Abstract
Ewing sarcoma is a bone and soft-tissue tumor that depends on the activity of the EWS-FLI1 transcription factor for cell survival. Although a number of compounds have been shown to inhibit EWS-FLI1 in vitro, a clinical EWS-FLI1-directed therapy has not been achieved. One problem plaguing drug development efforts is the lack of a suitable, non-invasive, pharmacodynamic marker of EWS-FLI1 activity. Here we show that 18F-FLT PET (18F- 3′-deoxy-3′-fluorothymidine positron emission tomography) reflects EWS-FLI1 activity in Ewing sarcoma cells both in vitro and in vivo. 18F-FLT is transported into the cell by ENT1 and ENT2, where it is phosphorylated by TK1 and trapped intracellularly. In this report, we show that silencing of EWS-FLI1 with either siRNA or small-molecule EWS-FLI1 inhibitors suppressed the expression of ENT1, ENT2, and TK1 and thus decreased 18F-FLT PET activity. This effect was not through a generalized loss in viability or metabolic suppression, as there was no suppression of 18F-FDG PET activity and no suppression with chemotherapy. These results provide the basis for the clinical translation of 18F-FLT as a companion biomarker of EWS-FLI1 activity and a novel diagnostic imaging approach for Ewing sarcoma.
Similar content being viewed by others
Introduction
Ewing sarcoma has a somatic mutation frequency that is among the lowest of human cancers, with few recurrent traditional drug targets1,2,3. The defining molecular feature of this tumor, in 85% of cases, is the balanced chromosomal translocation between the EWSR1 gene and the FLI1 gene, which generates the EWS-FLI1 oncogenic transcription factor4. Silencing of EWS-FLI1 has been shown to limit anchorage-independent growth, to put the cell in a more dedifferentiated state, and to limit the ability of the cells to continue to proliferate5. Therefore, efforts have been focused on developing small-molecule inhibitors of EWS-FLI1 transcriptional activity with the hope of reversing the malignant phenotype in patients6,7,8,9,10,11.
Three compounds that suppress the gene signature of EWS-FLI1, cytarabine, trabectedin, and mithramycin, have all been evaluated in patients12,13. In all three cases, it is unclear if the compounds achieved the exposure necessary to block EWS-FLI1 activity, either due to patient selection, schedule of administration, or toxicity12. As more compounds and approaches to target EWS-FLI1 move to the clinic, there is a great need for a pharmacodynamic marker of EWS-FLI1 activity to determine whether the therapy is suppressing the desired target. Ideally, this biomarker would be imaging-based to avoid the need for costly, time-limited, and invasive biopsies. This would allow adaptive clinical trial designs, and perhaps even patient-specific dosing, that may improve the therapeutic window.
In this report, we characterize the PET tracer 18F-FLT as a pharmacodynamic marker of EWS-FLI1 activity. We show that the proteins responsible for activity of this tracer, ENT1, ENT2 and TK1 are driven by EWS-FLI1 in Ewing sarcoma cells. We demonstrate that expression of these proteins correlates with the activity of 18F-FLT by positron emission tomography both in vitro and vivo. We link suppression of PET activity to EWS-FLI1 and exclude non-specific causes to establish 18F-FLT as a non-invasive biomarker of EWS-FLI1 activity.
Results
EWS-FLI1 induces expression of the proteins responsible for 18F-FLT PET activity in Ewing sarcoma cells
We reasoned that because EWS-FLI1 alters the expression of a percentage of the transcriptome, a commercially available PET tracer could likely be used to monitor EWS-FLI1 activity14. We hypothesized that we could identify a surface protein driven by EWS-FLI1 in Ewing sarcoma cells that could be labeled with a radioactively tagged transport substrate, ligand, or antibody (Fig. 1A). This would allow the measurement of expression before and after drug treatment to quantify EWS-FLI1 suppression. We linked available tracers to the proteins responsible for their PET activity and used qPCR to see if any were suppressed upon siRNA silencing of EWS-FLI1 in three Ewing sarcoma cell lines. We found limited suppression of the majority of these targets, except for three proteins responsible for 18F-FLT activity, all of which were significantly suppressed with silencing of EWS-FLI1 (see arrows, Fig. 1B; statistics in Supplementary Table 1).
(A) Schema showing theory of study. Suppression of an EWS-FLI1 target expressed on the cell surface will allow quantitation of EWS-FLI1 activity based on loss of transport of a labeled substrate or loss of binding of a labeled antibody or ligand. (B) Heat map summarizing screen results. EWS-FLI1 was silenced in three different cell lines (TC32, TC71 and EW8) and the induction (red) or suppression (blue) of expression was measured by qPCR. ENT1, ENT2, and TK1 (arrows) were all suppressed with silencing of EWS-FLI1. (C) Confirmatory PCR in 4 cell lines showing suppression of ENT1, ENT2, and TK1 with siRNA silencing of EWS-FLI1 as measured by qPCR. (P for all treatments vs. no change) (D) Western blots confirm suppression of ENT1, ENT2, and TK1 with siRNA silencing of EWS-FLI1.
18F-FLT is a labeled thymidine analog that was first identified by Shields et al. as a marker of tumor proliferation15. The tracer is transported into cells by ENT1 and ENT2 and to a lesser extent by CNT1 and CNT316. Once inside the cell, the tracer is phosphorylated by TK1 and thus trapped. It is not incorporated into newly synthesized DNA and is less prone to catabolism due to the replacement of the hydroxyl group at the 3′ position with the fluorine tag17. Therefore, the tracer is retained in cells and the tumor is measureable on PET.
The tracer reflects proliferation because TK1 activity increases during S-phase, and several studies link Ki67 staining to 18F-FLT activity18,19. Indeed, this relationship has been exploited to follow tumor proliferation for targeted agents including mTOR, EGFR, and BRAF, among others20,21,22,23. Although FLT activity is multifactorial and clearly dependent on nucleoside transport, the major determinant of its PET activity is likely related to TK1 activity, because passive diffusion out of the cell would be possible in the absence of TK1. Further, there is a correlation between 18F-FLT PET activity and TK1 activity but not 18F-FLT PET activity and nucleoside transport protein expression23,24. One study established the importance of context for the activity of the tracer, showing that it is most sensitive in the setting of high nucleoside transporter expression, high TK1, and an agent that induced cell cycle arrest23.
We confirmed that silencing of EWS-FLI1 repressed the mRNA expression of ENT1, ENT2, and TK1 in four Ewing sarcoma cell lines (Fig. 1C; statistics in Supplementary Table 2). TK1, the best indicator of 18F-FLT PET activity, was the most suppressed in all lines tested. Furthermore, TK1 has been independently linked to EWS-FLI1 in a previous study25. We confirmed suppression of ENT1, ENT2, and TK1 at the protein level in four Ewing sarcoma cell lines (Figs 1D and 2A). Again, TK1 was the most suppressed; ENT2 and ENT1 showed only modest suppression.
(A) Treatment of TC32 Ewing sarcoma cells with 50 nM mithramycin (MMA) phenocopied the suppression of ENT1, ENT2, and TK1 expression that occurs with siRNA silencing of EWS-FLI1. (B) Treatment of TC32 Ewing sarcoma cells with the mithramycin analogs EC-8042 and EC-8105 suppressed ENT1, ENT2, and TK1, but 5-FU induced their expression (P for all treatments vs. no change) (C) Expression of ENT1, ENT2, and TK1 was unchanged or induced in 4 control cell lines that do not express EWS-FLI1 with treatment with mithramycin or 5-FU.
Small molecule EWS-FLI1 inhibitors block expression of ENT1, ENT2 and TK1 in Ewing sarcoma cells but not in EWS-FLI1 negative control cell lines
In order for the clinical potential of this assay to be realized, we next needed to show that small molecules targeting EWS-FLI1 suppressed expression of these targets. We previously identified mithramycin as an EWS-FLI1 inhibitor in a high-throughput screen9. Mithramycin suppressed the expression of ENT1, ENT2, and TK1 in TC32 Ewing sarcoma cells to a similar extent as siRNA silencing of EWS-FLI1 (Fig. 2A). In addition, we showed that the second-generation mithramycin analogs EC-8042 and EC-8105 also suppressed expression of those three molecules in a highly significant manner (Fig. 2B; statistics in Supplementary Table 3a)26. In order to exclude nonspecific cytotoxicity as a cause of the suppression, we also showed that the chemotherapeutic agent 5FU did not suppress expression of TK1, despite a known sensitivity of Ewing sarcoma cells to the cytotoxic effects of the drug27.
In order to exclude a general suppression of transcription as the cause and to strengthen the link to EWS-FLI1, we evaluated how treatment with mithramycin or 5-FU affected expression of these proteins in four cell lines (MCF7, A2058, RH30, and RD) that do not express EWS-FLI1. We found no suppression of ENT1, ENT2, or TK1 with either agent in any of those cell lines (Fig. 2C).
EWS-FLI1 blockade leads to suppression of 18F-FLT activity but not 18F-FDG activity in vitro independent of effects on the cell cycle
Next, we showed that the suppression of EWS-FLI1 and of ENT1, ENT2, and TK1 expression translated into suppression of 18F-FLT activity in Ewing sarcoma cells in vitro. 18F-FLT activity was markedly suppressed with silencing of EWS-FLI1 by siRNA, but not when a non-targeting siRNA (control) was used (Fig. 3A). As additional evidence for the specificity of this effect for EWS-FL1, we showed that the suppression of 18F-FLT was phenocopied with EWS-FLI1-directed small molecules. We showed a highly significant suppression of 18F-FLT activity upon treatment with 50 nM mithramycin, 50 nM EC-8042, and 15 nM EC-8105 (Fig. 3A; statistics in Supplementary Table 4a). As further evidence of specificity, we treated cells with the chemotherapeutic agent 5-FU. Again, no suppression of 18F-FLT activity was found; in fact, its PET activity was induced consistent with known effects of 5-FU28 (Fig. 3A).
(A) 18F-FLT PET activity decreased in vitro in TC32 Ewing sarcoma cells with siRNA silencing of EWS-FLI1(siEWS) but not with a non-targeting control (siNeg); EWS-FLI1 suppression with mithramycin (MMA), EC-8042, or EC-8105 decreases 18F-FLT activity (B) 18F-FDG activity does not change with siRNA silencing of EWS-FLI1 and with EWS-FLI1 suppression with mithramycin (MMA), EC-8042, or EC-8105. 5FU suppresses 18F-FDG activity likely due to a loss of cell viability.
As an additional control for cell viability and non-specific metabolic effects as a cause for the suppression of 18F-FLT activity, we also evaluated the effect of siRNA silencing of EWS-FLI1 on the activity of the commonly employed PET tracer 18F-labeled fluorodeoxyglucose (18F-FDG). We found no significant suppression of 18F-FDG activity with EWS-FLI1 silencing by siRNA. Additionally, treatment of TC32 cells with 50 nM MMA, 50 nM EC8042 and 15 nM EC8105 for the same duration utilized above caused no significant suppression of 18F-FDG activity (Fig. 3B; statistics in Supplementary Table 4b). In contrast, since the 5-FU treatment suppressed cell viability, there was suppression of 18F-FDG activity.
In other cell contexts, 18F-FLT has been used to follow cell cycle progression because TK1 activity increases in late G1 and S phase29,30. Indeed it has been shown that in A549 carcinoma cells, 18F-FLT PET activity increases with an increasing percentage of cells in S-phase24. Since a major function of EWS-FLI1, particularly for the induced gene targets, is to modulate the cell cycle we needed to control for non-specific cell cycle changes as the cause of 18F-FLT suppression14,31. In this setting more rapid progression through the cell cycle would be associated with a loss of 18F-FLT PET activity.
Therefore, we needed to show that the suppression of FLT activity did not reflect a nonspecific more rapid progression through the cell cycle. In all cases, we found cell cycle arrest. Silencing of EWS-FLI1 with siRNA caused a marked arrest of TC32 cells in G1, from 40.8% (SEM +/− 3.0) to 60.8 (SEM +/− 3.1) (Fig. 4A,B). This G1 arrest was phenocopied with mithramycin and EC-8042 in particular (Fig. 4B). In contrast, EC-8105 did not cause a G1 arrest and instead increased the percentage of cells in S-phase from 17.4 (SEM +/− 1.7) to 25.8 (SEM +/− 3). One would therefore predict an increase in 18F-FLT activity with these treatments based on the accumulation of cells in late G1 or S that would be expected to increase TK1 expression and activity. However, in every instance where EWS-FLI1 was suppressed 18F-FLT activity decreased regardless of which phase in the cell cycle the cells were arrested. These results show that in Ewing sarcoma cells, blockade of EWS-FLI1 leads to an inability of cells to take up and retain 18F-FLT and this effect is the major determinant of activity in this cell type in the setting of EWS-FLI1 directed therapy.
The suppression of 18F-FLT activity only occurs with EWS-FLI1 suppression and can not be explained by non-specific changes in cell cycle progression (A) Cell cycle changes in TC32 Ewing sarcoma cells that occur with siRNA silencing of EWS-FLI1 or treatment with 50 nM mithramycin (MMA) or the mithramycin analogs EC8105 (15 nM) or EC8042 (50 nM). (B) Cell counts as percentage of total (+/− SD). Grey boxes highlight treatments that increase the number of cells in G1 or S. Treatments identical to Fig. 3A. (C) Treatment of TC32 Ewing sarcoma cells with non-specific chemotherapy: 2 nM vincristine (VCR), 80 nM etoposide (etop) or 50 nM doxorubicin (doxo) does not suppress 18F-FLT activity in comparison to mithramycin (MMA). Cells treated equitoxic concentrations.
Finally, as an additional control, we treated cells with chemotherapy that does not target EWS-FLI1 and is used in the clinic to treat Ewing sarcoma. We treated TC32 Ewing sarcoma cells with 2 nM vincristine, 80 nM etoposide and 50 nM, or concentrations around the IC50 (but before substantial cell death) and found no significant suppression of 18F-FLT PET activity (Fig. 4C). This suppression correlated with the small suppression of viability at this concentration and time of exposure. It is notable that at higher doses, treatment is associated with a loss of viability and therefore non-specific suppression of 18F-FLT (data not shown).
EWS-FLI1 blockade causes a loss of 18F-FLT activity that correlates with TK1 supression in xenograft models of Ewing sarcoma
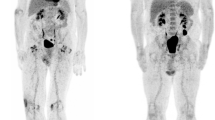
To demonstrate the clinical utility of this assay, we showed that these effects translated to the in vivo setting in our xenograft model of Ewing sarcoma. Mice were implanted with TC71 xenografts in the left gastrocnemius muscle. Once the tumors were established, they were treated with 1 mg/kg mithramycin, 24 mg/kg EC-8042, or 1.5 mg/kg of EC-8105, all by the intraperitoneal route. Again, marked suppression of 18F-FLT activity was observed in vivo (particularly at 24 h) and was most prominent with EC-8042, consistent with an EWS-FLI1-specific effect (Fig. 5A; Supplementary Figs 1 and 2). Cross-section imaging indicated that similar amounts of tracer were administered to the animals (Fig. 5A). Suppression was not observed with 50 mg/kg of intraperitoneal 5-FU, which markedly induced 18F-FLT activity in these Ewing sarcoma xenografts, particularly at 6 h of treatment (Supplementary Fig. 3). Recovery to normal FLT activity occurred at 48 h following treatment (data not shown). Staining of tissue for TK1 showed its suppressed expression upon drug treatment, consistent with in vitro data (Fig. 5B). Interestingly, EC-8042 showed the greatest degree of suppression of 18F-FLT activity and the most marked changes in staining, which correlates with the most profound response to treatment that we recently reported26. In contrast, the staining was more variable with EC-8105 and mithramycin, consistent with the relative activity of these drugs by the intraperitoneal route (Supplementary Fig. 4). Although this correlation with response would need to be tested in a prospective fashion preferably in patients.
(A) Treatment of mice bearing a Ewing sarcoma xenograft with EC-8042 suppressed PET avidity of the xenograft at 24 h of treatment. The bladder in cross section (lower panel) shows activity consistent with similar amounts of tracer administered. (B) Suppression of activity correlates with TK1 staining (top) in EC-8042-treated tissue but not control or 5-FU treated tissues (10× view). H&E staining confirms tumor tissue (bottom). Size bar indicates 200 micrometer.
Discussion
18F-FLT is a PET tracer originally identified by Shields et al. that has found widespread usefulness in the clinic as a pharmacodynamic marker of cell cycle progression and perhaps even a surrogate for ki67 staining15,18,32. This tracer is complementary to 18F-FDG and works particularly well in tumors that show high TK1 activity or in areas of the body associated with high 18F-FDG activity such as the brain that limit the use of 18F-FDG33.
In this report, we propose an alternative use of this tracer. We capitalize on the fact that in Ewing sarcoma cells EWS-FLI1 drives expression of the proteins responsible for the transport and retention of the 18F-FLT tracer within the cell. We therefore utilize the tracer to reflect expression of these targets and as a result EWS-FLI1 activity. We show that silencing of EWS-FLI1 leads to a loss in expression of TK1, ENT2 and ENT1. This effect occurred in multiple Ewing sarcoma cell lines with siRNA targeting EWS-FLI1 and with multiple small molecules that inhibit EWS-FLI134. The effect was not the result of nonspecific suppression of transcription of ENT1, ENT2 or TK1 by mithramycin as it did not occur in 4 control cell lines. Additionally, we exclude general cytotoxicity or a general suppression of metabolism as the mechanism of 18F-FLT suppression and show no effect on 18F-FDG at the same concentration and time of treatment with either siRNA directed against EWS-FLI1 or small molecules that inhibit EWS-FLI1. Finally, we exclude cell cycle perturbations as the cause of suppression by showing that mithramycin and its analogs arrest the cell cycle which would in theory increase 18F-FLT activity but instead cause marked suppression of 18F-FLT PET activity.
This tracer will provide investigators with an important tool to facilitate the development of new EWS-FLI1-directed therapies. It will afford the opportunity to determine if a given targeted compound is able to achieve EWS-FLI1 suppression without the need for invasive biopsies. This could potentially allow for patient-specific dosing regimens that broaden the therapeutic window to maximize target suppression and minimize toxicity. Alternatively, the tracer may be used to optimize the schedule of drug administration to either maintain EWS-FLI1 suppression or optimize combination therapies. We have previously reported a combination therapy that capitalizes on EWS-FLI1 suppression to target the toxicity of irinotecan to Ewing sarcoma cells based on the suppression of expression of the WRN gene35. It is likely that 18F-FLT could be used to optimize the schedule of administration of this combination such that irinotecan could be given at the point of maximal EWS-FLI1 suppression to yield the maximum clinical benefit.
The use of PET imaging to quantitate target suppression has substantial advantages over biopsy-based approaches. Although pre- and post treatment tissues sections could be used to evaluate EWS-FLI1 downstream target expression, it is difficult to obtain biopsies for research purposes because of perceived risks to the patient. Furthermore, the process of tissue collection and staining is time and labor intensive and requires several days to yield the final result. This limits the utility of the biopsy-based approach as a method to make patient-specific treatment decisions.
Finally, this tracer may also find utility as a diagnostic agent for Ewing sarcoma. Since EWS-FLI1 is the dominant oncogene of Ewing sarcoma, one would expect these tumors to be extremely 18F-FLT PET-avid. Therefore, 18F-FLT may ultimately serve as a sensitive biomarker for relapse to complement 18F-FDG PET. This tracer would be particularly sensitive for metastases in areas such as the brain that show high 18F-FDG activity that limits the use of this tracer in these locations. In addition, since tumor relapse can often be confused with processes like infection that also increase in 18F-FDG activity, 18F-FLT might be used as a complementary agent to help distinguish these two clinical scenarios (although this question would need to be formally studied). Finally, in cases where obtaining a diagnostic specimen may require an invasive procedure, 18F-FLT may provide additional evidence to justify the procedure.
In summary, this report provides the basis for the clinical evaluation of 18F-FLT as a biomarker for Ewing sarcoma. The tracer will aid in the development of therapies that target the dominant oncogene of Ewing sarcoma, EWS-FLI1. In addition, 18F-FLT may emerge as an important diagnostic tool for the detection of Ewing sarcoma in patients.
Methods
Cell Lines, Cell Culture, and Reagents
TC32, TC71, and A673 Ewing sarcoma cells were the gift of Dr. T. Triche (The Saban Research Hospital, Children’s Hospital of Los Angeles, CA). EW8 Ewing sarcoma cells were the gift of Peter Houghton (Nationwide Children’s Hospital, Columbus, OH). The MCF7 breast carcinoma cell line was the gift of Dr. Patricia Steeg (National Cancer Institute, Bethesda, MD). The RH30 and RD rhabdomyosarcoma cells were the gift of Lee Helman (National Cancer Institute, Bethesda, MD). The A2058 melanoma cell line was obtained from the American Type Culture Collection (ATCC, Manassas, VA). The identity of all cells was independently authenticated by short tandem repeat genotyping. All cells were maintained in culture in RPMI 1640 (Invitrogen, Carlsbad, CA). The medium was supplemented with 10% fetal bovine serum (Gemini Bio-Products, West Sacramento, CA), 2 mM L-Gln, 100 U/mL of penicillin and 100 μg/mL of streptomycin (ThermoFisher, Waltham, MA).
Compounds
Mithramycin and all analogs were obtained from EntreChem SL (Oviedo, Spain). 5-Fluorouracil, vincristine, doxorubicin and etoposide were obtained from Sigma Aldrich (St. Louis, MO). All compounds were aliquoted, stored frozen, and thawed immediately before use.
siRNA silencing
RNAiMax Lipofectamine (Life Technologies, Carlsbad, CA) was added to siRNA targeting the EWS-FLI1 breakpoint site II36, complexed, and combined with cells. Cells were incubated for 24 h before collection for RT-PCR.
Quantitative RT-PCR
TC32 cells (0.3 × 106) were exposed to siRNA or the indicated compound. RNA was collected using the RNEasy Kit with QIAshredder (Qiagen, Valencia, CA), immediately reverse-transcribed using a High Capacity cDNA Synthesis Kit (Life Technologies, Carlsbad CA) on a Veriti thermocycler (Life Technologies), and PCR-amplified using SYBR green master mix (BioRad, Hercules, CA) and the CFX 384 Real Time System (BioRad), The program was 95 °C for 10 min, 95 °C for 30 s, 55 °C for 30 s, and 72 °C for 30 s, for 40 cycles. The expression of target genes was determined using standard ΔΔCT methods and normalized to GAPDH control.
Immunoblot Analysis
TC32 cells (1.5 × 106) were incubated with drug, collected, washed with PBS, lysed, and boiled in 4% LDS buffer (0.125 M Trizma hydrochloride buffer solution, pH 7.5, and 4% lithium dodecyl sulfate (Sigma-Aldrich). Protein concentrations were determined after diluting the detergent using the bicinchoninic acid (BCA) assay kit (Pierce Protein Biology Products, Carlsbad, CA). Thirty micrograms of protein was resolved on a 4–12% NuPAGE Bis-Tris Mini gels (Invitrogen, Carlsbad, CA) in 1 × 4-morpholinepropanesulfonic acid (MOPS) sodium dodecyl sulfate (SDS) buffer (Invitrogen), and transferred to nitrocellulose (GE Healthcare Life Sciences, Pittsburgh, PA). The membrane was probed with the following antibodies: mouse monoclonal anti-FLI1 (1:1,000, Abcam, Cambridge, MA), mouse polyclonal anti-ACTB (1:1,000, Cell Signaling, Danvers, MA), rabbit polyclonal anti-ENT1 (1:500, Abcam), rabbit polyclonal anti-ENT2 (1:1000, Abcam), and rabbit polyclonal anti-TK1 (1:000, Santa Cruz Biotechnology, Dallas, TX). The protein was visualized by using horseradish peroxidase (HRP)-conjugated secondary antibody and ECL (Amersham, Buckinghamshire, UK).
Cell-cycle analysis
TC32 cells were plated, allowed to recover overnight, and then incubated with drug at the specified concentration 24 h. Following collection in PBS, the cells were fixed in 70% ethanol for 3 h at 4 °C, washed with PBS, and stained with propidium iodide (0.02 mg/mL) in 0.1% Triton X-100 with RNAse A for 30 min at room temperature. The cells were filtered and analyzed by with the Fortessa analytical cytometer (BD Biosciences, San Jose, CA).
18F-FLT Radiosynthesis
18F-FLT was prepared as previously described either internally or at Spectron MRC (South Bend, IN)37. 18F-FLT was obtained with average radiochemical purity of 98.5%.
FLT activity in cell line
TC32 cells were plated, allowed to recover overnight, and then incubated with drug (mithramycin or chemotherapy) at the specified concentration for 18 h. A second set of TC32 cells was plated following the siRNA protocol and incubated for 24 h. The medium was aspirated and the cells were incubated with 25–30 μCi for 1 h at 37 °C. The cells were collected, washed, and suspended in PBS. 18F-FLT activity was measured using a gamma well counter (Capintec, Ramsey NJ).
Mouse PET Imaging
Two million TC71 cells were injected intramuscularly in the left gastrocnemius of female 6-week-old female homozygous nude mice (Crl; Nu-Foxn1Nu) (Charles River Laboratories, Fredrick, MD). Tumors were established to a minimum diameter of 0.5 cm. Seven cohorts of five mice wwere then treated with vehicle; 1 mg/kg of EC8105 or mithramycin, 24 mg/kg of EC8042, or 50 mg/kg of 5-fluorouracil, administered intraperitoneally immediately after baseline image. The mice were imaged at 6 h, 24 h, and 48 h after initial treatment. Mice were injected retro-orbitally with ~7 MBq/0.2 mL 18F-FLT immediately after treatment/sham, at 24 hours, and at 48 hours post treatment/sham. Forty minutes later, the mice were anesthetized with 2% isofluorane and positioned in an Inveon PET/CT (Siemens Preclinical, Knoxville TN). The mice were imaged for 20 min in static mode. Data from all possible lines of response (LOR) were saved in the list mode raw data format. The raw data was then binned into 3D sinograms with a span of 3 and ring difference of 47. The images were reconstructed into transaxial slices (128 × 128 × 95) with voxel sizes of 0.095 × 0.095 × 0.08 cm3 using an iterative ordered subsets expectation maximization (OS-EM 2D) algorithm with 16 subsets and 4 iterations. Immediately following the PET scans, a CT image was obtained for anatomical co-registration, with an x-ray voltage of 80 kVp and an x-ray beam intensity of 25 mAs. The CT images were reconstructed into 384 × 384 × 417 voxels at a voxel size of 0.114055 × 0.114055 × 0.114055 mm3. The mice were sacrificed after the last image was obtained and the tissue was fixed as below. All experiments were performed in accordance with the guidelines and regulation of, and were approved by, the Animal Care and Use Committee at Vanderbilt University, Nashville, TN, or in accordance with Animal Care and Use Committee of the Southern Research Institute. Investigators who completed the imaging analysis were blinded to the treatment groups; the remaining investigators were not.
Data Analysis
PET/CT data were co-registered in Amide (www.sourgeforge.com) based on bed position. Three dimensional regions-of-interest (ROIs) were drawn around the entire tumor and around muscle (opposite hind limb) in the CT images and superimposed onto the PET images.
Immunofluorescence or immunohistochemistry
Tumors were collected, immediately fixed in 10% formalin for 48 h, transferred to 70% ethanol, and embedded in paraffin. Antigen retrieval and primary staining was performed on the Ventana automated stainer using a rabbit anti-TK1 antibody (Abcam). The tissue sections were visualized using an Aperio imager (Leica, Buffalo Grove, IL).
Statistical Methods
All statistical analyses were performed using R v 3.2.2. To test for changes in gene expression, two-sided one-sample t-tests were used to determine if the fold-change of expression was significantly different from 1.0. False-discovery rate corrections were used to adjust for multiple testing. False coverage rate adjusted intervals are presented instead of confidence intervals so that interval estimates are also adjusted appropriately for multiple testing. The FDG experiment was analyzed via standard one-way ANOVA and, due to heteroscedasiticy identified via Bartlett’s test, the FLT experiment was analyzed via robust regression using an M-estimator. False discovery rate corrected linear contrasts used for post-hoc testing on both the FDG and FLT data. Normality was assessed visually, no concerning deviations were identified. Fold-change appeared to be more normally distributed than log fold-change, so these values were not transformed. The level of significance was set at 0.05. FLT chemotherapy response was analyzed via Kruskal-Wallis with Dunn’s multiple comparisons test relative to Medium.
Additional Information
How to cite this article: Osgood, C. L. et al.18F-FLT Positron Emission Tomography (PET) is a Pharmacodynamic Marker for EWS-FLI1 Activity and Ewing Sarcoma. Sci. Rep. 6, 33926; doi: 10.1038/srep33926 (2016).
References
Tirode, F. et al. Genomic Landscape of Ewing Sarcoma Defines an Aggressive Subtype with Co-Association of STAG2 and TP53 Mutations. Cancer Discov 4, 1342–1353 (2014).
Crompton, B. D. et al. The genomic landscape of pediatric ewing sarcoma. Cancer Discov 4, 1326–1341 (2014).
Brohl, A. S. et al. The genomic landscape of the Ewing Sarcoma family of tumors reveals recurrent STAG2 mutation. PLoS Genet 10, e1004475 (2014).
Delattre, O. et al. Gene fusion with an ETS DNA-binding domain caused by chromosome translocation in human tumours. Nature 359, 162–165 (1992).
Maksimenko, A. & Malvy, C. Oncogene-targeted antisense oligonucleotides for the treatment of Ewing sarcoma. Expert Opin Ther Targets 9, 825–830 (2005).
Stegmaier, K. et al. Signature-based small molecule screening identifies cytosine arabinoside as an EWS/FLI modulator in Ewing sarcoma. PLoS Med 4, e122 (2007).
Erkizan, H. V. et al. A small molecule blocking oncogenic protein EWS-FLI1 interaction with RNA helicase A inhibits growth of Ewing’s sarcoma. Nat Med 15, 750–756 (2009).
Grohar, P. J. et al. Ecteinascidin 743 interferes with the activity of EWS-FLI1 in Ewing sarcoma cells. Neoplasia 13, 145–153 (2011).
Grohar, P. J. et al. Identification of an Inhibitor of the EWS-FLI1 Oncogenic Transcription Factor by High-Throughput Screening. J Natl Cancer Inst 103, 962–978 (2011).
Boro, A. et al. Small-molecule screen identifies modulators of EWS/FLI1 target gene expression and cell survival in Ewing’s sarcoma. Int J Cancer 131, 2153–2164 (2012).
Chen, C., Wonsey, D. R., Lemieux, M. E. & Kung, A. L. Differential disruption of EWS-FLI1 binding by DNA-binding agents. PLoS One 8, e69714 (2013).
DuBois, S. G. et al. Phase II study of intermediate-dose cytarabine in patients with relapsed or refractory Ewing sarcoma: a report from the Children’s Oncology Group. Pediatr Blood Cancer 52, 324–327 (2009).
Baruchel, S. et al. A phase 2 trial of trabectedin in children with recurrent rhabdomyosarcoma, Ewing sarcoma and non-rhabdomyosarcoma soft tissue sarcomas: a report from the Children’s Oncology Group. Eur J Cancer 48, 579–585 (2012).
Kauer, M. et al. A molecular function map of Ewing’s sarcoma. PLoS One 4, e5415 (2009).
Shields, A. F. et al. Imaging proliferation in vivo with [F-18]FLT and positron emission tomography. Nat Med 4, 1334–1336 (1998).
Paproski, R. J. et al. The role of human nucleoside transporters in uptake of 3′-deoxy-3′-fluorothymidine. Mol Pharmacol 74, 1372–1380 (2008).
Grierson, J. R., Schwartz, J. L., Muzi, M., Jordan, R. & Krohn, K. A. Metabolism of 3′-deoxy-3′-[F-18]fluorothymidine in proliferating A549 cells: validations for positron emission tomography. Nucl Med Biol 31, 829–837 (2004).
Wagner, M. et al. 3′-[18F]fluoro-3′-deoxythymidine ([18F]-FLT) as positron emission tomography tracer for imaging proliferation in a murine B-Cell lymphoma model and in the human disease. Cancer Res 63, 2681–2687 (2003).
Chalkidou, A. et al. Correlation between Ki-67 immunohistochemistry and 18F-fluorothymidine uptake in patients with cancer: A systematic review and meta-analysis. Eur J Cancer 48, 3499–3513 (2012).
Jensen, M. M. & Kjaer, A. Monitoring of anti-cancer treatment with (18)F-FDG and (18)F-FLT PET: a comprehensive review of pre-clinical studies. Am J Nucl Med Mol Imaging 5, 431–456 (2015).
McKinley, E. T., Zhao, P., Coffey, R. J., Washington, M. K. & Manning, H. C. 3′-Deoxy-3′-[18F]-Fluorothymidine PET imaging reflects PI3K-mTOR-mediated pro-survival response to targeted therapy in colorectal cancer. PLoS One 9, e108193 (2014).
McKinley, E. T. et al. 3′-Deoxy-3′-18F-fluorothymidine PET predicts response to (V600E)BRAF-targeted therapy in preclinical models of colorectal cancer. J Nucl Med 54, 424–430 (2013).
Zhang, C. C. et al. [(18)F]FLT-PET imaging does not always “light up” proliferating tumor cells. Clin Cancer Res 18, 1303–1312 (2012).
Rasey, J. S., Grierson, J. R., Wiens, L. W., Kolb, P. D. & Schwartz, J. L. Validation of FLT uptake as a measure of thymidine kinase-1 activity in A549 carcinoma cells. J Nucl Med 43, 1210–1217 (2002).
Bilke, S. et al. Oncogenic ETS fusions deregulate E2F3 target genes in Ewing sarcoma and prostate cancer. Genome Res 23, 1797–1809 (2013).
Osgood, C. L. et al. Identification of mithramycin analogs with improved targeting of the EWS-FLI1 transcription factor. Clin Cancer Res (2016).
Deneen, B., Hamidi, H. & Denny, C. T. Functional analysis of the EWS/ETS target gene uridine phosphorylase. Cancer Res 63, 4268–4274 (2003).
Lee, S. J. et al. Induction of thymidine kinase 1 after 5-fluorouracil as a mechanism for 3′-deoxy-3′-[18F]fluorothymidine flare. Biochem Pharmacol 80, 1528–1536 (2010).
Wang, N., He, Q., Skog, S., Eriksson, S. & Tribukait, B. Investigation on cell proliferation with a new antibody against thymidine kinase 1. Anal Cell Pathol 23, 11–19 (2001).
Gasparri, F., Wang, N., Skog, S., Galvani, A. & Eriksson, S. Thymidine kinase 1 expression defines an activated G1 state of the cell cycle as revealed with site-specific antibodies and ArrayScan assays. Eur J Cell Biol 88, 779–785 (2009).
Kowalewski, A. A., Randall, R. L. & Lessnick, S. L. Cell Cycle Deregulation in Ewing’s Sarcoma Pathogenesis. Sarcoma 2011, 598704 (2011).
Bollineni, V. R., Kramer, G. M., Jansma, E. P., Liu, Y. & Oyen, W. J. A systematic review on [(18)F]FLT-PET uptake as a measure of treatment response in cancer patients. Eur J Cancer 55, 81–97 (2016).
Li, Z., Yu, Y., Zhang, H., Xu, G. & Chen, L. A meta-analysis comparing 18F-FLT PET with 18F-FDG PET for assessment of brain tumor recurrence. Nucl Med Commun 36, 695–701 (2015).
Osgood, C. L. et al. Identification of Mithramycin Analogues with Improved Targeting of the EWS-FLI1 Transcription Factor. Clin Cancer Res (2016).
Grohar, P. J. et al. Dual targeting of EWS-FLI1 activity and the associated DNA damage response with trabectedin and SN38 synergistically inhibits Ewing sarcoma cell growth. Clin Cancer Res 20, 1190–1203 (2014).
Dohjima, T., Lee, N. S., Li, H., Ohno, T. & Rossi, J. J. Small interfering RNAs expressed from a Pol III promoter suppress the EWS/Fli-1 transcript in an Ewing sarcoma cell line. Mol Ther 7, 811–816 (2003).
Cheung, Y. Y., Nickels, M. L., McKinley, E. T., Buck, J. R. & Manning, H. C. High-yielding, automated production of 3′-deoxy-3′-[(18)F]fluorothymidine using a modified Bioscan Coincidence FDG reaction module. Appl Radiat Isot 97, 47–51 (2015).
Acknowledgements
The authors thank David Nadziejka for technical editing of this manuscript. The authors would like to thank EntreChem SL for providing the mithramycin analogs. We thank Lisa Turner and Galen Hostetter of the VARI Pathology Core for assistance developing the immunohistochemistry assay. The work was supported by a grant from the Alex’s Lemonade Stand Foundation Reach Award (PJG). Funding was also provided by the T32 training grant conducting research in pediatric oncology, as well as the Vanderbilt Clinical Oncology Research Career Development K12 (CLO). Additional internal funds were provided by the Vanderbilt University Department of Pediatrics and the Lily’s Garden Foundation (PJG, CLO, KNM). Vanderbilt Center for Molecular Probes and the Kleburg Foundation (HCM).
Author information
Authors and Affiliations
Contributions
The following contributions were made by the authors of the manuscript: Study Conception and Design: P.J.G., C.L.O., M.N.T., J.B., M.E.W. and H.C.M. Data Gathering and Interpretation: P.J.G., C.L.O., N.M., A.P., E.B., J.J., M.N.T. and H.C.M. Manuscript Preparation: P.J.G., C.L.O., M.N.T., Z.B.M. and H.C.M. Manuscript review: All authors reviewed the manuscript. Manuscript Editing: P.J.G., C.L.O. and Z.B.M. Study Supervision: P.J.G.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Osgood, C., Tantawy, M., Maloney, N. et al. 18F-FLT Positron Emission Tomography (PET) is a Pharmacodynamic Marker for EWS-FLI1 Activity and Ewing Sarcoma. Sci Rep 6, 33926 (2016). https://doi.org/10.1038/srep33926
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep33926
This article is cited by
-
A phase I/II trial and pharmacokinetic study of mithramycin in children and adults with refractory Ewing sarcoma and EWS–FLI1 fusion transcript
Cancer Chemotherapy and Pharmacology (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.