Abstract
TDP-43 protein plays an important role in regulating transcriptional repression, RNA metabolism and splicing. Typically it shuttles between the nucleus and the cytoplasm to perform its functions, while abnormal cytoplasmic aggregation of TDP-43 has been associated with neurodegenerative diseases amyotrophic lateral sclerosis (ALS) and frontotemporal lobar degeneration (FTLD). For the purpose of this study we selected a set of proteins that were misregulated following silencing of TDP-43 and analysed their expression in a TDP-43-aggregation model cell line HEK293 Flp-in Flag-TDP-43-12x-Q/N F4L. Following TDP-43 sequestration in insoluble aggregates, we observed higher nuclear levels of EIF4A3 and POLDIP3β, whereas nuclear levels of DNMT3A, HNRNPA3, PABPC1 and POLDIP3α dropped and cytoplasmic levels of RANBP1 dropped. In addition, immunofluorescence signal intensity quantifications showed increased nuclear expression of HNRNPL and YARS and downregulation of cytoplasmic DPCD. Furthermore, cytoplasmic levels of predominantly nuclear protein ALYREF increased. In conclusion, by identifying a common set of proteins that are differentially expressed in a similar manner in these two different conditions, we show that TDP-43 aggregation has a comparable effect to TDP-43 knockdown.
Similar content being viewed by others
Introduction
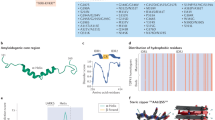
TDP-43 protein, encoded by the TARDBP gene, plays an important role in regulation of several processes, including microRNA processing, apoptosis, cell division, transcription, translation, splicing, axonal transport and neurite outgrowth1,2. Its major distinguishing features are the ability to bind RNA in a very specific manner through two RNA recognition motifs (RRM) and the C-terminal portion of the protein, which includes a glycine-rich domain that is involved in most of the protein interactions described3. This region contains a glutamine/asparagine (Q/N) prion-like domain that participates in protein–protein interactions and in the TDP-43 aggregation process4,5. Typically, TDP-43 is shuttled between the nucleus and the cytoplasm to perform its functions6,7,8. Depletion of TDP-43 is embryonic lethal at very early stages of development and its overexpression above normal levels is highly toxic to cells, especially neurons9,10,11.
Abnormal cytoplasmic and occasional intranuclear aggregation of TDP-43 has been associated with amyotrophic lateral sclerosis (ALS) and frontotemporal lobar degeneration (FTLD-TDP)12,13. The discovery of missense mutations of TARDBP in familial and sporadic ALS cases proved the essential role of abnormal TDP-43 in disease14. Wild-type TDP-43 itself is intrinsically aggregation-prone as well as toxic but a few ALS-causing mutations appear to significantly exaggerate the aggregation process15,16. From the point of view of the pathology, however, it is important to highlight that wild-type cytoplasmic TDP-43 positive inclusions can be found in 95% of all ALS and 60% of FTLD cases, which are now termed TDP-43 proteinopathies12,17,18. TDP-43 positive cytoplasmic inclusions have also been described in 57% of Alzheimer’s disease cases, 20% of Dementia with Lewy Bodies, Pick’s disease, hippocampal sclerosis, corticobasal degeneration, Huntington disease, Parkinson’s disease, argyrophilic grain disease and in a variety of other neurodegenerative conditions19,20. The histology in all these cases is similar, with TDP-43 present in cytoplasmic inclusions in glia and neurons, thus partially or totally cleared from the nucleus21,22. Taken together, aggregation of TDP-43 is most probably the root cause of ALS/FTLD either through a gain of toxic function (GOF) on its own or through a loss of function (LOF) with sequestration and depletion of nuclear TDP-4323,24 or both25. It is therefore of prime importance to better characterize its consequences at the cellular level. In this respect, previous studies demonstrated the effect of TDP-43 knockdown on the transcriptome2,26 and recently on proteome of SH-SY5Y cells27 and cytotoxicity has been observed to increase following cytoplasmic internalisation of TDP-43 containing inclusions bodies28. In general, aggregation-prone proteins that have been targeted to cytoplasm, show that cytoplasmic aggregates interfere with nuclear protein transport and inhibit mRNA transport29.
In this study, however, we have specifically investigated whether the aggregation/sequestration of TDP-43 correlated with its loss of function by comparing the expression changes of selected proteins responsive to silencing of TDP-43 in an aggregation and sequestration model cell line HEK293 Flp-in Flag-TDP-43-12x-Q/N F4L. Our results show that differential expression of proteins in TDP-12xQ/N-F4L cells correlated with proteomic results of TDP-43 knockdown in SH-SY5Y, revealing a common set of proteins whose expression is influenced via TDP-43 aggregation or knockdown.
Results
For the purpose of this study, we selected 13 proteins (Table 1) whose expression levels we previously demonstrated to be affected by TDP-43 knockdown27. These particular proteins where selected based on their function, association with ALS and FTLD-TDP and availability of the antibodies. We analyzed their expression in HEK293 Flp-in Flag-TDP-43-12x-Q/N F4L cell line, comparing protein expressions before and after induction of Flag-TDP-43-12x-Q/N expression. We chose the mutant overexpressing Flag-TDP-43-12x-Q/N F4L, with double-site mutations F147/149L and F229/231 in RRM1 and RRM2, due to its inability to bind RNA and thus downregulate the levels of soluble endogenous TDP-43 through the negative feedback loop6,30. In this manner, we sought to replicate the most likely disease-condition where aggregates form in the presence of fully functional endogenous TARDBP gene expression.
In HEK293 Flp-in Flag-TDP-43-12x-Q/N F4L cells (referred to as HEK TDP-12xQ/N-F4L hereafter) TDP-43 aggregation was induced by addition of doxycycline (DOX) to the growth medium (Fig. 1). The aggregates were observed in the nucleus and the cytoplasm and could be detected both by anti-TDP-43 and anti-Flag antibodies. After 72 hours, the nuclear levels of soluble TDP-43 significantly dropped to 37.4 ± 4.7% (mean ± s.e.m., n = 3) as determined by western blot (Fig. 2a,c).
Expression of selected proteins in nuclear (a) or cytoplasmic (b) fractions of HEK Flp-in Flag-TDP-43-12xQ/N F4L before (−DOX) and after (+DOX) induction of TDP-43 aggregation. Relative expression of proteins was quantified with western blot (c). Fibrillarin (FBL) and GAPDH were used as loading controls for nuclear and cytoplasmic fractions. Unpaired Student’s t-test was used to determine significant differences between samples (significance levels: *for p < 0.05, ***for p < 0.001). Error bars represent s.e.m.
Splicing of more than 1000 mRNAs is affected by decreased cellular levels of TDP-432,26. It has been previously shown that POLDIP3 (SKAR) and RANBP1 splicing is under control of TDP-432,26,32. TDP-43 knockdown or sequestration promoted splicing of POLDIP3 (SKAR) exon 3 and inclusion of RANBP1 exon 5 in both SH-SY5Y and HEK TDP-12xQ/N-F4L in the same pattern (Fig. S1), thus validating their applicability for this study.
In HEK TDP-12xQ/N-F4L western blot analyses showed differential expression of DNMT3A, HNRNPA3, EIF4A3, POLDIP3, PABPC1 and RANBP1, before and after induction of TDP-43 aggregation, which correlated with proteomic results of TDP-43 knockdown in SH-SY5Y (Table 1, Fig. 2 and Table S2). Namely, nuclear expression of DNMT3A, HNRNPA3 and PABPC1 dropped, whereas expression levels of POLDIPβ and EIF4A3 increased and cytoplasmic levels of RANBP1 dropped.
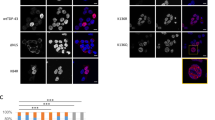
Next we quantified the expression changes of the selected proteins following immunofluorescence (IF) staining (Figs 3 and 4). Signal intensity quantifications showed that 12 out of 13 proteins had statistically significant expression level changes after induction of aggregation (Figs 3b and 4b, Table S3). Most importantly, for 10 out of 13 proteins, the change in expression levels correlated with knockdown data from Štalekar et al.27. After TDP-43 aggregation, relative levels of nuclear proteins HNRNPL, EIF4A3, POLDIP3 and YARS increased, whereas nuclear levels of DNMT3A, HNRNPA3 and PABPC1 dropped (Fig. 3b). Although ALYREF is localised mainly in the nucleus, we observed its upregulation in the cytoplasmic fraction of HEK TDP-12xQ/N-F4L after TDP-43 aggregation (Fig. 4). We have previously shown reduced levels of RANBP1 after TDP-43 silencing in SH-SY5Y27 and herein confirmed this occurrence also in HEK TDP-12xQ/N-F4L after induction of TDP-43 aggregation (Figs 2b,c and 4). Finally, also DPCD cytoplasmic levels dropped in HEK TDP-12xQ/N-F4L after TDP-43 aggregation (Fig. 4).
Expression of five nuclear proteins and one predominantly cytoplasmic protein (YARS) increased and of DNMT3A, HNRNPA3 and PABPC1 dropped in the nuclear fraction of HEK Flp-in Flag-TDP-43-12xQ/N F4L after induction of TDP-43 aggregation.
(a) Non-induced cells (−DOX) and cells after induction of TDP-43 aggregation (+DOX). Arrows mark protein aggregates, which in the case of TIAL1 colocalise with TDP-43 aggregates. Scale bars are 5 μm. (b) Immunofluorescence signal intensity quantification. Unpaired Student’s t-test was used to determine significant differences between samples (significance levels: *for p < 0.05, **for p < 0.01, ***for p < 0.001). Error bars represent s.e.m.
Cytoplasmic expression of RANBP1, STRA6 and DPCD dropped, while expression of, predominantly nuclear protein, ALYREF, increased in HEK Flp-in Flag-TDP-43-12xQ/N F4L after induction of TDP-43 aggregation.
(a) Non-induced cells (−DOX) and cells after induction of TDP-43 aggregation (+DOX). Scale bars are 5 μm. (b) Immunofluorescence signal intensity quantification. Unpaired Student’s t-test was used to determine significant differences between samples (significance levels: **for p < 0.01, ***for p < 0.001). Error bars represent s.e.m.
In addition, some discrepancies between the proteomic data obtained for HEK TDP-12xQ/N-F4L in comparison to RNAi in SH-SY5Y may be on account of the difference in experimental model used (knockdown vs. aggregation), the cell line (HEK vs. SH-SY5Y), the experimental conditions (induction vs. transfection, media), which in turn reflect a moderate difference in the regulation of expression of proteins in question. There is also some variation to be expected from the antibodies used, as some perform better in denaturing conditions of western blot, while others recognise better the proteins in cells (IF). The recognition of the latter can also be influenced by method of fixation of cells (for instance using paraformaldehyde or methanol) or the accessibility of the epitope against which the antibody was raised due to specific protein conformation.
Discussion
In this study, using a cellular model of TDP-43 sequestration, we have examined expression variations of a series of proteins previously shown to be differentially expressed following TDP-43 knockdown27. As has been previously demonstrated in HEK293 Flp-in Flag-TDP-43-12x-Q/N F4L cell line, loss of TDP-43 function was achieved because endogenous TDP-43 was able to interact fully with Flag-TDP-43-12xQ/N aggregates and ended up sequestered in both nuclear and cytoplasmic insoluble aggregates30. The aggregation was enforced by a Q/N-rich region of TDP-43, which itself is involved in aggregate formation and in the interaction of TDP-43 with inclusions4. Thus, cells had drastically reduced levels of active nuclear TDP-43 and presented a suitable model to test the impact of TDP-43 sequestration on expression and redistribution of a selection of proteins, whose functions have already been shown to be altered by TDP-43 knockdown27. As TDP-43 inclusions in HEK TDP-12xQ/N-F4L formed in the nucleus as well as the cytoplasm, it is not clear whether the change in expression of TDP-43 and other proteins was due to the GOF or LOF mechanisms, although our study suggests that the aggregation of TDP-43 causes proteomic changes in the cells akin to TDP-43 LOF.
TDP-43 is an RNA/DNA binding protein with multiple functions. For this reason, its downregulation influences a large number of RNA and protein targets. Decreased TDP-43 cellular levels affected splicing of 158 exons in neuroblastoma cells and altered 965 splicing events in adult mouse brain2,26,30. A remarkable alteration of splicing of polymerase delta-interacting protein 3 (POLDIP3/SKAR) has been previously noted as a result of the depletion of TDP-432,26,32,33. The decreased inclusion of exon 3 in POLDIP3 gene has been reported to favour the synthesis of the β isoform in respect to the main α isoform32,33,34. After TDP-43 sequestration we confirmed this splicing pattern, in both SH-SY5Y and HEK TDP-12xQ/N-F4L, detecting higher levels of β-isoform at the mRNA level30 and this observation was mirrored at the protein level. The increased expression of the shorter POLDIP3 mRNA variant was also observed in ALS patients’ spinal motor neurons32. In this study, we have employed an antibody that recognises both α and β isoforms of POLDIP 3. After densitometric quantification of western blot data we confirmed the upregulation of β isoform, 252.0 ± 6.2% (mean ± s.e.m., n = 3) and the concomitant downregulation of α isoform, 13.3 ± 17.6% (mean ± s.e.m., n = 3). However, immunofluorescence showed only a smaller increase, 121.6 ± 4.0% (mean ± s.e.m., n = 3), as it was a cumulative measurement of α and β POLDIP3 isoform.
Our comparative analysis of siRNA-mediated knockdown versus sequestration and aggregation of TDP-43 has allowed us to confirm two possible connections between loss of TDP-43 functional activity and disease. First of all, our analyses add further support to the possible link between TDP-43 and nonsense-mediated decay. In fact, POLDIP3 was also reported to associate with the exon junction complex (EJC), which is recruited to exon junctions during splicing35. Furthermore, after TDP-43 aggregation, one of the core proteins of EJC, eukaryotic translation initiation factor 4A3 (EIF4A3), was also upregulated in the nuclear fraction of HEK TDP-12xQ/N-F4L. In addition, EIF4A3 is connected to ALYREF, which is known to recruit export factors during the formation of export competent messenger ribonucleoprotein complexes (mRNPs), thus enabling mRNA export36. Although POLDIP3 and EIF4A3 were upregulated in the nucleus, suggestive of stalled mRNA export and protein production, ALYREF was upregulated in the cytoplasm. ALYREF shuttles between nucleus and cytoplasm and beside mRNA export it has been implicated in linking splicing with transcription37,38. This suggests that TDP-43 aggregation induces staling of ALYREF in the cytoplasm, compromising transcription, nuclear RNA stability and mRNA export.
Secondly, in a Drosophila study, ALYREF was singled out as potential modifier of G4C2 expansion related toxicity39. (G4C2)n hexanucleotide repeat expansion mutation in the C9orf72 gene, which can span from several hundred to several thousand repeats, is the major genetic cause of ALS and FTLD leading to TDP-43 proteinopathy40,41,42. As toxic G4C2 RNA are sequestered in nuclear foci, ALYREF may act as part of a control mechanism that retains unspliced or faulty RNAs in the nucleus36. The question arises weather the redistribution of ALYREF to the cytosol, as a consequence of TDP-43 sequestration, might therefore be a mechanism of inhibiting nuclear transport due to aggregation induced proteotoxicity of TDP-43.
Thirdly, TDP-43 sequestration impacts several HNRNP proteins by either increasing their concentration in the nucleus, or by nuclear clearance, thus altering RNA metabolism and potentially leading to ALS and FTLD. Following TDP-43 aggregation we observed altered levels of HNRNPA3 and HNRNPL. Previously, HNRNPA3 was found to be aggregated in cytosolic TDP-43 negative inclusions in the brains of patients with C9orf72 expanded repeats43. In FTLD-TDP patients, also increased expression of HNRNPA1/A2 was detected44. On the other hand, in ALS patient motor neurons loss of HNRNPA1 expression was concomitant with TDP-43 cytoplasmic inclusions45. The expression of HNRNP proteins is probably regulated not only by sequestration of TDP-43 but by network of several other RNA-binding proteins (RBP), whose expression might also depend on the cell/tissue type tested or the course of disease progression. In addition to HNRNP proteins, PABPC1 is another RBP whose expression was observed to be altered after TDP-43 aggregation. Namely, its nuclear expression dropped in HEK TDP-12xQ/N-F4L. PABPC1 is predominantly cytoplasmic protein that shuttles between the nucleus and the cytoplasm. Together with TDP-43 it accumulates in the stress granules leading to translational repression46,47. Its mislocalisation in robust cytoplasmic inclusions has been observed in ALS spinal cord motor neurons48. We did not observe aggregation of PABPC1 in HEK TDP-12xQ/N-F4L but detected colocalization of TDP-43 aggregates with another stress granule marker, TIAL1. After TDP-43 knockdown in SH-SY5Y TIAL1 nuclear expression increased, however, we have not observed this when quantifying fluorescence levels in HEK TDP-12xQ/N-F4L. This might be specific to the aggregates formed or the antibody used as it detects both TIAL1 and TIA1, thus, we were most likely quantifying fluorescence signals for both proteins.
Finally, we have previously shown in SH-SY5Y that depletion of TDP-43 influences intracellular transport through downregulation of RANBP1 and herein confirmed the expression drop in HEK TDP-12xQ/N-F4L model27. In addition, in both cell lines we confirmed that silencing/aggregation of TDP-43 enables inclusion of exon 5 in RANBP1 transcript, in turn decreasing the level of protein.
In conclusion, although the pathological relevance of these processes in vivo still needs to be determined, our cellular analysis has added further support that aggregation and sequestration model overlaps TDP-43 LOF following its knockdown by siRNA. As a consequence, the major hits from this comparison might represent good candidates to be followed up in further studies that aim to elucidate the causes of ALS/FTLD pathology.
Methods
Cell culture
HEK293 Flp-in Flag-TDP-43-12x-Q/N F4L cell line was established as previously described30. Cells were grown in DMEM-Glutamax-I (Gibco) supplemented with 10% tet-free fetal bovine serum (Biowest), 100 U/ml penicillin-streptomycin (Gibco), 100 μg/ml hygromycin B (Sigma) and 10 μg/ml blastycydin (Sigma). The induction of expression of Flag-TDP-43-12x-Q/N proteins was achieved by adding 1 μg/ml doxycycline (Sigma) to the culture medium for 72 hours.
Cell fractionation
Cells were grown in 6-well plates and were harvested in cold CLB buffer (50 mM Tris, pH 7.4, 10 mM NaCl, 0.5% Igepal Ca-630 (Sigma-Aldrich), 0.25% Triton X-100, cocktail of protease inhibitors (Roche)) and centrifuged for 5 min at 3000 g at 4 °C. Supernatants were transferred to fresh tubes and re-centrifuged at 16100 g at 4 °C for 10 min. Obtained supernatants were used as cytoplasmic fractions. The first pellets were washed three times in cold CLB, resuspended in 1X SDS loading buffer without bromophenol blue, sonicated, boiled for 5 min and re-centrifuged. Resulting supernatants were saved as nuclear fractions. The protein concentration in the fractions were determined by Bio-Rad DC Protein Assay.
Western blot
Reducing SDS-PAGE was run on 4–12% SDS precast gels (C.B.S. Scientific) loaded with 10–20 μg of protein samples in 1X SDS loading buffer with 100 mM dithiothreitol at 175 V. Wet transfer onto nitrocellulose membrane (GE Healthcare) was carried out at 200 mA for 90 min. Membranes were blocked with 5% non-fat dry milk in TBS with 0, 05% Tween-20 (TBST, Sigma) at room temperature for 1 hour. Primary antibodies (Table S1) diluted in blocking medium were incubated over night at 4 °C with gentle rocking. Membranes were washed three times with TBST and incubated in the dark with fluorescently-labelled secondary antibodies (Goat anti-Rabbit IgG (H + L) Cross Adsorbed Secondary Antibody, Dylight 650 conjugate and Goat anti-Mouse IgG (H + L) Cross Adsorbed Secondary Antibody, Dylight 550 conjugate, both Thermo Fischer Scientific) at room temperature for 1 hour. After three washes with TBST, images were acquired using GelDoc System (Bio-Rad). ImageLab software (Bio-Rad) was used for densitometric analysis. Each experiment was performed three times in triplicates (n = 3). Additionally, samples were loaded in triplicates. Loading controls were run on the same blot for each experiment.
Immunofluorescence
Cells were grown on poly-L-lysine (Sigma) coated glass coverslips and were washed once with PBS and fixed with 4% paraformaldehyde in PBS for 15 min. Then they were permeabilised with 0.1% Trition X-100 (Sigma) in PBS for 5 min. Blocking was performed with 3% BSA (Sigma) in PBS for 1 hour. Cells were incubated with primary antibodies (Table S1) diluted in blocking solution for 1 hour at room temperature, followed by washing with PBS and incubation in the dark with fluorescently-labelled secondary antibodies (Donkey Anti-Rabbit IgG (H + L) Alexa Fluor 488, Goat anti-Mouse IgG (H + L) Alexa Fluor 633, Goat Anti-Rat IgG (H + L) Alexa Fluor 555, all Invitrogen) for 1 hour. After washing with PBS cells were stained with DAPI (Sigma) for 7 min and washed with PBS. Coverslips were mounted using FluorSave reagent (Millipore). Images were acquired with Zeiss LSM 710 inverted confocal laser scanning microscope with a Plan-Apochromat 63 ×/1.4NA M27 oil immersion objective using immersion oil (Carl Zeiss). DAPI, Alexa Fluor 488, Alexa Fluor 555 and Alexa Fluor 633 were excited at 405, 488, 543 or 633 nm, respectively. The zoom factor was set to 1 × . X- and Y-scanning sizes were each 1024 pixels. All images were further cropped in ZEN 2010 B SP1 software and scale bars were added.
ImageJ was used to quantify immunofluorescence signal intensities as follows. Several regions of interest (ROI) were selected in the appropriate cellular location and the mean fluorescence of detected protein was measured, along with several adjacent background readings. The total corrected cellular fluorescence, (CTCF = integrated density−(area of selected cell × mean fluorescence of background readings), was calculated for each ROI. The averages and s.e.m. were determined for several images for each marker in induced cells compared to non-induced, where expression was set to 100%. Statistical analysis (2-sided unpaired Student’s t-tests) was performed.
Statistical analysis
Quantitative values of protein bands in western blots were normalized to GAPDH in cytoplasmic fractions and to fibrillarin (FBL) in nuclear fractions. Relative protein expression levels in TDP-43 aggregating HEK TDP-12xQ/N-F4L were calculated towards non-induced controls. For immunofluorescence quantification, immunofluorescence signal intensities of cells with induced TDP-43 aggregation and non-induced controls were compared and relative protein expression levels were calculated. Statistical significance of differential expression of proteins according to western blot and immunofluorescence was determined with unpaired Student’s t-test analysis. Student’s t-test was performed in Microsoft Excel Professional Plus 2013. A p-value of <0.05 was considered significant.
Additional Information
How to cite this article: Prpar Mihevc S. et al. TDP-43 aggregation mirrors TDP-43 knockdown, affecting the expression levels of a common set of proteins. Sci. Rep. 6, 33996; doi: 10.1038/srep33996 (2016).
References
Smethurst, P., Sidle, K. C. L. & Hardy, J. Review: Prion-like mechanisms of transactive response DNA binding protein of 43 kDa (TDP-43) in amyotrophic lateral sclerosis (ALS). Neuropathol. Appl. Neurobiol. 41, 578–597 (2015).
Tollervey, J. R. et al. Characterizing the RNA targets and position-dependent splicing regulation by TDP-43. Nat. Neurosci. 14, 452–458 (2011).
Buratti, E. & Baralle, F. E. TDP-43: gumming up neurons through protein-protein and protein-RNA interactions. Trends Biochem. Sci. 37, 237–247 (2012).
Budini, M. et al. Cellular model of TAR DNA-binding protein 43 (TDP-43) aggregation based on its C-terminal Gln/Asn-rich region. J. Biol. Chem. 287, 7512–7525 (2012).
Mompeán, M. et al. Structural characterization of the minimal segment of TDP-43 competent for aggregation. Arch. Biochem. Biophys. 545, 53–62 (2014).
Ayala, Y. M. et al. TDP-43 regulates its mRNA levels through a negative feedback loop. EMBO J. 30, 277–288 (2011).
Nishimura, A. L. et al. Nuclear import impairment causes cytoplasmic trans-activation response DNA-binding protein accumulation and is associated with frontotemporal lobar degeneration. Brain J. Neurol. 133, 1763–1771 (2010).
Winton, M. J. et al. Disturbance of nuclear and cytoplasmic TAR DNA-binding protein (TDP-43) induces disease-like redistribution, sequestration and aggregate formation. J. Biol. Chem. 283, 13302–13309 (2008).
Tsao, W. et al. Rodent models of TDP-43: recent advances. Brain Res. 1462, 26–39 (2012).
Wächter, N., Storch, A. & Hermann, A. Human TDP-43 and FUS selectively affect motor neuron maturation and survival in a murine cell model of ALS by non-cell-autonomous mechanisms. Amyotroph. Lateral Scler. Front. Degener. 16, 431–441 (2015).
Igaz, L. M. et al. Dysregulation of the ALS-associated gene TDP-43 leads to neuronal death and degeneration in mice. J. Clin. Invest. 121, 726–738 (2011).
Neumann, M. et al. Ubiquitinated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Science 314, 130–133 (2006).
Neumann, M. et al. TDP-43 in the ubiquitin pathology of frontotemporal dementia with VCP gene mutations. J. Neuropathol. Exp. Neurol. 66, 152–157 (2007).
Sreedharan, J. et al. TDP-43 mutations in familial and sporadic amyotrophic lateral sclerosis. Science 319, 1668–1672 (2008).
Lim, L., Wei, Y., Lu, Y. & Song, J. ALS-Causing Mutations Significantly Perturb the Self-Assembly and Interaction with Nucleic Acid of the Intrinsically Disordered Prion-Like Domain of TDP-43. PLoS Biol. 14 (2016).
Johnson, B. S. et al. TDP-43 is intrinsically aggregation-prone and amyotrophic lateral sclerosis-linked mutations accelerate aggregation and increase toxicity. J. Biol. Chem. 284, 20329–20339 (2009).
Arai, T. et al. TDP-43 is a component of ubiquitin-positive tau-negative inclusions in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Biochem. Biophys. Res. Commun. 351, 602–611 (2006).
Kwong, L. K., Neumann, M., Sampathu, D. M., Lee, V. M.-Y. & Trojanowski, J. Q. TDP-43 proteinopathy: the neuropathology underlying major forms of sporadic and familial frontotemporal lobar degeneration and motor neuron disease. Acta Neuropathol. (Berl.) 114, 63–70 (2007).
Amador-Ortiz, C. et al. TDP-43 immunoreactivity in hippocampal sclerosis and Alzheimer’s disease. Ann. Neurol. 61, 435–445 (2007).
Josephs, K. A. et al. TDP-43 is a key player in the clinical features associated with Alzheimer’s disease. Acta Neuropathol. (Berl.) 127, 811–824 (2014).
Baloh, R. H. TDP-43: the relationship between protein aggregation and neurodegeneration in amyotrophic lateral sclerosis and frontotemporal lobar degeneration. FEBS J. 278, 3539–3549 (2011).
Prpar Mihevc, S. et al. Nuclear trafficking in amyotrophic lateral sclerosis and frontotemporal lobar degeneration. Brain J. Neurol. doi: 10.1093/brain/aww197 (2016).
Buratti, E. & Baralle, F. E. The molecular links between TDP-43 dysfunction and neurodegeneration. Adv. Genet. 66, 1–34 (2009).
Ratti, A. & Buratti, E. Physiological Functions and Pathobiology of TDP-43 and FUS/TLS proteins. J. Neurochem. doi: 10.1111/jnc.13625 (2016).
Cascella, R. et al. Quantification of the relative contributions of loss-of-function and gain-of-function mechanisms in TDP-43 proteinopathies. J. Biol. Chem. doi: 10.1074/jbc.M116.737726 (2016).
Polymenidou, M. et al. Long pre-mRNA depletion and RNA missplicing contribute to neuronal vulnerability from loss of TDP-43. Nat. Neurosci. 14, 459–468 (2011).
Štalekar, M. et al. Proteomic analyses reveal that loss of TDP-43 affects RNA processing and intracellular transport. Neuroscience 293, 157–170 (2015).
Capitini, C. et al. TDP-43 Inclusion Bodies Formed in Bacteria Are Structurally Amorphous, Non-Amyloid and Inherently Toxic to Neuroblastoma Cells. PLOS ONE 9, e86720 (2014).
Woerner, A. C. et al. Cytoplasmic protein aggregates interfere with nucleocytoplasmic transport of protein and RNA. Science 351, 173–176 (2016).
Budini, M., Romano, V., Quadri, Z., Buratti, E. & Baralle, F. E. TDP-43 loss of cellular function through aggregation requires additional structural determinants beyond its C-terminal Q/N prion-like domain. Hum. Mol. Genet. 24, 9–20 (2015).
König, J. et al. iCLIP reveals the function of hnRNP particles in splicing at individual nucleotide resolution. Nat. Struct. Mol. Biol. 17, 909–915 (2010).
Shiga, A. et al. Alteration of POLDIP3 Splicing Associated with Loss of Function of TDP-43 in Tissues Affected with ALS. PLoS ONE 7, e43120 (2012).
Fiesel, F. C., Weber, S. S., Supper, J., Zell, A. & Kahle, P. J. TDP-43 regulates global translational yield by splicing of exon junction complex component SKAR. Nucleic Acids Res. 40, 2668–2682 (2012).
Colombrita, C. et al. From transcriptomic to protein level changes in TDP-43 and FUS loss-of-function cell models. Biochim. Biophys. Acta 1849, 1398–1410 (2015).
Le Hir, H., Izaurralde, E., Maquat, L. E. & Moore, M. J. The spliceosome deposits multiple proteins 20–24 nucleotides upstream of mRNA exon-exon junctions. EMBO J. 19, 6860–6869 (2000).
Gromadzka, A. M., Steckelberg, A.-L., Singh, K. K., Hofmann, K. & Gehring, N. H. A short conserved motif in ALYREF directs cap- and EJC-dependent assembly of export complexes on spliced mRNAs. Nucleic Acids Res. doi: 10.1093/nar/gkw009 (2016).
Bruhn, L., Munnerlyn, A. & Grosschedl, R. ALY, a context-dependent coactivator of LEF-1 and AML-1, is required for TCRalpha enhancer function. Genes Dev. 11, 640–653 (1997).
Stubbs, S. H. & Conrad, N. K. Depletion of REF/Aly alters gene expression and reduces RNA polymerase II occupancy. Nucleic Acids Res. 43, 504–519 (2015).
Freibaum, B. D. et al. GGGGCC repeat expansion in C9orf72 compromises nucleocytoplasmic transport. Nature 525, 129–133 (2015).
Cooper-Knock, J., Shaw, P. J. & Kirby, J. The widening spectrum of C9ORF72-related disease; genotype/phenotype correlations and potential modifiers of clinical phenotype. Acta Neuropathol. (Berl.) 127, 333–345 (2014).
DeJesus-Hernandez, M. et al. Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p-linked FTD and ALS. Neuron 72, 245–256 (2011).
Renton, A. E. et al. A hexanucleotide repeat expansion in C9ORF72 is the cause of chromosome 9p21-linked ALS-FTD. Neuron 72, 257–268 (2011).
Mori, K. et al. hnRNP A3 binds to GGGGCC repeats and is a constituent of p62-positive/TDP43-negative inclusions in the hippocampus of patients with C9orf72 mutations. Acta Neuropathol. (Berl.) 125, 413–423 (2013).
Mohagheghi, F. et al. TDP-43 functions within a network of hnRNP proteins to inhibit the production of a truncated human SORT1 receptor. Hum. Mol. Genet. doi: 10.1093/hmg/ddv491 (2015).
Honda, H. et al. Loss of hnRNPA1 in ALS spinal cord motor neurons with TDP-43-positive inclusions. Neuropathol. Off. J. Jpn. Soc. Neuropathol. 35, 37–43 (2015).
Colombrita, C. et al. TDP-43 is recruited to stress granules in conditions of oxidative insult. J. Neurochem. 111, 1051–1061 (2009).
Freibaum, B. D., Chitta, R., High, A. A. & Taylor, J. P. Global analysis of TDP-43 interacting proteins reveals strong association with RNA splicing and translation machinery. J. Proteome Res. 9, 1104–1120 (2010).
Kim, H.-J. et al. Therapeutic modulation of eIF2α-phosphorylation rescues TDP-43 toxicity in amyotrophic lateral sclerosis disease models. Nat. Genet. 46, 152–160 (2014).
Acknowledgements
This work was supported by the Slovenian Research Agency (grants P4-0127, J3-6789, J3-5502, J7-5460), AriSLA grant “TARMA”, Thierry Latran Foundation (REHNPALS) and the EU Joint Programme-Neurodegenerative Diseases JPND (RiMod-FTD, Italy, Ministero della Sanita’, MIUR). We would like to thank Francisco E. Baralle for critical reading of the manuscript.
Author information
Authors and Affiliations
Contributions
B.R., E.B., M.B. and S.P.M. designed research. S.P.M. performed the experiments. M.B. and E.B. developed the inducible cell line. S.P.M., M.B., E.B. and B.R. wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Prpar Mihevc, S., Baralle, M., Buratti, E. et al. TDP-43 aggregation mirrors TDP-43 knockdown, affecting the expression levels of a common set of proteins. Sci Rep 6, 33996 (2016). https://doi.org/10.1038/srep33996
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep33996
This article is cited by
-
The era of cryptic exons: implications for ALS-FTD
Molecular Neurodegeneration (2023)
-
Emerging Therapies and Novel Targets for TDP-43 Proteinopathy in ALS/FTD
Neurotherapeutics (2022)
-
Loss of aly/ALYREF suppresses toxicity in both tau and TDP-43 models of neurodegeneration
GeroScience (2022)
-
Cell environment shapes TDP-43 function with implications in neuronal and muscle disease
Communications Biology (2022)
-
Characterization of the human TARDBP gene promoter
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.