Abstract
The bacterial stringent response is a key regulator of bacterial virulence, biofilm formation and antibiotic tolerance, and is a promising target for the development of new antibacterial compounds. The intracellular nucleotide (p)ppGpp acts as a messenger orchestrating the stringent response. A synthetic peptide 1018 was recently proposed to specifically disrupt biofilms by inhibiting the stringent response via direct interaction with (p)ppGpp (de la Fuente-Núñez et al. (2014) PLoS Pathogens). We have interrogated the specificity of the proposed molecular mechanism. When inhibition of Pseudomonas aeruginosa planktonic and biofilm growth is tested simultaneously in the same assay, peptides 1018 and the control peptide 8101 generated by an inversion of the amino acid sequence of 1018 are equally potent, and, importantly, do not display a preferential activity against biofilm. 1018 inhibits planktonic growth of Escherichia coli equally efficiently either when the alleged target, (p)ppGpp, is essential (MOPS media lacking amino acid L-valine), or dispensable for growth (MOPS media supplemented with L-valine). Genetic disruption of the genes relA and spoT responsible for (p)ppGpp synthesis moderately sensitizes – rather than protects – E. coli to 1018. We suggest that the antimicrobial activity of 1018 does not rely on specific recognition of the stringent response messenger (p)ppGpp.
Similar content being viewed by others
Introduction
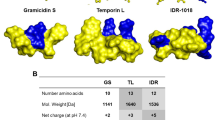
IDR-1018 (or just 1018) is a small cationic synthetic peptide (VRLIVAVRIWRR-NH2) that has been developed based on bactenecin, a peptide antibiotic isolated from bovine neutrophil granules1,2,3. This peptide has numerous biological activities that target both eukaryotic and bacterial cells (reviewed by Mansour and colleagues4). By targeting the eukaryotic host it acts as a modulator of the immune system, affecting macrophage polarization5,6 and reducing levels of lipopolysaccharide-induced cytokine production3,7,8. By targeting both Gram-negative (e.g. Pseudomonas aeruginosa and Escherichia coli) and Gram-positive (e.g. Staphylococcus aureus) bacteria 1018 acts as a potent antibacterial: it kills bacteria, disperses biofilms and inhibits bacterial swarming9,10. Potential practical applications of 1018 range from neuroprotection8 and wound healing11 to potentiation of antimalarial12, antiviral13 and antibacterial compounds14.
It has been proposed that the dispersal of biofilms by 1018 is mediated by a complex formation between the peptide and the intracellular alarmone nucleotides guanosine tetraphosphate (ppGpp) and pentaphosphate (pppGpp)9. The complex formation, in turn, triggers degradation of the nucleotides via a presently unidentified mechanism. The two alarmones, collectively known as (p)ppGpp, are key physiological regulators of the stringent response15, which, in turn, is a key regulator of bacterial virulence and antibiotic tolerance16,17,18. In this study we analyse the specificity of peptide 1018 in its effect against P. aeruginosa PAO1 planktonic and biofilm growth, as well as its interaction with ppGpp.
Results
Peptide 1018 is equally efficient against biofilm and planktonic populations, as judged by crystal violet staining assay
In the original report by de la Fuente-Núñez and colleagues, 1018 was shown to be relatively inefficient against planktonic P. aeruginosa, with a minimal inhibitory concentration (MIC) of 41 μM (64 μg/ml)9 or 19 μM (29.7 μg/ml)3,19, depending on the exact method used for the determination of the MIC. It was, however, considerably more potent against biofilms, completely inhibiting their establishment at 6.4 μM (10 μg/ml) (MBIC100, minimal biofilm inhibitory concentration 100%), as well as killing an established biofilm at the same concentration. Moreover, 1018 dispersed biofilms at a concentration as low as 0.5 μM (0.8 μg/ml). Since, in general, biofilms are much more resistant to antibacterial compounds than planktonic bacteria20, the authors have concluded that an unusual molecular mechanism of action of 1018 is responsible for the peptide specifically targeting biofilms.
An alternative explanation is that 1018 was performing better in the biofilm assay simply due to the way the assay was set up. Effects on biofilms were studied on glass in flow cell chambers under conditions of constant medium flow10. This would allow for the sorption and accumulation of the peptides on the cell surface over time, and surface-specific sorption is a well-known property of cationic peptides21. The MIC was determined using a broth microdilution method, which does not involve a continuous flow of liquid and is performed with 96-well plates or Eppendorf tubes made of laboratory plastic22. Finally, the antibacterial efficiency is strongly affected by bacterial density23, which can potentially differ in two separate assays.
In order to circumvent the above-mentioned issues we opted for a robust and widely adopted crystal violet (CV) staining assay that can measure both planktonic growth and biofilm formation in the same well of a 96-well plate24,25. First, the planktonic population is measured by removing half of the culture form the well, measuring the absorbance at 600 nm. Second, the remaining liquid is aspirated, the biofilm is stained with CV dye and the absorbance at 595 nm is quantified. Similarly to de la Fuente-Núñez and colleagues9, we used P. aeruginosa strain PAO1. We do not observe a specific anti-biofilm effect: both planktonic and biofilm growth are completely inhibited by 1018 at 6.4 μM (Fig. 1a).
Peptide 1018 (a) and its inverted version 8101 (b) inhibit P. aeruginosa PAO1 growth and biofilm formation equally efficiently. Planktonic growth at 600 nm (filled cycles) was measured from the cultures taken from the same plates that were used later for biofilm measurements. Biofilm formation (open circles) was determined using a crystal violet staining assay24 where the intensity of the violet colour corresponds to the biofilm thickness measured spectrophotometrically at 595 nm (open cycles). Highlighted area indicates peptide concentration range where it becomes lethal. Cell count measurements are expressed in colony forming units, CFU, per ml. The results are shown as mean values ± SD of two biological replicates, each estimated from three technical replicates.
Inversion of the amino acid sequence of 1018 improves its antimicrobial properties
Peptide 1018 was suggested to exert its anti-biofilm effect by means of entering the cell, directly binding to the intracellular messenger ppGpp − a highly complex and chiral molecule − and targeting it for degradation via a presently unidentified mechanism9. In the follow-up report, de la Fuente-Núñez and colleagues tested several stereoisomeric variants of 101826. Several of the compounds were equally as or more efficient than the parental compound, which is surprising given the importance of stereospecificity in biomolecular interactions in general and drug design in particular27,28. To further test the structure–activity relationship of 1018, we tested its inverted version (RRWIRVAVILRV-NH2), which was designated “8101”. The inverted peptide 8101 inhibits planktonic and biofilm formation even more efficiently then the parental compound, completely preventing bacterial growth at 3.2 μM (Fig. 1b). Similar to 1018, 8101 shows no preferential anti-biofilm activity in the CV assay. Cell count measurements expressed in colony forming units (CFU) per ml corroborate the OD600 estimates of cell density. A possible explanation for the similar functional activity of the two compounds is that inverted peptide 8101 retained the 3D structure of the parental 1018, just like in the context of larger proteins where short helical fragments, upon inversion, tend to retain the original structure29,30.
Antimicrobial activity of 1018 against E. coli does not depend on conditional essentiality of its alleged molecular target (p)ppGpp
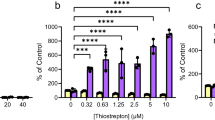
Severity of bacterial phenotype upon genetic disruption of the RSH genes mediating the (p)ppGpp production is dramatically affected by the growth medium31,32. The growth curves of wild type and ppGpp0 ΔrelAΔspoT in MOPS minimal medium containing L-valine are virtually indistinguishable; conversely, in MOPS medium lacking L-valine ppGpp0 stain is unable to grow due to valine auxotrophy32,33 (Fig. 2a).
Antimicrobial activity of 1018 against E. coli is insensitive to conditional essentiality of the peptide’s alleged molecular target, (p)ppGpp.
(a) Growth curves of wild type and ppGpp0 (ΔrelAΔspoT) BW25113 E. coli in conditions in which (p)ppGpp is essential (MOPS-based minimal medium lacking L-valine) or dispensable with no significant growth defect (MOPS-based minimal medium containing the full set of 20 amino acids). (b) Effects of increasing concentrations of 1018 on bacterial survival of wild type BW25113 E. coli in MOPS minimal medium with or without addition of L-valine and ppGpp0 strain in MOPS supplemented with L-valine. The results are shown as geometric mean values ± SD of two biological replicates, each estimated from two technical replicates. Inhibition efficiency (IC50) was calculated using 4-parameter logistic model (Hill equation) as per Sebaugh48.
We exploited this conditionality of (p)ppGpp’s essentiality for bacterial growth to directly test the proposed mechanism of action of 1018: if, as per de la Fuente-Núñez and colleagues9, 1018 acts via direct degradation of intracellular (p)ppGpp, its efficacy should be dramatically different in conditions where the alleged molecular target is essential (MOPS medium without L-valine) or disposable without significant effects on growth rate (MOPS medium supplemented with L-valine). By the same argument, ppGpp0 ΔrelAΔspoT strain lacking (p)ppGpp grown in MOPS medium supplemented with L-valine should be protected from 1018 due to the complete absence of the alleged molecular target of the peptide. However, the killing efficiency of 1018 is similar in all three cases: while at 1 μM the peptide does not affect the cell count, already at 5 μM we detect no viable bacteria (Fig. 2b). The ppGpp0 strain is moderately sensitized to 1018 as compare to wild type (IC50 1.3 ± 0.1 μM vs IC50 1.7 ± 0.2 μM) despite the complete lack on the alleged target, (p)ppGpp. Analogous auxotrophy-based tests performed using Gram-positive Bacillus subtilis also detect no difference in the killing efficiency of 1018 in conditions when (p)ppGpp is essential or not34. Taken together, these results reinforce the idea that the antimicrobial activity of 1018 does not rely on a specific recognition of stringent response messenger (p)ppGpp.
Peptides 1018 and 8101 co-precipitate with ppGpp in a buffer-specific manner
The direct and specific interaction between 1018 and ppGpp was suggested on the bases of co-precipitation of the two compounds9. The original report used 1018 at 250 μM, which is almost two orders of magnitude higher than the concentration used in microbiological assays. We reproduced the experiments with more physiologically relevant concentrations of the peptide by taking advantage of more sensitive detection of the precipitation of tritium-labeled ppGpp (3H-ppGpp) by using liquid scintillation counting.
First, we titrated both 1018 and 8101 in the presence of 4.5 μM 3H-ppGpp in the buffer that was used in the original report (50 mM Tris pH 7.5 buffer9) (Fig. 3a). The buffer lacks the essential components that ensure the solubility of biological samples, i.e. inorganic salts that provide the ionic strength, as well as divalent metal ions (e.g. Mg2+) and polyamines (e.g. spermidine) that serve as counter-ions and ligands. Almost 40% of 3H-ppGpp precipitates at a 1:1 nucleotide-peptide ratio for both peptides and a further increase in peptide concentrations resulted in the precipitation of most of the nucleotides from the solution. However, when we repeat the experiment in the more physiological realistic HEPES-Polymix buffer used to study bacterial protein synthesis35,36, the peptides are dramatically less efficient in co-precipitating with ppGpp, although there is, again, no difference between the two peptides (Fig. 3b).
Peptides 1018 and 8101 co-precipitate with ppGpp in a buffer-specific manner.
Tritium-labelled 4.5 μM H3-ppGpp was mixed with increasing concentrations of peptide 1018 or 8101 in 50 mM Tris-HCl pH 7.5 (a) or in HEPES-Polymix (b) buffer. After incubation for 10 minutes at room temperature the insoluble material was removed by centrifugation and the radioactivity was quantified by liquid scintillation counting. The results are shown as mean values ± SD of three technical replicates.
Discussion
Peptide 1018 and its inverted version 8101 are equally efficient in eliminating both planktonic and biofilm P. aeruginosa as judged by crystal violet (CV) staining assays (Fig. 1). This suggests the possibility that the preferential effect against biofilms observed by de la Fuente-Núñez and colleagues9 could be the result of a flow-chamber detection method favouring inhibition of biofilms by hydrophobic cationic peptides. Current High Throughput Screening protocols for the discovery of anti-biofilm compounds rely on either CV staining in a 96-well format37 or flow chambers10 for detection. It is possible that the latter approach could overestimate the real potency of these compounds.
While several biochemical and microbiological interactions between 1018 and ppGpp are well documented, one should be cautious in putting forward a specific molecular mechanism of action. In the test tube both 1018 and its inverted version 8101 precipitate ppGpp equally well (Fig. 2). Therefore, specificity towards ppGpp is questionable, and it is likely that general physicochemical properties such as charge and hydrophobicity are at play. Moreover, peptide 1018 co-precipitates with other nucleotides with an efficiency correlating with the number of phosphate groups present in the nucleotide molecule, but not with the nature of the base, i.e. it co-precipitates GTP equally efficiently as ATP, and GDP equally efficiently as ADP9.
Induction of the stringent response renders P. aeruginosa more tolerant to 10189, which was interpreted as an indication of a specific mechanistic connection between the two. However, the stringent response renders P. aeruginosa more resilient in general: it becomes tolerant not only to various antibiotics38,39,40, but also to environmental challenges such as hydrogen peroxide38 and UVA41 exposure. However, this does not suggest a mechanistic interaction between ppGpp and the stress factors: while accumulation of (p)ppGpp protects from β-lactam ampicillin, (p)ppGpp is not the molecular target of ampicillin – cell wall synthesis is. Degradation of cellular ppGpp upon exposure to 1018 observed by de la Fuente-Núñez and collegues9 is, again, not necessarily decisive evidence of the two compounds interacting inside the cell since the alarmone nucleotide is highly labile and rapidly degraded by SpoT hydrolase upon antibacterial treatment with, for example, the antibiotic chloramphenicol42.
Conclusions
While peptide 1018 is a potent antimicrobial, it does not specifically disrupt biofilms via a direct and specific interaction with the intracellular messenger nucleotide (p)ppGpp.
Methods
Peptides
Peptides 1018 (VRLIVAVRIWRR-NH2) and its inverted version 8101 (RRWIRVAVILRV-NH2) were ordered in lyophilized format from Storkbio Ltd and Nordic BioSite AB, respectively (both >95% pure). Peptides were dissolved in water and stored at −80 °C in glass vials. Prior usage peptide concentrations were re-measured according to tryptophan fluorescence (extinction coefficient 5690 M−1 cm−1) to make sure non-inverted and inverted version of the peptide was used in same concentrations.
Bacterial strains and growth media
E. coli BW25113 wild type (lacIq rrnBT14 ΔlacZWJ16 hsdR514 ΔaraBADAH33 ΔrhaBADLD78) and isogenic ppGpp0 ΔspoTΔrelA strain were described in Jõers and Tenson43. P. aeruginosa wild type PAO1 was described earlier by Holloway44. MOPS supplemented 0.4% glucose was prepared as per Neidhardt et al.45, except for the omission of thiamine and supplementation with 20 common amino acids (600 μg/ml L-serine, 100 μg/ml L-aspartate, 100 μg/ml L-glutamic acid, 40 μg/ml for others).
Crystal violet staining assay
Biofilm formation was determined using a crystal violet staining assay as per O’Toole24. Wells of a 96-well plate were inoculated with 105 cells in BM2-glucose minimal medium46 and increasing concentrations of peptides. After 24 hours at 37 °C, 100 μl of cell suspension was removed from the well and was used for the evaluation of planktonic growth at 600 nm. Adherent biofilms were washed with distilled water and stained with 0.1% crystal violet solution for 20 minutes. Unbound stain was removed with distilled water and biofilm-attached crystal violet was dissolved with ethanol:acetone solution (4:1; vol:vol) for 10 minutes. The intensity of the violet colour corresponding to the biofilm thickness was measured spectrophotometrically at 595 nm.
Determination of peptide 1018 efficiency against E. coli
For 1018 efficiency measurements against E. coli BW25113 strains fresh colonies were suspended in MOPS medium lacking L-valine, OD600 of the suspension was adjusted to 1.0, and diluted 1000 times for inoculating MOPS medium, with or without L-valine. 104 CFUs per well were seeded in 96-well plates (Sarstedt) in presence of increasing concentrations of 1018 and grown aerobically at 37 °C for 8 hours followed by the CFU measurements.
Colony forming unit (CFU) measurements
10x serial dilution series of the bacterial cultures were prepared in PBS buffer and 5 μl of each suspension was spotted on dry LB plates. After an overnight incubation at 37 °C colonies were counted and CFU/ml calculated. Detection limit for CFU measurements was ≈500 CFU/ml.
ppGpp precipitation assay
Tritium-labelled 4.5 μM ppGpp was mixed with increasing concentrations of peptide 1018 or 8101 in 50 mM Tris-HCl pH 7.5 or in HEPES-Polymix35,36 buffer. After incubation for 10 minutes at room temperature the insoluble material was collected by centrifugation for 5 minutes at 16,000 g and the radioactivity was quantified by liquid scintillation counting. H3-labelled ppGpp was prepared as described in Shyp et al.47 using H3-GDP as a substrate (Hartmann Analytic). The experiments were performed in three technical replicates.
Additional Information
How to cite this article: Andresen, L. et al. Cationic bactericidal peptide 1018 does not specifically target the stringent response alarmone (p)ppGpp. Sci. Rep. 6, 36549; doi: 10.1038/srep36549 (2016).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Romeo, D., Skerlavaj, B., Bolognesi, M. & Gennaro, R. Structure and bactericidal activity of an antibiotic dodecapeptide purified from bovine neutrophils. J Biol Chem 263, 9573–9575 (1988).
Wu, M. & Hancock, R. E. Interaction of the cyclic antimicrobial cationic peptide bactenecin with the outer and cytoplasmic membrane. J Biol Chem 274, 29–35 (1999).
Wieczorek, M. et al. Structural studies of a peptide with immune modulating and direct antimicrobial activity. Chem Biol 17, 970–980 (2010).
Mansour, S. C., de la Fuente-Nunez, C. & Hancock, R. E. Peptide IDR-1018: modulating the immune system and targeting bacterial biofilms to treat antibiotic-resistant bacterial infections. J Pept Sci 21, 323–329 (2015).
Niyonsaba, F. et al. The innate defense regulator peptides IDR-HH2, IDR-1002, and IDR-1018 modulate human neutrophil functions. J Leukoc Biol 94, 159–170 (2013).
Pena, O. M. et al. Synthetic cationic peptide IDR-1018 modulates human macrophage differentiation. PLoS One 8, e52449 (2013).
Scott, M. G. et al. An anti-infective peptide that selectively modulates the innate immune response. Nat Biotechnol 25, 465–472 (2007).
Bolouri, H. et al. Innate defense regulator peptide 1018 protects against perinatal brain injury. Ann Neurol 75, 395–410 (2014).
de la Fuente-Núñez, C., Reffuveille, F., Haney, E. F., Straus, S. K. & Hancock, R. E. W. Broad-Spectrum Anti-biofilm Peptide That Targets a Cellular Stress Response. PLoS Pathog 10, e1004152 (2014).
de la Fuente-Nunez, C. et al. Inhibition of bacterial biofilm formation and swarming motility by a small synthetic cationic peptide. Antimicrob Agents Chemother 56, 2696–2704 (2012).
Steinstraesser, L. et al. Innate defense regulator peptide 1018 in wound healing and wound infection. PLoS One 7, e39373 (2012).
Achtman, A. H. et al. Effective adjunctive therapy by an innate defense regulatory peptide in a preclinical model of severe malaria. Sci Transl Med 4, 135ra64 (2012).
Shestakov, A., Jenssen, H., Hancock, R. E., Nordstrom, I. & Eriksson, K. Synthetic analogues of bovine bactenecin dodecapeptide reduce herpes simplex virus type 2 infectivity in mice. Antiviral Res 100, 455–459 (2013).
Reffuveille, F., de la Fuente-Nunez, C., Mansour, S. & Hancock, R. E. A broad-spectrum antibiofilm peptide enhances antibiotic action against bacterial biofilms. Antimicrob Agents Chemother 58, 5363–5371 (2014).
Hauryliuk, V., Atkinson, G. C., Murakami, K. S., Tenson, T. & Gerdes, K. Recent functional insights into the role of (p)ppGpp in bacterial physiology. Nat Rev Microbiol 13, 298–309 (2015).
Poole, K. Bacterial stress responses as determinants of antimicrobial resistance. J Antimicrob Chemother 67, 2069–2089 (2012).
Dalebroux, Z. D. & Swanson, M. S. ppGpp: magic beyond RNA polymerase. Nat Rev Microbiol 10, 203–212 (2012).
Dalebroux, Z. D., Svensson, S. L., Gaynor, E. C. & Swanson, M. S. ppGpp conjures bacterial virulence. Microbiol Mol Biol Rev 74, 171–199 (2010).
Rivas-Santiago, B. et al. Ability of innate defence regulator peptides IDR-1002, IDR-HH2 and IDR-1018 to protect against Mycobacterium tuberculosis infections in animal models. PLoS One 8, e59119 (2013).
Romling, U. & Balsalobre, C. Biofilm infections, their resilience to therapy and innovative treatment strategies. J Intern Med 272, 541–561 (2012).
Kristensen, K., Henriksen, J. R. & Andresen, T. L. Adsorption of cationic peptides to solid surfaces of glass and plastic. PLoS One 10, e0122419 (2015).
Wiegand, I., Hilpert, K. & Hancock, R. E. Agar and broth dilution methods to determine the minimal inhibitory concentration (MIC) of antimicrobial substances. Nat Protoc 3, 163–175 (2008).
Udekwu, K. I., Parrish, N., Ankomah, P., Baquero, F. & Levin, B. R. Functional relationship between bacterial cell density and the efficacy of antibiotics. J Antimicrob Chemother 63, 745–757 (2009).
O’Toole, G. A. Microtiter dish biofilm formation assay. J Vis Exp (2011).
O’Toole, G. A. & Kolter, R. Flagellar and twitching motility are necessary for Pseudomonas aeruginosa biofilm development. Mol Microbiol 30, 295–304 (1998).
de la Fuente-Nunez, C. et al. D-enantiomeric peptides that eradicate wild-type and multidrug-resistant biofilms and protect against lethal Pseudomonas aeruginosa infections. Chem Biol 22, 196–205 (2015).
Agranat, I., Caner, H. & Caldwell, J. Putting chirality to work: the strategy of chiral switches. Nat Rev Drug Discov 1, 753–768 (2002).
Burke, D. & Henderson, D. J. Chirality: a blueprint for the future. Br J Anaesth 88, 563–576 (2002).
Lorenzen, S., Gille, C., Preissner, R. & Frommel, C. Inverse sequence similarity of proteins does not imply structural similarity. FEBS Lett 545, 105–109 (2003).
Sridhar, S. & Guruprasad, K. Can natural proteins designed with ‘inverted’ peptide sequences adopt native-like protein folds? PLoS One 9, e107647 (2014).
Varik, V., Oliveira, S. R., Hauryliuk, V. & Tenson, T. Composition of the outgrowth medium modulates wake-up kinetics and ampicillin sensitivity of stringent and relaxed Escherichia coli. Sci Rep 6, 22308 (2016).
Ross, W. et al. ppGpp Binding to a Site at the RNAP-DksA Interface Accounts for Its Dramatic Effects on Transcription Initiation during the Stringent Response. Mol Cell (2016).
Xiao, H. et al. Residual guanosine 3′,5′-bispyrophosphate synthetic activity of relA null mutants can be eliminated by spoT null mutations. J Biol Chem 266, 5980–5990 (1991).
Andresen, L. et al. Auxotrophy-based High Throughput Screening assay for the identification of Bacillus subtilis stringent response inhibitors. Sci Rep 6, 35824 (2016).
Jelenc, P. C. & Kurland, C. G. Nucleoside triphosphate regeneration decreases the frequency of translation errors. Proc Natl Acad Sci USA 76, 3174–3178 (1979).
Antoun, A., Pavlov, M. Y., Tenson, T. & Ehrenberg, M. M. Ribosome formation from subunits studied by stopped-flow and Rayleigh light scattering. Biol Proced Online 6, 35–54 (2004).
Junker, L. M. & Clardy, J. High-throughput screens for small-molecule inhibitors of Pseudomonas aeruginosa biofilm development. Antimicrob Agents Chemother 51, 3582–3590 (2007).
Khakimova, M., Ahlgren, H. G., Harrison, J. J., English, A. M. & Nguyen, D. The stringent response controls catalases in Pseudomonas aeruginosa and is required for hydrogen peroxide and antibiotic tolerance. J Bacteriol 195, 2011–2020 (2013).
Nguyen, D. et al. Active starvation responses mediate antibiotic tolerance in biofilms and nutrient-limited bacteria. Science 334, 982–986 (2011).
Viducic, D. et al. Functional analysis of spoT, relA and dksA genes on quinolone tolerance in Pseudomonas aeruginosa under nongrowing condition. Microbiol Immunol 50, 349–357 (2006).
Pezzoni, M., Pizarro, R. A. & Costa, C. S. Protective effect of low UVA irradiation against the action of lethal UVA on Pseudomonas aeruginosa: role of the relA gene. J Photochem Photobiol B 116, 95–104 (2012).
Murray, K. D. & Bremer, H. Control of spoT-dependent ppGpp synthesis and degradation in Escherichia coli. J Mol Biol 259, 41–57 (1996).
Jõers, A. & Tenson, T. Growth resumption from stationary phase reveals memory in Escherichia coli cultures. Sci Rep 6, 24055 (2016).
Holloway, B. W. Genetic recombination in Pseudomonas aeruginosa. J Gen Microbiol 13, 572–581 (1955).
Neidhardt, F. C., Bloch, P. L. & Smith, D. F. Culture medium for enterobacteria. J Bacteriol 119, 736–747 (1974).
Overhage, J., Bains, M., Brazas, M. D. & Hancock, R. E. Swarming of Pseudomonas aeruginosa is a complex adaptation leading to increased production of virulence factors and antibiotic resistance. J Bacteriol 190, 2671–2679 (2008).
Shyp, V. et al. Positive allosteric feedback regulation of the stringent response enzyme RelA by its product. EMBO Rep 13, 835–839 (2012).
Sebaugh, J. L. Guidelines for accurate EC50/IC50 estimation. Pharm Stat 10, 128–134 (2011).
Acknowledgements
The authors are grateful to Vallo Varik for providing E. coli strains and media. This work was supported by the grant IUT2-22 from the Estonian Research Council (TT); European Regional Development Fund through the Centre of Excellence for Molecular Cell Technology (VH and TT); Estonian Science Foundation (PUT37 to VH); Umeå University, the Swedish Research council (Vetenskapsrådet; grant 2013-4680) and the Kempe and Ragnar Söderberg foundations (VH). The work was performed as part of the Umeå Centre for Microbial Research (UCMR) Linnaeus Program supported by Umeå University and the Swedish Research Council (349-2007-8673). The funders had no role in study design, data collection and interpretation or the decision to submit the work for publication.
Author information
Authors and Affiliations
Contributions
L.A., T.T. and V.H. conceived the project and designed the experiments. L.A. performed experiments. L.A. and V.H. wrote the paper with contributions from T.T.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Andresen, L., Tenson, T. & Hauryliuk, V. Cationic bactericidal peptide 1018 does not specifically target the stringent response alarmone (p)ppGpp. Sci Rep 6, 36549 (2016). https://doi.org/10.1038/srep36549
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep36549
This article is cited by
-
Awakening sleeper cells: a narrative review on bacterial magic spot synthetases as potential drug targets to overcome persistence
Current Genetics (2022)
-
Many birds with one stone: targeting the (p)ppGpp signaling pathway of bacteria to improve antimicrobial therapy
Biophysical Reviews (2021)
-
Supreme activity of gramicidin S against resistant, persistent and biofilm cells of staphylococci and enterococci
Scientific Reports (2019)
-
Construction of a genetically modified T7Select phage system to express the antimicrobial peptide 1018
Journal of Microbiology (2019)
-
Inhibition and Eradication of Pseudomonas aeruginosa Biofilms by Host Defence Peptides
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.