Abstract
The presenilin-mediated Notch1 cleavage pathway plays a critical role in controlling pancreatic beta cell fate and survival. The aim of the present study was to investigate the role of Notch1 activation in glucotoxicity-induced beta cell impairment and the contributions of miR-375, miR-30a, and miR-34a to this pathway. We found that the protein levels of presenilins (PSEN1 and PSEN2), and NOTCH1 were decreased in INS-1 cells after treatment with increased concentrations of glucose, whereas no significant alteration of mRNA level of Notch1 was observed. Targeting of miR-375, miR-30a, and miR-34a to the 3′utr of Psen1, Psen2, and Notch1, respectively, reduced the amounts of relevant proteins, thereby reducing NICD1 amounts and causing beta cell apoptosis. Overexpression of NICD1 blocked the effects of glucotoxicity as well as miRNA overabundance. Downregulating the expression of miR-375, miR-30a, and miR-34a restored PSEN1, PSEN2, and NICD1 production and prevented glucotoxicity-induced impairment of the beta cells. These patterns of miRNA regulation of the Notch1 cleavage pathway were reproduced in GK rats as well as in aged rats. Our findings demonstrated that miRNA-mediated suppression of NICD1 links the presenilin/Notch1 pathway to glucotoxicity in mature pancreatic beta cells.
Similar content being viewed by others
Introduction
Decreases in pancreatic beta cell function and mass are hallmarks of the progression of type 2 diabetes1,2, with glucotoxicity serving as a critical contributor to beta cell impairment observed in diabetic patients as well as in rodents3,4. Prolonged exposure of beta cells to elevated concentrations of glucose leads to decreases in glucose-stimulated insulin secretion (GSIS), inhibition of insulin gene expression, and induction of beta cell death by apoptosis. These effects of glucotoxicity are thought to involve several mechanisms, including oxidative stress, endoplasmic reticulum (ER) stress, and inflammation5,6,7. Numerous functional genes related to glucotoxicity-induced beta cell failure have been identified, including Neurod1, Pdx1, and Bcl2, but additional genes still require investigation. One of these is the Notch1 gene.
Notch1 is a transmembrane receptor that plays a critical role in controlling cell fate during developmental processes, including those occurring in pancreatic tissues8. Notch1 is activated by interacting with its ligands (Delta or Jagged) located on adjacent cell surfaces and then undergoes intercellular proteolytic cleavage to generate the Notch1 intercellular domain (NICD1), which regulates cell differentiation, proliferation, and apoptosis. The complete cleavage and activation of Notch1 is mediated by the γ-secretase enzyme complex, consisting of presenilins, nicastrin (NCSTN), presenilin enhancer 2 (PEN2), and anterior pharynx-defective 1 (APH1)9,10.
The presenilins are critical catalytic subunits of γ-secretase and are implicated in vesicular trafficking, calcium homeostasis, and the regulation of apoptosis11,12. Their functional role is not well defined, but their presence of both presenilins and γ-secretase has been reported in beta cells12,13. Current evidence indicates that PSEN1 can promote beta cell survival via the cleavage of Notch1 in both adult human and mouse pancreatic islet cells14. Moreover, members of the Notch pathway are upregulated by the cytokine IL-1β in both rat primary islets and INS-1E cells15. The existing evidence therefore supports the retention of an intact presenilin/Notch1 pathway in mature pancreatic beta cells. However, the mechanisms that regulate the presenilin/Notch1 pathway in beta cells remain poorly understood.
One potential regulatory mechanism is via microRNAs (miRNAs). These are endogenous noncoding RNAs (~22 nucleotides) that regulate gene expression by binding to the 3′utr of their target mRNAs, resulting in degradation and/or translational inhibition of potentially hundreds of target mRNAs16. RNA sequencing and expression studies have identified several miRNAs expressed in pancreatic islets, including miR-375, miR-30a, and miR-34a16,17. Genetic studies have shown that miR-375 knockout mice develop hyperglycemia and show reduced beta cell mass18, whereas miR-375 overabundance suppresses GSIS and beta cell survival19,20. MiR-30a has been associated with glucotoxicity-induced defects in insulin secretion17. A role for miR-34a has also been repeatedly reported in both type 1 and type 2 diabetes as well as in age-associated diabetes21,22,23. The effects of miR-375, miR-30a, and miR-34a have been examined in pancreatic islet cells under diabetic conditions, but their contributions to a specific pathway have never been reported. The present study investigated the potential role of miR-375, miR-30a, and miR-34a on regulation of the presenilin/NOTCH1 pathway in mature pancreatic beta cells.
Results
Decreased γ-secretase-mediated Notch1 cleavage by glucotoxicity
An involvement of notch1 signaling in glucotoxicity-induced beta cell impairment was explored by determining the mRNA levels and protein levels of Notch1 and the main components of γ-secretase (Psen1, Psen2, Ncstn, and Psenen) (Fig. 1A,B). The mRNA levels of Notch1, Psen1, and Psen2 were not altered following the 11.1 mmol/l glucose treatment and were only slightly decreased after 24 h exposure to 25 and 33.3 mmol/l glucose. The gene expression levels of Ncstn, Psenen, and Ins2 were significantly decreased in a dose-dependent manner, (except for the alteration in Ncstn expression observed with 11.1 mmol/l glucose) (Fig. 1A). The protein levels were significantly decreased in a dose-dependent manner, with a particularly dramatic decrease observed in the level of NICD1, the active form of Notch1 (Fig. 1B).
Notch1 cleavage was decreased by glucotoxicity.
The mRNA (A) and protein (B) levels were analyzed in INS-1 cells treated for 24 h with different glucose concentrations (mmol/l): 5.5 (control, white bar), 11.1 (black bar), 25 (light grey bar), and 33.3 (heavy grey bar). Quantitative measurements were obtained by gray intensity analysis relative to β-TUBULIN. Relative luciferase activity of the Hes-1 promoter (C) in INS-1 cells treated with 5.5 mmol/l glucose (control) or the indicated glucose concentration. Data shown are means ± SEM and are representative of three separate experiments. *p < 0.05 and **p < 0.01 vs. control.
The effect of high glucose on Notch1 signaling was further examined using a luciferase reporter carrying the active site of Hes1, a major downstream regulator of Notch1. The Hes1 activity was decreased after a 12 h treatment with glucose concentrations greater than 25 mmol/l (Fig. 1C). All the data suggest that Notch1 signaling can be repressed by glucotoxicity.
Decreased Notch1 cleavage mediated glucotoxicity-induced beta cell apoptosis
The glucotoxic condition has been associated with a gradually increased impairment of pancreatic beta cells, characterized as dysfunction in insulin secretion and defects in insulin biosynthesis in both rat primary islets and INS-1 cells (Supplementary Fig. 1). Moreover, prolong exposure of INS-1 cell to high concentrations of glucose enforced cellular apoptosis in a dose- (Fig. 2A) and time- (Fig. 2B) dependent manner.
Inactivation of Notch1 caused beta cell apoptosis.
INS-1 cells were treated with low glucose (5.5 mmol/l glucose, control), or high glucose (11.1, 25, 33.3 mmol/l glucose) for 60 h, followed by PI staining and flow cytometry analysis (A). INS-1 cells were treated with low glucose (5.5 mmol/l glucose, control, white bar), or high glucose (25 mmol/l glucose, black bar) for the indicated times, followed by PI staining and flow cytometry analysis (B). After transient transfection with vector (white bar) or an NICD (black bar) plasmid for 24 h, INS-1 cells were treated with glucose for 24 h when the protein levels of cleaved Notch1 were analyzed (C) and for 72 h the INS-1 cells were stained with propidium iodide (PI) and analyzed by flow cytometry (D). Data shown are means ± SEM and representative of three separate experiments. (A,B) **p < 0.01 vs. control. (D) **p < 0.01 vs. Vector +5.5 mmol/l glucose, and ##p < 0.01 vs. Vector +25 mmol/l glucose.
NICD1 was also overexpressed to restore the cellular level of active Notch1 following its decrease by glucotoxicity (Fig. 2C). The overabundance of NICD1 resulted in a partial, but dramatic, inhibition of glucotoxicity-induced cellular apoptosis (Fig. 2D); however, the recovery of the glucotoxicity-induced insulin secretion defect was not statistically significant (Supplementary Fig. 2). All the data suggest that restoration of NICD1 levels can prevent glucotoxicity-induced apoptosis of beta cells without improvements in defective insulin secretion.
MiR-375 targeted Psen1, miR-30a targeted Psen2, and miR-34a targeted Notch1
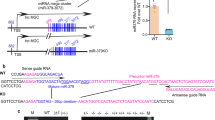
The protein levels of NOTCH1, PSEN1, and PSEN2 following cell culture with 11.1 mmol/l glucose were significantly decreased while that of mRNA levels were not altered suggesting the action of miRNAs. The combined use of bioinformatics prediction softwares (TargetScan and miRanda) and literature screening pointed to miR-375, miR-30a, and miR-34a as potential targets for further exploration. As shown in Fig. 3A,D,G, the miRNA response element (MRE) was well matched between miR-375 and Psen1, between miR-30a and Psen2, and between miR-34a and Notch1. The luciferase activity assays showed that miR-375 inhibited the activity of the wt-Psen1 construct when compared with a vector control, whereas no alteration was observed with the mt-Psen1 construct (Fig. 3B). Neither overexpression nor knockdown of miR-375 altered the Psen1 gene level (data not shown), whereas elevated miR-375 repressed and downregulated miR-375 enhanced PSEN1 protein level, consequently leading to the same protein expression profile for NICD1 (Fig. 3C). No alterations were observed with PSEN2 (Fig. 3C).
Target identification for miRNAs.
The 3′utr sequence of Psen1 (A), Psen2 (D), and Notch1 (G), predicted to include relative miRNA MREs, were aligned and both wt and mt sequences are listed. Wt and mt plasmids were cotransfected with relative miRNAs for 24 h, and luciferase reporter activities were analyzed for miR-375 toward Psen1 (B), miR-30a toward Psen2 (E), and miR-34a toward Notch1 (H). Protein levels were determined following overexpression or knockdown of miR-375 (C), miR-30a (F), or by miR-34a (I). Data shown are means ± SEM and representative of three separate experiments. (B,E,H) **p < 0.01 vs. vector control, and ##p < 0.01 vs. relative wt plasmid.
Luciferase activity assays, qRT-PCR, and western blotting were also performed to verify the relationships between miR-30a and Psen2 and between miR-34a and Notch1. MiR-30a could target to the Psen2 3′utr region (Fig. 3E) and reduce the PSEN2 protein level (Fig. 3F) without impacting its mRNA level (data not shown). The reduction in PSEN2 also caused a reduction in NICD1 (Fig. 3F) without affecting PSEN1 level. Further, miR-34a could regulate the Notch1 3′utr region (Fig. 3H) and reduce the NOTCH1 protein level, without affecting PSEN1 and PSEN2 levels (Fig. 3I). No alteration of Notch1 mRNA level by miR-34a overexpression was observed (data not shown). Taken together, these results reveal that miR-375 targets Psen1, miR-30a targets Psen2, and miR-34a targets Notch1 by regulating the relevant 3′utr regions and repressing their translation, thereby suppressing the NICD1 level in INS-1 cells.
miR-375, miR-30a, and miR-34a mimicked glucotoxicity-induced defects
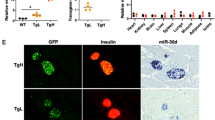
The expression of miR-375, miR-30a and miR-34a was significantly induced by high glucose in both rat pancreatic islets (Fig. 4A) and INS-1 cells (Fig. 4B). The miRNA mimics were introduced to enforce cellular levels of miR-375, miR-30a, and miR-34a in INS-1 cells (Fig. 4C). Elevated levels of miR-375, miR-30a, and miR-34a decreased the K+-stimulated insulin secretion index (KSI) (SEM Fig. 3A), insulin content (SEM Fig. 3B), and insulin gene expression (SEM Fig. 3B), and increased the numbers of apoptotic cells (Fig. 4D). Interestingly, the overabundance of NICD1 resulted in a significant inhibition of miRNAs-induced cellular apoptosis (Fig. 4E–G); however, the recovery of these miRNAs-induced insulin secretion defect was not observed (data not shown). Taken together, these results suggest that miR-375, miR-30a, and miR-34a are capable of mimicking glucotoxicity-induced beta cell impairment and NOTCH1 inactivation associated with these miRNAs-induced beta cell apoptosis.
Effects of miR-375, miR-30a, and miR-34a expression of INS-1 cells.
The expression levels of miRNAs were analyzed by qRT-PCR assays in islets treated with 11.1 (control) and 25 mmol/l glucose for 48 h (A), as well as in INS-1 cells treated with 5.5 (control) and 25 mmol/l glucose for 12 h (B). The expressions of miR-375, miR-30a and miR-34a were determined after 24 h miRNA transfection (C). Cellular apoptosis was analyzed 48 h after transfection (D). A vector (white bar) or NICD1 (black bar) plasmid was co-transfected with the indicated miRNAs or NC mimics for 48 h and the cells were then collected for PI staining and analyzed by flow cytometry (E). A vector or NICD1 plasmid was co-transfected with the indicated miRNAs or NC mimics for 48 h and TUNEL assay was carried out to detect dead cells. Scale bars = 40 μm (F). Quantitative result of TUNEL assay was analyzed. Data were obtained from 8 scans in each group (G). Data shown are means ± SEM and representative of three separate experiments. (A–D) *p < 0.05 or **p < 0.01 vs. control. (E,G) **p < 0.01 vs. NC + vector, and #p < 0.05 or ##p < 0.01 vs. relative miRNA + vector.
Inhibiting miR-375, miR-30a, and miR-34a rescued glucotoxicity-induced defects
The function of these three miRNAs was further clarified by knocking down their expression levels in INS-1 cells using miRNA inhibitors (Fig. 5A). The expression level of miR-375 was downregulated to 22.9%, that of miR-30a was downregulated to 28.1%, and that of miR-34a to 22.7% after anti-miRNAs transfection. The repression of PSEN1 by high glucose was recovered by anti-miR-375, repression of PSEN2 was recovered by anti-miR-30a, and repression of NOTCH1 was recovered by anti-miR-34a; all of these responses resulted in the recovery of NICD1 protein level (Fig. 5B). Moreover, all three anti-miRNAs effectively rescued the apoptosis induction (Fig. 5C) caused by high glucose. These data demonstrated that inhibition of miR-375, miR-30a, and miR-34a can partially rescue the beta cell apoptosis caused by glucotoxicity.
Inhibition of each of the three miRNAs recovered beta cell apoptosis.
INS-1 cells were transfected with anti-NC (white bar, control) or the indicated anti-miRNAs (black bar) for 24 h, and the knockdown efficiency was determined by qRT-PCR (A). Transfected cells were allowed to equilibrate with 5.5 mmol/l glucose medium for 12 h, followed by an additional treatment with 5.5 (control) or 25 mmol/l glucose for the indicated times. After 24 h treatment, protein levels were determined (B). After 48 h treatment, apoptotic cells were analyzed by PI staining and flow cytometry (C). Data shown are means ± SEM and representative of three separate experiments. **p < 0.01 vs. control, and #p < 0.05 or ##p < 0.01 vs. Anti-NC + 25 mmol/l glucose.
Reciprocal regulation between miRNAs and Notch1 signaling in diabetic GK rats and aged Wistar rats
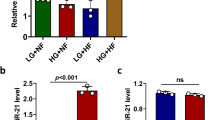
The possibility that this regulation model also exists in vivo was evaluated using primary islets isolated from diabetic GK rats and aged Wistar rats. The random blood glucose levels were higher in GK rats than in age-matched control Wistar rats, confirming that the GK rats were diabetic (Fig. 6A). Analysis of the beta cell specific genes Ins2, Pdx1, Mafa, and Neurod1 confirmed that the mRNA expression were significantly decreased in GK rats (Fig. 6B). The protein levels of PDX1, MAFA, and NEUROD1 were also dramatically reduced in the GK rats (Fig. 6C). The deficiencies in beta cell markers further confirmed a molecular basis for the diabetic phenotype of the GK rats. These GK rats also showed that miR-375 and miR-30a expressions were decreased, while miR-34a expression was significantly increased (Fig. 6D). The mRNA level of Notch1 was increased, while no alterations were observed in Psen1, Psen2, and Ncstn in the GK rats (Fig. 6E). Although the protein levels of PSEN1 and PSEN2 were significantly increased, that of NCSTN, another component of γ-secretase was dramatically decreased (Fig. 6F), suggesting a defective γ-secretase in islets isolated from GK rats. The decreased NOTCH1 protein level (Fig. 6F) further confirmed this defect in the γ-secretase in GK rat islets.
Alterations of genes and miRNAs in GK rats.
Blood glucose levels in Wistar rats (white bar, control) and GK rats (black bar) (A). mRNA (B) and protein (C) levels of genes specific for beta cells. miR-375, miR-30a and miR-34a expression levels (D). mRNA (E) and protein (F) levels of NOTCH1 and components of γ-secretase. Data shown are means ± SEM. *p < 0.05 or **p < 0.01 vs. control. n = 6 per group.
The random plasma insulin levels were significantly lower in aged rats than in young Wistar rats, suggesting an impairment of insulin secretion in islets from aged rats (Fig. 7A). Aged rats also showed defective expressions of beta cell specific markers (Fig. 7B,C). The levels of miR-375 and miR-34a were modestly elevated (Fig. 7D), in parallel with the decreased gene expression levels of components of γ-secretase and Notch1 (Fig. 7E). The protein levels were also significantly decreased (Fig. 7F), indicating a defective γ-secretase in islets from aged rats. Taken together, the data demonstrated that alteration of miRNA expression disrupted the γ-secretase-mediated Notch1 cleavage which was associated with beta cell impairment.
Alterations of genes and miRNAs in aged Wistar rats.
Plasma insulin levels in young rats (white bar, control) and aged rats (black bar) (A). mRNA (B) and protein (C) levels of genes specific in beta cells were examined in islets isolated from young rats (white bar, control) and aged rats (black bar). miR-375 and miR-34a expression levels (D). mRNA (E) and protein (F) levels of NOTCH1 and components of γ-secretase. Data shown are means ± SEM. *p < 0.05 or **p < 0.01 vs. control. n = 6 per group.
Discussion
This study was undertaken to elucidate the regulatory networks operating between miRNAs (miR-375, miR-30a, and miR-34a) and the presenilin/Notch1 pathway in the context of glucotoxicity-induced pancreatic beta cell impairment. Our research generated three major findings. First, we demonstrated that exposure to high glucose levels decreases the amount of NICD1, the active form of Notch1, by disruption of the γ-secretase-mediated Notch1 proteolytic cleavage pathway. Second, miRNAs function at the posttranscriptional level to play key roles in regulating Notch1 signaling. Specifically, miR-375 targets Psen1, miR-30a targets Psen2, and miR-34a targets Notch1 itself, but they all decrease the NICD1 level and inactivate Notch1 signaling. The third main observation is that a reciprocal modulation pattern occurs between miRNAs and the presenilin/Notch1 pathway in diabetic GK rats as well as in aged rats, suggesting that the in vitro regulatory network is likely to be functional in vivo and may contribute to the development of type 2 diabetes.
Glucotoxicity is an important factor that contributes to advancing beta cell failure and development of overt diabetes. We have demonstrated that glucotoxicity induces beta cell failure by inhibition of the γ-secretase-mediated Notch1 cleavage pathway. Early studies of Notch1 signaling focused on the regulation of pancreatic cell fate determination during development24, with later reports showing that Notch1 signaling regulates apoptosis in several mature cell types25,26,27. Interestingly, Darville et al. has pointed that the Notch1 signaling pathway induction by cytokines is likely to contribute to beta cell de-differentiation15. Dror et al. documented that the Notch cleavage pathway mediated by γ-secretase remained intact and functional in mature islets14. Suppression of Notch signaling, as indicated by decreased NICD1 levels, results in an increase in the Bax/Bcl-2 ratio, which consequently leads to cellular apoptosis through the caspase 3-dependent pathway28. An increase in the Bax/Bcl-2 ratio has a well-known association with glucotoxicity-induced apoptosis in several cell types, including beta cells29. Therefore, a decreased level of NICD1 will potentially act as a trigger of glucotoxicity-induced impairment in beta cells.
NICD1, the active form of Notch1, is generated by γ-secretase-mediated proteolytic cleavage10. Therefore, blocking either this cleavage pathway or Notch1 itself results in decreased NICD1 levels. Indeed, the components of the γ-secretase proteolytic complex were significantly decreased by high glucose in our study. The effects of presenilins have been comprehensively studied in genetic Alzheimer disease because of their contribution to the processing of the amyloid precursor protein (APP)30. Diabetes and Alzheimer disease share risk factors, and it is possible that these two diseases may share common molecular defects.
Although presenilins are key mediators in neurodegeneration, their role in beta cell degeneration remains unclear. Previous studies have shown that presenilin levels can be increased by low concentration of glucose (2 mmol/l glucose for PSEN1, and 5 mmol/l glucose for PSEN2), resulting in defective intercellular Ca2+ homeostasis and beta cell death due to low metabolic activity12. Consistent with these findings, the protein levels of presenilins are also most abundant at low glucose levels. However, our results suggest that the decreased levels of presenilins due to high glucose lead to reductive cleavage of Notch1 and ultimately to beta cell apoptosis.
The protein levels of presenilins and NOTCH1 are significantly decreased by high glucose, and yet their mRNA levels are only slightly altered, supporting a role for miRNAs as the key regulators underlying the deficiency in the Notch1 signaling pathway. Previously, upregulation of miR-375 and miR-30a expression by glucotoxicity has been reported in rat islets and INS-1 cells17. Elevated serum levels of miR-375 and miR-30a are also observed in diabetic patients31,32. In the present study, miR-375 expression increased initially but was significantly decreased by high glucose over time. The alteration in this expression pattern for miR-375 is related to the transcriptional activity of PDX1, since miR-375 is one of its target genes33. Several reports have demonstrated that miR-34a expression is increased in primary islets from high-fat-diet (HFD) mice and genetic diabetic db/db mice, as well as in beta cells treated with cytokines and palmitate16,23. Elevated levels of miR-34a were also observed in the present study in INS-1 cells stimulated by high glucose and in primary islets from diabetic GK rats and aged rats. The regulatory network between miR-34a and p53 has been confirmed by several groups34,35,36. The transcriptional activity of p53 is promoted by high glucose in INS-1 cells (Our unpublished data). Therefore, the elevated levels of miR-34a observed are likely an effect of triggering of p53 activity by high glucose in our study, as miR-34a is a conventional target gene of p53.
The present study and other studies have shown that elevation of miR-375, miR-30a, or miR-34a individually results in beta cell dysfunction, decreases cell survival, and increases apoptosis, thereby mimicking the cellular phenotype provoked by glucotoxicity. A combination of literature screening, bioinformatics analysis, and experimental assays confirms that miR-375, miR-30a, and miR-34a regulate NICD1 protein expression via diverse targets involving γ-secretase-mediated Notch1 cleavage. However, Notch1 signaling is only associated with glucotoxicity-induced beta cell apoptosis and its effects are unlike those evoked by miR-375, miR-30a, and miR-34a.
These differences are likely to result from the differences in the target proteins they modulate. Indeed, previous research has elucidated that miR-375 regulates the expression of MTPN and PDK1 proteins19,20, while Kim et al. have claimed that miR-30a silencing of NEUROD1 expression is an important initial event of glucotoxicity-induced beta cell dysfunction17. MiR-34a targets VAMP2 and BCL2 expression, leading to beta cell damage23,37. In the present study, we showed that miR-375 targets Psen1, miR-30a targets Psen2, and miR-34a targets Notch1, all of which are newly identified targets that converge onto one signaling pathway. To our knowledge, this effective modulation network has never been reported before.
Previous studies have elucidated the role of single miRNAs regulating different targets. For instance, we have reported that miR-24 synchronously regulates four MODY (Mature Onset of Diabetes of the Young) genes16. Other reports have shown the regulation of one target by two or more miRNAs. Guttilla et al. have shown that FOXO1 can be coordinately regulated by miR-27a, miR-96, and miR-18238. Nevertheless, our study is the first to demonstrate that multiple miRNAs hierarchically regulate one pathway through diverse targets. This regulatory network would somehow insure that the cells receive the correct signal for committing to a cellular response. The use of INS-1 cells might raise a cautionary concern in interpreting our results, but a reciprocal correlation was noted in vivo between miRNAs and NICD1 in both diabetic GK rats and aged rats.
In conclusion, we have shown that miR-375, miR-30a, and miR-34a orchestrate the regulation of γ-secretase-mediated Notch1 cleavage in a glucotoxicity setting in pancreatic beta cells. This miRNA/Notch1 signaling pathway is likely to be related to beta cell impairment in mature pancreatic islets in rodents. This study provides a more comprehensive view of miRNAs, broadens our understanding of the pathophysiological development of diabetes, and identifies a fine-tuned regulatory network of miRNAs in disease.
Materials and Methods
Isolation of pancreatic islets
All animal studies performed were approved by the Research Animal Care Committee of Nanjing Medical University. Animals were treated humanely, using approved procedures in accordance with the guidelines of the Institutional Animal Care and Use Committee at Nanjing Medical University, China. Male Sprague-Dawley (SD) rats were purchased from Shanghai Laboratory Animal Centre (Chinese Academy of Sciences, China). Male Goto-Kakizaki (GK) and Wistar rats were purchased from Cavens Laboratory Animal Co., Ltd. (Chang Zhou, China). Islet isolation and culturing techniques have been described previously16.
Cell culture
The rat insulinoma INS-1 cell line was cultured in RPMI-1640 (Invitrogen, Carlsbad, CA) supplemented with 10% FBS, 1% penicillin-streptomycin, 1 mM pyruvic acid sodium, 10 mM HEPES, and 0.05 mM β-mercaptoethanol (Sigma-Aldrich, St. Louis, MO) and incubated in a Thermo tissue-culture incubator at 37° with a 95% O2/5% CO2 atmosphere. Cells between passages 16 and 32 were used for experiments39.
Plasmid construction
The primers for generating wide-type (wt) and mutant (mt) 3′utr-luciferase constructs of Psen1, Psen2, and Notch1 are given in Supplementary information. To generate the wide-type (wt) and mutant (mt) 3′UTR-luciferase constructs of Psen11, Psen2 and Notch1, the primers in Supplementary Table 1 were designed and synthesized by Generay Biotech Co., Ltd (Shanghai, China). Primers were annealed by incubating at 95° for 5 min, then heated at 70° for 10 min and allowed to cool down to room temperature. The generated double-stranded oligonucleotides were treated with T4 polynucleotide kinase (Promega) by incubating at 37° for 30 min, then heated at 70° for 10 min and transfered on ice. Wt or mt sequences were subcloned into the SpeI and HindIII sites of the pMIR-REPORT vector.
Transient transfection and luciferase reporter assay
INS1 cells (1 × 104 cells/well) were cultured in 48-well plates and co-transfected with luciferase reporter plasmids (either a WT or mutant plasmid) and miRNAs or the NC (negative control) mimics (GenePharma, Shanghai, China) using Lipofecatamine 2000 (Invitrogen, Carlsbad, CA) according to the manufacturer’s instructions. Twenty-four hours after transient transfection, the cells were harvested for dual luciferase reporter assays as described previously. The Firefly luciferase activity was normalized with the Renilla activity of the PRL-CMV plasmid (Promega, Madison, WI).
Insulin secretion assay
Insulin secretion was determined in INS-1 cells and islets from SD rats. INS-1 cells (1 × 104 cells/well) and isolated islet from SD rats (8 islets/well) in 48-well plates were cultured in the presence of different concentration of glucose (5.5, 11.1, 25 and 33.3 mmol/l) for 24 h in INS-1 cells and for 72 h in islets. INS1 cells were used for potassium-stimulated insulin secretion (KSIS) assays, islets were used for GSIS assays. The insulin levels were measured using a [125I] Insulin Radioimmunoassay Kit (North Institute of Biological Technology, Beijing, China)40.
Flow cytometry analysis
INS1 cells (1 × 106 cells/well) were cultured in 35 mm dishes and treated as described previously. The cells were harvested and fixed with 1 mL 75% ice-cold ethanol at −20° overnight. Cells were stained with propidium iodide (PI) solution (50 mg/ml) containing RNase (25 mg/ml) for 30 min in the dark at 37° and analyzed by using a FACSCalibur Flow Cytometer (Becton Dickinson, San Jose, CA).
TUNEL assay
A TUNEL assay was performed with the commercially available in situ cell apoptosis detection kit (Vazyme, Nanjing, china), according to the manufacturer’s instructions. The TUNEL signals were observed with laser scanning confocal microscope (FV1200, Olympus, Japan). In each sample, eight scans were selected and the cells positive with TUNEL staining (green signals) and hoechst33342 staining (blue signals) were counted. Quantitative assessment of apoptosis was determined as a ratio of the number of TUNEL-positive cells to total nuclear number in each field.
qRT-PCR analysis
Total RNA was extracted using Trizol reagent (Invitrogen). For miRNA quantitation, stem-loop primers designed to have a short single-stranded part that is complementary to the 3′-end of miRNA, a double-stranded part (the stem) and the loop that contains the universal primer-binding sequence were used as reverse primers. For mRNA determination, oligo-dT was used as reverse primer. Then, first-strand cDNA synthesis was performed using 1 μg of total RNA (Promega). All Reverse transcriptase reactions included no-template controls and RT minus controls. Quantitative RT-PCR was performed using the SYBR Green PCR Master Mix and LighteCycler480 Sequence Detection System (Roche). MiRNAs were normalized with u6 and mRNA was normalized with β-actin. Primers for quantitative RT-PCR were available in Supplemental Table 2.
Western blot analysis
Cells or primary islets were lysed with ice-cold lysis buffer containing: 50 mmol/l Tris-HCl, pH 7.4, 1% NP-40, 150 mmol/l NaCl, 1 mmol/l EDTA, 1 mmol/l phenylmethylsulphonyl fluoride, and complete proteinase inhibitor (one tablet/10 ml; Roche). After protein content determination, western blot was performed as described before. Individual immunoblots were probed with antibodies to NOTCH1, NCSTN, PEN2, PSEN1 and PSEN2 (Cell Signalling, Danvers, MA) diluted 1:1000; to PDX1 (Upstate) diluted 1:3000; to MAFA (Santa cruz,) diluted 1:800; to NEUROD1 (abcam) diluted 1:3000. β-tubulin antibody (Sigma-Aldrich, St. Louis, MO) diluted 1:5000 was used as a standard.
Statistical analysis
Comparisons were performed using the Student’s t test between two groups or ANOVA in multiple groups. The results are presented as means ± SEM. A p value of less than 0.05 was considered to be statistically significant.
Additional Information
How to cite this article: Li, Y. et al. A Presenilin/Notch1 pathway regulated by miR-375, miR-30a, and miR-34a mediates glucotoxicity induced-pancreatic beta cell apoptosis. Sci. Rep. 6, 36136; doi: 10.1038/srep36136 (2016).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Hara, M., Fowler, J. L., Bell, G. I. & Philipson, L. H. Resting beta-cells - A functional reserve? Diabetes & metabolism, doi: 10.1016/j.diabet.2016.01.001 (2016).
Esterhazy, D. et al. Bace2 is a beta cell-enriched protease that regulates pancreatic beta cell function and mass. Cell metabolism 14, 365–377, doi: 10.1016/j.cmet.2011.06.018 (2011).
Chen, F. et al. Transcription factor Ets-1 links glucotoxicity to pancreatic beta cell dysfunction through inhibiting PDX-1 expression in rodent models. Diabetologia 59, 316–324, doi: 10.1007/s00125-015-3805-3 (2016).
Masini, M. et al. Prevention by metformin of alterations induced by chronic exposure to high glucose in human islet beta cells is associated with preserved ATP/ADP ratio. Diabetes research and clinical practice 104, 163–170, doi: 10.1016/j.diabres.2013.12.031 (2014).
Qureshi, F. M., Dejene, E. A., Corbin, K. L. & Nunemaker, C. S. Stress-induced dissociations between intracellular calcium signaling and insulin secretion in pancreatic islets. Cell calcium 57, 366–375, doi: 10.1016/j.ceca.2015.03.002 (2015).
Wali, J. A. et al. The proapoptotic BH3-only proteins Bim and Puma are downstream of endoplasmic reticulum and mitochondrial oxidative stress in pancreatic islets in response to glucotoxicity. Cell death & disease 5, e1124, doi: 10.1038/cddis.2014.88 (2014).
Maedler, K. et al. Glucose-induced beta cell production of IL-1beta contributes to glucotoxicity in human pancreatic islets. The Journal of clinical investigation 110, 851–860, doi: 10.1172/JCI15318 (2002).
Apelqvist, A. et al. Notch signalling controls pancreatic cell differentiation. Nature 400, 877–881, doi: 10.1038/23716 (1999).
De Strooper, B. & Annaert, W. Novel research horizons for presenilins and gamma-secretases in cell biology and disease. Annual review of cell and developmental biology 26, 235–260, doi: 10.1146/annurev-cellbio-100109-104117 (2010).
Zhang, X., Li, Y., Xu, H. & Zhang, Y. W. The gamma-secretase complex: from structure to function. Frontiers in cellular neuroscience 8, 427, doi: 10.3389/fncel.2014.00427 (2014).
Kim, T. W., Pettingell, W. H., Jung, Y. K., Kovacs, D. M. & Tanzi, R. E. Alternative cleavage of Alzheimer-associated presenilins during apoptosis by a caspase-3 family protease. Science 277, 373–376 (1997).
Dror, V. et al. Glucose and endoplasmic reticulum calcium channels regulate HIF-1beta via presenilin in pancreatic beta-cells. The Journal of biological chemistry 283, 9909–9916, doi: 10.1074/jbc.M710601200 (2008).
Figueroa, D. J., Shi, X. P., Gardell, S. J. & Austin, C. P. Abetapp secretases are co-expressed with Abetapp in the pancreatic islets. Journal of Alzheimer’s disease: JAD 3, 393–396 (2001).
Dror, V. et al. Notch signalling suppresses apoptosis in adult human and mouse pancreatic islet cells. Diabetologia 50, 2504–2515, doi: 10.1007/s00125-007-0835-5 (2007).
Darville, M. I. & Eizirik, D. L. Notch signaling: a mediator of beta-cell de-differentiation in diabetes? Biochemical and biophysical research communications 339, 1063–1068, doi: 10.1016/j.bbrc.2005.11.111 (2006).
Zhu, Y. et al. MicroRNA-24/MODY gene regulatory pathway mediates pancreatic beta-cell dysfunction. Diabetes 62, 3194–3206, doi: 10.2337/db13-0151 (2013).
Kim, J. W. et al. miRNA-30a-5p-mediated silencing of Beta2/NeuroD expression is an important initial event of glucotoxicity-induced beta cell dysfunction in rodent models. Diabetologia 56, 847–855, doi: 10.1007/s00125-012-2812-x (2013).
Poy, M. N. et al. miR-375 maintains normal pancreatic alpha- and beta-cell mass. Proceedings of the National Academy of Sciences of the United States of America 106, 5813–5818, doi: 10.1073/pnas.0810550106 (2009).
Poy, M. N. et al. A pancreatic islet-specific microRNA regulates insulin secretion. Nature 432, 226–230, doi: 10.1038/nature03076 (2004).
El Ouaamari, A. et al. miR-375 targets 3′-phosphoinositide-dependent protein kinase-1 and regulates glucose-induced biological responses in pancreatic beta-cells. Diabetes 57, 2708–2717, doi: 10.2337/db07-1614 (2008).
Tugay, K. et al. Role of microRNAs in the age-associated decline of pancreatic beta cell function in rat islets. Diabetologia, doi: 10.1007/s00125-015-3783-5 (2015).
Nesca, V. et al. Identification of particular groups of microRNAs that positively or negatively impact on beta cell function in obese models of type 2 diabetes. Diabetologia 56, 2203–2212, doi: 10.1007/s00125-013-2993-y (2013).
Roggli, E. et al. Involvement of microRNAs in the cytotoxic effects exerted by proinflammatory cytokines on pancreatic beta-cells. Diabetes 59, 978–986, doi: 10.2337/db09-0881 (2010).
Murtaugh, L. C., Stanger, B. Z., Kwan, K. M. & Melton, D. A. Notch signaling controls multiple steps of pancreatic differentiation. Proceedings of the National Academy of Sciences of the United States of America 100, 14920–14925, doi: 10.1073/pnas.2436557100 (2003).
Jena, N. et al. CDK6-mediated repression of CD25 is required for induction and maintenance of Notch1-induced T-cell acute lymphoblastic leukemia. Leukemia, doi: 10.1038/leu.2015.353 (2015).
Jundt, F. et al. Activated Notch1 signaling promotes tumor cell proliferation and survival in Hodgkin and anaplastic large cell lymphoma. Blood 99, 3398–3403 (2002).
Kong, R. et al. Silencing NACK by siRNA inhibits tumorigenesis in non-small cell lung cancer via targeting Notch1 signaling pathway. Oncology reports, doi: 10.3892/or.2016.4552 (2016).
Gao, F. et al. Notch pathway is involved in high glucose-induced apoptosis in podocytes via Bcl-2 and p53 pathways. Journal of cellular biochemistry 114, 1029–1038, doi: 10.1002/jcb.24442 (2013).
Kim, W. H. et al. Exposure to chronic high glucose induces beta-cell apoptosis through decreased interaction of glucokinase with mitochondria: downregulation of glucokinase in pancreatic beta-cells. Diabetes 54, 2602–2611 (2005).
Xia, W. Role of presenilin in gamma-secretase cleavage of amyloid precursor protein. Experimental gerontology 35, 453–460 (2000).
Nielsen, L. B. et al. Circulating levels of microRNA from children with newly diagnosed type 1 diabetes and healthy controls: evidence that miR-25 associates to residual beta-cell function and glycaemic control during disease progression. Experimental diabetes research 2012, 896362, doi: 10.1155/2012/896362 (2012).
Higuchi, C. et al. Identification of circulating miR-101, miR-375 and miR-802 as biomarkers for type 2 diabetes. Metabolism: clinical and experimental 64, 489–497, doi: 10.1016/j.metabol.2014.12.003 (2015).
Keller, D. M. et al. Characterization of pancreatic transcription factor Pdx-1 binding sites using promoter microarray and serial analysis of chromatin occupancy. The Journal of biological chemistry 282, 32084–32092, doi: 10.1074/jbc.M700899200 (2007).
Lovis, P. et al. Alterations in microRNA expression contribute to fatty acid-induced pancreatic beta-cell dysfunction. Diabetes 57, 2728–2736, doi: 10.2337/db07-1252 (2008).
Raver-Shapira, N. et al. Transcriptional activation of miR-34a contributes to p53-mediated apoptosis. Molecular cell 26, 731–743, doi: 10.1016/j.molcel.2007.05.017 (2007).
Sen, N., Satija, Y. K. & Das, S. PGC-1alpha, a key modulator of p53, promotes cell survival upon metabolic stress. Molecular cell 44, 621–634, doi: 10.1016/j.molcel.2011.08.044 (2011).
Lin, X. et al. Downregulation of Bcl-2 expression by miR-34a mediates palmitate-induced Min6 cells apoptosis. Journal of diabetes research 2014, 258695, doi: 10.1155/2014/258695 (2014).
Guttilla, I. K. & White, B. A. Coordinate regulation of FOXO1 by miR-27a, miR-96, and miR-182 in breast cancer cells. The Journal of biological chemistry 284, 23204–23216, doi: 10.1074/jbc.M109.031427 (2009).
Merglen, A. et al. Glucose sensitivity and metabolism-secretion coupling studied during two-year continuous culture in INS-1E insulinoma cells. Endocrinology 145, 667–678, doi: 10.1210/en.2003-1099 (2004).
Chen, F. et al. Resveratrol prevents interleukin-1beta-induced dysfunction of pancreatic beta-cells. Journal of biomedical research 24, 381–388, doi: 10.1016/S1674-8301(10)60051-6 (2010).
Acknowledgements
This study was supported by research grants from (1) the National Natural Science Foundation of China (Nos 81420108007 and 81130013) to XH, (2) the National Natural Science Foundation of China (81670703) and China Postdoctoral Science Foundation (2016M590479) to YZ. XH is a Fellow at the Collaborative Innovation Center for Cardiovascular Disease Translational Medicine of Jiangsu Province.
Author information
Authors and Affiliations
Contributions
Yunxia Zhu and X.H. designed research; Yunxia Zhu, Y.L., T.Z., Yuncai Zhou, Y.S. and Y.C. performed research; X.C. contributed new reagents; Yunxia Zhu and X.H. analyzed data and wrote the paper. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Li, Y., Zhang, T., Zhou, Y. et al. A Presenilin/Notch1 pathway regulated by miR-375, miR-30a, and miR-34a mediates glucotoxicity induced-pancreatic beta cell apoptosis. Sci Rep 6, 36136 (2016). https://doi.org/10.1038/srep36136
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep36136
This article is cited by
-
A reduced proteomic signature in critically ill Covid-19 patients determined with plasma antibody micro-array and machine learning
Clinical Proteomics (2024)
-
Comparative study of microRNA profiling in one Chinese Family with PSEN1 G378E mutation
Metabolic Brain Disease (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.