1a) in response to CR (Fig. 2e). Since antibacterial and antiviral peptides influence the composition of the gut microbiota, and intestinal bacteria modify body fat content42,43,44,45,46, we then performed molecular profiling of the faecal microbiota, which revealed significant differences in the bacterial composition of WT vs. iePPARγKO mice fed ad libitum (Fig. 2f, Supplementary Fig. S4, and Supplementary Table S2). Interestingly, CR triggered a strong shift in the faecal microbiota resulting in a similar final microbiota composition in both WT and iePPARγKO mice. These data indicated that the effects of CR on microbiota were stronger than those caused by PPARγ deletion (Fig. 2f). We predicted that these changes would be reflected in faecal metabolites, which we then analysed using nuclear magnetic resonance (NMR). Confirming the sequencing results, there were differences in the faecal metabolite profiles of WT vs. iePPARγKO mice fed ad libitum (e.g. lower levels of lactate and succinate in iePPARγKO mice; data not shown). These differences disappeared under conditions of CR, which triggered a large shift in the metabolite profiles in both strains (Fig. 2g).
Integrating the sequencing and NMR datasets using canonical correlation confirmed that CR had a much stronger effect than intestinal PPARγ deletion on gut microbiota (Fig. 2h). In contrast to the significant differences in microbiota composition between WT and iePPARγKO mice fed ad libitum, there were no significant differences in either bacterial composition or faecal metabolic profiles in WT vs. iePPARγKO CR mice, suggesting that the gut microbiota was most likely not an important determinant of the iePPARγKO CR adiposity phenotype.
iePPARγKO mice fail to adjust their body temperature and locomotor activity appropriately during CR
Next we tested whether the iePPARγKO CR adiposity phenotype was caused by defective regulation of whole body metabolism by measuring metabolic parameters and recording physical activity using indirect calorimetry. Under CR, mice of both genotypes decreased their respiratory exchange ratio values (Supplementary Fig. S4), reflecting the switch from carbohydrate to lipid as a main energy source. WT mice responded to CR by decreasing VCO2, although changes to VO2 levels were not statistically significant (Supplementary Fig. S4). We detected slight differences in CO2 production between iePPARγKO and WT mice only in ad libitum conditions, but there were no data to suggest differences in either energy usage or its dissipation between CR WT and CR iePPARγKO mice. Animals of both genotype decreased their heat production upon CR (Supplementary Fig. S4), confirming previous findings for body temperature adjustments to low energy availability47. Even though we did not detect differences in heat production between WT and iePPARγKO mice using indirect calorimetry, telemetry revealed decreased body temperatures in iePPARγKO CR compared to WT CR mice (Fig. 3a). Importantly, we neither observed differences in brown adipose tissue (BAT) weight (Fig. 3b), nor Ucp1 expression (Fig. 3c) when comparing iePPARγKO CR and WT CR mice. Furthermore, Pparγ was normally expressed in the BAT of iePPARγKO CR mice, consistent with an absence of Pparγ gene deletion (Fig. 3c, Supplementary Fig. S4).
iePPARγKO mice subject to caloric restriction (CR) show altered body temperature, locomotor activity, sympathetic nervous system (SNS) signalling, and WAT lipase expression.
Body temperature (n = 7 mice) was recorded before and after CR (a). BAT was dissected from ad libitum and CR mice with weight (b) and gene expression profiles (c) subsequently measured. Mice locomotor activity was recorded for ad libitum fed mice, and after 14 days of CR (n = 12 mice per group) (d). During the daily food portion delivery, the time from when the pellets were placed in the cage to the initiation of feeding was measured each day for the 2-week CR protocol (n = 8–13 mice) (e). The frequency of sympathetic nerve firing in WT and iePPARγKO mice fed ad libitum or subject to CR was measured (n = 6–7 mice) (f,g). Phospho-HSL, HSL, and β-tubulin protein levels in epididymal WAT were assayed by western blot for ad libitum (h) and CR (i) mice. Signals were quantified, with the data submitted to the Student’s t-test and presented as means with STD (j,k, respectively). Phospho-HSL activity was determined by normalisation of phospho-HSL protein expression relative to HSL protein expression. *p < 0.05. One-way ANOVA followed by the Bonferroni post-hoc test was used to compare the experimental groups from panels (a–f). Symbols #, # #, * correspond to statistically significant differences between the WT and WT CR datasets, KO and KO CR datasets, and WT CR and KO CR datasets, respectively. Data are presented as means ± SEM.
Mice showed increased locomotor activity at night compared to day-time (Fig. 3d). Under conditions of CR, WT mice increased their activity with lesser day/night differences compared to mice feeding ad libitum. Movement counts of iePPARγKO CR mice revealed that their mobility was significantly lower than that of WT CR mice and was comparable to that of animals fed ad libitum. To evaluate whether this difference in physical activity reflected differences in hunger perception, or a decreased motivation to eat, we measured how fast CR mice reached for food after daily chow portion delivery. As the duration of CR increased, the time taken for all mice to initiate food intake decreased (Fig. 3e), with no difference between iePPARγKO CR and WT CR mice. Furthermore, the expression of hunger-related genes in mouse hypothalami under CR were similar in iePPARγKO and WT mice (Supplementary Fig. S4). Thus, the perception of hunger and the drive to feed do not appear to underlie the differences in physical activity between iePPARγKO CR and WT mice.
We subsequently verified that there was no hypothalamic Pparγ deletion in iePPARγKO mice under CR, given that stress could promote ectopic expression of the villin promoter, resulting in a central nervous system-associated phenotype. We confirmed no difference in Pparγ expression compared to WT mice (Supplementary Fig. S4), with expression unaffected in the hypothalamus of iePPARγKO mice, and a reproducible absence of Pparγ deletion in the hypothalamus (Supplementary Fig. S4).
iePPARγKO mice show dysfunctional sympathetic nervous system (SNS) stimulation during CR
In seeking a connection between lipolysis stimulation, physical activity, and body temperature, we then recorded autonomic nervous system activity to evaluate the main body function control centre. Interestingly, CR increased SNS firing in WT mice but not in iePPARγKO mice (Fig. 3f,g). SNS sympathetic activity remained at comparable levels in iePPARγKO mice irrespective of CR or ad libitum feeding conditions. Furthermore, parasympathetic signalling (PSNS) was upregulated in iePPARγKO CR compared to WT CR mice (Supplementary Fig. S4), which is in line with their less active behaviour.
As the SNS stimulates β-adrenergic signalling in adipocytes, which promotes lipolysis48, we then determined levels of hormone-sensitive lipase (HSL) and activated HSL (Phospho-HSL) in epididymal WAT. Although we observed no differences in Hsl mRNA expression levels between WT CR and iePPARγKO CR mice (Supplementary Fig. S3), there were decreased levels of HSL and phosphorylated HSL (p-HSL) in the WAT of iePPARγKO CR vs. WT CR (Fig. 3i,k). These differences were not detected in mice fed ad libitum (Fig. 3h,j). These results imply that iePPARγKO mice fail to adjust their autonomic nervous system activity in response to CR, which manifests as deficient lipolysis in their WAT.
Discussion
Investigating the role of PPARγ in the intestine, we observed that iePPARγKO mice retained more fat than WT mice when subject to a two-week period of CR. Since iePPARγKO and WT mice both lost a substantial portion of their body fat during this period, we propose that the higher WAT mass in iePPARγKO mice after CR results from diminished fat loss, rather than from increased fat accumulation. Supporting this hypothesis, our data showed that iePPARγKO mice manifested no signs of either aberrant lipid uptake or delivery to peripheral tissues, but instead exhibited decreased levels of a key lipolytic enzyme, HSL, and its active form p-HSL, in WAT. From these data we deduced that the iePPARγKO mouse adiposity phenotype results from an altered release of lipids stored in WAT during CR-stimulated energy mobilization. Importantly, the SNS is a major regulator of lipolysis via β-adrenergic receptor activation49. Our observations suggest that lipolysis activation is impaired in iePPARγKO mice due to a failure in the SNS response to CR (Fig. 4). Interestingly, the adiposity phenotype is similar for iePPARγKO mice subject to either CR or a NSD, suggesting that some of the benefits of CR might be replicated when the nutritional composition of the diet is changed.
Model of adiposity regulation by intestinal PPARγ during caloric restriction.
Transcriptional activity of PPARγ in enterocytes is changed during caloric restriction. This results in decreased expression of incretins and lower plasma level of GLP1. Incretins exported to the bloodstream signal to central nerve system (CNS) and affect the transmission of information to the autonomic nerve system (ANS). In response to shortage of nutrients sympathetic nerve system (SNS) is activated. SNS stimulates β-adrenergic receptors in adipocytes resulting in lipase phosphorylation and lipolysis activation.
Based on the plasma lipoprotein lipid content, liver weight, and hepatic gene expression data, we conclude that liver lipid metabolism is not affected in iePPARγKO mice and is unlikely to contribute to the adiposity phenotype. Liver glycogen release upon fasting is stimulated by the SNS50. However, glycogen stores are activated and depleted in advance of fat deposits, which most likely explains why we saw no differences in these stores when comparing WT CR and iePPARγKO CR mice (i.e. after 2 weeks of CR).
Other than the intestinal epithelium, the villin gene is also selectively expressed in the kidney, in the epithelium of proximal tubules41. There are no reports showing villin expression in the juxtaglomerular cells that produce and store renin. Thus, despite our observation of a very limited deletion of Pparγ in the kidney, it is extremely unlikely that a renin-angiotensin-aldosterone system failure would contribute to the iePPARγKO adiposity phenotype.
Applying the 2-week CR protocol, we did not observe differences in intestinal epithelium PPARγ protein levels. However, the stimulation of PPARγ target genes indicated an increased transcriptional activity of the receptor during CR, most likely induced by an increase in its endogenous ligands. Under CR, WT mice showed elevated locomotor activity, which is interpreted as natural food-seeking behaviour. Surprisingly, iePPARγKO mice did not show this basic behavioural response to CR. We investigated several possible reasons for this phenotype. First, we tested muscular integrity, since iePPARγKO mice might utilize muscle protein as a source of energy under conditions of CR, which could lead to sarcopenia and diminished locomotive function. However, this was ruled out based on skeletal muscle weight and gene expression data. Second, we tested hunger perception and the motivation to eat. We could find no differences in hypothalamic gene expression, or the time taken to approach food during daily feedings in iePPARγKO CR vs. WT CR mice, demonstrating that neither hunger perception, nor the drive to feed were altered in iePPARγKO mice under CR. Third, we tested activation of the SNS to mobilize energy stores under stress situations. A primitive SNS-related fight-or-flight response is associated with increased mobility to locate food and ensure survival. Critically, the iePPARγKO mice showed a disturbed SNS activation in response to hunger and manifested higher parasympathetic activity compared to WT mice. In addition, their body temperature was significantly lower than that of WT mice. Thus, a decreased perception of energy deprivation might be linked to reduced fat store depletion in iePPARγKO mice. The lack of SNS stimulation in iePPARγKO CR mice and their higher PSNS activity most likely explains why they are less active than WT CR mice.
Lipolysis in adipose tissue is activated by a cascade of phosphorylation events derived from a SNS-stimulated activation of β-adrenergic receptors, leading to lipase activation48. Decreased levels of HSL, as well as p-HSL in the WAT of iePPARγKO vs. WT mice under conditions of CR, correlated with reduced SNS activity and the iePPARγKO adiposity phenotype.
Looking for possible traits to link the intestine and autonomic nervous system, we considered extrinsic bacteria-derived, as well as intrinsic, host-specific causes. As bacterial-derived acetate affects the activation of the parasympathetic nervous system51, we determined the metabolite composition of mice faeces. However, we could not observe any differences between metabolite generation in WT vs. iePPARγKO mice, including acetates. As a possible host-specific cause of the adiposity phenotype, we then considered intestinal hormones that can influence SNS activity and signalling to the brain52,53. Again, we could not demonstrate a direct link between intestinal PPARγ and the SNS. However, the observed PPARγ-related modulation of plasma concentrations of intestinal hormones suggests that PPARγ could influence the SNS via intestinal hormone production. This will be the focus of a follow-up investigation.
Previously, Festuccia et al., proposed a link between PPARγ agonist administration and a tissue-specific reduction in BAT sympathetic activity54. Here we show, for the first time, that intestinal PPARγ signals the brain, and influences SNS activity. Our results suggest that PPARγ-triggered SNS activation may be WAT specific, as it neither affected BAT, skeletal muscle, nor liver. However, we observed at least two other phenotypes, a decreased body temperature, and reduced physical activity, suggesting the involvement of the central nervous system. Consequently, we anticipate that the iePPARγKO mouse will emerge as a valuable model for research on the gut-brain axis.
Until now, there has been no clear link between PPARγ and CR in any tissue, although a connection has been previously suggested55,56,57,58,59. We now provide, for the first time, evidence to suggest that intestinal PPARγ is provoked by CR to activate the SNS, triggering a response to energy deprivation. The pathway through which intestinal PPARγ regulates sympathetic signalling remains obscure and is presently under investigation.
Materials and Methods
Animal care and experimental procedures
All experiments were performed in accordance with institutional guidelines and were approved by the Vaud Cantonal Authority in Switzerland, and by the Institutional Animal Care and Use Committee in Singapore. PPARγfloxVillinCre+ (iePPARγKO) and PPARγfloxVillinCre- (WT) C57/Bl6/SV129 male mice were kept under a 12 h light/12 h dark cycle. Mice were fed a standard laboratory diet (Diet 3436; Provimi Kliba AG), and housed at a maximum of 5 animals per cage at 23 °C.
The experimental WT and iePPARγKO mice were divided into two groups, the ad libitum and CR groups. Mice were housed one per cage to avoid fighting due to food restriction and to exclude the possibility of dominant males taking the majority of the available food. The CR animals were subject to a two-week CR that involved a reduction to 75% of their normal daily food intake. Food was provided daily 1–2 h before the dark phase. All animals had free access to water. Mouse body composition was measured under anaesthesia before and after the CR period using an EchoMRI whole-body composition analyzer (EchoMRI, Huston, TX, USA). Similarly, metabolic parameters (VO2, VCO2, heat, locomotor activity) were monitored before and after 2 weeks of CR using the Comprehensive Lab Animal Monitoring System (CLAMS, Columbus Instruments, Columbus, OH, USA). Body temperature was assayed using the DSI PhysioTel telemetry system (Data Sciences International, St. Paul, MN, USA). Mice were euthanized using CO2, with blood drawn by cardiac puncture. Blood was mixed with 2% aprotinin-EDTA (Sigma Aldrich, St. Louis, MO, USA), centrifuged for 10 min at 8,000 ×g, and stored at −20 °C. White adipose tissue fat pads, liver, and muscle weights were recorded. Tissues were snap frozen and stored at −80 °C until use. The lipid oral load test was performed on CR mice by gavaging 200 μl of oil. Blood was subsequently drawn from the tail at selected time points for analyses.
For the re-feeding experiments, mice were subject to CR for 2 weeks, followed by 2 weeks of free access to food. The weight and body composition of each mouse was measured before, and after 2 weeks of CR, and again after 2 weeks of re-feeding following CR.
For the diet/feeding experiments, 5-week-old mice were switched from standard chow to a sucrose-free diet (no-sucrose diet, NSD; #D12450K), or to a high fat diet from which 60% of energy requirements are derived from fat (#D12492). Both feeds were from Research Diets, Inc. (New Brunswick, NY, USA). Body-fat content was measured by EchoMRI, 16 weeks after the diet regimens began, with mice sacrificed one week later.
Western blotting
Upon harvest, adipose and duodenal tissues were immediately snap-frozen in liquid nitrogen. Tissues were stored at −80 °C for subsequent lysis and western blot analyses with the indicated antibodies. Adipose tissues were lysed in HNTG buffer comprising 50 mM Hepes, pH 7.5, 150 mM NaCl, 10% glycerol, 1% trition-X-100, supplemented with the cOmplete Mini Protease (Roche #4693159001) and PhosSTOP phosphatase inhibitor cocktails (Roche #4906837001). Tissues were homogenized using a TissueLyser (Qiagen). Lysates were then centrifuged at 18,000 × g, at 4 °C, for 20 min. The fat cake was removed prior to collecting the supernatant. Duodenal tissues were lysed in cold RIPA buffer (Thermofisher Scientific # 89900), again supplemented with the cOmplete™ Mini Protease (Roche #4693159001) and PhosSTOP phosphatase inhibitor cocktails (Roche #4906837001). Tissues were homogenized using a TissueLyser (Qiagen), then subject to centrifugation at 14,000 × g, at 4 °C, for 15 min, before collecting the supernatant. Comparable concentrations of total protein were loaded onto acrylamide/bis-acrylamide gels and, after their electrophoretic resolution, transferred to polyvinylidene fluoride membranes for detection with the indicated antibodies (anti Phospho-HSL (Ser660) antibody, Cell Signaling #4126; anti HSL antibody, Cell Signaling, #4107; anti beta-tubulin antibody, AbCam #ab6046; anti PPARγ antibody (H-100), Santa Cruz #sc-7196; and the anti U2AF65 antibody (H-300), Santa Cruz #sc-48804). Briefly, membranes were incubated with primary antibodies (used at a 1: 1,000 dilution for antibodies purchased from Cell Signaling, or a 1:200 dilution for reagents purchased from Santa Cruz) in 5% BSA/TBST, overnight. Horse radish peroxidase-conjugated secondary antibodies (Goat anti-rabbit IgG-HRP, Santa Cruz #sc-2054) (1: 5,000 dilution) were then added for 1 h. Membrane stripping was with Restore PLUS Western Blot Stripping Buffer (Thermofisher Scientific #46430). Western blots were developed using the Luminata Crescendo Western HRP Substrate (Millipore, #WBLUR0500), with signal detection by X-ray film.
Plasma and marker analyses
Plasma glucose, lipid, and cholesterol levels were measured using a Hitachi robot (Roche Diagnostics, Basel, Switzerland) according to the manufacturer’s instructions. Plasma lipase activity was analysed using the LPL activity Assay Kit (Roar Biomedical, Inc., New York, NY, USA). Plasma leptin and ghrelin concentrations were estimated using Bioplex (Luminex Corporation, Austin, TX, USA). Liver glycogen content was quantified using a glycogen assay kit (Sigma) according to the manufacturer’s instructions. Faecal energy load was measured using the IKA C200 (IKA Werke, Staufen, Germany) for direct bomb calorimetry. For plasma lipoprotein analyses, the lipoproteins in 2 μL of plasma were separated by size-exclusion chromatography, followed by an online determination of total cholesterol and triglycerides TG as described previously60.
Autonomic nervous system activity
The firing rates of the thoracic branch of the vagal and sympathetic nerves were recorded along the carotid artery as described previously61,62. The sympathetic and vagal nerves, which lie close to the carotid artery, were dissected free of underlying tissues, to a distance of approximately 5 mm. The nerves were then covered with mineral oil to avoid dehydration and carefully placed on a pair of silver-wire recording electrodes (0.6-mm diameter). The electrodes were connected to a high-impedance probe, and the action potentials displayed and saved by computer after initial amplification through a low-noise amplifier (BIO amplifier, AD Instruments, Oxford, UK). Unipolar nerve activity was recorded continuously for 30 minutes using the LabChart 8 software (AD Instruments). Data were digitized with PowerLab 16/35 (AD Instruments). Signals were amplified 105 times and filtered using a 200/1000-Hz band pass filter. Firing rate analyses were performed using LabChart 8. All animal experimentation protocols were approved by the Vaud Cantonal Authority (authorization VD 2440), Switzerland, and by the Institutional Animal Care and Use Committee (authorization 2015/SHS/1023) in Singapore. The Supplementary Methods provide details for qPCR, the sequencing of 16 S rDNA genes, 1 H NMR metabolomics, and data analyses.
Additional Information
How to cite this article: Duszka, K. et al. Intestinal PPARγ signaling is required for sympathetic nervous system activation in response to caloric restriction. Sci. Rep. 6, 36937; doi: 10.1038/srep36937 (2016).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Masoro, E. J. Caloric restriction and aging: an update. Experimental gerontology 35, 299–305 (2000).
Weindruch, R. & Sohal, R. S. Seminars in medicine of the Beth Israel Deaconess Medical Center. Caloric intake and aging. The New England journal of medicine 337, 986–994, doi: 10.1056/NEJM199710023371407 (1997).
Abalan, F., Mayo, W., Simon, H. & Le Moal, M. Paradoxical effect of severe dietary restriction on Long-Evans rat life span. International journal for vitamin and nutrition research. Internationale Zeitschrift fur Vitamin- und Ernahrungsforschung. Journal international de vitaminologie et de nutrition 80, 386–393, doi: 10.1024/0300-9831/a000027 (2010).
Speakman, J. R. & Mitchell, S. E. Caloric restriction. Molecular aspects of medicine 32, 159–221, doi: 10.1016/j.mam.2011.07.001 (2011).
Anghel, S. I. & Wahli, W. Fat poetry: a kingdom for PPAR gamma. Cell Res 17, 486–511, doi: 10.1038/cr.2007.48 (2007).
Auwerx, J. Nuclear receptors. I. PPAR gamma in the gastrointestinal tract: gain or pain? American journal of physiology. Gastrointestinal and liver physiology 282, G581–G585, doi: 10.1152/ajpgi.00508.2001 (2002).
Dubuquoy, L. et al. PPARgamma as a new therapeutic target in inflammatory bowel diseases. Gut 55, 1341–1349, doi: 10.1136/gut.2006.093484 (2006).
Leonardini, A., Laviola, L., Perrini, S., Natalicchio, A. & Giorgino, F. Cross-Talk between PPARgamma and Insulin Signaling and Modulation of Insulin Sensitivity. PPAR research 2009, 818945, doi: 10.1155/2009/818945 (2009).
Tontonoz, P. & Spiegelman, B. M. Fat and beyond: the diverse biology of PPARgamma. Annual review of biochemistry 77, 289–312, doi: 10.1146/annurev.biochem.77.061307.091829 (2008).
Escher, P. et al. Rat PPARs: quantitative analysis in adult rat tissues and regulation in fasting and refeeding. Endocrinology 142, 4195–4202, doi: 10.1210/endo.142.10.8458 (2001).
Harmon, G. S. et al. Pharmacological correction of a defect in PPAR-gamma signaling ameliorates disease severity in Cftr-deficient mice. Nature medicine 16, 313–318, doi: 10.1038/nm.2101 (2010).
Mansen, A., Guardiola-Diaz, H., Rafter, J., Branting, C. & Gustafsson, J. A. Expression of the peroxisome proliferator-activated receptor (PPAR) in the mouse colonic mucosa. Biochemical and biophysical research communications 222, 844–851, doi: 10.1006/bbrc.1996.0832 (1996).
Marion-Letellier, R., Dechelotte, P., Iacucci, M. & Ghosh, S. Dietary modulation of peroxisome proliferator-activated receptor gamma. Gut 58, 586–593, doi: 10.1136/gut.2008.162859 (2009).
Schwab, M. et al. Involvement of different nuclear hormone receptors in butyrate-mediated inhibition of inducible NF kappa B signalling. Molecular immunology 44, 3625–3632, doi: 10.1016/j.molimm.2007.04.010 (2007).
Wachtershauser, A., Loitsch, S. M. & Stein, J. PPAR-gamma is selectively upregulated in Caco-2 cells by butyrate. Biochemical and biophysical research communications 272, 380–385, doi: 10.1006/bbrc.2000.2793 (2000).
Voltan, S. et al. Lactobacillus crispatus M247-derived H2O2 acts as a signal transducing molecule activating peroxisome proliferator activated receptor-gamma in the intestinal mucosa. Gastroenterology 135, 1216–1227, doi: 10.1053/j.gastro.2008.07.007 (2008).
Dubuquoy, L. et al. Impaired expression of peroxisome proliferator-activated receptor gamma in ulcerative colitis. Gastroenterology 124, 1265–1276 (2003).
Cerbone, A. et al. 4-Hydroxynonenal and PPARgamma ligands affect proliferation, differentiation, and apoptosis in colon cancer cells. Free radical biology & medicine 42, 1661–1670, doi: 10.1016/j.freeradbiomed.2007.02.009 (2007).
Martinasso, G. et al. Involvement of PPARs in Cell Proliferation and Apoptosis in Human Colon Cancer Specimens and in Normal and Cancer Cell Lines. PPAR research 2007, 93416, doi: 10.1155/2007/93416 (2007).
Theocharis, S., Margeli, A., Vielh, P. & Kouraklis, G. Peroxisome proliferator-activated receptor-gamma ligands as cell-cycle modulators. Cancer treatment reviews 30, 545–554, doi: 10.1016/j.ctrv.2004.04.004 (2004).
Xu, W. P., Zhang, X. & Xie, W. F. Differentiation therapy for solid tumors. Journal of digestive diseases 15, 159–165, doi: 10.1111/1751-2980.12122 (2014).
Chen, G. G. et al. Apoptosis induced by activation of peroxisome-proliferator activated receptor-gamma is associated with Bcl-2 and NF-kappaB in human colon cancer. Life sciences 70, 2631–2646 (2002).
Chen, G. G. et al. 15-hydroxy-eicosatetraenoic acid arrests growth of colorectal cancer cells via a peroxisome proliferator-activated receptor gamma-dependent pathway. International journal of cancer. Journal international du cancer 107, 837–843, doi: 10.1002/ijc.11447 (2003).
Lee, C. J. et al. Pioglitazone, a synthetic ligand for PPARgamma, induces apoptosis in RB-deficient human colorectal cancer cells. Apoptosis: an international journal on programmed cell death 11, 401–411, doi: 10.1007/s10495-006-4003-z (2006).
Sharma, C., Pradeep, A., Wong, L., Rana, A. & Rana, B. Peroxisome proliferator-activated receptor gamma activation can regulate beta-catenin levels via a proteasome-mediated and adenomatous polyposis coli-independent pathway. The Journal of biological chemistry 279, 35583–35594, doi: 10.1074/jbc.M403143200 (2004).
Lewis, J. D. et al. Rosiglitazone for active ulcerative colitis: a randomized placebo-controlled trial. Gastroenterology 134, 688–695, doi: 10.1053/j.gastro.2007.12.012 (2008).
Lewis, J. D. et al. An open-label trial of the PPAR-gamma ligand rosiglitazone for active ulcerative colitis. The American journal of gastroenterology 96, 3323–3328, doi: 10.1111/j.1572-0241.2001.05333.x (2001).
Liang, H. L. & Ouyang, Q. A clinical trial of combined use of rosiglitazone and 5-aminosalicylate for ulcerative colitis. World journal of gastroenterology: WJG 14, 114–119 (2008).
Bassaganya-Riera, J. & Hontecillas, R. CLA and n-3 PUFA differentially modulate clinical activity and colonic PPAR-responsive gene expression in a pig model of experimental IBD. Clinical nutrition 25, 454–465, doi: 10.1016/j.clnu.2005.12.008 (2006).
Hontecillas, R. et al. Nutritional regulation of porcine bacterial-induced colitis by conjugated linoleic acid. The Journal of nutrition 132, 2019–2027 (2002).
Sanchez-Hidalgo, M., Martin, A. R., Villegas, I. & de la Lastra, C. A. Rosiglitazone, a PPARgamma ligand, modulates signal transduction pathways during the development of acute TNBS-induced colitis in rats. European journal of pharmacology 562, 247–258, doi: 10.1016/j.ejphar.2007.01.047 (2007).
Sato, N. et al. Peroxisome proliferator-activated receptor gamma mediates protection against cyclooxygenase-2-induced gut dysfunction in a rodent model of mesenteric ischemia/reperfusion. Shock 24, 462–469 (2005).
Saubermann, L. J. et al. Peroxisome proliferator-activated receptor gamma agonist ligands stimulate a Th2 cytokine response and prevent acute colitis. Inflammatory bowel diseases 8, 330–339 (2002).
Su, C. G. et al. A novel therapy for colitis utilizing PPAR-gamma ligands to inhibit the epithelial inflammatory response. The Journal of clinical investigation 104, 383–389, doi: 10.1172/JCI7145 (1999).
Rousseaux, C. et al. Intestinal antiinflammatory effect of 5-aminosalicylic acid is dependent on peroxisome proliferator-activated receptor-gamma. J Exp Med 201, 1205–1215, doi: 10.1084/jem.20041948 (2005).
Shah, Y. M., Morimura, K. & Gonzalez, F. J. Expression of peroxisome proliferator-activated receptor-gamma in macrophage suppresses experimentally induced colitis. American journal of physiology. Gastrointestinal and liver physiology 292, G657–G666, doi: 10.1152/ajpgi.00381.2006 (2007).
Peyrin-Biroulet, L. et al. Peroxisome proliferator-activated receptor gamma activation is required for maintenance of innate antimicrobial immunity in the colon. Proceedings of the National Academy of Sciences of the United States of America 107, 8772–8777, doi: 10.1073/pnas.0905745107 (2010).
Imai, T. et al. Peroxisome proliferator-activated receptor gamma is required in mature white and brown adipocytes for their survival in the mouse. Proceedings of the National Academy of Sciences of the United States of America 101, 4543–4547, doi: 10.1073/pnas.0400356101 (2004).
el Marjou, F. et al. Tissue-specific and inducible Cre-mediated recombination in the gut epithelium. Genesis 39, 186–193, doi: 10.1002/gene.20042 (2004).
Kucherlapati, M. H., Nguyen, A. A., Bronson, R. T. & Kucherlapati, R. S. Inactivation of conditional Rb by Villin-Cre leads to aggressive tumors outside the gastrointestinal tract. Cancer Res 66, 3576–3583, doi: 10.1158/0008-5472.CAN-05-2699 (2006).
Pinto, D., Robine, S., Jaisser, F., El Marjou, F. E. & Louvard, D. Regulatory sequences of the mouse villin gene that efficiently drive transgenic expression in immature and differentiated epithelial cells of small and large intestines. J Biol Chem 274, 6476–6482 (1999).
Armougom, F. & Raoult, D. Use of pyrosequencing and DNA barcodes to monitor variations in Firmicutes and Bacteroidetes communities in the gut microbiota of obese humans. BMC genomics 9, 576, doi: 10.1186/1471-2164-9-576 (2008).
Bervoets, L. et al. Differences in gut microbiota composition between obese and lean children: a cross-sectional study. Gut pathogens 5, 10, doi: 10.1186/1757-4749-5-10 (2013).
van Hylckama Vlieg, J. E., Veiga, P., Zhang, C., Derrien, M. & Zhao, L. Impact of microbial transformation of food on health - from fermented foods to fermentation in the gastro-intestinal tract. Current opinion in biotechnology 22, 211–219, doi: 10.1016/j.copbio.2010.12.004 (2011).
Zhang, C. et al. Interactions between gut microbiota, host genetics and diet relevant to development of metabolic syndromes in mice. The ISME journal 4, 232–241, doi: 10.1038/ismej.2009.112 (2010).
Aronsson, L. et al. Decreased fat storage by Lactobacillus paracasei is associated with increased levels of angiopoietin-like 4 protein (ANGPTL4). PloS one 5, doi: 10.1371/journal.pone.0013087 (2010).
Soare, A., Cangemi, R., Omodei, D., Holloszy, J. O. & Fontana, L. Long-term calorie restriction, but not endurance exercise, lowers core body temperature in humans. Aging 3, 374–379 (2011).
Carmen, G. Y. & Victor, S. M. Signalling mechanisms regulating lipolysis. Cellular signalling 18, 401–408, doi: 10.1016/j.cellsig.2005.08.009 (2006).
Collins, S., Cao, W. & Robidoux, J. Learning new tricks from old dogs: beta-adrenergic receptors teach new lessons on firing up adipose tissue metabolism. Molecular endocrinology 18, 2123–2131, doi: 10.1210/me.2004-0193 (2004).
Lautt, W. W. Autonomic neural control of liver glycogen metabolism. Medical hypotheses 5, 1287–1296 (1979).
Perry, R. J. et al. Acetate mediates a microbiome-brain-beta-cell axis to promote metabolic syndrome. Nature 534, 213–217, doi: 10.1038/nature18309 (2016).
Lambert, E. et al. Ghrelin modulates sympathetic nervous system activity and stress response in lean and overweight men. Hypertension 58, 43–50, doi: 10.1161/HYPERTENSIONAHA.111.171025 (2011).
Skibicka, K. P. The central GLP-1: implications for food and drug reward. Front Neurosci 7, 181, doi: 10.3389/fnins.2013.00181 (2013).
Festuccia, W. T. et al. Peroxisome proliferator-activated receptor-gamma-mediated positive energy balance in the rat is associated with reduced sympathetic drive to adipose tissues and thyroid status. Endocrinology 149, 2121–2130, doi: 10.1210/en.2007-1553 (2008).
Masternak, M. M. et al. Caloric restriction results in decreased expression of peroxisome proliferator-activated receptor superfamily in muscle of normal and long-lived growth hormone receptor/binding protein knockout mice. The journals of gerontology. Series A, Biological sciences and medical sciences 60, 1238–1245 (2005).
Masternak, M. M. et al. Effects of caloric restriction and growth hormone resistance on the expression level of peroxisome proliferator-activated receptors superfamily in liver of normal and long-lived growth hormone receptor/binding protein knockout mice. The journals of gerontology. Series A, Biological sciences and medical sciences 60, 1394–1398 (2005).
Masternak, M. M. & Bartke, A. PPARs in Calorie Restricted and Genetically Long-Lived Mice. PPAR research 2007, 28436, doi: 10.1155/2007/28436 (2007).
Weindruch, R., Kayo, T., Lee, C. K. & Prolla, T. A. Microarray profiling of gene expression in aging and its alteration by caloric restriction in mice. The Journal of nutrition 131, 918S–923S (2001).
Sung, B., Park, S., Yu, B. P. & Chung, H. Y. Modulation of PPAR in aging, inflammation, and calorie restriction. The journals of gerontology. Series A, Biological sciences and medical sciences 59, 997–1006 (2004).
Parini, P., Johansson, L., Broijersen, A., Angelin, B. & Rudling, M. Lipoprotein profiles in plasma and interstitial fluid analyzed with an automated gel-filtration system. Eur J Clin Invest 36, 98–104, doi: 10.1111/j.1365-2362.2006.01597.x (2006).
Magnan, C. et al. Lipid infusion lowers sympathetic nervous activity and leads to increased beta-cell responsiveness to glucose. The Journal of clinical investigation 103, 413–419, doi: 10.1172/JCI3883 (1999).
Tarussio, D. et al. Nervous glucose sensing regulates postnatal beta cell proliferation and glucose homeostasis. The Journal of clinical investigation 124, 413–424, doi: 10.1172/JCI69154 (2014).
Acknowledgements
This study was funded by the following: the Lee Kong Chian School of Medicine; Nanyang Technological University Start-Up Grant (WW); the Swiss National Science Foundation (WW, BT); the 7th EU program TORNADO (WW, AN); the European Research Council (BT); the Région Midi-Pyrénées (HG, Mathusalem program; WW, Chaire d’Excellence Pierre de Fermat); the Bonizzi-Theler-Stiftung (WW); and the Etat de Vaud (WW, BT). The authors would like to acknowledge the staff at the Metabolic Evaluation Facility (MEF), the Phenotyping Facility, and the Genomic Technologies Facility (all at the Center for Integrative Genomics, University of Lausanne). The authors are especially grateful to Frederic Preitner for helpful discussions about the experimental design, to Anabela Cristina Da Costa for performing the bomb calorimetry, Zoltan Spiró and Catherine Moret for assistance with microscopy. The authors would also like to thank the French National Infrastructure of Metabolomics and Fluxomics (MetaboHUB-ANR-11-INBS-0010) for their support.
Author information
Authors and Affiliations
Contributions
K.D. designed and performed the animal experiments, measured gene expression, plasma lipase and liver glycogen levels, and wrote the manuscript. Al.P. recorded autonomic nervous system activity. S.E.S., C.C. and H.G., performed the NMR experiments, analysed the results, and performed the statistical analyses. J.C. prepared the western blots. M.D. performed the statistical analysis of the microbiota sequences, and merged the sequencing and NMR data. E.P. assisted with mice care and sample handling. A.N. and An.P. prepared the faecal material and performed microbiota sequencing and analyses. P.P. analysed the lipoprotein TG content. L.V. derived the iePPARγKO mouse strain. B.T. contributed expert advice, and helped write the manuscript. W.W. designed the study, analysed the data, and wrote the manuscript. All authors corrected and approved the final manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Duszka, K., Picard, A., Ellero-Simatos, S. et al. Intestinal PPARγ signalling is required for sympathetic nervous system activation in response to caloric restriction. Sci Rep 6, 36937 (2016). https://doi.org/10.1038/srep36937
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep36937
This article is cited by
-
Complementary intestinal mucosa and microbiota responses to caloric restriction
Scientific Reports (2018)
-
Klebsiella oxytoca expands in cancer cachexia and acts as a gut pathobiont contributing to intestinal dysfunction
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.