Abstract
Genome-wide studies on high-altitude adaptation have received increased attention as a classical case of organismal evolution under extreme environment. However, the current genetic understanding of high-altitude adaptation emanated mainly from autosomal analyses. Only a few earlier genomic studies paid attention to the allosome. In this study, we performed an intensive scan of the X chromosome of public genomic data generated from Tibetan Mastiff (TM) and five other dog populations for indications of high-altitude adaptation. We identified five genes showing signatures of selection on the X chromosome. Notable among these genes was angiomotin (AMOT), which is related to the process of angiogenesis. We sampled additional 11 dog populations (175 individuals in total) at continuous altitudes in China from 300 to 4,000 meters to validate and test the association between the haplotype frequency of AMOT gene and altitude adaptation. The results suggest that AMOT gene may be a notable candidate gene for the adaptation of TM to high-altitude hypoxic conditions. Our study shows that X chromosome deserves consideration in future studies of adaptive evolution.
Similar content being viewed by others
Introduction
Adaptation to high-altitude environments, characterised by extreme conditions like hypoxia, low temperature, and high ultraviolet radiation, has been of great interest in evolutionary biology research1,2,3,4,5,6. Numerous candidate genes which may be contributing to high-altitude adaptation, for instance those related to hypoxia (e.g., EPAS1, EGLN1, PPARA and HBB identified from human populations), energy metabolism (e.g., PKLR, ENO3 and DNAH9 identified from non-human highland animals) and DNA repair (e.g., BCL3, ERCC4 and ERCC6 identified from non-human highland animals), have been identified from studies of native highlanders with different time depths of exposure to high-altitude. These included human populations (e.g., Tibetans, Andeans and Ethiopians)1,3,5,6,7,8,9,10, wild animals (e.g., Tibetan antelopes, Tibetan wild boars, Andean waterfowls, Gray wolves, ground tits and the snub-nosed monkeys)2,11,12,13,14,15,16 and domesticated animals (e.g., Tibetan Mastiffs, Tibetan chickens and yaks)4,17,18,19,20.
Notably, most of the current understanding of genetic basis of high-altitude adaptation is based on selective scans of autosomes, despite the chromosomes of diploid organisms usually harbouring both autosomes and allosomes. Relative to autosomes, X chromosome shows reduced population size and hemizygote state in male individuals. These characters can make it be more sensitive to demographic history and selective effects21,22. Selective scans on X chromosome require a rigorous analysis design to exclude any noise caused by demographic factors from real selective signals. To date, only a few genomic studies have analysed the allosomes of highland organisms8. No detailed and systematic studies of allosomes in relation to high-altitude adaptation have been reported. A recent genome-wide study of Drosophila melanogaster populations revealed that some X-linked genes had undergone positive selection and were responsible for hypoxia tolerance23. Therefore, a detailed study of X chromosome, with respect to the genetic mechanism underlying the adaptation to high-altitude, is of great significance. This knowledge will boost the understanding of the complex biological feature of high-altitude adaptation.
Recently, the high-altitude adaptation of Tibetan Mastiff (TM), an ancient dog breed that migrated to the Tibetan Plateau (typically 4,500 m) with humans, has received wide attention4,17,18,24. Assessment of autosomal single nucleotide polymorphisms (SNPs) identified candidate genes, whose associated functions signified adaptation to high-altitude hypoxia. Here, we performed an X-chromosome-wide scan to investigate the adaptive strategies of the TM. By reanalysing publicly available genomic data from a TM population (10 individuals) and five other dog populations (50 individuals) from lower altitudes17, we identified five genes showing signatures of selection (ALG13, AMOT, DCX, LHFPL1 and TRPC5) on the X chromosome of TM population. Notable among these genes is angiomotin (AMOT) gene, which was indicated to regulate the process of angiogenesis25,26,27,28. We then sampled additional 11 indigenous Chinese dog populations (175 individuals in total) from continuous altitudes ranging from 300 to 4,000 m to validate and test the association between the haplotype frequency of AMOT gene and altitude adaptation. The results corroborated the initial findings that AMOT gene may be an important target for TM’s adaptation to hypoxic conditions at their high-altitude habitat.
Results
Summary of mapping and SNPs calling
In the present study, the publicly available genome-wide re-sequencing data sets from six populations of 60 dogs, including a Tibetan Mastiff (TM) population (10 individuals; 4380 m altitudes) and five other dog populations (Diqing indigenous dogs, DQ; Lijiang indigenous dogs, LJ; Kunming dogs, KM; German Shepherds, GS and Yingjiang indigenous dogs, YJ; 50 individuals in total from 800–3300 m altitudes), were used17. Of the six dog populations, KM and GS are working dogs and the other four are indigenous dogs in China (Supplementary Tables S1 and S2).
The average raw reads of each individual from the six populations was 383, 754, 395 and the average mapping ratio was 99.73%. After filtering out duplicated reads and low-quality bases, an average of 33.96G mapped bases (~14.19 and ~11.38 folds coverage for autosomes and X chromosome respectively) for each dog were obtained (Supplementary Table S3). After SNP calling and quality control, we obtained 435, 757 high-quality SNPs from X chromosome as well as about 14.3 M from autosomes for the six populations. Compared to autosomes, X chromosome had lower SNP density (112 in X chromosome vs. 207 in autosomes per 50 kb) and transition/transversion (Ts/Tv) ratio (1.76 in X chromosome vs. 2.15 in autosomes) (Supplementary Fig. S1; Supplementary Tables S4 and S5). Based on annotations, most of the variants on X chromosome were located in the intergenic regions (about 76%), and only about 1.4% in exon regions (Supplementary Table S6). In subsequent selective scans, only the variants (435, 757 SNPs) from X chromosome were used.
Genetic backgrounds of the six dog populations
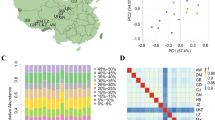
The genetic diversity or pairwise nucleotide diversity29 (π) assessment showed that autosomes have higher levels of genetic diversity relative to X chromosome across all the six dog populations (1.0E-3 in X chromosome vs. 1.3E-3 in autosomes) (Fig. 1). Among these populations, the two working dog populations, i.e., KM and GS, were excluded from further selective analyses for four main reasons. First, KM and GS populations showed the lowest genetic diversity values in both autosomes and X chromosome compared to other populations, implying likely serious genetic bottlenecks faced by the two modern breeds. This finding is similar to that reported from autosomes analyses17. Second, principle components analysis (PCA) results based on X chromosome showed that the six populations separated into two major groups, one group included KM and GS, which are modern dog breeds, and the other group included TM, DQ, LJ and YJ, which are ancient breed and indigenous dogs in China. Therefore, the population structure estimations of X chromosome indicated that KM and GS have significantly different genetic backgrounds relative to the other four dog populations (Supplementary Fig. S2). Third, we performed phylogenetic analyses of the autosomes data sets and observed an evident split separating East Asian and European dogs (Supplementary Fig. S3). Interestingly, this kind of divergence was also reported by Frantz et al.30. In our present tree, KM and GS populations clustered together with the Western Eurasian core group. In contrast with the two modern breeds, the four dog populations (TM, DQ, LJ and YJ) showed closer relationships with East Asian core group. Fourth, from our Fst assessment of all the autosomal SNPs among different paired population comparisons (GS vs. YJ, GS vs.TM, KM vs. YJ, KM vs. TM and TM vs. YJ), we found that the introduction of GS or KM both resulted in a significantly higher Fst level as compared to TM vs. YJ pair (P < 2.2E-16) (Supplementary Fig. S4). These results may reflect the significant genetic difference between Chinese and European dogs, indicating that dogs of GS and KM belong to European lineage while dogs of TM and YJ belong to East Asia Linage. Taking all evidence together, as Tibetan Mastiff is a Chinese aboriginal breed, to avoid bias from the divergent genetic backgrounds of Chinese and European dogs, KM and GS populations were excluded from further selective analyses.
Boxplot of pairwise nucleotide diversity (π) for the six dog populations from Gou et al.17.
Genetic diversity surveys were carried out on each population with non-overlapping 100 k windows across the whole X chromosome. The divergence level of nucleotide diversity on X chromosome between four dog populations and two working breeds was assessed using T-test.
Selective footprints of X chromosome in Tibetan Mastiff
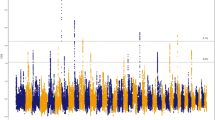
The X-chromosome-wide selection scans were conducted on TM with the highest altitude of 4,380 m and on YJ population with the lowest altitude of 800 m. Based on a total of 306,678 SNPs from the two populations, using the threshold of 1% top level (equivalent to 0.68 and 1.96 after FDR correction for Fst test and Fisher’s exact test respectively), we identified 3869 (~1.2% of total SNPs) and 4049 (~1.3% of total SNPs) SNPs as outliers (Fig. 2a,b). According to the dog genome annotation (Canfam3.1)31, 34 and 36 genes were identified from the two respective SNPs sets. An overlap was observed between the two sets for a total of 32 genes. Moreover, 24 of these genes were characterised as protein-coding genes (Supplementary Table S7). We calculated the densities of outlier SNPs located in the 24 genes and found that six genes (ALG13, AMOT, DCX, LHFPL1, TRPC5 and ZCCHC16) showed highest densities (Fig. 2c). Furthermore, the six genes were located continuously on X chromosome, and occupied a large region (~1.4 M bp). ALG13, DCX, TRPC5 and ZCCHC16 have functions in brain or neuron developments and cognitive competence32,33,34,35, whereas the functions of LHFPL1 are presently unknown. Interestingly, AMOT (~47 kb) has been reported to code for angiomotin protein, which is located on the surface of vascular endothelial cells (VECs), and has been thought to be crucial for angiogenesis25,26,27,28.
Selection scans on X chromosome for TM and YJ populations.
(a) Fst test on X chromosome. The cutoff was set at 0.67 (1% top level). Points representing outlier values are coloured red; (b). Fisher’s exact test on X chromosome. The cutoff was set at 1.96 (P ≈ 0.01, after FDR corrections). Points with outlier values are coloured red. (c) Bar graph shows SNP densities of 24 overlapping genes between the Fst and Fisher’s tests.
Since both TM and DQ are high-altitude populations, 4,380 m and 3,300 m respectively, we combined the two populations and compared them against YJ, a low-altitude group, and rescanned the X chromosome following the process and criteria stated above. Interestingly, we identified 64 genes with outlier SNPs (Supplementary Table S8). Moreover, 21 of the 64 genes had featured in our previous scans (P < 1.00E-5). Except for ZCCHC16, the other five genes ALG13, AMOT, DCX, LHFPL1, and TRPC5 with the highest outlier SNPs densities in the previous analyses were also observed among the 21 shared genes. Therefore, we considered the five genes as candidate positively selected genes (PSGs).
It has been reported that angiogenesis could be activated by hypoxia-inducible factor (HIF) pathway in hypoxic conditions36. Additionally, some genes involved in angiogenesis have been identified as targets of natural selections in high-altitude animals11,13,19. Thus, following the consistent featuring of AMOT in our analyses and its cited importance in angiogenesis, we sought to attentively assess AMOT with respect to high-altitude adaptation. From annotations of outlier SNPs in AMOT, we observed two nonsynonymous mutations (p.Ser971Ala and p.Leu1025Pro) and two synonymous mutations. Interestingly, for the two nonsynonymous mutations, the reference alleles showed significant divergent distributions between TM and YJ (86.7% in TM vs. 13.3% in YJ, P = 4.03E-6).
Genetic diversity and haplotype analyses around AMOT
The genetic diversity (π) scans on about 1 million bp region, covering the AMOT gene for four dog populations (TM, DQ, LJ and YJ), were calculated. TM and YJ had the lowest genetic diversity compared to DQ and LJ populations (Fig. 3a). This pattern may imply a gradual change of allele frequency among the four populations. A further haplotype analysis based on a region about 500 kb covering AMOT showed that TM and YJ populations exhibited obvious divergence in haplotypes, and a gradual change from YJ to TM (Fig. 3b), which is consistent with the results from the initial genetic diversity analysis. Moreover, in the same region, LD analysis showed that TM population had a strong LD block across the whole region and the strongest tendency located in AMOT (Fig. 3c; Supplementary Fig. S5).
Genetic diversity surveys and haplotype analyses of the regions around AMOT.
(a) Genetic diversity survey on about 1 million bp regions around AMOT for the four dog populations. Candidate gene regions are marked by red vertical lines. Different populations are represented with different colours. (b) Haplotype clusters analyses of ~500 kb around AMOT for the four dog populations. Size and locations of LHFPL1 and AMOT are marked by red vertical lines. For each polymorphic locus of a chromosome or haplotype, point with yellow colour represents high-altitude allele, and that with light blue colour represents low-altitude allele. (c) Linkage disequilibrium (LD) analysis on TM population from the same region which is used for haplotype cluster analyses.
Given that genetic drift can also generate similar genetic patterns as positive selection did, we performed simulations for 1,000,000 times to generate allele frequency data sets of TM and YJ populations with ms tools37 for the AMOT region (sequence length 47,544 bp, including 108 SNPs). Then we used these data to test whether the observed TM-YJ genetic divergence could be due to genetic drift instead of selection. For each run, we calculated Fst values of the 108 SNPs from the simulated data sets, and compared the result to the Fst values from the real data. Our analyses showed that the probability of the real Fst values attributable to genetic drift was only 2.32E-3, implying that the genetic pattern observed on AMOT was likely not caused by genetic drift, and AMOT may be a target of positive selection during adaptation to hypoxia.
Correlation between AMOT haplotype frequency and altitude
Inspired by the observed gradual variation in genetic diversity and haplotype distributions among the four dog populations, we additionally sampled 175 individuals of 11 aboriginal dog populations from different areas in China (altitude range: 300 m to 4,000 m) to further validate the selective signatures of AMOT (Supplementary Tables S9 and S10; Supplementary Fig. S6). In this part, we genotyped two fragments around AMOT to test the correlation between haplotype frequency and altitude. The two fragments were about 112 kb apart and included 11 SNPs. Combining the four dog populations used in X-chromosome-wide selective analyses, we built haplotypes of the 11 SNPs from all the 15 dog populations (215 individuals in total). Interestingly, one kind of haplotype (‘CGTTCGCTTTG’) occupied a very high ratio (about 82% on average) in Tibetan Mastiff populations (Table 1). Moreover, there was a significant correlation between the TM’s major haplotype frequency and altitude (P = 1.03E-3) (Fig. 4).
Taken together, the scans of X chromosome of TM and other dog populations, and the subsequent analyses of the genetic diversity, haplotype distributions, as well as the correlation between haplotype frequency and altitude, all provide evidence suggesting that AMOT could be a key target gene of positive selections, and could be associated with high-altitude hypoxia tolerance among Tibetan Mastiff.
Discussion
Several selective scans on X chromosome have been reported in previous animal domestications and breeding studies. Specific regions or genes have shown likely involvement in the process of domestications of pig and sheep38,39,40. However, the present study is the first to perform detailed X-chromosome-wide scans to address the genetic basis of high-altitude adaptation, a notable contribution in filling up the knowledge gaps in the molecular mechanisms of high-altitude adaptation. In addition, relative to previous analyses of X chromosome in animal domestications studies, in which the genome-wide data were mainly generated from microarray or pooling re-sequencing platforms, our present work has used more high-quality data sets with independent re-sequenced libraries and good sequencing coverage, increasing the possibility of identifying high-quality SNPs and more signatures of positive selection (Supplementary Tables S3 and S5).
Here, we chose Tibetan Mastiff, which is a typical representative of domesticated animal that migrated to the high-altitude plateau with humans, as our study model. TM has been investigated in several studies for its high-altitude adaptation via autosome genome analyses4,17,18. From these analyses, two PSGs, i.e., EPAS1 and HBB, which are associated with HIF pathway and oxygen transportation, and one PSG, i.e., PLXNA4, which functions in angiogenesis by interacting with VEGF41, have been reported as targeted candidate genes for adaptation to high-altitude18. In our current selective scans on X chromosome in TM and other dog populations from different elevation gradients (from 800 to 4380m), we have identified five new PSGs. Among these PSGs, AMOT gene has been reported to act in the process of angiogenesis25,26,27,28. With simulation analysis and genotyping experiments on extra 11 dog populations, we propose that this gene may have undergone selective effects during the adaptation to hypoxic conditions in TM. The genetic diversity levels of AMOT in TM and YJ populations based on a large genetic region were similar low. As high and low elevations represent two different environments, divergent selective pressures may have acted on AMOT in TM and YJ resulting in the similar genetic diversity patterns, although a narrowed scan showed a marked haplotype divergence between the two populations. Interestingly, studies on Atlantic Salmons and dog populations have observed signals of divergent selections in some regions, in which the selective effects caused increased genetic differentiation between populations and reduced genetic diversity at linked loci42,43.
Although several angiogenesis-related genes, including VEGF, VAV3, NOS3, SRF, TXNRD2, ADAM17 and RNASE4, have been proposed as candidate genes in the adaptation of highland human and wild animals to high-altitude conditions8,10,11,13,16,19, our study for the first time identified that AMOT, which is also an angiogenesis related gene, may have played a key role in the high-altitude adaptation of TM. Hence, it is likely that, besides the HIF pathway, angiogenesis also played a similarly critical role in the adaptation of TM populations to their high-altitude hypoxic habitat.
This study presents important insights for understanding the contribution of sex chromosomes to hypoxia adaptation, as most previous genome-wide studies have often excluded sex chromosome because it represents a different effective population size. Relative to autosomes, our genetic diversity assessments demonstrated that X chromosome had lower genetic diversity across all six dog populations (Fig. 1), consistent with the findings of a recent genome-wide study among human populations22. Although population structures estimated from X chromosomal SNPs showed similar patterns with Gou et al.17, PCA based on allosomes could not clearly distinguish each dog population. On the other hand, the Fst distributions of autosomes and X chromosome between YJ and TM showed that allosome had greater genetic divergence compared to autosomes (P = 2.2E-16) (Supplementary Fig. S7). Taken together, these findings lead to the postulation that X chromosome might have suffered stronger selection pressure or genetic drift effects than autosomes, and that allosome might be more sensitive to selection pressure, resulting in a different population structure.
In summary, our study suggests that X chromosome, like autosomes, may have a contribution in the adaptation of Tibetan Mastiff to hypoxic conditions at high-altitudes, indicating that X chromosome warrants a more attention in future studies of adaptive evolution. In addition, the identification of AMOT as a target candidate gene provides a foundation for future functional studies to further elucidate its role in adaptation to hypoxia.
Methods
All methods were performed in accordance with the guidelines approved by the Kunming Institute of Zoology, Chinese Academy of Sciences. All experimental protocols were approved by the Kunming Institute of Zoology, Chinese Academy of Sciences.
Reads mapping and SNPs calling
Raw sequence reads were mapped to the dog reference genome (version: Canfam3.1) using BWA-MEM (version: 0.7.8-r455)44, and Bam files generated by SAMtools (version: 0.1.18)45. PICARD software packages (version: 1.87; http://picard.sourceforge.net) was used to remove duplicate sequences in each individual, then all the Bam files were processed with a standard pipeline for local re-alignment and base-recalibration in the Genome Analysis Tool Kits (GATK) (version: 2.5-2-gf57256b)46.
Genome-wide SNPs calling were handled by UnifiedGenotypeCaller module in GATK. Raw SNPs were recalibrated with the Variant Quality Score Recalibration (VQSR) module, where a list of known or verified SNPs/INDELs was downloaded from Ensembl Database and used as the training data set. Then the recalibrated SNPs were filtered with rules as described below: 1) SNPs that were within 5 base pairs proximity to inserts and deletions (INDELs) were filtered out, and the INDELs also abandoned; 2) SNP locus at which the ratio of individuals missing SNP detection was greater than 20% of all the samples was excluded; 3) Only bi-allelic SNPs were retained; 4) The remaining SNPs were sorted by their depths, with depths ranging from 2.5% to 97.5% retained; and 5) SNPs with Quality value greater than 40 were retained. Finally, all high-quality SNPs were annotated by ANNOVAR47.
Genetic background survey of the six populations
With the availability of complete gender information for the six dog populations, we made corrections to the allele frequencies of the study populations before assessing population genetic diversity and structure. Here, we first computed pairwise nucleotide diversity (π)29 of autosomes and X chromosome among the six dog populations using custom Perl scripts (non-overlapped 100 k window-size), then principal component analysis (PCA) and population structure estimations were conducted with EIGENSOFT (version: 6.0.1)48 and ADMIXTURE (version: 1.23)49. Before population structure estimations, some SNPs were filtered out, in a process referred to as SNPs thinning, using PLINK (version: v1.07; http://pngu.mgh.harvard.edu/purcell/plink/; Parameter: –indep-pairwise 50 10 0.1), to make sure that each SNP evolved independently. Based on the whole autosomal SNPs from our downloaded genome-wide data sets17, and the recently published genome-wide data sets of dogs and wolves50, we employed SNPhylo51 to construct the phylogenetic relationships of the six dog populations. In addition, Fst surveys on the five population pairs (KM vs. YJ, KM vs. TM, GS vs. YJ, GS vs. TM, and YJ vs. TM) were also conducted.
Selection scans on X chromosome
Two approaches were applied for positive selection scans: Fst test, which reflects the genetic differences among populations52. The other is Fisher’s exact test, which does not need biological hypothesis and can estimate the probability of divergence in allele frequency among populations. Only the top 1% of SNPs with high Fst values in Fst test, or small P-values (after FDR correction) in Fisher’s exact test, were considered as candidate loci of selection.
Genetic diversity and haplotype surveys in candidate genes
The genetic diversity (π) scans of regions around candidate genes were performed with our custom Perl scripts, applying 10 k-bp non-steps windows. Prior to our haplotype analyses, haplotypes of the four dog populations (TM, DQ, LJ and YJ) were inferred by SHAPEIT software53. To increase the haplotypes estimation accuracy, the genetic map of domestic dogs downloaded from a recent work of Auton et al.54 was used in the haplotype construction. Linkage disequilibrium (LD) and haplotype clusters of regions around candidate genes were then analysed using Haploview (version: 4.2)55 and R (version: 3.1.0) software respectively.
Simulation tests of Candidate genes
We applied dense computational simulations to test whether the selective signals of AMOT gene were caused by genetic drift instead of selection. First, using G-PhoCS tools56, 1463 neutral loci of X chromosome from ten female individuals of the TM and YJ populations were chosen for demographic history inference. It has been reported that mutation rate is 1.3 times higher in the autosome than in the X chromosome57, and that the mutation rate in the autosome of dogs is 2.2 × 10−9 per year58. Therefore, for our simulation, we used 1.692 × 10−9 per year as the mutation rate in the X chromosome. Our results showed that TM diverged from YJ about 3,400 years ago, and the effective population size of their common ancestor was approximately 37,800. On the other hand, the effective population size of extant YJ and TM was 8,681 and 4,058 respectively. With parameters drawn from previous analyses, we used ms tools37 to simulate the allele frequency of 108 SNPs spanning the whole AMOT region (about 47 k bps) for both TM and YJ populations in a neutral model. In each simulation, we calculated the Fst values of simulated data sets, and compared them to those computed from the real data sets, with the significance of the divergence measured by t-test. After 1,000,000 simulations, the distribution of all P-values was evaluated to determine whether selective footprints that were produced by the real data sets were generated by genetic drift.
Genotyping and AMOT haplotype correlation tests
To verify the selective signals of AMOT identified from X-chromosome-wide scans, we sampled additional 175 dogs from 11 different places in China (altitude: 300 m to 4000 m) and added them into our present study (Supplementary Tables S9 and S10; Supplementary Fig. S6). Total genomic DNA was extracted from peripheral venous blood using the Phenol/Chloroform method. Polymerase Chain Reaction (PCR) was performed using ExTaq (TaKaRa; Dalian, China). We genotyped two fragments around AMOT. To genotype the first fragment, which is located in the sixth intron of AMOT, the following primers, F1: 5′-CACCATACCCAATCTCTGTCTA-3′and R1: 5′-GAAGGTATGAAGGTGCCTAGGA-3′, and PCR conditions, 94 °C for 30 sec, 57 °C for 30 sec, 72 °C for 1 min, and 32 cycles, were used. For the second fragment, a flanking sequence upstream of AMOT, primer pair of F2: 5′-AAGTCAGAGCAATGGGAAGC-3′ and R2: 5′-GTTTGGTGGGCTGTGAAGAC-3′, as well as PCR conditions of 94 °C for 30 sec, 65 °C for 30 sec, 72 °C for 1 min, and 32 cycles were used. After quality control procedures, we obtained 350 sequences with high-quality. The newly determined sequences have been deposited in GenBank: KX245540-KX245889. We then joined the two fragments into one continuous sequence (containing 11 SNPs) for each individual, and used PHASE (version: 2.1)59,60 to build haplotypes of the all SNPs for the overall dataset of 15 dog populations. Based on the gender information for all samples, we corrected the frequency of the TM major haplotype in each dog population. Finally, the correlation analysis between haplotype frequency and altitude was assessed via R (version: 3.1.0).
Additional Information
Accession codes: All newly generated sequences of the 11 dog populations have been deposited in GenBank (KX245540-KX245889).
How to cite this article: Wu, H. et al. Identifying molecular signatures of hypoxia adaptation from sex chromosomes: A case for Tibetan Mastiff based on analyses of X chromosome. Sci. Rep. 6, 35004; doi: 10.1038/srep35004 (2016).
References
Beall, C. M. et al. Natural selection on EPAS1 (HIF2 alpha) associated with low hemoglobin concentration in Tibetan highlanders. Proc. Natl. Acad. Sci. USA 107, 11459–11464 (2010).
Natarajan, C. et al. Convergent Evolution of Hemoglobin Function in High-Altitude Andean Waterfowl Involves Limited Parallelism at the Molecular Sequence Level. PLoS Genet. 11, e1005681 (2015).
Simonson, T. S. et al. Genetic Evidence for High-Altitude Adaptation in Tibet. Science 329, 72–75 (2010).
Wang, G. D. et al. Genetic Convergence in the Adaptation of Dogs and Humans to the High-Altitude Environment of the Tibetan Plateau. Genome Biol. Evol. 6, 2122–2128 (2014).
Xu, S. H. et al. A Genome-Wide Search for Signals of High-Altitude Adaptation in Tibetans. Mol. Biol. Evol. 28, 1003–1011 (2011).
Yi, X. et al. Sequencing of 50 Human Exomes Reveals Adaptation to High Altitude. Science 329, 75–78 (2010).
Alkorta-Aranburu, G. et al. The Genetic Architecture of Adaptations to High Altitude in Ethiopia. Plos Genet. 8, e1003110 (2012).
Bigham, A. et al. Identifying Signatures of Natural Selection in Tibetan and Andean Populations Using Dense Genome Scan Data. Plos Genet. 6, e1001116 (2010).
Peng, Y. et al. Genetic Variations in Tibetan Populations and High-Altitude Adaptation at the Himalayas. Mol. Biol. Evol. 28, 1075–1081 (2011).
Scheinfeldt, L. B. et al. Genetic adaptation to high altitude in the Ethiopian highlands. Genome Biol. 13, R1 (2012).
Ge, R. L. et al. Draft genome sequence of the Tibetan antelope. Nat. Commun. 4, e1858 (2013).
Li, M. Z. et al. Genomic analyses identify distinct patterns of selection in domesticated pigs and Tibetan wild boars. Nat. Genet. 45, 1431–1438 (2013).
Qu, Y. et al. Ground tit genome reveals avian adaptation to living at high altitudes in the Tibetan plateau. Nat. Commun. 4, e2071 (2013).
Zhang, W. P. et al. Hypoxia Adaptations in the Grey Wolf (Canis lupus chanco) from Qinghai-Tibet Plateau. Plos Genet. 10, e1004466 (2014).
Galen, S. C. et al. Contribution of a mutational hot spot to hemoglobin adaptation in high-altitude Andean house wrens. Proc. Natl. Acad. Sci. USA 112, 13958–13963 (2015).
Yu, L. et al. Genomic analysis of snub-nosed monkeys (Rhinopithecus) identifies genes and processes related to high-altitude adaptation. Nat. Genet. 48, 947–952 (2016).
Gou, X. et al. Whole-genome sequencing of six dog breeds from continuous altitudes reveals adaptation to high-altitude hypoxia. Genome Res. 24, 1308–1315 (2014).
Li, Y. et al. Population Variation Revealed High-Altitude Adaptation of Tibetan Mastiffs. Mol. Biol. Evol. 31, 1200–1205 (2014).
Qiu, Q. et al. The yak genome and adaptation to life at high altitude. Nat. Genet. 44, 946–949 (2012).
Wang, M. S. et al. Genomic Analyses Reveal Potential Independent Adaptation to High Altitude in Tibetan Chickens. Mol. Biol. Evol. 32, 1880–1889 (2015).
Meisel, R. P. & Connallon, T. The faster-X effect: integrating theory and data. Trends Genet. 29, 537–544 (2013).
Gottipati, S., Arbiza, L., Siepel, A., Clark, A. G. & Keinan, A. Analyses of X-linked and autosomal genetic variation in population-scale whole genome sequencing. Nat. Genet. 43, 741–743 (2011).
Zhou, D. et al. Experimental selection of hypoxia-tolerant Drosophila melanogaster. Proc. Natl. Acad. Sci. USA 108, 2349–2354 (2011).
Li, Y. & Zhang, Y. P. High genetic diversity of Tibetan Mastiffs revealed by mtDNA sequences. Chinese Sci. Bull. 57, 1483–1487 (2012).
Bratt, A. et al. Angiomotin regulates endothelial cell-cell junctions and cell motility. J. Biol. Chem. 280, 34859–34869 (2005).
Ernkvist, M. et al. The Amot/Patj/Syx signaling complex spatially controls RhoA GTPase activity in migrating endothelial cells. Blood 113, 244–253 (2009).
Troyanovsky, B., Levchenko, T., Mansson, G., Matvijenko, O. & Holmgren, L. Angiomotin: an angiostatin binding protein that regulates endothelial cell migration and tube formation. J. Cell Biol. 152, 1247–1254 (2001).
Aase, K. et al. Angiomotin regulates endothelial cell migration during embryonic angiogenesis. Genes Dev. 21, 2055–2068 (2007).
Kimura, M. Genetic variability maintained in a finite population due to mutational production of neutral and nearly neutral isoalleles. Genet. Res. 11, 247–269 (1968).
Frantz, L. A. et al. Genomic and archaeological evidence suggest a dual origin of domestic dogs. Science 352, 1228–1231 (2016).
Lindblad-Toh, K. et al. Genome sequence, comparative analysis and haplotype structure of the domestic dog. Nature 438, 803–819 (2005).
Bissar-Tadmouri, N. et al. X Chromosome Exome Sequencing Reveals a Novel ALG13 Mutation in a Nonsyndromic Intellectual Disability Family With Multiple Affected Male Siblings. Am. J. Med. Genet. A 164, 164–169 (2014).
Gleeson, J. G. et al. Doublecortin, a brain-specific gene mutated in human X-linked lissencephaly and double cortex syndrome, encodes a putative signaling protein. Cell 92, 63–72 (1998).
Sossey-Alaoui, K. et al. Molecular cloning and characterization of TRPC5 (HTRP5), the human homologue of a mouse brain receptor-activated capacitative Ca2+ entry channel. Genomics 60, 330–340 (1999).
Irie, M. et al. Cognitive Function Related to the Sirh11/Zcchc16 Gene Acquired from an LTR Retrotransposon in Eutherians. Plos Genet. 11 (2015).
Rey, S. & Semenza, G. L. Hypoxia-inducible factor-1-dependent mechanisms of vascularization and vascular remodelling. Cardiovasc. Res. 86, 236–242 (2010).
Hudson, R. R. Generating samples under a Wright-Fisher neutral model of genetic variation. Bioinformatics 18, 337–338 (2002).
Rubin, C.-J. et al. Strong signatures of selection in the domestic pig genome. Proc. Natl. Acad. Sci. USA 109, 19529–19536 (2012).
Ma, Y., Zhang, H., Zhang, Q. & Ding, X. Identification of Selection Footprints on the X Chromosome in Pig. Plos One 9, e94911 (2014).
Moradi, M. H., Nejati-Javaremi, A., Moradi-Shahrbabak, M., Dodds, K. G. & McEwan, J. C. Genomic scan of selective sweeps in thin and fat tail sheep breeds for identifying of candidate regions associated with fat deposition. BMC Genet. 13, 10; 1186/1471-2156-13-10 (2012).
Kigel, B., Rabinowicz, N., Varshavsky, A., Kessler, O. & Neufeld, G. Plexin-A4 promotes tumor progression and tumor angiogenesis by enhancement of VEGF and bFGF signaling. Blood 118, 4285–4296 (2011).
Tezuka, A. et al. Divergent selection for opsin gene variation in guppy (Poecilia reticulata) populations of Trinidad and Tobago. Heredity (Edinb) 113, 381–389 (2014).
Pilot, M. et al. Diversifying Selection Between Pure-Breed and Free-Breeding Dogs Inferred from Genome-Wide SNP Analysis. G3 Bethesda 6, 2285–2298 (2016).
Li, H. & Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv (1303.3997). Eprint Arxiv 1303 (2013).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Patterson, N., Price, A. L. & Reich, D. Population structure and eigenanalysis. Plos Genet. 2, 2074–2093 (2006).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
Wang, G. D. et al. Out of southern East Asia: the natural history of domestic dogs across the world. Cell Res. 26, 21–33 (2016).
Lee, T. H., Guo, H., Wang, X., Kim, C. & Paterson, A. H. SNPhylo: a pipeline to construct a phylogenetic tree from huge SNP data. BMC Genomics 15, 162 (2014).
Akey, J. M., Zhang, G., Zhang, K., Jin, L. & Shriver, M. D. Interrogating a high-density SNP map for signatures of natural selection. Genome Res. 12, 1805–1814 (2002).
Delaneau, O., Marchini, J. & Zagury, J. F. A linear complexity phasing method for thousands of genomes. Nat. Methods 9, 179–181 (2012).
Auton, A. et al. Genetic Recombination Is Targeted towards Gene Promoter Regions in Dogs. Plos Genet. 9, e1003984 (2013).
Barrett, J. C., Fry, B., Maller, J. & Daly, M. J. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21, 263–265 (2005).
Gronau, I., Hubisz, M. J., Gulko, B., Danko, C. G. & Siepel, A. Bayesian inference of ancient human demography from individual genome sequences. Nat Genet 43, 1031–1034 (2011).
Ebersberger, I., Metzler, D., Schwarz, C. & Paabo, S. Genomewide comparison of DNA sequences between humans and chimpanzees. Am. J. Hum. Genet. 70, 1490–1497 (2002).
Kumar, S. & Subramanian, S. Mutation rates in mammalian genomes. Proc. Natl. Acad. Sci. USA 99, 803–808 (2002).
Stephens, M., Smith, N. J. & Donnelly, P. A new statistical method for haplotype reconstruction from population data. Am. J. Hum. Genet. 68, 978–989 (2001).
Stephens, M. & Donnelly, P. A comparison of bayesian methods for haplotype reconstruction from population genotype data. Am. J. Hum. Genet. 73, 1162–1169 (2003).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Nos 91531303 and 31321002), the Breakthrough Project of Strategic Priority Program of the Chinese Academy of Sciences (No. XDB13000000), the Key Research Program of the Chinese Academy of Sciences, the Youth Innovation Promotion Association, Chinese Academy of Sciences (to G.-D.W.), the CAS-TWAS President’s Fellowship Program for Doctoral Candidates to N.O.O., the grants from the National Youth Talent Support Program to L.Y., and Program for Innovative Research Team (in Science and Technology) in University of Yunnan Province.
Author information
Authors and Affiliations
Contributions
H.W., C.-T.Y. and F.L. performed the experiments. H.W., Y.-H.L. and G.-D.W. analyzed the data. S.-F.W. helped with the sample collection. H.W., L.W., Y.-H.L., N.O.O. and L.Y. wrote the paper. L.Y., Y.-P.Z. and L.W. designed and managed this project.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Wu, H., Liu, YH., Wang, GD. et al. Identifying molecular signatures of hypoxia adaptation from sex chromosomes: A case for Tibetan Mastiff based on analyses of X chromosome. Sci Rep 6, 35004 (2016). https://doi.org/10.1038/srep35004
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep35004
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.