Abstract
The cytochrome (cyt) bc1 complex is an integral component of the respiratory electron transfer chain sustaining the energy needs of organisms ranging from humans to bacteria. Due to its ubiquitous role in the energy metabolism, both the oxidation and reduction of the enzyme’s substrate co-enzyme Q has been studied vigorously. Here, this vast amount of data is reassessed after probing the substrate reduction steps at the Qi-site of the cyt bc1 complex of Rhodobacter capsulatus using atomistic molecular dynamics simulations. The simulations suggest that the Lys251 side chain could rotate into the Qi-site to facilitate binding of half-protonated semiquinone – a reaction intermediate that is potentially formed during substrate reduction. At this bent pose, the Lys251 forms a salt bridge with the Asp252, thus making direct proton transfer possible. In the neutral state, the lysine side chain stays close to the conserved binding location of cardiolipin (CL). This back-and-forth motion between the CL and Asp252 indicates that Lys251 functions as a proton shuttle controlled by pH-dependent negative feedback. The CL/K/D switching, which represents a refinement to the previously described CL/K pathway, fine-tunes the proton transfer process. Lastly, the simulation data was used to formulate a mechanism for reducing the substrate at the Qi-site.
Similar content being viewed by others
Introduction
To maintain diverse and complex cellular functions such as reproduction, growth or movement, all living organisms rely on constant supply of energy. The fundamentals of this life-sustaining energy metabolism or bioenergetics are known to a large extent, but the mechanistic details of relevant enzymatic reactions are still being actively studied and debated on. The membrane-embedded cytochrome (cyt) bc1 complex (or complex III; Fig. 1A) is a crucial component of the respiratory and photosynthetic electron transfer chains sustaining the energy requirements of both eukaryotes and bacteria.
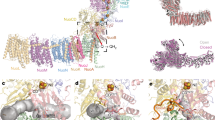
The cytochrome bc1 complex, proton/electron transfers of the Q-cycle and the CL/K proton transfer pathway.
(A) The dimer complex includes the cyt b (red/blue), cyt c1 (yellow/orange) and iron sulfur protein (ISP; cyan/magenta) subunits (PDB: 1ZRT)22. The Qo-site is located between the 2-iron 2-sulfur (Fe2S2) cluster and the low potential heme (bL). The Qi-site is adjacent to the high potential heme (bH). The arrows indicate the routes of the e− (orange) and H+ transfers (green). (B) The arrows indicate the direction of the H+/e− transfers during oxidation/reduction of the non-protonated Q, the half-protonated radical SQ and the fully protonated QH2 at the Qo- or Qi- sites, respectively. (C) In the CL/K pathway, lysine acquires a H+ from a cardiolipin (CL) molecule (PDB: 3CX5)9 and (D) passes it via a string of interconnected water molecules into the Qi-site to reduce the substrate (PDB: 1PP9)20. The H-bonds (≤3.4 Å) and possible bonds (≤3.6 Å) are shown with magenta and orange dotted lines, respectively. The CL-Lys251 interactions taking place at the membrane-protein periphery were considered in a previous MD simulation study8. The amino acid residues are shown as black sticks, substrate as yellow ball-and-stick representation and the heme bH group is shown as a CPK model.
The cyt bc1 complex operation or Q-cycle (named for the substrate co-enzyme Q or ubiquinone) begins when the fully protonated substrate quinol (QH2; Fig. 1B) binds into the Qo-site, where it is oxidized, i.e. it gives away two electrons (e−) and protons (H+). One e− is transferred to the prosthetic 2-iron 2-sulfur cluster of the iron sulfur protein subunit, which then passes it on to the heme c1 group in the cyt c1 subunit (Fig. 1A). Meanwhile, the other e− is routed towards the heme bL cluster and the heme bH group in the cyt b subunit (Fig. 1A). At the Qi-site of the cyt bc1 complex, the non-protonated substrate quinone (Q; Fig. 1B) acquires consecutively two electrons and, in total, two protons from the negative (N) side of the bioenergetic membrane1,2,3.
In our prior study, the binding modes of two substrate forms, QH2 and Q, were determined at the Qo-site of the cyt bc1 complex of purple photosynthetic bacterium Rhodobacter capsulatus using atomistic molecular dynamics (MD) simulations4. A highly coordinated water molecule was found to serve in a dual role both as a potential proton acceptor and as a structural gating mechanism for the short-circuit suppression. Similar arrangement was reported in a follow-up modelling study5. Likewise, coordinated water could also affect H+ transfers of the cyt c oxidase6,7.
The simulations also supported the X-ray crystallographic results by showing that cardiolipin (CL) has a conserved binding position close to the Qi-site (Fig. 1C)8,9. From this position the dianionic CL has been suggested to act as a H+ attracting antenna that feeds protons to Q being reduced at the Qi-site (Fig. 1C)10. The CL would donate protons first to Lys251 of the cyt b subunit, which then passes them to a string of interconnected water molecules leading up to the Qi-site (Fig. 1D). Utilizing the protons supplied by the CL/K pathway (Fig. 1C,D), a Q molecule is reduced to the fully protonated QH2 (Fig. 1B) via a reaction intermediate radical semiquinone (SQ; Fig. 1B). Again, lipids have been suggested to play a similar role in the H+ transfers of subunit III in the cyt c oxidase11.
There exist plenty of X-ray crystal structures showing the bound substrate at the Qi-site (Figs 1D and S1; Table S1; see Supplementary Information (SI)), but the exact reduction steps are unknown due to lack of structural data on explicit protons (Fig. 1B). To address this issue, the bacterial cyt bc1 complex was studied afresh using explicitly set up MD simulations (Table 1). Because only the anionic SQ has been detected using frozen electron paramagnetic resonance experiments at the Qi-site12,13,14,15,16, the substrate has been presumed to acquire both of the electrons (dianionic state) before accepting the two protons concomitantly17. While this scenario is possible, the other option is that the proton transfers are tightly coupled to separate e− transfers (Fig. 1B) as recently suggested by quantum mechanics calculations16. Accordingly, the half-protonated SQ (Fig. 1B) could be formed prior to the second e− transfer, but it has not been detected so far for example due to its inherent lability or short duration.
The results presented and discussed in this paper suggest that the binding of half-protonated/neutral SQ would be more stable and coordinated than that of Q and that there would be a clear advantage in reducing the C1-group for coordinating the C4-carbonyl H-bonding with His217. Importantly, the conserved residues Lys251 and Asp252 (Figs S1 and S2; Table S1) can form a salt bridge that meets the geometry criteria for a direct H+ transfer. Thus, instead of relying on an interconnected string of water molecules for the H+ transfers (Fig. 1C,D), the simulations indicate that the CL/K pathway ends in Lys251 rotating into the Qi-site and forming a salt bridge with Asp252. The empirical pKa calculations indicate that a direct H+ transfer could happen between the two ionizable residues and the lysine rotation-based H+ transfer would be pH-dependent. Moreover, the e− transfer from the heme bH group to Q slows down when Lys251 is substituted with residues unable to participate in H+ transfers18,19.
The explicitly set up simulations provide an unprecedented opportunity for reviewing prior site-directed mutagenic and X-ray crystallographic results (Table S1) regarding the Qi-site. Mechanistically the main finding of this work is the discovery of Lys251 rotation-based proton shuttle, which is a tangible refinement to the prior CL/K pathway hypothesis10. However, based on the simulations the inward rotation of Lys251 not only fine-tunes the H+ transfer process into the Qi-site, but potentially influences also the substrate binding/unbinding, reduction and could even curb the aging-related superoxide generation. Accordingly, the accumulated data was used to formulate a novel Q reduction model in which the e−/H+ transfers are tightly coupled and sequential by nature as opposed to the previously suggested concomitant model17.
Figuring out the details of the reduction reaction at the Qi-site is a necessary step for achieving complete understanding of the cyt bc1 complex Q-cycle and, on a larger scale, the electron transfer chain itself.
Results and Discussion
The substrate binding requirements at the Qi-site
The SQ/Q binding at the Qi-site is described thoroughly based on the MD simulations (Fig. 2) and X-ray crystallographic data in the SI (Fig. S1; Table S1); however, the binding requirements are summarized here:
The binding modes of semiquinone and quinone at the Qi-site of cytochrome bc1 complex.
(A) On the A side of the dimer, the C1-hydroxyl of neutral SQ (shown with yellow ball-and-stick representation) forms a water bridge with the Asp252COO−, while the C4-carbonyl H-bonds with the epsilon protonated His217. A Lys251NH3+-Asp252COO− salt bridge is formed (conf1 in Table 1). The Asn221NH2 stabilizes the His217 positioning by H-bonding. (B) On the B side, the C4-hydroxyl of SQ H-bonds with both Lys251 and Asp252 that are forming a salt bridge (conf1 in Table 1). Both His217 and Asn221 H-bond with the C4-carbonyl of SQ. (C) On the B side, Q forms a water bridge with the Asp252COO− and H-bonds with the His217 and Asn221 side chains that are also bonded to each other (conf3 in Table 1). The lysine assumes the outward rotamer pose. (D) On the B side, the C1-carbonyl of Q H-bonds with the Asp252COOH and the As221NH2 H-bonds with the C4-carbonyl (conf4 in Table 1). Neutral Lys251 and Asp252 side chains are not bonding. For clarity only the polar protons are shown (cyan color). For further details see Fig. 1.
(1) The C1-group of the substrate needs to form a H-bond or a water bridge with the Asp252 side chain (Figs 2; and S3A; Table S2). With bound neutral SQ at the Qi-site, the C1-hydroxyl would donate a hydrogen to the carboxylate group of Asp252 (or Asp252COO−; Fig. 2B). The opposite would happen with bound Q (or explicitly identical anionic SQ) when its C1-carbonyl would accept a hydrogen from the neutral Asp252 side chain (or Asp252COOH; Fig. 2D) or from a water molecule neighboring the Asp252COO− (Fig. 2C).
(2) The His217 and/or Asn221 side chains H-bond with the C4-carbonyl of Q, in order to orient the substrate’s quinone ring to H-bond with the Asp252COOH. Unlike the X-ray crystallographic data (Fig. S1; Table S1), the simulations suggest that the Asn221 could H-bond with the C4-carbonyl at least momentarily in addition to interacting with the substrate’s C5-methoxy group (Figs 2C vs. D and S4).
(3) If the C1-group is reduced first (to produce neutral/half-protonated SQ), Lys251 side chain can rotate inward to form a lasting salt bridge with the Asp252COO− (Figs 3A,B and S3; Table S2) and participate in the substrate binding (Fig. 2A,B and S3; Table S2). The Lys251NH3+ at the Qi-site helps to orient the quinone ring to assure H-bonding between the C4-carbonyl and the epsilon protonated His217 side chain. Accordingly, the neutral Lys251 points out of the Qi-site similarly as would happen with stably binding Q (Figs 2C,D and 3C,D).
The Lys251-Asp252 salt bridge formation in the simulations.
(A) The Lys251NH3+-Asp252COO− salt bridge is formed on both A (blue line) and (B) B (red line) dimer sides in the conf1 (Table 1) simulation with neutral SQ bound at the Qi-site. (C) On the B side, Lys251 side chain assumes a clearly outward rotamer pose during the conf3 (Table 1) simulation with bound Q, although the side chain is initially forming a salt bridge with Asp252COO− after the equilibration time. (D) On the A side, the neutral Lys251 and Asp252 side chains are not within bonding distance from the very beginning of the conf4 (Table 1) simulation with bound Q at the Qi-site. The H-bonding distance of 3.4 Å is indicated with a black line. For clarity, the results are shown as 10-point moving averages. The trajectory data shown in the A–D panels correspond to the substrate binding mode snapshots shown in Fig. 2A–D, where the salt bridge forming residues are shown.
(4) The C4-carbonyl of neutral SQ (and Q; see Step 2) needs to form a continuous H-bond with His217 (Figs 2A,B and S3; Tables S2–S3). This interaction, which would presumably be needed for the second set of sequential e−/H+ transfers (Fig. 1A,B), could be re-enforced by a H-bond between the amine group of Asn221 side chain and the delta nitrogen of His217 side chain (Fig. 2A,C).
Lys251 functions as a switch-like proton shuttle between cardiolipin and Asp252
The Lys251NH3+ can rotate directly into the Qi-site, form a salt bridge with the Asp252COO− (Figs 3A,B and 4A and S3; Table S2) and participate in the half-protonated SQ binding either directly (Fig. 2B) or via a water bridge (Fig. 2A).
The switch-like operation of CL/K/D proton shuttle is pH-dependent.
(A) The Lys251NH3+ forms a salt bridge with the Asp252COO−, when SQ is bound at the Qi-site. The rotation is extensive, if compared to the structure with peripheral cardiolipin (CL)9 or the initial pose22. If the inward pose of Lys251NH3+ is compared to the Lys251NH2 pose, it shows that the neutral lysine turns outwards. See Fig. 1 for further details. (B) The empirical pKa calculations indicate that the Asp252 side chain would be neutral, when the Lys251 side chain is out of the Qi-site (pKa value from 1SQP; Table S7). The Lys251NH3+ keeps the outward pose, when the Asp252 side chain is neutral, i.e. the switching does not happen, when the Qi-site is acidic. (C) The CL/K/D switching is triggered by the deprotonation of Asp252 side chain. The Lys251NH3+ rotates inwards and forms a salt bridge with the Asp252COO−. The empricial pKa calculations indicate that Lys251 and Asp252 are charged when forming a salt bridge (pKa value from 2FYN, G chain; Table S7). After a direct H+ proton transfer (*) from the Lys251NH3+ to the Asp252COO−, the neutral lysine rotates out to accept another H+ from the CL (*). The Asp252COOH donates the newly acquired H+ to the solvent, if the pH rises at the Qi-site. This back-and-forth rotation of Lys251 would happen as long as the Qi-site was basic; ensuring also continuous protonation of His217.
In the inward pose, the positive charge of Lys251 side chain matches the opposite properties of the Asp252COO−. The lysine forms a water bridge with the substrate in an X-ray crystal structure (1PPJ in Figs 1D and S1)20, but, notably, the salt bridge is seen in only one substrate-free mutant structure (PDB: 2FYN; chain G)21. The missing structure factors make it difficult to evaluate the PDB entry regarding the bridge. The inward pose of Lys251 is in marked contrast to the rotamer pose visible in the yeast cyt bc1 complex (PDB: 3CX5; Figs 1C and 4A)9, where the lysine is positioned close to the dianionic cardiolipin (CL; Fig. 4A). The close CL-lysine arrangement is found in altogether 17 X-ray crystal structures (Table S5). If Lys251 and Asp252 are set neutral, the lysine side chain turns more outwards in the simulation than in any prior structures (Figs 4A and S4B; Table S6).
When considering the Lys251 rotation (Figs 4A and S3; Tables S2–S3) in more depth, it seems that it is not only relevant for the substrate binding (Fig. 2A,B; Table S2) but that it could be of mechanistic importance as well (Fig. 4). In the substrate-bound structures and with the nonprotonated substrate Q in the simulations (Fig. 2C,D), the lysine resides out of the Qi-site (Fig. S1; Table S1) and it would therefore not be needed at least for the initial binding (Figs 2C,D and 3C,D; Table S1). Although the CL/K pathway could very well rely on water-mediated H+ transfers (Fig. 1C,D), we propose an alternative, simpler and more efficient mechanism. The lysine rotation could facilitate H+ transfers from the CL positioned in the periphery of the protein surface into the buried active site (Fig. 4B,C). The transfers into the Qi-site would rely solely on changing the protonation states of Lys251 and Asp252 and the lysine rotamer (Figs 4 vs. C; and 2A,B vs. C,D).
After the first e−/H+ transfers (Fig. 1A,B), there is no apparent reason for either Lys251 or Asp252 to be neutral, if neutral SQ is formed at the Qi-site. If Lys251 donates the first H+ to the Asp252COO− in order to reduce Q, the neutral side chain should be able to rotate out and then return back in a fully protonated and positively charged state to reinforce the SQ binding (Figs 2A,B and 3A,B; Table S2). At any rate, the lysine side chain could reside at the site with the bound SQ long enough to ensure proper H-bonding between the C4-carbonyl and His217 (Fig. 2A,B) right after the first e−/H+ transfers (Fig. 1B). Then again, the salt bridge formation should not depend on the neutral SQ binding20. Thus, we propose that this salt bridge is formed and broken when a H+ is transferred into the substrate-free Qi-site or to promote stable SQ binding or even subsequent QH2 unbinding.
The back-and-forth repetition of the rotation would be consistent in situ, because the protonation of Lys251 and Asp252 would alternate between the two rotamers (Fig. 4B vs. C). A switch-like CL/K/D proton shuttle would function between the membrane periphery and the Qi-site without intermediary water molecules (Figs 1C,D vs. 4B,C). Although the Lys251 rotation probably happens at a subnanosecond time scale, even multiple successive water-mediated H+ transfers could be faster as a whole (0.93 ps/transfer)22. However, any potential timescale disadvantage of the switching (Fig. 4B–D) against the water transport model (Fig. 1C,D) would be overcome by the consistency and precise coordination of the Lys251 rotation inside the Qi-site (Fig. 4). The water-mediated H+ transfers are random or incoherent by nature, which reduces their overall efficiency.
Negative feedback: pH-dependent switching shuttles protons to the Qi-site
Proton transfers require short distance between the ionizable groups and comparable pKa values for the residues23. The first requirement is met by the Lys251-Asp252 salt bridge formation (Figs 2A,B and 3A,B)21. The pKa values, derived using empirical calculations, indicate, quite reasonably, that Lys251 and Asp252 are charged, when forming the bridge (Table S7). The second requirement of comparable pKa values is observed in a majority of the X-ray crystal structures, furthermore, the residue pair has alternative titration states suggesting coupling (Tables S5 and S7)24. The outward pointing Lys251 side chain is generally charged; but the close presence of the peripheral CL increases this tendency.
The empirical pKa calculations predict that the Asp252 side chain is neutral with the bound substrate (Table S5), but the charged state would be possible without the substrate (Table S7). Both the charged and neutral Lys251 ultimately assume rotamer poses pointing out of the active site, when nonprotonated substrate Q binding was relatively coordinated (Figs 2C,D and S4; Tables S4 and S6). Based on these observations, it seems likely that in the substrate-bound X-ray crystal structures, where the lysine is pointing outward (Fig. S1; Table S1), the Qi-sites are occupied by non-protonated Q (or anionic SQ) together with the Asp252COOH (Fig. S1; Table S1). Thus, the H+ transfer to the Q’s C1-carbonyl probably originates directly from the Asp252 (Fig. 2D) and without a bound substrate, the protons would be accepted by the solvent (Fig. 4B,C).
His217 is a logical proton source for the second reduction reaction with bound SQ (Fig. 1A,B) based on both the simulations (Fig. 2) and X-ray crystallographic data (Fig. S1; Table S1). The double protonation of His217 side chain (Table 1) would conveniently avoid the completely deprotonated state of imidazole ring during substrate reduction. However, this arrangement is not supported by the empirical pKa calculations (Tables S5 and S7) and, generally, it led to instability and/or the substrate unbinding in the simulations. The neutral SQ binding continued with the double protonated His217, only when the Asn221 side chain moved far away from its original position close to the bound substrate (conf2 at the B side in Fig. S3B; Table S3). This simulation result does not exclude the possibility that the double protonated His217 exist at least transiently in situ.
Overall, the empirical pKa calculations indicate that Lys251 could donate a H+ to the carboxylic acid of Asp252 side chain, which could then pass it along to the Q and via water to His217 (Fig. 4B,C). If the active site becomes basic enough, the Lys251NH3+ would rotate inward and donate a H+ to the Asp252COO−. This pH-dependent negative feedback is possible due to the unique position of the Qi-site at the protein-lipid interface (Fig. 1A).
The CL/K/D switching hypothesis in the context of mutagenic studies
Site-directed mutagenesis experiments can produce precise data on the importance of specific residues for even the subtlest ligand-receptor interactions25,26,27,28. The conserved cyt b residues His217, Lys251 and Asp252 (Fig. S2) have been studied in prior mutagenic studies18,19 and this data is reassessed here in the light of the simulation results.
With H217A and D252A mutants for the cyt bc1 complex of R. sphaeroides (Fig. S2), the e− transfer from the heme bH to Q induced by a flash of light was blocked and, overall, the photosynthetic growth halted18. The loss of activity with H217A mutant results from the inability of alanine to form a direct H-bond with the substrate’s C4-carbonyl, although the CL/K/D switching should remain unaffected (Fig. S5A). Similarly, the D252A mutation should effectively prevent H-bonding between the residue and the substrate’s C1-group. The CL/K/D switching (Fig. 4B,C) should also be deactivated in the D252A mutant (Fig. S5B), because there is no polar attraction forcing the Lys251 side chain to rotate into the Qi-site (Fig. S5B).
The D252N, K251M and K251I mutations did not affect the photosynthetic growth but slowed down the e− transfer from the heme bH to Q18,19. With D252N mutant, this minor effect on the complex operation suggests that asparagine is able to H-bond with the substrate’s C1-group and also assist in water-mediated proton transfers to Q (Fig. S5C). The lysine could still rotate inward and even participate in the neutral SQ binding (Fig. S5C), albeit there is no strong electrostatic incentive for this. In contrast, the side chains of the mutated residues in the K251M and K251I mutants are hydrophobic and, thus, likely remain outside the Qi-site (Fig. S5D), where they cannot participate in the SQ binding or promote H+ transfers. In the absence of the Lys251 rotation-based proton shuttle (Fig. 4), the H+ transport from CL molecule to the Asp252 side chain would likely happen via interconnected water molecules (Fig. S5D).
On the one hand, the mutagenic studies clearly indicate that the CL/K/D switching would not be an on/off process but a subtler mechanism. When the proton shuttle is disabled (e.g. K251M mutant), water seems to be able to compensate for the lost function (Fig. 1C,D). On the other hand, the slowdown of the e− transfer from the heme bH to Q with the Lys251 mutants, the involvement of Lys251 in the neutral SQ binding in the simulations (Fig. 2A,B) and titration coupling (Tables S5 and S7) and overall conservation of the KD residue pair (Fig. S2) indicate that the mechanism would be an integral part of the Qi-site operation. The importance of protonable groups at positions 251 and 252 is emphasized by the observation that the photosynthetic growth is not blocked even, if the residues are swapped to produce K251D/D252K double mutant19. In fact, our recent studies employing combinations of single and double mutants suggest that a protonable residue is needed either at the 251 or 252 position to assure activity29.
Although the proposed mechanism is not an on/off process, it fulfills several hallmarks of a molecular switch. Firstly, the lysine side chain reversibly shifts between two stable states in and out of the Qi-site (Fig. 4A). Secondly, based on the empirical pKa calculations, the shifting happens in response to the pH level change (Fig. 4B,C). Thirdly, presence of a ligand (half-protonated SQ in Fig. 2A,B) at the site promotes the inward pose of Lys251 (Fig. 2A,B). Fourthly, although both the CL/K/D switching (Fig. 4B,C) and water (Fig. 1C,D) can shuttle protons to Q, only the lysine rotation and pairing/unpairing with the aspartate would be unequivocally directional and coordinated.
From evolutionary perspective, the inherent randomness of water-mediated H+ transfers is the logical reason why the lysine rotation-based proton shuttle involving dianionic CL would have evolved in the first place. The CL/K/D switching would increase the efficiency of the energy metabolism by fine-tuning the Qi-site operation of the cyt bc1 complexes (Fig. S2) and perhaps even boost the process in aberrantly low H+ concentration. It is likely that similar pH-dependent (K/D, D/K, K/D/K, K/D/K/D etc.) pathways involving protonable residues/lipids exist elsewhere as well.
Proposed order of sequential quinone reduction at the Qi-site
Based on the simulations (Fig. 2), prior mutagenesis experiments18 and X-ray crystallographic data (Tables S1,S5 and S7), the sequential Q reduction is suggested to happen accordingly (Fig. 5):
The proposed sequential quinone reduction mechanism at the Qi-site of the cyt bc1 complex.
Those protons (H+) that are subject to transport (*) are shown in orange. For cardiolipin (CL) residing at the periphery is shown only the phosphate group. Note that the proton transfers between the peripheral CL and Lys251 do not necessarily involve water molecules. H-bonds are shown as magenta dotted lines. It is noteworthy that, if the two electrons are acquired separate from the H+ transfers (according to the concomittand proton transfer theory)17, it is unlikely that Lys251 rotation would play a role in the substrate binding (Fig. 5).
(1) Q binding is driven by the polar interactions between the quinone ring and the residues flanking the Qi-site.
(2) Upon Q binding, before e−/H+ transfers, the C1- and C4-carbonyl groups H-bond with Asp252 and Asn221 side chains, respectively. Note that the X-ray crystallographic data suggest that here His217 would eventually be H-bonding with the C4-carbonyl instead of the Asn221 side chain. At this point Lys251 would point out of the Qi-site (Fig. S1; Table S1).
(3) The first e− transfer from the heme bH to the substrate (producing anionic SQ) drives the H+ transfer to happen between the C1-carbonyl and the Asp252COOH. Alternatively, the H+ transfer could involve a water molecule residing between the aspartic acid and the C1-carbonyl (Fig. 2C). The Lys251NH3+ rotates into the Qi-site and forms a salt bridge with the Asp252COO−, which in turn stabilizes the positioning for the quinone ring of now neutral SQ by facilitating H-bonding and water bridge formation via the C1-hydroxyl and the C6-methoxy groups. At this point, the half-protonated SQ would be H-bonding stably with both the Asp252COO− and the epsilon (or double) protonated His217, in other words, the quinone ring positioning would be fully coordinated.
(4) The H+ transfer could be reversible; i.e. the proton could move back-and-forth between Asp252COO− and C1-carbonyl until the Lys251 side chain rotates inward and/or the second e−/H+ transfer ensues. In fact, only the deprotonated SQ has been detected at the Qi-site in previous experiments12,13,14,15. The stable binding of SQ could be required to wait for the second e−/H+ transfer and to prevent otherwise potentially rampant SQ-fuelled superoxide generation at the Qi-site. When the final e− comes from the heme bH, the C4-carbonyl of half-protonated SQ takes a H+ from the His217 side chain. As a result, a fully protonated substrate QH2 is formed at the Qi-site.
(5) Finally, the reaction product QH2 unbinds. H-bonding between the Asp252COO− and the C1-hydroxyl might postpone the H+ transfer from Lys251 to Asp252 for a while, but eventually the Lys251NH3+ could act as a “bouncer” that throws QH2 out of the Qi-site by donating a H+ to the Asp252COO−. The potential deprotonated state of His217 would also likely end quickly via water-mediated H+ transfers that also could promote the QH2 unbinding.
(6) At the outset, both His217 and Asp252 side chains are protonated and ready for Q binding. Lys251 would rotate in and out of the substrate-free Qi-site to upkeep this arrangement; transferring one H+ at a time from the peripheral CL molecule to the Asp252 side chain, which passes them to the solvent and via water also to His217 (Fig. 4B,C).
Conclusions
The simulations indicate that the binding of half-protonated semiquinone (SQ) would acquire more coordinated binding pose than the deprotonated quinone (Q) at the Qi-site, because the Lys251 side chain participates in the neutral SQ binding (Fig. 2A,B). Thus, the substrate binding and H-bonding coordination could benefit from acquiring the protons sequentially shortly after each electron transfer. This sequential mechanism suggesting that the e− transfers to be tightly coupled to H+ transfers should be tested in the future using for example quantum mechanics/molecular mechanics (QM/MM) calculations utilizing the new binding geometry seen in the simulations. The firmly coordinated binding of neutral SQ at the Qi-site, involving the inward rotamer of the Lys251 side chain, could be in part needed to curb the superoxide generation linked to aging-related cellular damage.
More importantly, in the fully bent rotamer pose (Fig. 4A), the positive lysine side chain forms a salt bridge with the negative Asp252 side chain (Figs 2A,B and 3A,B). The peripheral cardiolipin (CL) and the two ionizable residues are here suggested to function as a switch-like proton shuttle that transports protons from the membrane periphery directly into the Qi-site (Fig. 4B,C). Lys251 would acquire the protons from the CL molecule (Fig. 4A), but instead of relying on a string of interconnected water molecules (Fig. 1C,D)10, the lysine side chain rotation alone would shuttle the protons into the Qi-site (Fig. 4B,C). The Asp252 side chain would acquire the protons and pass them to the solvent. Upon Q binding the neutral Asp252 could H-bond with the C1-carbonyl and donate the H+ in response to the e− transfer (see Step 3 in Fig. 5). The proposed CL/K/D switching and involvement of Lys251 in proton transfers in general is supported by the observation that the K251M mutation slows down the e− transfer from the heme bH group to Q18,19. The switching could assure constant protonation of Asp252 and via water also of His217 (Fig. 4B,C) that are likely primary proton sources during Q reduction (Fig. 5). In addition, the switching would be pH-dependent based on empirical pKa calculations and thus the proton shuttle is activated only after the Qi-site becomes basic enough and protons are needed.
Methods
The simulation protocol, force field derivation and system build-up are presented in prior publications4,8,30,31. The dimer interface (PDB: 1ZRT)32 was filled with lipids that entered the cavity in a previous study (conf4 in ref. 8). The system compositions regarding the Qo- and Qi-site occupancies, redox centers states and the substrate/residue protonation are shown in Table 1.
The molecular dynamics (MD) simulations were performed with NAMD2.933 using the CHARMM 22 force-field and CMAP for the protein34 and the CHARMM 36 force field parameters for lipids35. The PME method was used for long-range electrostatics with a 12 Å cut-off36 for real vs. reciprocal space calculations. The time step was 1 fs. The target temperature and pressure were 310 K and 1 atm, respectively. Before running the 90 ns production simulations, specific angle and distance constraints were used during 10 ns equilibration simulations to keep the substrate’s C1- and C4-groups (Fig. 1B) within a H-bonding range with His217 and Asp252, respectively. The substrate binding was regarded as coordinated, if these two canonical interactions are formed (Fig. S1; Table S1).
The trajectory analysis and the 3D representations were prepared using BODIL37 and VMD1.938. The 2D representations were made with MARVINSKETCH15.8.10.0 (2015, ChemAxon; http://www.chemaxon.com). The pKa predictions were done at pH 7.4 with default settings for the PDB structures using PROPKA3.139,40 which uses an empirical approach to rapidly estimate the ionization state of protein groups. The predictive power of the software tool on the Qi-site residues was verified by analyzing selected snapshot structures with known protonation states (Table 1) extracted from the MD trajectories.
Additional Information
How to cite this article: Postila, P. A. et al. Atomistic determinants of co-enzyme Q reduction at the Qi-site of the cytochrome bc1 complex. Sci. Rep. 6, 33607; doi: 10.1038/srep33607 (2016).
References
Mitchell, P. Protonmotive redox mechanism of the cytochrome b-c″1 complex in the respiratory chain: Protonmotive ubiquinone cycle. FEBS Letters. 56, 1–6 (1975).
Brandt, U. & Trumpower, B. The protonmotive Q cycle in mitochondria and bacteria. Crit. Rev. Biochem. Mol. Biol. 29, 165–197 (1994).
Berry, E. A., Guergova-Kuras, M., Huang, L. S. & Crofts, A. R. Structure and function of cytochrome bc complexes. Annu. Rev. Biochem. 69, 1005–1075 (2000).
Postila, P. A. et al. Key role of water in proton transfer at the Qo-site of the cytochrome bc1 complex predicted by atomistic molecular dynamics simulations. Biochim. Biophys. Acta -Bioenergetics. 1827, 761–768 (2013).
Barragan, A. M., Crofts, A. R., Schulten, K. & Solov’yov, I. A. Identification of Ubiquinol Binding Motifs at the Q o -Site of the Cytochrome bc1 Complex. J. Phys. Chem. B. 119, 433–447 (2015).
Sharma, V., Enkavi, G., Vattulainen, I., Róg, T. & Wikström, M. Proton-coupled electron transfer and the role of water molecules in proton pumping by cytochrome c oxidase. Proc. Natl. Acad. Sci. USA 112, 2040–2045 (2015).
Wikström, M., Verkhovsky, M. I. & Hummer, G. Water-gated mechanism of proton translocation by cytochrome c oxidase. Biochim. Biophys. Acta. 1604, 61–65 (2003).
Pöyry, S. et al. Atomistic simulations indicate cardiolipin to have an integral role in the structure of the cytochrome bc1 complex. Biochim. Biophys. Acta - Bioenergetics 1827, 769–778 (2013).
Solmaz, S. & Hunte, C. Structure of complex III with bound cytochrome c in reduced state and definition of a minimal core interface for electron transfer. J. Biol. Chem. 283, 17542–17549 (2008).
Lange, C., Nett, J. H., Trumpower, B. L. & Hunte, C. Specific roles of protein-phospholipid interactions in the yeast cytochrome bc1 complex structure. EMBO J. 20, 6591–6600 (2001).
Sharma, V., Ala-Vannesluoma, P., Vattulainen, I., Wikström, M. & Róg, T. Role of subunit III and its lipids in the molecular mechanism of cytochrome c oxidase. Biochim. Biophys. Acta - Bioenergetics 1847, 690–697 (2015).
Robertson, D. E. et al. Thermodynamic properties of the semiquinone and its binding site in the ubiquinol-cytochrome c (c2) oxidoreductase of respiratory and photosynthetic systems. J. Biol. Chem. 259, 1758–1763 (1984).
Jünemann, S., Heathcote, P. & Rich, P. R. On the mechanism of quinol oxidation in the bc1 complex. J. Biol. Chem. 273, 21603–21607 (1998).
Cape, J. L., Bowman, M. K. & Kramer, D. M. A semiquinone intermediate generated at the Qo site of the cytochrome bc1 complex: importance for the Q-cycle and superoxide production. Proc. Natl. Acad. Sci. USA 104, 7887–7892 (2007).
Zhang, H., Osyczka, A., Dutton, P. L. & Moser, C. C. Exposing the complex III Qo semiquinone radical. Biochim. Biophys. Acta. 1767, 883–887 (2007).
Hong, S. et al.The Semiquinone at the Qi Site of the bc1 Complex Explored Using HYSCORE Spectroscopy and Specific Isotopic Labeling of Ubiquinone in Rhodobacter sphaeroides via 13C Methionine and Construction of a Methionine Auxotroph. Biochemistry 53, 6022−6031 (2014).
Kolling, D. R. J. et al.Exploration of Ligands to the Qi Site Semiquinone in the bc1 Complex Using High-resolution EPR. J. Biol. Chem. 278, 39747–39754 (2003).
Hacker, B., Barquera, B., Crofts, A. R. & Gennis, R. B. Characterization of mutations in the cytochrome b subunit of the bc1 complex of Rhodobacter sphaeroides that affect the quinone reductase site (Qc). Biochemistry. 32, 4403–4410 (1993).
Crofts, A. R. et al. Structure and function in the bc1-complex of Rh. sphaeroides. Photosynthesis: from light to biosphere (Kluwer Academic Publ., 1995).
Huang, L. S., Cobessi, D., Tung, E. Y. & Berry, E. A. Binding of the respiratory chain inhibitor antimycin to the mitochondrial bc1 complex: a new crystal structure reveals an altered intramolecular hydrogen-bonding pattern. J. Mol. Biol. 351, 573–597 (2005).
Esser, L. et al. Surface-modulated motion switch. Capture and release of iron-sulfur protein in the cytochrome bc1 complex. Proc. Natl. Acad. Sci. USA 103, 13045–13050 (2006).
Wraight, C. A. Chance and design–proton transfer in water, channels and bioenergetic proteins. Biochim. Biophys. Acta. 1757, 886–912 (2006).
Ishikita, H. & Saito, K. Proton transfer reactions and hydrogen-bond networks in protein environments. J. R. Soc. Interface. 11, 20130518 (2013).
Klingen, A. R., Palsdottir, H., Hunte, C. & Ullmann, G. M. Redox-linked protonation state changes in cytochrome bc1 identified by Poisson–Boltzmann electrostatics calculations. Biochim. Biophys. Acta – Bioenergetics 1767, 204–221 (2007).
Postila, P. A., Swanson, G. T. & Pentikäinen, O. T. Exploring kainate receptor pharmacology using molecular dynamics simulations. Neuropharmacology. 58, 515–527 (2010).
Frydenvang, K. et al. Full Domain closure of the ligand-binding core of the ionotropic glutamate receptor iGluR5 induced by the high affinity agonist dysiherbaine and the functional antagonist 8,9- dideoxyneodysiherbaine. J. Biol. Chem. 284, 14219–14229 (2009).
Lash, L. L. et al. Novel analogs and stereoisomers of the marine toxin neodysiherbaine with specificity for kainate receptors. J. Pharmacol. Exp. Therapeutics. 324, 484–496 (2008).
Lash-Van Wyhe, L. L. et al. T.Pharmacological activity of C10-substituted analogs of the highaffinity kainate receptor agonist dysiherbaine. Neuropharmacology 58, 640–649 (2010).
Kuleta, P., Sarewicz, M., Postila, P. A., Róg, T. & Osyczka, A. Identifying involvement of Lys251/Asp252 pair in electron transfer and associated proton transfer at the quinone reduction site of Rhodobacter capsulatus cytochrome bc1. Biochim. Biophys. Acta - Bioenergetics 10.1016/j.bbabio.2016.07.003 (2016).
Kaszuba, K., Róg, T., Bryl, K., Vattulainen, I. & Karttunen, M. Molecular Dynamics Simulations Reveal Fundamental Role of Water As Factor Determining Affinity of Binding of β-Blocker Nebivolol to β2-Adrenergic Receptor. J. Phys. Chem. B. 114, 8374–8386 (2010).
Kaszuba, K. et al. Parameterization of the prosthetic redox centers of the bacterial cytochrome bc1 complex for atomistic molecular dynamics simulations. Theor. Chem. Accounts. 132 (2013).
Berry, E. A. et al. X-Ray Structure of Rhodobacter Capsulatus Cytochrome bc (1): Comparison with its Mitochondrial and Chloroplast Counterparts. Photosyn. Res. 81, 251–275 (2004).
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005).
Mackerell, A. D., Feig, M. & Brooks, C. L. Extending the treatment of backbone energetics in protein force fields: limitations of gas-phase quantum mechanics in reproducing protein conformational distributions in molecular dynamics simulations. J. Comput. Chem. 25, 1400–1415 (2004).
Taylor, J., Whiteford, N. E., Bradley, G. & Watson, G. W. Validation of all-atom phosphatidylcholine lipid force fields in the tensionless NPT ensemble, Biochim. Biophys. Acta 1788, 638–649 (2009).
Essmann, U. et al. A smooth particle mesh Ewald method. J. Chem. Phys. 103, 8577 (1995).
Lehtonen, J. V. et al. BODIL: a molecular modeling environment for structure-function analysis and drug design. J. Comput. Aided Mol. Des. 18, 401–419 (2004).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33-8, 27-8 (1996).
Søndergaard, C. R., Olsson, M. H. M., Rostkowski, M. & Jensen, J. H. Improved Treatment of Ligands and Coupling Effects in Empirical Calculation and Rationalization of pKa Values. J. Chem. Theory Comput. 7, 2284–2295 (2011).
Olsson, M. H. M., Søndergaard, C. R., Rostkowski, M. & Jensen, J. H. PROPKA3. Consistent Treatment of Internal and Surface Residues in Empirical p Ka Predictions. J. Chem. Theory Comput. 7, 525–537 (2011).
Acknowledgements
We wish to thank CSC – IT Centre for Science (Espoo, Finland) for computational resources. For financial support, we wish to thank the Academy of Finland (TR, IV and PAP; Center of Excellence in Biomembrane Research (IV, TR)), the Finnish Doctoral Programme in Computational Sciences (KK), the Sigrid Juselius Foundation (IV), the Paulo Foundation (PAP) and the European Research Council (IV, TR; Advanced Grant project CROWDED-PRO-LIPIDS). AO acknowledges The Wellcome Trust International Senior Research Fellowship.
Author information
Authors and Affiliations
Contributions
K.K. prepared the simulated systems. P.A.P. performed the simulations and data analysis. P.A.P., A.O., M.S., P.K., I.V. and T.R. participated in the design of the project, the interpretation of the data and the writing of the article. T.R. supervised the study.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Postila, P., Kaszuba, K., Kuleta, P. et al. Atomistic determinants of co-enzyme Q reduction at the Qi-site of the cytochrome bc1 complex. Sci Rep 6, 33607 (2016). https://doi.org/10.1038/srep33607
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep33607
This article is cited by
-
The evolution of the human mitochondrial bc1 complex- adaptation for reduced rate of superoxide production?
Journal of Bioenergetics and Biomembranes (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.