Abstract
Large blue butterflies, Phengaris (Maculinea), are an important focus of endangered-species conservation in Eurasia. Later-instar Phengaris caterpillars live in Myrmica ant nests and exploit the ant colony’s resources, and they are specialized to specific host-ant species. For example, local extinction of P. arion in the U. K. is thought to have been due to the replacement of its host-ant species with a less-suitable congener, as a result of changes in habitat. In Japan, Myrmica kotokui hosts P. teleius and P. arionides caterpillars. We recently showed, however, that the morphological species M. kotokui actually comprises four genetic clades. Therefore, to determine to which group of ants the hosts of these two Japanese Phengaris species belong, we used mitochondrial COI-barcoding of M. kotokui specimens from colonies in the habitats of P. teleius and P. arionides to identify the ant clade actually parasitized by the caterpillars of each species. We found that these two butterfly species parasitize different ant clades within M. kotokui.
Similar content being viewed by others
Introduction
Several orders of animals are found in ant nests. Some of them depend in some way on ants during their life cycle, which are known as myrmecophiles1,2. In lepidopteran insects, more than half of Lycaenidae species have associations with ants that range from facultative association to obligate nest parasitism3. In order to communicate ants, lycaenid caterpillars and pupae have some myrmecophilous organs, such as dorsal nectary organs, pore cupola organs and tentacle organs, producing nectars and other substances, and organs for sound production3. By using these myrmecophilous organs, lycaenids emit chemical and acoustic cues to manipulate their host ants.
Large blue Phengaris (Maculinea) butterflies (Lepidoptera: Lycaenidae) are widely distributed in Europe and Asia, and all known species (about 10) are considered to be obligately myrmecophilous. Phengaris butterflies are the best-known example of parasitic myrmecophily, and they exhibit a high degree of host-ant specificity4. Early instar caterpillars feed on specific host plants (flowers of Lamiaceae, Gentianaceae, or Rosaceae). When they reach the fourth instar, they drop from their host plant to the ground and gain entry to a nest of Myrmica ants (Myrmicinae) by using chemical mimicry to cause themselves to be recognized as ant larvae by worker ants, who then carry them into their nest4,5,6. Ant nests are strongly protected by their ant inhabitants. Therefore, if an organism can enter a nest without being attacked by the ants, the nest becomes a safe shelter against natural enemies1.
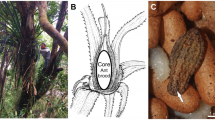
Once they gain entry into an ant nest, the caterpillars grow by exploiting the resources of the ant colony (Fig. 1). Phengaris uses two parasitic strategies: “predatory” caterpillars prey on the ant brood, and “cuckoo” caterpillars are fed by the ants via regurgitation7,8. Phengaris teleius and P. arionides, which are widely distributed in East Asia, including Japan, are predatory species. Caterpillars following both strategies gain more than 98% of their biomass in the ant nest; thus, these butterfly species are obligate parasites9. By the time the fourth-instar caterpillars pupate, the host-ant colonies have suffered serious damage, yet the ants transport these parasites into their nest in their own mandibles.
Because of the high specificity of parasitic Phengaris butterflies toward their host plants and ants, both must occur together for a habitat to be suitable for the butterflies. As a result, these butterflies are vulnerable to environmental change, and all species of Phengaris are endangered worldwide10. Two Phengaris species, P. teleius and P. arionides, are distributed in the Japanese archipelago, and geographic sub-species of P. teleius are classified as “Near Threatened” or “Critically Endangered”, and P. arionides is classified as “Near Threatened” in the 4th (latest) version of the Japanese red lists11. In addition, in March, 2016, the government of Japan’s Ministry of Environment added a sub-species P. teleius kazamoto living in Chubu area of central Honshu to the list of “National Endangered Species”, and prohibited the collecting and transferring of it.
To develop a conservation strategy for endangered Phengaris butterflies, it is essential to identify their host-ant species. In the United Kingdom, P. arion became extinct after its host-ant species was replaced by unsuitable congeners5,12. Before the 1980s, it was thought that Phengaris caterpillars could parasitize any Myrmica ant species13, but in a comprehensive investigation of host specificity among eight Myrmica species and five Phengaris species, Thomas, et al.5 found a one-to-one association between each ant and butterfly species. For example, they found that the survival rate of P. arion caterpillars in nests of M. sabuleti and M. scabrinodis was on average 15% and 2%, respectively5. Thus, the major host-ant species of P. arion is M. sabuleti, and it is difficult for the caterpillars to mature in a nest of M. scabrinois.
In past morphological studies, the host-ant species of Japanese Phengaris species was identified as Myrmica kotokui2,14,15,16. However, Ueda, et al.17 showed that the species recognized as M. kotokui on the basis of morphology actually consisted of four genetic clades. Therefore, the host-ant specificity of Phengaris needs to be determined not just at the species level but also at the genetic level. Moreover, Ueda, et al.18 showed that each cryptic clade prefers a different habitat and nesting microhabitat. Thus, P. teleius, which inhabits grasslands, and P. arionides, which lives in woodlands, might parasitize different ant clades within M. kotokui. To determine the true host ant of P. teleius and P. arionides, we (1) investigated M. kotokui colonies in the habitats of P. teleius and P. arionides, (2) used DNA barcoding to estimate the frequencies of the different ant clades in each habitat, and (3) then identified the ant clade that the caterpillars of each butterfly species actually parasitized.
Results
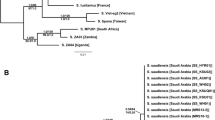
The DNA clade of each of the 99 ant colonies collected from the six Phengaris habitats was identified by neighbor-joining (NJ) analysis of 470-bp sequences of the mitochondrial COI gene (Fig. S1). We found that four belonged to the L1 clade, 67 to the L2 clade, and 28 to the L3 clade (Table 1). Thus, L2 was the dominant clade in the P. teleius grassland habitats (86.2–100%), and in the woodland P. arionides habitat, all ant colonies belonged to the L3 clade (Table 1). These habitat preferences of the ant clades are congruent with the findings of Ueda, et al.18.
Next we identified the DNA clade of each ant colony parasitized by Phengaris caterpillars. The four ant colonies parasitized by P. teleius belonged to L2, and the three ant colonies parasitized by P. arionides belonged to L3 (Table 1). Although the sample size is too small for statistical testing, based on the habitat preferences of ants and the parasitic frequency of Phengaris caterpillars, we tentatively conclude that P. teleius parasitizes L2 colonies and P. arionides parasitizes L3 colonies under natural conditions. To determine the specificity of the Japanese Phengaris—Myrmica interaction more definitively, additional sampling is essential. During this study, however, we decided not to collect more specimens because we judged that additional collections risked excessively depleting the populations of both butterflies and ants.
Discussion
We showed that two Japanese Phengaris butterfly species apparently parasitize the nests of different ant clades within the M. kotokui morphological species. This finding raises the question, does this apparent specificity represent an adaptation on the part of the butterfly, or did the dominance of L2 and L3 clades in grasslands and woodlands, respectively, lead to this apparent one-to-one correspondence without adaptation? It is possible that Japanese Phengaris—Myrmica interactions are an example of parasitic adaptation, because preliminary tests indicate that P. teleius caterpillars and their host ants have some cuticular hydrocarbons (CHCs) in common (R. Seki, personal communication). To confirm that the adaptation has occurred, in addition to a CHC analysis, acoustic measurements should also be performed to compare the sounds produced by caterpillars and butterfly pupae to those of worker and queen ants, because both chemical and acoustical mimicry by P. rebeli caterpillars of their host Myrmica ants have been demonstrated6,19,20,21,22.
In this study, we found cryptic host-ant specificity in Phengaris butterflies for the first time. Therefore, to preserve these East Asian butterflies, it is important to maintain their particular host ant clades. Do declines in the number of host-ant colonies in appropriate butterfly habitats drive the extinction of the butterflies? We investigated the ant species composition in the area of the most endangered population of P. teleius in habitat E (Table 1). We found that because the soil had acidified, become drier and swampy meadows, suitable micro-habitat to L2 clade, decreased (Table 1), probably as a result of changing agricultural practices, only Lasius japonicus (Formicidae) and Myrmica jessensis, neither of which are suitable host ants, occurred beneath or near Sanguisorba officinalis (Rosaceae), the host plant of early-stage P. teleius caterpillars. Because all previous studies showed that P. teleius and P. arionides in Japan parasitize the nest of M. kotokui, we determined that M. jessensis may not be a suitable host ant species. There were some reports that the caterpillars of the Japanese Phengaris species parasitize the nest of Aphaenogaster japonica14,23, but the ant’s name was mistake for M. kotokui15. In this area, we were able to fine only two ant nests of the L1 clade, and it was located at the edge of a forest and far from any suitable host plants for the caterpillars. Given the suitable host ant of P. teleius is L2 clade, the caterpillars cannot live in any of the ant nests in habitat E. The displacement of ant species in Phengaris habitats in the UK has been shown to lead to the extinction of native populations of P. arion5,12. Thomas et al.11 showed that this high specificity triggered the local extinction of P. arion. Myrmica scabrinodis prefers to nest in tall grass, whereas M. sabuleti prefers areas where grass height is kept low by herbivory. When herbivores were excluded from the P. arion habitat and the grass became high, M. sabuleti replaced M. scabrinodis and, as a result, the P. arion population declined sharply12,24. On the basis of this finding, in UK sanctuaries for P. arion, a suitable environment for the host ant was produced by controlled burning and grazing. Then, once M. sabuleti was re-established in the restoration sites, the butterflies were successfully re-introduced from Sweden5,12. This finding suggests the displacement of ant species in Japan could cause P. teleius and P. arionides to become extinct. To save Japanese Phengaris butterflies from extinction, Phengaris populations and their habitats should be surveyed, interactions between the butterflies and ants should be investigated, and the anthropogenic impact on their habitats and hosts should be evaluated.
Methods
Parasitization rates of Japanese Phengaris on Myrmica kotokui
It is not necessary to acquire government permission to collect the ant samples in the concerned regions. However, we got approval to collect the samples from the managers of each butterfly sanctuary. We searched for M. kotokui nests in four P. teleius habitats (A–E) separated from one another by more than 100 km, and in one P. arionides habitat (F). To protect the butterflies, we do not show the detailed collecting site locations here, but each habitat area has a large population of butterflies except for habitat E. The population size of the butterfly may relate to colony density of M. kotokui (Table 1). The butterfly population in the habitat (E) with the lowest colony density was much smaller than the others. And the colony density of the ants may relate to soil moisture (Table 1). In each P. teleius habitat A–E, M. kotokui nested in the muddy soil of a moist grassland, and the swampy meadows decrease may lead to decreasing the colony density of the ants (Table 1). To determine whether P. teleius caterpillars were present in a nest, we removed all soil to a depth of 0.5 m within a radius of 1.0 m of the nest entrance. We left all ants, including queen ants, in the colony, except for some worker specimens removed for DNA analysis. In all, we investigated 29 ant colonies in habitat A, 14 in habitat B, 18 in habitat C, 11 in habitat D and 2 in habitat E (Table 1); we found caterpillars in three habitat A colonies and one caterpillar in a habitat B colony, but no caterpillars in habitats C and D (Table 1). Thus, the parasitization rate of P. teleius in ant nests was 5.4%, which is lower than rates reported this species in Poland and France5,25. The lower parasitizing rate in Japan may indicate that the P. teleius population is small, or it may be an underestimate, because in our survey we did not completely excavate the ant colonies.
In P. arionides habitat F, M. kotokui nested in decayed logs in a forest. We opened decayed wood from around each nest to determine whether P. arionides caterpillars were present in the nest. We investigated 25 ant colonies in habitat E, and found caterpillars in three of them (Table 1). Thus, the parasitization rate of P. arionides in the ant nests was 12.0%. This rate is higher than the P. teleius rate (5.4%), but we cannot compare it with rates in other regions because, to our knowledge, this is the first report of the parasitization rate of P. arionides in ant nests.
DNA barcoding of ants
During the nest survey, we collected 10 to 20 worker ants from each colony for DNA barcoding and preserved them in 99.5% EtOH until the analysis. We deposited voucher specimens at the Faculty of Science, Shinshu University, Matsumoto, Japan. We extracted DNA from the whole body of each ant using a DNeasy Blood & Tissue Kit (Qiagen, Hilden, Germany) following the manufacturer’s protocols. Then we amplified the mitochondrial COI gene by polymerase chain reaction (PCR) using Takara Ex Taq polymerase (Takara Bio, Shiga, Japan), and the primers MyrCOI-F1 (5′-TA GGR TCR CCT GAT ATA GC-3′) and MyrCOI-R1 (5′-CC AGG TAY YAT TAA AAT ATA AAC TTC-3′)18. The reaction was carried out for 30 cycles of 95 °C for 30 s, 50 °C for 30 s, and 72 °C for 40 s. After amplification, the PCR products were purified with ExoSap-IT reagent (USB, Cleveland, Ohio, USA). Both strands were sequenced with a BigDye Terminator v1.1 Cycle Sequencing Kit (ABI, Weiterstadt, Germany) on an ABI 3130 Genetic Analyzer.
The mitochondrial COI sequences were edited and aligned with SeqScape v. 2.5 software (ABI, Weiterstadt, Germany). We imported the obtained COI dataset into the COI dataset of Ueda et al.17, and then determined the clade of the ants in each colony by a neighbor-joining NJ analysis, performed with MEGA6 software26. Although Ueda et al.17 used both COI and LwRh sequences to infer the molecular phylogeny, in this study we analyzed only the COI sequences because the mutation rate of the LwRh gene is slow and it is possible to determine the clade by using only the COI gene data. The GenBank accession numbers of the COI gene sequences are listed in Table S1.
Additional Information
How to cite this article: Ueda, S. et al. Host-ant specificity of endangered large blue butterflies (Phengaris spp., Lepidoptera: Lycaenidae) in Japan. Sci. Rep. 6, 36364; doi: 10.1038/srep36364 (2016).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Kistner, D. H. Social and evolutionary significance of social insect symbionts. Vol. 1 (Academic Press, 1979).
Maruyama, M., Komatsu, T., Kudo, S., Shimada, T. & Kinomura, K. The Guests of Japanese Ants. 222 (Tokai University Press, 2013).
Pierce, N. E. et al. The ecology and evolution of ant association in the Lycaenidae (Lepidoptera). Annu. Rev. Entomol. 47, 733–771, doi: 10.1146/annurev.ento.47.091201.145257 (2002).
Als, T. D. et al. The evolution of alternative parasitic life histories in large blue butterflies. Nature 432, 386–390, doi: 10.1038/nature03020 (2004).
Thomas, J., Elmes, G., Wardlaw, J. & Woyciechowski, M. Host specificity among Maculinea butterflies in Myrmica ant nests. Oecologia 79, 452–457 (1989).
Akino, T., Knapp, J. J., Thomas, J. A. & Elmes, G. W. Chemical mimicry and host specificity in the butterfly Maculinea rebeli, a social parasite of Myrmica ant colonies. Proc R Soc B 266, 1419–1426 (1999).
Thomas, J. A. & Wardlaw, J. C. The capacity of a Myrmica ant nest to support a predacious species of Maculinea butterfly. Oecologia 91, 101–109 (1992).
Thomas, J. A. et al. Effects of latitude, altitude and climate on the habitat and conservation of the endangered butterfly Maculinea arion and its Myrmica ant hosts. J. Insect Conserv. 2, 39–46, doi: 10.1023/A:1009640706218 (1998).
Elmes, G. W., Thomas, J. A., Munguira, M. L. & Fiedler, K. Larvae of lycaenid butterflies that parasitize ant colonies provide exceptions to normal insect growth rules. Biol. J. Linn. Soc. 73, 259–278, doi: 10.1006/bijl.2001.0534 (2001).
Wynhoff, I. The recent distribution of the European Maculinea species. J. Insect Conserv. 2, 15–27, doi: 10.1023/A:1009636605309 (1998).
Ministry of the Environment, Governmet of Japan the 4th version of the Japanese red lists - Insect, http://www.env.go.jp/press/files/jp/21555.pdf (2012).
Thomas, J. A., Simcox, D. J. & Clarke, R. T. Successful conservation of a threatened Maculinea butterfly. Science 325, 80–83, doi: 10.1126/science.1175726 (2009).
Cottrell, C. Aphytophagy in butterflies: its relationship to myrmecophily. Zool. J. Linn. Soc. 80, 1–57 (1984).
Yamaguchi, S. The Life Histories of Five Myrmecophilous Lycaenid Butterflies of Japan. 262 (Kodansha, 1988).
Hiraga, S. Reidentifying of host ants of Maculinea arionides. Yadoriga 196, 31–34 (2003).
Shirôzu, T. The Standard of Butterflies in Japan. (Gakken, 2006).
Ueda, S. et al. Phylogeny and Phylogeography of Myrmica rubra Complex (Myrmicinae) in the Japanese Alps. Psyche 2012, 1–7, doi: 10.1155/2012/319097 (2012).
Ueda, S. et al. Ecological and morphological differentiation between two cryptic DNA clades in the red ant Myrmica kotokui Forel 1911 (Myrmicinae). New Entomol. 62, 1–10 (2013).
Devries, P. J., Cocroft, R. B. & Thomas, J. Comparison of acoustical signals in Maculinea butterfly caterpillars and their obligate host Myrmica ants. Biol. J. Linn. Soc. 49, 229–238, doi: 10.1111/j.1095-8312.1993.tb00902.x (1993).
Elmes, G. W., Akino, T., Thomas, J. A., Clarke, R. T. & Knapp, J. J. Interspecific differences in cuticular hydrocarbon profiles of Myrmica ants are sufficiently consistent to explain host specificity by Maculinea (large blue) butterflies. Oecologia 130, 525–535, doi: DOI 10.1007/s00442-001-0857-5 (2002).
Schlick-Steiner, B. C. et al. A butterfly’s chemical key to various ant forts: intersection-odour or aggregate-odour multi-host mimicry? Naturwissenschaften 91, 209–214, doi: 10.1007/s00114-004-0518-8 (2004).
Barbero, F., Thomas, J. A., Bonelli, S., Balletto, E. & Schonrogge, K. Queen ants make distinctive sounds that are mimicked by a butterfly social parasite. Science 323, 782–785, doi: 10.1126/science.1163583 (2009).
Watanabe, H. Life history of Maculinea arionides. Chôken Field 2, 22–26 (1987).
Elmes, G. W. et al. The ecology of Myrmica ants in relation to the conservation of Maculinea butterflies. J. Insect Conserv. 2, 67–78, doi: 10.1023/A:1009696823965 (1998).
Witek, M. et al. Local host ant specificity of Phengaris (Maculinea) teleius butterfly, an obligatory social parasite of Myrmica ants. Ecol. Entomol. 35, 557–564, doi: 10.1111/j.1365-2311.2010.01213.x (2010).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 30, 2725–2729, doi: 10.1093/molbev/mst197 (2013).
Acknowledgements
We thank T. Ando, S. Shimamoto, M. Hattori, K. Abe, K. Maruyama and T. Kobayashi for sampling support; T. Itoh, T. Nakatani, Y. Hagiwara, S. Hiraga, M. Yago and S. Yamaguchi for providing information about sampling sites; T. Akino and R. Seki for providing information about chemical mimicry; and H. Mohri for the initiation of this research project.
Author information
Authors and Affiliations
Contributions
S.U., T.I. and H.S. conceived and designed the experiments. S.U., T.K., R.A. and H.S. performed the sampling and experiments. S.U. and T.K. analyzed the DNA data. S.U. and T.I. contributed reagents/materials/analysis tools. S.U., R.A. and H.S. wrote the paper and T.K. prepared Fig. 1. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Ueda, S., Komatsu, T., Itino, T. et al. Host-ant specificity of endangered large blue butterflies (Phengaris spp., Lepidoptera: Lycaenidae) in Japan. Sci Rep 6, 36364 (2016). https://doi.org/10.1038/srep36364
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep36364
This article is cited by
-
Trophic host-parasitoid interactions of two Neotropical butterfly species in southeastern Mexico
International Journal of Tropical Insect Science (2022)
-
Multi-year occupancy of the hops blue butterfly (Celastrina humulus): habitat patch colonization and extinction
Journal of Insect Conservation (2020)
-
Host specificity pattern and chemical deception in a social parasite of ants
Scientific Reports (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.