Abstract
In 2010, a pathogenic flavivirus termed duck Tembusu virus (DTMUV) caused widespread outbreak of egg-drop syndrome in domesticated ducks in China. Although the glycoprotein E of DTMUV is an important structural component of the virus, the B-cell epitopes of this protein remains uncharacterized. Using phage display and mutagenesis, we identified a minimal B-cell epitope, 374EXE/DPPFG380, that mediates binding to a nonneutralizing monoclonal antibody. DTMUV-positive duck serum reacted with the epitope, and amino acid substitutions revealed the specific amino acids that are essential for antibody binding. Dot-blot assays of various flavivirus-positive sera indicated that EXE/DPPFG is a cross-reactive epitope in most flaviviruses, including Zika, West Nile, Yellow fever, dengue, and Japanese encephalitis viruses. These findings indicate that the epitope sequence is conserved among many strains of mosquito-borne flavivirus. Protein structure modeling revealed that the epitope is located in domain III of the DTMUV E protein. Together, these results provide new insights on the broad cross-reactivity of a B-cell binding site of the E protein of flaviviruses, which can be exploited as a diagnostic or therapeutic target for identifying, studying, or treating DTMUV and other flavivirus infections.
Similar content being viewed by others
Introduction
Flaviviruses are positive-sense RNA viruses classified in the genus Flavivirus, family Flaviviridae1, which include several important vector-borne viruses of zoonotic nature. Many flaviviruses play a considerable role in human health and disease, such as dengue virus (DENV), West Nile virus (WNV), Japanese encephalitis virus (JEV), and Zika virus (ZIKV). In April 2010, a severe outbreak of a duck viral infection, which led to a devastating drop in egg production (i.e. egg-drop syndrome), spread throughout the major duck-producing regions in China. Postmortem examination of infected ducks indicated severe ovarian hemorrhage, ovaritis, and regression. Genomic sequencing revealed that the outbreak was caused by the duck Tembusu virus (DTMUV), which is a mosquito-borne Ntaya group flavivirus2,3,4,5,6,7.
The DTMUV genome, like that of other flaviviruses, encodes three structural proteins (C, prM/M, and E) and seven nonstructural proteins (NS1, NS2A, NS2B, NS3, NS4A, NS4B, and NS5)1,3,8. In most flaviviruses, the glycosylated E protein is located on the virion surface and plays an important role in virus receptor binding, host-cell entry, and antigenicity9. The flavivirus E protein contains three structurally distinct domains: DI, DII, and DIII. The DI domain is composed predominantly of type-specific nonneutralizing epitopes9. The DII domain contributes to virus-mediated membrane fusion and contains many cross-reactive epitopes, eliciting neutralizing and nonneutralizing antibodies9,10,11,12,13. The DIII domain contains multiple type- and subtype-specific epitopes, which elicit only virus-neutralizing antibodies10,14,15,16. Although the biologic characteristics of most flaviviruses are well defined, no specific antiviral drugs are available for clinical use against flavivirus infections.
Studies on the immunopathogenesis of severe dengue fever suggest that induction of cross-reactive nonneutralizing antibodies may increase the likelihood of acute disease during subsequent infections with different serotypes17,18,19. Preexisting cross-reactive antibodies form nonneutralized complexes that allow the virus to enter Fc receptor–expressing cells more efficiently. Therefore, the presence of cross-reactive nonneutralized antibodies most likely plays a role in antibody-dependent enhancement of the infection process. Manipulation of a virus to limit antigen exposure and dominant recognition of cross-reactive nonneutralizing sites, while augmenting the induction of protective neutralizing antibodies directed to epitopes, is a useful strategy for vaccine development.
In this study, we performed phage display and structure modeling to map a new epitope within the DIII domain of the DTMUV E protein that generates broad cross-reactivity to other flavivirus-positive sera. These findings better our understanding of the structure–function relations of E protein epitopes that are important for flavivirus biology, which can lead to improved sero-diagnosis, inform vaccine design, and knowledge of flavivirus pathogenesis.
Results
Epitope prediction
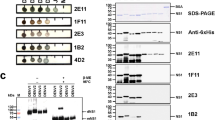
To map the location of the epitope within the DTMUV E protein, we screened a phage-displayed 12-mer random peptide library using the monoclonal antibody (mAb) 3B6. After three rounds of biopanning, we selected and evaluated several phage clones for their reactivity with the 3B6 mAb and with the negative control anti-porcine interferon-c (IFN-c) mAb. Of 30 clones, 26 (B1–B26) reacted with the 3B6 mAb (OD450 ≥ 1.20), whereas the remaining clones were less reactive (OD450 < 0.36). None of the selected clones reacted with the IFN-c mAb (OD450 < 0.27) (Supplementary Figure S1). Sequencing of the phage clones with the highest OD values revealed the consensus sequence EXE/DPPFG (Table 1). This amino acid sequence is identical to amino acids 374 to 380 (EVEPPFG) of the E protein of DTMUV.
Mapping of the minimal epitope
To determine the minimal epitope required for binding to the 3B6 mAb, we first expressed and purified several variants of the EXE/DPPFG fragment (Table 2). In a dot blot assay, we found that the 3B6 mAb recognized the EVE/DPPFG fragment variants and full-length E protein (Fig. 1) but not the control peptide YIRTPACWD. This result suggests that the EVE/DPPFG fragment is a B-cell epitope of the DTMUV E protein. To define the epitope more precisely, we next generated EXE/DPPFG fragments containing substitutions or C- or N-terminal deletions (Table 2). The V375A, V375L, or E376D substitutions did not abolish 3B6 mAb– binding activity (Fig. 1), suggesting that the 375X and 376E/D residues in the 374EXE/DPPFG380 epitope fragment are indiscriminate. In contrast, deletions of the 374E or 380G residues at the N or C terminus of 374EXE/DPPFG380 abolished 3B6 mAb binding, indicating that the 374EXE/DPPFG380 fragment is the minimal epitope mapped by the 3B6 mAb.
Sequence analysis of the identified epitope in DTMUV and other flaviviruses
To determine whether the EXE/DPPFG sequence is conserved among the E proteins of DTMUV and other flaviviruses, we aligned the E protein amino acid sequences, including the EXE/DPPFG epitope region, of several flaviviruses. Specifically, we aligned the E protein sequences of three strains of DTMUV, four strains of DENV, two strains of WNV, two strains of JEV, ZIKV, yellow fever virus (YFV), Murray Valley encephalitis virus (MVEV), Saint Louis encephalitis virus (SLEV), and Kunjin virus (KJV) (Table 3). We found that the amino acids in the E protein minimal B-cell epitope of many mosquito-borne flaviviruses (Fig. 2) were essentially identical. The EXE/DPPFG motif was highly conserved (approximately 85%) in all the viruses tested, with variability occurring only in the indiscriminate 375X and 376E/D residues. The considerable sequence homology of the EXE/DPPFG motif suggests that it is a wide-spectrum epitope of many flaviviruses (Fig. 2).
Sequence alignment of a minimal epitope in the E protein of flavivirus strains.
The EXE/DPPFG epitope of various strains of DTMUV and other mosquito-borne flaviviruses were aligned using Lasergene software. Amino acid positions for each sequence are numbered on both sides. Dashes indicate identical amino acids. The identified epitope region is indicated by grey shading.
Epitope binding by duck DTMUV anti-serum
Dot-blot and western blot assays were used to test whether duck DTMUV anti-serum could recognize various EXE/DPPFG epitopes. Dot blots of EVEPPFG, EAEPPFG, EADPPFG, ELEPPFG, and ELDPPFG peptides, in addition to the full-length E protein, demonstrated positive reactivity to duck DTMUV anti-serum (Fig. 3), but the negative control peptide (i.e. YIRTPACWD) did not. This is further support that EXE/DPPFG most likely represents the minimal B-cell epitope of the DTMUV E protein, and the 375X and 376E/D residues are indiscriminate amino acids in these two positions. Western blot assays also confirmed the reactivity of the EXE/DPPFG epitope to duck DTMUV anti-serum (Supplementary Figure S2), suggesting that both methods reliably identify the minimal B-cell epitope of the DTMUV E protein.
Epitope reactivity to ZIKV-, DENV-, JEV-, WNV-, and YFV-positive sera
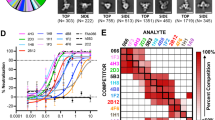
To determine the epitope cross-reactivity to other flavivirus, dot blots of the purified EVEPPFG fragment were incubated with ZIKV-, DENV-, JEV-, WNV-, and YFV-positive sera. All flavivirus-positive sera reacted with the EVEPPFG peptide and full-length E protein, but did not react with the negative control peptide (Fig. 4). Because the EXE/DPPFG epitope in the E proteins of MVEV, SLEV, and KJV is highly conserved (Table 3), it is likely that positive sera produced by these flaviviruses might also positively react. However, this could not be confirmed because MVEV-, SLEV-, and KJV-positive sera were unavailable.
Cross-reactivity of the E protein minimal epitope with flavivirus-positive sera.
The EVEPPFG epitope was probed for reactivity with various flavivirus-positive sera, including Zika virus (ZIKV), dengue virus (DENV), Japanese encephalitis virus (JEV), West Nile virus (WNV), and yellow fever virus (YFV), by dot-blot assay. YIRTPACWD (N) and full-length E protein (E) were used as negative and positive controls, respectively.
Location of the epitope on the E protein 3D structure
We evaluated the molecular structure and stereochemical quality of the DTMUV E protein with ProSA and PROCHECK. Using GlycoEP glycosylation prediction, we found that the DTMUV E protein most likely contains two N-glycosylation sites, 154N and 314N. Prediction scores for 154N and 314N were 0.838 and 0.438, respectively. However, using the NGlycPred algorithm to analyze the structural and residue pattern information of the epitope for potential N-glycosylation sites, we found that 154N, and not 314N, is likely the only site glycosylated. Modeling of the molecular structure of the DTMUV E protein revealed that the EXE/DPPFG epitope possesses a loop conformation and is located in the DIII domain but is not close to the DIII lateral ridge (Fig. 5).
The EVEPPFG epitope of the DTMUV E protein is located in domain III.
The protein-dimer structure of the DTMUV E protein was modeled using the crystal structure of the JEV E protein as a template. The domain I (DI), domain II (DII), and domain III (DIII) in one E protein monomer are coloured magenta, yellow, and blue, respectively. The other monomer is coloured grey. The location of the epitope is depicted as spheres and labeled. Two predicted N-glycosylation sites are coloured sky blue.
Discussion
Various mAbs have been used to identify flavivirus-specific epitopes and to investigate the antigenic structure of these epitopes. The DIII domain of the flavivirus E protein may play an important role in viral replication and infection. However, molecular information about the epitope in the DIII domain of the DTMUV E protein is lacking. The aim of this study was to investigate the antigenic site within the DIII domain using the E protein-specific 3B6 mAb. Using a 12-mer random peptides phage display system and mutagenesis, we precisely mapped a minimal B-cell epitope to amino acids 374 through 380 of the E protein. Duck DTMUV anti-serum positively reacted to the epitope, indicating the importance of these amino acids for antibody-epitope binding and the fact that detectable levels of antibody are generated to it following natural infection.
Sequence alignment revealed that the EXE/DPPFG epitope is highly conserved in DTMUV, ZIKV, WNV, DENV, MVEV, SLEV, KJV, and JEV, suggesting that the epitope has a similar function in these viruses. Cross-reactivity of ZIKV-, DENV-, JEV-, WNV-, and YFV-positive sera suggests that EXE/DPPFG is an immunodominant epitope among flaviviruses; however, the cross-reactivity of MVEV-, SLEV-, and KJV-positive sera is yet to be confirmed. Sequences alignments indicated that E of DTMUV showed 46.43-48.81% to DENV1-4, 61.48% to WNV, 64.87% to JEV, 43.65% to YFV, 64.47% to SLEV, 65.67% to KJV, 63.87% to MVEV, and 53.17% to ZIKV, but the EXE/DPPFG motif reacted to almost all flavivirus anti-serum, which suggests that the wide-spectrum epitope are unusual. In future vaccine development, it may be prudent to take effort to reduce induction of antibodies to this wide-spectrum immunodominant site to limit the formation of nonneutralized antibody complexes and antibody-dependent enhancement of the infection process. In contrast, design of optimal antigen peptides that specifically recognize the EXE/DPPFG epitope could improve the broad detection of DTMUV and other flaviviruses.
Using 3D-structure modeling of the DTMUV E protein, we observed that the EXE/DPPFG epitope is not located in the DII domain, as previously predicted9,10,11,12,13. In contrast, we found that the epitope is located in the DIII domain and does not protrude from the surface of the E protein. Previous studies have demonstrated that binding of some E reactive antibodies relies on the dynamic motion of protein molecules (i.e. “breathing”) in the virion particle, leading to transient exposure of buried epitopes20,21,22. Whether the 3B6 mAb requires such “breathing” for the epitope to become exposed to permit antibody binding remains to be resolved.
In conclusion, we have defined a novel epitope of the DTMUV E protein, its cross-reactivity to other flavivirus-positive sera, and its location within the 3D structure of the E protein. Our findings provide new insights into the structure and organization of the DTMUV E protein and valuable epitope information for the development of diagnostic assays and potential vaccines to prevent DTMUV and other flavivirus infections.
Methods
Generation of DTMUV virus, 3B6 mAb, and flavivirus-positive sera
DTMUV TA was grown in duck embryo fibroblasts cells or embryonated chicken eggs, as previously described5. Development and characterization of the E-specific 3B6 mAb has been described previously23. JEV- and WNV-positive rabbit sera were donated by Dr. Hua, and DENV-positive serum was donated by Dr. Qi Xian. ZIKV- and YFV-positive sera were obtained from the Human Zika Virus IgM ELISA Kit (MyBioSource) and the Human Yellow Fever Virus Antibody IgG ELISA Kit (MyBioSource), respectively.
Affinity purification of the 3B6 mAb
The 3B6 mAb was purified from mouse ascites fluid using protein G agarose (Invitrogen), according to manufacturer instructions. The concentration of purified IgG was determined by measuring absorbance at 278 nm.
Epitope mapping
The epitope was mapped with purified 3B6 mAb using the Ph.D-12TM Phage Display Peptide Library Kit (New England BioLabs), as previously described24,25. Three rounds of biopanning were performed. Briefly, each well of a 96-well plate was coated with 10 μg/mL of purified 3B6 mAb and incubated with blocking buffer. The phage library was then added to the plate and incubated for 1 hour. After five washes with TBS buffer, 1 M Tris-HCl was added to the plate to elute the bound phages24,25. The phages were then amplified and titred on LB/IPTG/Xgal plates for selection. The ratio of output to input was calculated as the titre of the amplified output phages to the titre of the input phages.
ELISA and sequencing of phage clones
After three rounds of biopanning, as described above and elsewhere24,25, individual phage clones were selected for target binding in an ELISA. Briefly, 96-well plates were coated with 100 ng of 3B6 mAb or mouse anti–porcine IFN-c (Sigma-Aldrich) as a negative control. The coated wells were then blocked, and the selected phages (1010 pfu/well, 100 μL/well) were added. The coated plates were then washed ten times with TBST, and bound phages were detected with horse radish peroxidase (HRP)-conjugated sheep anti–M13 antibody (Pharmacia), as described previously24,25. Colour development was achieved by adding a substrate solution containing o-phenylenediamine. Positive phage clones were sequenced as previously described24,25.
Sequence analysis
To assess the level of conservation of the epitope among DTMUVs and other representative flaviviruses, we performed sequence alignments of the epitopes and their corresponding locations in the E protein of three DTMUV strains and several strains of other flaviviruses using Lasergene software (DNASTAR)26.
Identification of the minimal epitope
To precisely define the minimal B-cell epitope of the DTMUV E protein, we designed and generated fragments corresponding to the roughly mapped epitope. Complementary oligonucleotide primers that were specific for each peptide fragment were designed as previously described27. Nucleotide segments with sticky ends were produced by EcoRI/XhoI digestion and direct annealing. The oligonucleotide fragments were then cloned into the pGEX6p-1 vector (GE Healthcare). The expressed peptides were purified using a GST Purification Kit (TaKaRa). Dot-blot assays were performed by spotting the purified peptide solutions onto a nitrocellulose membrane (Millipore). Approximately 1 μg of each synthesized peptide diluted with TNE buffer was spotted onto the membrane and incubated with the 3B6 mAb (diluted 1:2000 in PBS) or with duck DTMUV anti-serum (1:150 in PBS) at 37 °C for 1 hour. After washing three times with PBST, the membrane was probed with either HRP-conjugated goat anti–mouse IgG (1:500 dilution, KPL) or HRP-conjugated goat anti–duck IgG (1:500 dilution, KPL) at 37 °C for 1 hour. Western blots were performed by electrophoresis of purified GST-peptides in 10% polyacrylamide gels and transfer to nitrocellulose membranes. Membranes were incubated with duck anti–DTMUV serum diluted 1:150 in PBST, followed by reaction with HRP-conjugated goat anti–duck IgG (1:500 dilution, KPL) for 90 min at room temperature.
Epitope cross-reactivity to ZIKV-, YFV-, WNV-, JEV-, and DENV-positive sera
Epitope cross-reactivity to sera infected with other flaviviruses was determined by dot blot, as described above. Briefly, approximately 1 μg of each synthesized epitope peptide diluted with TNE buffer was spotted onto a nitrocellulose membrane and incubated with ZIKV-, YFV-, WNV-, JEV-, and DENV-positive sera at 37 °C for 1 hour. After washing three times with PBST, the membrane was probed with a species-specific HRP-conjugated IgG (KPL) at 37 °C for 1 hour.
Protein E modeling and prediction
To ascertain the location of the epitope within the 3D molecular structure of the DTMUV E protein, we performed in silico modeling. Because the DTMUV E protein shares 62% sequence identity with that of JEV, we used the crystal structure of JEV E protein (PDB ID: 3P54)28 as a modeling template using MODELLER software29. ProSA30 and PROCHECK31 software was used to validate the stereochemical quality of the final model. GlycoEP software32 and the NGlycPred algorithm33 were used to predict the N-glycosylation sites on the DTMUV E protein. The final structure was visualized and analyzed with PyMOL (v1.5.0.4, Schrodinger)29.
Additional Information
How to cite this article: Li, C. et al. Identification of a New Broadly Cross-reactive Epitope within Domain III of the Duck Tembusu Virus E Protein. Sci. Rep. 6, 36288; doi: 10.1038/srep36288 (2016).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Lindenbach, B. D., Thiel, H. J. & Rice, C. M. Flaviviridae: the viruses and their replication. In Fields Virology, 5th edn (eds Knipe, D. M., Howley, P. M., Griffin, D. E., Lamb, R. A., Martin, M. A., Roizman, B., Straus, S. E. ) 1101–1152 (Lippincott, Williams & Wilkins 2007).
Su, J. et al. Duck egg drop syndrome caused by BYD virus, a new Tembusu related virus. PLoS One. 6, e18106, doi: 10.1371/journal.pone.0018106 (2011).
Liu, M. et al. Complete genomic sequence of duck flavivirus from China. J Virol. 86, 3398–3399 (2012).
Cao, Z. et al. Tembusu virus in ducks, China. Emerg. Infect Dis. 17, 1873–1875 (2011).
Liu, M. et al. Adapted Tembusu-like virus in chickens and geese in China. J Clin Microbiol. 50, 2807–2809 (2012).
Yan, P. et al. An infectious disease of ducks caused by a newly emerged Tembusu virus strain in mainland China. Virology 417, 1–8 (2011).
Li, X. et al. Development of a Blocking ELISA for Detection of Serum Neutralizing Antibodies against Newly Emerged Duck Tembusu Virus. PLoS ONE. 7, e53026 (2012).
Bai, X. et al. Molecular characterization of a duck Tembusu virus from China. Virus Genes 47, 478–482 (2013).
Crill, W. D. & Chang, G. J. Localization and characterization of flavivirus envelope glycoprotein cross-reactive epitopes. J Virol. 78, 13975–13986 (2004).
Rey, F. A., Heinz, F. X., Mandl, C., Kunz, C. & Harrison, S. G. The envelope glycoprotein from tick-borne encephalitis virus at 2 A° resolution. Nature 375, 291–298 (1995).
Goncalvez, A. P., Purcell, R. H. & Lai, C. J. Epitope determinants of a chimpanzee Fab antibody that efficiently cross-neutralizes dengue type 1 and type 2 viruses map to inside and in close proximity to fusion loop of the dengue type 2 virus envelope glycoprotein. J Virol. 78, 12919–12928 (2004).
Oliphant, T. et al. Antibody recognition and neutralization determinants on domains I and II of West Nile virus envelope protein. J Virol. 80, 12149–12159 (2006).
Stiasny, K., Kiermayr, S., Holzmann, H. & Heinz, F. X. Cryptic properties of a cluster of dominant flavivirus cross-reactive antigenic sites. J Virol. 80, 9557–9568 (2006).
Oliphant, T. et al. Development of a humanized monoclonal antibody with therapeutic potential against West Nile virus. Nat Med. 11, 522–530 (2005).
Roehrig, J. T., Bolin, R. A. & Kelly, R. G. Monoclonal antibody mapping of the envelope glycoprotein of the dengue 2 virus, Jamaica. Virology 246, 317–328 (1998).
Gromowski, G. D., Barrett, N. D. & Barrett, A. D. Characterization of dengue virus complex-specific neutralizing epitopes on envelope protein domain III of dengue 2 virus. J Virol. 82, 8828–8837 (2008).
Dejnirattisai, W. et al. Cross-reacting antibodies enhance dengue virus infection in humans. Science 328, 745–748 (2010).
Scott, A. S. et al. Human monoclonal antibodies derived from memory B cells following live attenuated dengue virus vaccination or natural infection exhibit similar characteristics. J Infect Dis. 207, 1898–908 (2013).
Burke, D. S., Nisalak, A., Johnson, D. E. & Scott, R. M. A prospective study of dengue infections in Bangkok. Am J Trop Med Hyg. 38, 172–180 (1988).
Wahala, W. M. & Silva, A. M. The human antibody response to dengue virus infection. Viruses 3, 2374–2395 (2011).
Austin, S. K. et al. Structural basis of differential neutralization of DENV-1 genotypes by an antibody that recognizes a cryptic epitope. PLoS Pathol. 8, e1002930, doi: 10.1371/journal.ppat.1002930 (2012).
Fibriansah, G. et al. Structural changes in dengue virus when exposed to a temperature of 37 degrees C. J Virol. 87, 7585–7592 (2013).
Bai, X. et al. Characterization of monoclonal antibodies against duck Tembusu virus E protein: an antigen-capture ELISA for the detection of Tembusu virus infection Arch Virol. 160, 757–764 (2015).
Xue, M. et al. Identification of a conserved B-cell epitope on reticuloendotheliosis virus envelope protein by screening a phage-displayed random peptide library. PLoS One. 7, e49842, doi: 10.1371/journal.pone.0049842 (2012).
Wu, X. et al. Identification of a conserved B-cell epitope on duck hepatitis A type 1 virus VP1 protein. PLoS One. 10, e0118041, doi: 10.1371/journal.pone.0118041 (2015).
Burland, T. G. DNASTAR’s Lasergene sequence analysis software. Meth Mol Biol. 132, 71–91 (2000).
Li, Y. et al. Antigenic analysis monoclonal antibodies against different epitopes of σB protein of Muscovy duck reovirus. Virus Res. 163, 546–551 (2012).
Luca, V. C., AbiMansour, J., Nelson, C. A. & Fremont, D. H. Crystal structure of the Japanese encephalitis virus envelope protein. J Virol. 86, 2337–2346 (2012).
Eswar, N. et al. Comparative protein structure modeling using MODELLER. Curr Protoc Protein Sci 50, 1–31 (2007).
Wiederstein, M. & Sippl, M. J. ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res. 35, 407–410 (2007).
Laskowski, R. A., Macarthur, M. W., Moss, D. S. & Thornton, J. M. PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Cryst. 26, 283–291 (1993).
Chauhan, J. S., Rao, A. & Raghava, G. P. In silico platform for prediction of N-, O- and C-glycosites in eukaryotic protein sequences. PLoS One 8, e67008, doi: 10.1371/journal.pone.0067008 (2013).
Chuang, G. Y. et al. Computational prediction of N-linked glycosylation incorporating structural properties and patterns. Bioinformatics 28, 2249–2255 (2012).
Acknowledgements
This work was funded by China’s Important Research and Developments Program (2016YFD0500106) and the Natural Science Foundation of China (31670153). Yun Zhang supervised and provided the funding for the study.
Author information
Authors and Affiliations
Contributions
M.L. and Y.Z. designed the study. C.L. and X.B. performed experiments. R.M., W.S., Q.Z., R.H. and J.L. analyzed the data. M.L., C.L. and Y.Z. wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Li, C., Bai, X., Meng, R. et al. Identification of a New Broadly Cross-reactive Epitope within Domain III of the Duck Tembusu Virus E Protein. Sci Rep 6, 36288 (2016). https://doi.org/10.1038/srep36288
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep36288
This article is cited by
-
Molecular analysis and serological survey of Tembusu virus infection in Zhejiang, China, 2010-2016
Archives of Virology (2018)
-
Mapping a Type-specific Epitope by Monoclonal Antibody against VP3 Protein of Duck Hepatitis A Type 1 Virus
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.