Abstract
EGFP oligomers are convenient standards for experiments on fluorescent protein-tagged biomolecules. In this study, we characterized their hydrodynamic and fluorescence properties. Diffusion coefficients D of EGFP1–4 were determined by analytical ultracentrifugation with fluorescence detection and by fluorescence correlation spectroscopy (FCS), yielding 83.4…48.2 μm2/s and 97.3…54.8 μm2/s from monomer to tetramer. A “barrels standing in a row” model agreed best with the sedimentation data. Oligomerization red-shifted EGFP emission spectra without any shift in absorption. Fluorescence anisotropy decreased, indicating homoFRET between the subunits. Fluorescence lifetime decreased only slightly (4%) indicating insignificant quenching by FRET to subunits in non-emitting states. FCS-measured D, particle number and molecular brightness depended on dark states and light-induced processes in distinct subunits, resulting in a dependence on illumination power different for monomers and oligomers. Since subunits may be in “on” (bright) or “off” (dark) states, FCS-determined apparent brightness is not proportional to that of the monomer. From its dependence on the number of subunits, the probability of the “on” state for a subunit was determined to be 96% at pH 8 and 77% at pH 6.38, i.e., protonation increases the dark state. These fluorescence properties of EGFP oligomeric standards can assist interpreting results from oligomerized EGFP fusion proteins of biological interest.
Similar content being viewed by others
Introduction
Protein complexes in live cells are often investigated by observing fluorescent protein (FP) tags. Intracellular concentration, aggregation state and mobility of the labeled protein of interest can be quantified by fluorescence microscopic tools: fluorescence correlation spectroscopy (FCS), number and brightness analysis (N&B), Förster resonance energy transfer (FRET), fluorescence recovery after photobleaching (FRAP), etc.1,2,3,4,5,6. Proteins can be present in various oligomerization states, which may differ in their mobility and brightness. The measured values of these parameters are influenced by the photophysical behavior of the FP, which depends on the measurement conditions (excitation light intensity, duration of illumination, pH, etc.). Different photophysical processes of individual EGFP and other fluorescent proteins have already been described (photobleaching, triplet state formation, protonation- or light-induced “blinking”)7,8,9,10,11,12,13. There are still open questions how these processes contort the measured parameters (apparent brightness, apparent diffusion coefficient) of protein complexes containing more than one fluorescent protein. Earlier Pack and coworkers14 and Dross in our lab5 reported FCS-derived diffusion coefficients of EGFP oligomers in live cells and crude cell lysates. Here we characterize the fluorescence and hydrodynamic properties of recombinant covalently linked EGFP oligomers composed of 1 to 4 EGFP subunits in solution in order to validate them as natural standards for higher protein complexes. We test the effect of illumination power on the fluorescence parameters of the EGFP oligomers at two pH values and interpret how they relate to the behavior of the constituting monomeric subunits. We compare FCS-derived diffusion coefficients of EGFP oligomers with values yielded by analytical ultracentrifugation. Finally we propose a simple procedure to quantify the probability of long-lived dark states of EGFP by analyzing the apparent brightness of the oligomers as a function of the number of subunits.
Results
Characterization of EGFP oligomers by analytical ultracentrifugation
To use EGFP oligomers of 1 to 4 covalently linked subunits as calibration standards, we determined their molar masses and hydrodynamic parameters in solution by analytical ultracentrifugation with fluorescence detection system (AU-FDS). Sedimentation equilibrium runs were recorded at several speeds and protein concentrations. Molar masses were determined by globally fitting the radial distribution of the fluorescence signals at all rotor speeds for each oligomer with a single species model (Supplementary Fig. S1) using the software BPCFIT15. For EGFP1–3 the molar masses, Meq, (Table 1) agreed within 5% with the sequence-based calculation Mcalc of the EGFP monomer (26.9 kg/mol) and its multiples. Our Meq value for the EGFP monomer agrees well with that reported earlier using a similar setup16,17. For EGFP4 the Meq value was 10% smaller than Mcalc. This deviation and the systematic divergence of the fit residuals observed for EGFP4 (Supplementary Fig. S1) indicate an additional pool of smaller particles. Indeed, some minor faster, fluorescent species were also present in the lanes of EGFP4 and also EGFP1 in the native polyacrylamide gel (Supplementary Fig. S2B), which may be degradation products or, in the case of the tetramer, a lower oligomer. The amount of neither impurity exceeds 5% in these preparations.
The homogeneity of the EGFP oligomers in size and shape was then characterized by sedimentation velocity runs at 50,000 rpm using four different protein concentrations between 50 and 400 nM. Figure 1 shows the sedimentation velocity analysis at 400 nM EGFP subunits. The normalized distributions of the sedimentation coefficients for EGFP1–3 could be fitted with a single Gaussian peak, while EGFP4 had a second minor component (5%) in the lower s value range as seen in the equilibrium runs and in the gel. The single peaks suggest that each oligomer has a unique arrangement of the subunits, which may be due to the short length of the linkers limiting the possible conformations. svel values of the main peaks are listed in Table 1. Diffusion coefficients, Dvel, were assessed from peak broadening18,19 (Table 1). Molar masses, Mvel, were determined from svel and Dvel values (Table 1). The excellent agreement between the Mvel, Meq and Mcalc values for EGFP1–3 indicates that these samples were highly homogeneous.
Sedimentation velocity centrifugation of the EGFP oligomers with fluorescence detection.
Normalized representative g(s*) plots are displayed. Sedimentation coefficients were converted to 20 °C and water as solvent. Distributions were fitted with a single Gaussian peak for EGFP1–3 and with two Gaussians for EGFP4; the second minor component (5%) present in the EGFP4 sample is shown at a lower s* value. The peaks yielded svel, and peak broadening Dvel values.
The possible shape of the oligomers may also be inferred from the sedimentation coefficients. The monomeric subunits can assume various spatial arrangements in the oligomers. We calculated a sedimentation coefficient scalc = 2.6 S for monomeric EGFP based on the crystal structure with the HYDROPRO software, which agreed well with the measured s value of 2.52 S. For the oligomers, the sedimentation coefficients of some possible spatial arrangements (Fig. 2) were also estimated by this software. Here, a constraint was taken into account: both the C and N-termini of a barrel-shaped monomer are on the top of the barrel20, and two neighboring barrels are connected by 5-amino-acid linkers. In the case of a side-by-side linear arrangement the deviation of scalc from the measured svel values was between 2–5% for all the oligomers (Table 1). A star- or square-shaped conformation of the tetramer yielded larger deviations between the model-based calculated s value and and the measured s values (9 and 11%, respectively). Thus, the oligomers are probably arranged in an elongated rather than a compact shape. Pack et al.14. also suggested an elongated rather than spherical shape for EGFP2–5 oligomers connected with 25 aa long linkers, much longer than our 5 aa linkers.
Effect of oligomerization on the fluorescence properties of EGFP
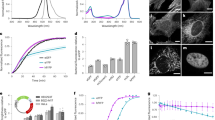
We tested the influence of EGFP oligomerization on the absorption and fluorescence emission spectra, anisotropy and lifetime. Normalized absorption spectra overlapped perfectly, but emission spectra were red-shifted by ~2 nm from monomer to tetramer (Supplementary Fig. S3). This slight bathochromic shift may be due to direct long-range interaction between EGFP fluorophores. Fluorescence lifetime decay curves could be well fitted to single exponentials. Lifetimes decreased monotonously from the monomer to the tetramer by 4% (Table 2).
This suggests that no or only little energy transfer takes place from fluorescent to non-emitting dark subunits in an oligomer. The steady-state fluorescence anisotropy decreased progressively from 0.32 for the monomer to 0.23 for the tetramer, which provides evidence for homo-FRET between the subunits21,22.
As shown, the bulk fluorescence parameters: the emission peak wavelength, lifetime and anisotropy changed progressively with the size of the oligomer. Next, we investigated the fluorescence properties of individual oligomers by using fluorescence correlation spectroscopy.
Fluorescence correlation spectroscopy of EGFP oligomers
The state of aggregation of a fluorescently tagged protein can be assessed from the diffusion coefficient and the molecular brightness. However, the measured values of these parameters are influenced by the photophysical behavior of the fluorophore, which in turn depends on the experimental conditions (excitation intensity, pH, etc.). In our fluorescence correlation spectroscopy (FCS) experiments, we varied the illumination power of the laser and the pH of the buffer solution, and recorded fluorescence intensity and autocorrelation functions (ACFs). ACFs were fitted to obtain the apparent diffusion coefficients, concentrations and molecular brightness values for each oligomer. With this model system we could show how to estimate the “real” parameters from the measured apparent values.
Fluorescence intensity
First we analyzed the FCS-derived average fluorescence intensities, F, of the oligomers as a function of the illumination power, P (Fig. 3). Measurements were done in Tris buffer, pH 8 at 1, 4 and 10% laser power corresponding to 5, 20 and 50 μW at the microscope objective. As a comparison, the behavior of the Alexa Fluor 488 dye (A488) is also shown. Fluorescence intensities (normalized at 5 μW) increased with increasing laser power (Fig. 3a), but not linearly and not uniformly for the different molecules. The highest rate of increase was observed for A488, followed by EGFP1 and got less as the size of the oligomer increased. To visualize the deviation from linearity, F was divided by the illumination power and normalized to 1 at 5 μW (Fig. 3b). The decreasing tendency of the graphs means that at higher illumination power fewer photons were emitted per incoming photons. The decrease became greater with increasing oligomer size. An obvious reason for this effect is a light-induced transition to a dark state, i.e., photobleaching or blinking, which becomes more prominent for larger molecules moving more slowly and residing longer in the focal volume. This means that a simple intensity measurement with strong illumination may lead to an underestimation of the concentration of fluorescent molecules to an extent depending on the mobility and the laser power.
Dependence of the relative fluorescence intensities of EGFP1–4 and Alexa 488 on the illumination power.
(a) Raw fluorescence intesities normalized to 1 at the lowest illumination power. (b) Fluorescence intensities divided by the illumination power and normalized as above. Fits are single exponentials. Measurements were done in 20 mM Tris-HCl, 100 mM NaCl, pH 8.
ACF amplitude
A principally different method to determine the concentrations from the same data is the analysis of autocorrelation functions, which in addition yields the diffusion coefficients. As an example, Fig. 4a shows the ACF curves of EGFP1 and EGFP4 at different illumination powers (curves for Alexa 488, EGFP2 and EGFP3 are not shown). ACFs were fitted to a model assuming triplet state formation and a single free diffusion component. The fast decay at the beginning of the curve (in the μs range) is related to triplet state formation. The fit yields N, the average number of molecules in the focal volume (the reciprocal of the amplitude) and the diffusion time (τD, the mean dwell time) or the related diffusion coefficient, D (see Materials and Methods). With increasing illumination power the amplitudes of ACF curves decreased for EGFP2–4 and for A488, but remained constant (or decreased slightly) for the EGFP1. Accordingly, the apparent normalized particle numbers increased: Fig. 4b shows an approximately linear relationship of Nnorm with the laser power. The slope increased with size of the oligomers. With increasing illumination power, excitation at the center of the illuminated area may be saturated because EGFP cannot absorb another photon during its nanosecond fluorescence lifetime. This flattens the detection efficiency profile and widens the observation volume, i.e., photons from an area of larger radius will contribute to the signal23. This may lead to an increase of the apparent particle number. On the other hand, the apparent particle number may also decrease due to photobleaching with increasing illumination power. These two effects were separately simulated for EGFP1–4 by using Monte-Carlo method, and resulted in the expected dependence of the apparent particle number on the illumination power (Supplementary Fig. S4a for saturation and Supplementary Fig. S4b for photobleaching). In these simulations the different oligomers behaved similarly, contrary to the divergence seen in the measurements (Fig. 4b). This discrepancy suggests that further photophysical processes could play a role. In our measurements saturation seemed to dominate over bleaching for EGFP2–4, whereas for EGFP1 these effects canceled each other. The A488 dye is very photostable, its bleaching was not significant at these laser powers; therefore, saturation dominated and the fitted N value increased with the laser power. These effects must be taken into account for exact concentration measurements by FCS to avoid significant errors (here, ~50% error in N would occur at a 10 fold variation of the laser power).
Analysis of autocorrelation curves and their parameters as a function of oligomer size and illumination power.
(a) Representative autocorrelation functions for EGFP1 and EGFP4 at different illumination powers at pH 8. (b) Normalized particle numbers obtained from fits of the ACFs of EGFP1–4. Alexa 488 is shown as a comparison. (c) Normalized representative ACF curves from part a). (d) Diffusion times obtained from fitting ACFs of EGFP1–4 and Alexa 488. (e) Apparent diffusion coefficients of EGFP1–4 calculated from the diffusion times according to equation (9). Intercepts of linear fits define the zero light extrapolated DFCS0 values. (f) Comparison of diffusion coefficients obtained by different methods. Dcalc was calculated from scalc with the Svedberg equation, assuming side-by-side linear arrangement of barrels (see Fig. 2). Dvel was obtained from peak broadening of sedimentation velocity runs (see Fig. 1). DFCS0 values were extrapolated from FCS-derived D values to zero light conditions. Experimental D values vs. n were fitted to power functions yielding similar powers. D dependence for a sphere is given as comparison.
Diffusion time and diffusion coefficient
The other principal parameter of the ACF is the correlation time τD. To analyze its behavior at different illumination powers, we normalized the curves at the plateau amplitudes to 1, and presented the time interval of interest for EGFP1 and EGFP4 as examples in Fig. 4c (EGFP2 and EGFP3 behaved similarly and the curves lied between those of the EGFP1 and EGFP4). EGFP1–4 curves were all shifted to the left, decaying at shorter times, contrary to Alexa 488 curves, which were shifted to the right with increasing illumination power24. Apparent diffusion times (τD) are shown in Fig. 4d. The apparent increase of τD for A488 probably stems from the same saturation effect as the apparent increase of its N: the observation volume for this dye is increased and therefore the dwell time is longer. For EGFP1–4, on the contrary, we observed an apparent decrease of τD, to a similar extent for each oligomer. Simulation of the dependence of τD on the illumination power (Supplementary Fig. S4c,d) demonstrated that saturation and bleaching influenced τD in opposing ways. Contrary to the particle number, the effect of bleaching on the diffusion time seemed to dominate over the effect of saturation for all EGFP oligomers.
From τD we calculated the diffusion coefficient D according to equation (9). D values increased strongly with the illumination power for each oligomer (Fig. 4e). At the highest illumination power (50 μW) D would be overestimated by ~90%. The apparent change of D with illumination power results from the disparate photophysical behavior of Alexa 488 and EGFP as discussed above. To empirically eliminate the effect of photophysical processes, we linearly extrapolated the D vs. P graph to zero illumination power to get DFCS0. DFCS0 values corrected for temperature and viscosity differences were compared with those obtained from ultracentrifugation (Dvel) or model prediction (Dcalc) (Table 1, Fig. 4f). As expected, DFCS0 decreased as the oligomer length increased. DFCS0 or Dvel vs. n (number of subunits) was fitted to a power function, yielding exponents of ca. −0.4 in both cases, which is larger than would be expected for spheres (−0.333), in agreement with our assumption of an elongated shape. DFCS0 values were ~3.6–13.5% larger than Dcalc values, and by ~10–16% larger than Dvel. Possible reasons for the discrepancies include uncertainties in the assumptions about the molecular shape for Dcalc, and in the fitted FCS model function.
Molecular brightness
The molecular brightness, F/N, characterizes the average fluorescence intensity detected per particle, where F is measured directly and N is derived from the fit to the ACF. F/N can be an indicator of the association state of fluorophores or labeled molecules, but its value may be prone to photophysical artefacts. Figure 5a presents the molecular brightness as a function of illumination power for the EGFP oligomers. The increase of F/N was not linear especially for the higher oligomers. Correspondingly, the F/N/P brightness per unit illumination power decreased as a function of P (Fig. 5b). According to our simulations both saturation and bleaching lead to a decrease of brightness (Supplementary Fig. 4e,f). In the simulation of bleaching, the normalized brightness decreased more strongly with increasing oligomer size due to the longer dwell times. On the other hand, the effect of excitation saturation is size independent as it only depends on the fluorescence lifetime relative to the time lag between two excitation photons reaching a fluorophore. In our measurements we observed a very strong decrease of F/N/P at higher P values, which was larger than 3 fold for the tetramer, and less than two-fold for the monomer. This length dependence was also present in our simulations. To estimate the value of F/N/P free of light-induced artifacts, we made an exponential extrapolation of the graph to P = 0, which we considered as the most artifact-free brightness values, and used them in subsequent analyses.
Dependence of FCS-derived molecular brightness on the oligomer size and the illumination power.
(a) Raw molecular brightness data of EGFP1–4. (b) Molecular brightness normalized by the illumination power as a function of the illumination power. Data were fitted to exponential decays. (c,d) Molecular brightness normalized by the illumination power as a function of the number of subunits at pH 8 and 6.38. The data for zero illumination power were obtained as the amplitudes of the fits in F/N/P vs. P. From the slope and intercept of linear fits the long-lived dark fractions of EGFP were derived according to equation (5).
Assessment of long-lived dark fraction from the apparent brightness of oligomers
In addition to light-induced processes (triplet, bleaching, light-induced blinking), fluorescent proteins can also be found in dark states due to other reasons. If a monomeric EGFP is in a dark state lasting longer than the diffusion time, it is not seen at all, leading to underestimation of the number of fluorophores or fluorescently tagged proteins. On the other hand, if one or several EGFP subunits are dark in an oligomer, but there is at least one in a bright state, the particle will still be observed, although the apparent number of particles, i.e., the 1/G(0) reciprocal amplitude of the ACF, will decrease (see Supplementary Information, equations S16–17). We calculated the effect of long-lived dark states on the measurable brightness of oligomers as shown below.
In an oligomer of fluorescent proteins (FPs), each subunit can be in a bright “on” or a long-lived dark “off” state. For the calculation of the apparent brightness we need the following parameters:
p: probability of an FP molecule to be in the “on” state (involving the fluorescent state, the triplet state and other short-lived dark states having an off-time shorter than τdiff).
1-p: probability of an FP molecule to be in a non-fluorescent “off” state lasting longer than the diffusion time (τoff>>τdiff).
N: mean number of particles in the detection volume.
F: mean total fluorescence.
Ψ: molecular brightness of a monomeric FP.
n: number of subunits in the FP oligomer.
k: number of FPs in the “on” state in a given oligomer.
In an FP oligomer composed of n subunits, k = 0, 1, …, n subunits can be “on”. For a mixture of different species the mean fluorescence intensity is

where Nk is the number of molecules with k subunits in the “on” state and kΨ is the molecular brightness of this species. The amplitude of the autocorrelation function is

where γ is a factor depending on the geometry of the detection volume. For simplicity we set γ = 1. Napp is the reciprocal of the G(0) amplitude, which for samples with heterogeneous brightness is an underestimation of the total number of molecules N. The denominator of equation (2) is the square of the mean fluorescence intensity. The apparent molecular brightness is

where pk = Nk/N is the fraction of molecules with k subunits in the “on” state. If the “on” or “off” state of a subunit is independent of the state of the others, we can assume that k follows a binomial distribution. It can then be derived (see Supplementary Information, Chapter 2) that the apparent molecular brightness is

F/Napp is a linear function of n with a nonzero intercept if p is not unity, i.e., if there are FPs in the long-lived dark state. From the parameters of the linear fit to the F/Napp vs. n graph, we get p as

To our knowledge, this is the first time that the fraction of long-lived dark states of freely diffusing FPs are determined. The above combinatorial approach is made possible by the application of oligomers composed of different numbers of subunits.
This model was applied to fit our measurements on EGFP oligomers to assess the amount of long-lived dark species under different conditions. We were interested in the value of p in the absence of light-induced processes, thus we used the F/N/P values extrapolated for P = 0 μW as a measure of artefact-free brightness. In Fig. 5c F/N/P is shown as a function of n for different illumination powers. Only the results for EGFP1–3 are presented because the EGFP4 sample contained a few percent of lower oligomers (Fig. S2b), which could corrupt the linear fit. The graphs were linear for all illumination powers. The slopes decreased while the intercepts increased with increasing illumination power. The apparent probability p of the dark state was obtained for each illumination power the way described above. In the absence of excitation light, almost all EGFP subunits were in the “on” state, p = 0.96, i.e., 4% of the subunits were in long-lived dark states. In the presence of light the value of p decreased dramatically with the illumination power (Table 3). In this case initially bright subunits could transit to light-induced dark states (e.g. due to bleaching) during their dwell time. Thus, the assumption of the model that a given subunit is either on or off throughout its dwell time, is not fulfilled. This way only the p values derived from data extrapolated to zero illumination are valid indicators of the “on” state fractions.
Protonation induced dark state
Previous studies used EGFP monomers to investigate protonation dependent dark state formation at different pH values8,13. We used our approach based on different oligomers and extrapolation to zero light to assess the effect of a lower pH on the apparent “on” fraction. As an example, we measured the autocorrelation functions of EGFP1–4 at pH 6.38 with different illumination powers. Curves were fitted with equation (8) without taking the protonation process into account because a second non-fluorescent component had an insignificant fraction and did not improve the fit. The apparent probability p of the “on” state was calculated as detailed above for pH 8. The value of p was 0.77 for zero light extrapolation. This corresponds to a dark state fraction of 23%, which includes the long-lived dark states and the additional protonation-induced dark states. Haupts et al.8 also reported ~3–4% and ~20–25% protonated EGFP fraction from excitation spectra at pH 8 and pH 6.4. Widengren at al13 measured ~35% protonation induced dark state fraction by FCS at pH 6 at buffer conditions similar to ours. The characteristic time constants of protonation-deprotonation processes are in the hundred μs to ms range8,13, which are close to the dwell times (100–200 μs). Hence, transition between the protonated and deprotonated states could take place during the dwell time (such as for the case of bleaching) thereby affecting the ACF amplitude and the apparent brightness. Thus, our model assumptions are not fulfilled, and the derived p values are not exact. Apparent values of p decreased with increasing illumination power probably due to bleaching and saturation such as for pH 8. To determine the fraction of long-lived dark states, measurements at several low illumination powers and extrapolation to zero light are recommended.
Discussion
Oligomers of fluorescent proteins were first introduced to increase the brightness of tagged proteins in live cells25,26,27. Our group mapped the intracellular mobility of EGFP1–4 oligomers by single-point FCS5 and to study accessibility of nuclear regions. Different fluorescence methods (FCS, Number and Brightness Analysis or FRET) allowed the identification of oligomerization of various FP-tagged proteins (e.g. in refs 3,6,28,29). We propose that investigation of the fluorescence properties of pure EGFP oligomeric standards would help to interpret results obtained with oligomerized EGFP fusion proteins of biological interest.
The hydrodynamic and photophysical properties of EGFP have been studied by several labs. The corrected sedimentation coefficient of the EGFP monomer derived from AU-FDS was 2.68 S in ref. 30, and 2.52 S for our measurement, whereas the calculated value was 2.6 S. Macgregor et al.16 compared results from analytical ultracentrifugation, AUC, (2.81 S) and AU-FDS (2.73 S) on His tagged EGFP. Petrasek et al.31 used scanning FCS and received 92.3 μm2/s (corrected for 20°), and Widengren et al. measured 85 ± 4 μm2/s by FCS at room temperature12. FRAP yielded 87 ± 16 μm2/s (at 20 °C)32. In this study, we applied AU-FDS, a frequently used method for diffusion coefficient determination, which resulted in D(w,20 °C) = 83.4 ± 0.5 μm2/s, and FCS, which yielded D(w,20 °C) = 97.3 ± 5 μm2/s after extrapolation to zero excitation light. It seems that FCS-determined D values are systematically higher than those obtained from AUC. Fluorescence of EGFP monomers is affected by several light and protonation induced processes. For triplet correlation times we obtained values in the microsecond range, which decreased with increasing illumination power. The increase of illumination power resulted in the increase of the apparent diffusion coefficient as a net effect of photobleaching (increasing the apparent D) and excitation saturation (reducing the apparent D)11,12. The dominance between these two processes is dye dependent: for fluorescein, Rh6G33, similarly to EGFP, bleaching seems to dominate, contrary to the more photostable Alexa 488 (Fig. 4d). Due to this disparate behavior, measuring the Alexa 488 standard at the same laser power does not compensate the error of D due to bleaching; on the contrary, it strengthens this effect.
The hydrodynamic and fluorescence properties of purified EGFP oligomers have been systematically investigated in this study. The obtained sedimentation coefficients allowed us to identify the most probable, “barrels standing in a row”, arrangement of the subunits. Whereas the decrease of diffusion coefficients with EGFP oligomerization degree was similar in AU-FDS and FCS detection, FCS-derived D values were systematically higher for each oligomer. FCS-derived D values taken from earlier studies5,14 are within 5–20% of ours after correction for differences in temperature, and show a similar dependency on oligomer length.
The dependence of fluorescence properties of the oligomers on the illumination power can only partially be explained by the properties of the individual subunits. The measured apparent N values (representing the concentration) did not vary significantly for the monomers, and increased with increasing P for oligomers, to an extent proportional to the oligomer size. Excitation saturation and bleaching of the subunits was not sufficient to explain this size dependent behavior. The normalized molecular brightness (F/N/P) decreased more strongly for larger oligomers, in accordance with our simulations. Apparent D values increased substantially for monomers with illumination power, and less strongly for oligomers (in agreement with simulations) because if not all subunits are bleached, the oligomer is still detectable.
By now we considered only light-induced dark states of the subunits. Fluorescent proteins can also be in dark states due to other reasons such as imperfect dye maturation or protonation. Since the individual subunits in an oligomer may be in “on” (bright) or “off” (dark) states, the FCS-determined apparent brightness (F/N) is not an integer multiple of that of the monomer. From the length dependence of F/N we determined the p “on” state probability of the subunits. The apparent value of p decreased with illumination power. To eliminate the effect of light induced dark states, we extrapolated to zero excitation light, and found that only 4% of the subunits were in long-lived dark states at pH 8. Madl et al. reported 12% dark state (not corrected for the effect of illumination) for GFP tagging CRAC channels expressed in the cell membrane28.
pH-dependence of excitation spectra was used earlier to calculate the protonated nonfluorescent fraction of EGFP monomers, e.g. in ref. 8. Widengren et al.12 introduced a model function to account for protonation induced blinking in FCS. Our results on the long lasting dark fractions are in accurate agreement with their data. Since our method detects long lasting dark states, the subunits in the purified recombinant oligomers probably do not reside in dark states other than the protonated state. However, in live cells the expression of FP-labeled proteins is continuous, and immature, nonfluorescent species are also present and participate in protein complexes, which could not be detected by using the earlier methods. Our proposed procedure can assess the fraction of “on” and “off” states (by expressing FP oligomers in cells under similar conditions as the FP-tagged proteins of interest), and make the quantitation of oligomerization more accurate.
Materials and Methods
Preparation and purification of EGFP1–4 oligomers
EGFP1–4 are multimeric fluorescent proteins, with 5-amino-acid (G-P-V-A-T) linkers connecting the EGFP units. The plasmids encoding for EGFP1–4 were a kind gift of Dr. M. M. Nalaskowski (Universitätsklinikum Hamburg-Eppendorf)25,26. For stable transfection, multimeric EGFP inserts, EGFPn5 were cloned into a pSV vector originating from promoterless pECFP-1 (Clontech). SV40 promoter sequence was inserted at the Hind III restriction site to drive overexpression. HeLa cells were cultivated in DMEM without phenol red, containing 10% fetal calf serum and 200 mM L-glutamine in humidified 5% CO2 atmosphere at 37 °C. The cells were transfected with the pSV-EGFPn vector according to the Fugene HD user manual (Roche, Germany). Stably transfected cells were treated with G418 (Geneticin, Invitrogen, Life Technology, CA, USA) at a final concentration of 50 μg/ml and clones with strong EGFP expression were selected. For protein purification cells were seeded in petri dishes 72 h before harvesting. Cells were collected, homogenized (“dounced”), treated with 0.2% Nonidet P40 (Shell Chemicals) and centrifuged at 1200 × g for 10 min. The white pellet was removed and discarded, the green supernatant was incubated with DNase (5 μg/ml) and RNase (30 μg/ml) for 15 min. EGFP1–4 proteins were purified on 1 ml DEAE Sepharose Fast Flow columns (GE Healthcare). The column was washed with 10 ml 20 mM Tris pH 8, 50 mM NaCl. Proteins were eluted in 1 ml 20 mM Tris pH 8, 150 mM NaCl. In the SDS gel the oligomers appear as unique bands according to their expected masses (Fig. S2A). Homogeneity of the oligomers was checked by fluorescence detection in native polyacrylamide gel electrophoresis (Fig. S2B). Proteins were stored in this buffer with protease inhibitor (complete mini, EDTA free, Roche) and 0.01% Nonidet P40 at −80 °C. In all experiments 20 mM Tris-HCl, 100 mM NaCl, pH 8 buffer was used, unless specified otherwise.
Absorption and fluorescence characteristics
Absorption spectra of the purified EGFP1–4 samples were measured on Varian Cary-4E spectrophotometers. The fluorophore concentration was calculated using an extinction coefficient of 56,000 M−1cm−1 at 489 nm supposing additivity34.
Fluorescence spectra were recorded using an SLM 810 spectrofluorimeter equipped with a 450 W Xenon lamp. Samples were excited at 470 nm and spectra recorded at a bandwidth of 4 nm in the 480–650 nm wavelength range. The presented spectra were corrected for the instrument characteristics. Anisotropy was measured in the same setup using Glan-Thompson prisms to polarize and analyze the light. Fluorescence lifetime measurements were performed on a FluoTime 100 (Picoquant, Germany) instrument.
Structural models of the EGFP1–4
The molar masses Mcalc and partial specific volume (Vspec = 0.732 l/kg of EGFP1–4 according to the amino acid sequence) were calculated with the program SEDNTERP ver. 1.0935. The three-dimensional structure of the EGFP monomer was taken from the Protein Data Bank20. The 5 amino-acid-linkers between the monomers were taken into account in the calculation. Different arrangements of the oligomers were built with the help of the program Wincoot36. Sedimentation coefficients scalc and diffusion coefficients Dcalc for the cylinder-shaped EGFP and its different oligomeric arrangements were calculated with the program HYDROPRO ver. 5a37.
Analytical ultracentrifugation with fluorescence detection system (AU-FDS)
AU-FDS runs were carried out in a ProteomeLab XL-I analytical ultracentrifuge (Beckman Coulter, Indianapolis, IN) equipped with a fluorescence detection system (AU-FDS; Aviv Biomedical Inc., Lakewood, NJ), using An-50 Ti and An-60 Ti rotors. Excitation was at 488 nm using a 10 mW solid-state laser; emission was detected through a dichroic mirror and a 505–565 nm bandpass filter. The centrifuge was programmed and data were recorded using AOS software (Aviv Biomedical Inc., Lakewood, NJ). Special cell housings (Nanolytics, Potsdam, Germany) allowed the placement of 3 mm double sector centerpieces directly beneath the upper window of the cell. All runs were done in 20 mM Tris/HCl pH 8.0, 150 mM NaCl, 1 mM DTT, 0.01% NP40.
Sedimentation velocity runs were done in charcoal-filled epon double-sector centerpieces at 50,000 rpm and 20 °C using a sample volume of 100 μl. Scans were recorded with a radial increment of 20 μm and were averaged over 5 rotor revolutions. Data were analyzed with the program DCDT + (ver. 2.3.2)19,38, which implements the algorithms described in ref.18,39 to obtain the sedimentation and diffusion coefficients (svel and Dvel). All values were corrected to water as solvent. The molar masses (Mvel) of the EGFP oligomers were calculated through the Svedberg equation as:

where R is the gas constant, T is the absolute temperature, ρ0 is the density of the buffer and Vspec is the partial specific volume of EGFP1–4.
Sedimentation equilibrium runs were done at 4 °C in titanium double-sector centerpieces filled with 40 μl sample at 7000, 11000 and 18000 rpm and three different concentrations of the proteins (20, 50 and 150 nM). Data were measured with a radial increment of 5 μm and were averaged over 10 rotor revolutions. After no further change in the concentration distribution could be observed, scans at equilibrium were averaged and molar masses were determined by globally fitting the data at all rotor speeds with a single species model using the program package BPCFIT15:

Here x is the distance from the center of rotation, F(x) and F(x0) are the fluorescence intensities at positions x and x0, ω the angular velocity of the rotor and Meq the molar mass. Baseline offset (Foffset) was set to the buffer fluorescence measured in each cell near the meniscus after sedimenting the proteins for 16 h at 44000 rpm at the end of the experiment.
Fluorescence Correlation Spectroscopy
To measure diffusion properties of EGFP oligomers, we used fluorescence correlation spectroscopy (FCS). The FCS microscope is described in ref. 6. The instrument is based on an Olympus FluoView 1000 confocal microscope (UPlanAPO 60× water immersion objective, NA 1.2) equipped with an FCS extension (Steinbeis Center for Biophysical Analysis, Heidelberg, Germany). For excitation, the 488 nm line of an Ar ion laser was used at different intensities (5, 20, 50 μW at the objective corresponding to ~40–400 MW/m2); fluorescence emission was detected through a 500–550 nm band-pass filter by an avalanche photodiode (Perkin-Elmer). The autocorrelation function of the signal was calculated by an ALV-5000E correlator (ALV-Laser GmbH, Langen, Germany). Runs of 5 × 20 seconds were recorded. Autocorrelation curves were fitted by using the software Quickfit 3.040 to a model assuming 3D diffusion of a single component plus triplet state formation:

where τ is the lag time, T is the equilibrium fraction of dyes in the triplet state, τtr is the triplet correlation time, τd is the diffusion time (the average dwell time of the molecule in the detection volume), N is the average number of molecules in the detection volume and S is the axial ratio of the ellipsoidal detection volume. The diffusion coefficient was calculated using the following expression:

where wxy is the lateral radius of the detection volume (where detection efficiency is e−2 times that of the maximum); wxy was calibrated (using equation (9)) with a solution of Alexa-488 having a known diffusion coefficient (435 μm2/s at 22.5 °C)31, yielding wxy~208 nm when extrapolating its diffusion time to zero laser power.
Additional Information
How to cite this article: Vámosi, G. et al. EGFP oligomers as natural fluorescence and hydrodynamic standards. Sci. Rep. 6, 33022; doi: 10.1038/srep33022 (2016).
References
Bag, N., Huang, S. & Wohland, T. Plasma Membrane Organization of Epidermal Growth Factor Receptor in Resting and Ligand-Bound States. Biophysical journal 109, 1925–1936, doi: 10.1016/j.bpj.2015.09.007 (2015).
Beaudouin, J., Mora-Bermudez, F., Klee, T., Daigle, N. & Ellenberg, J. Dissecting the contribution of diffusion and interactions to the mobility of nuclear proteins. Biophysical journal 90, 1878–1894, doi: 10.1529/biophysj.105.071241 (2006).
Chen, Y., Wei, L. N. & Muller, J. D. Probing protein oligomerization in living cells with fluorescence fluctuation spectroscopy. Proceedings of the National Academy of Sciences of the United States of America 100, 15492–15497, doi: 10.1073/pnas.2533045100 (2003).
Costantini, L. M., Fossati, M., Francolini, M. & Snapp, E. L. Assessing the tendency of fluorescent proteins to oligomerize under physiologic conditions. Traffic 13, 643–649, doi: 10.1111/j.1600-0854.2012.01336.x (2012).
Dross, N. et al. Mapping eGFP oligomer mobility in living cell nuclei. PloS one 4, e5041, doi: 10.1371/journal.pone.0005041 (2009).
Szaloki, N., Krieger, J. W., Komaromi, I., Toth, K. & Vamosi, G. Evidence for Homodimerization of the c-Fos Transcription Factor in Live Cells Revealed by Fluorescence Microscopy and Computer Modeling. Molecular and cellular biology 35, 3785–3798, doi: 10.1128/MCB.00346-15 (2015).
Dean, K. M. et al. Analysis of red-fluorescent proteins provides insight into dark-state conversion and photodegradation. Biophysical journal 101, 961–969, doi: 10.1016/j.bpj.2011.06.055 (2011).
Haupts, U., Maiti, S., Schwille, P. & Webb, W. W. Dynamics of fluorescence fluctuations in green fluorescent protein observed by fluorescence correlation spectroscopy. Proceedings of the National Academy of Sciences of the United States of America 95, 13573–13578 (1998).
Moerner, W. E., Peterman, E. J., Brasselet, S., Kummer, S. & Dickson, R. M. Optical methods for exploring dynamics of single copies of green fluorescent protein. Cytometry 36, 232–238 (1999).
Peterman, E. J., Brasselet, S. & Moerner, W. E. The Fluorescence Dynamics of Single Molecules of Green Fluorescent Protein. J. Phys. Chem. A 103, 10553–10560 (1999).
Visser, A. J. W. G. & Hink, M. A. New perspectives of fluorescence correlation spectroscopy. J Fluoresc 9, 81–87, doi: 10.1023/A:1020595926133 (1999).
Widengren, J., Mets, U. & Rigler, R. Photodynamic properties of green fluorescent proteins investigated by fluorescence correlation spectroscopy. Chem Phys 250, 171–186, doi: 10.1016/S0301-0104(99)00255-4 (1999).
Widengren, J., Terry, B. & Rigler, R. Protonation kinetics of GFP and FITC investigated by FCS - aspects of the use of fluorescent indicators for measuring pH. Chem Phys 249, 259–271, doi: 10.1016/S0301-0104(99)00256-6 (1999).
Pack, C., Saito, K., Tamura, M. & Kinjo, M. Microenvironment and effect of energy depletion in the nucleus analyzed by mobility of multiple oligomeric EGFPs. Biophysical journal 91, 3921–3936, doi: 10.1529/biophysj.105.079467 (2006).
Witte, G., Urbanke, C. & Curth, U. Single-stranded DNA-binding protein of Deinococcus radiodurans: a biophysical characterization. Nucleic acids research 33, 1662–1670, doi: 10.1093/nar/gki310 (2005).
MacGregor, I. K., Anderson, A. L. & Laue, T. M. Fluorescence detection for the XLI analytical ultracentrifuge. Biophysical chemistry 108, 165–185, doi: 10.1016/j.bpc.2003.10.018 (2004).
Kroe, R. R. & Laue, T. M. NUTS and BOLTS: applications of fluorescence-detected sedimentation. Analytical biochemistry 390, 1–13, doi: 10.1016/j.ab.2008.11.033 (2009).
Stafford, W. F. Sedimentation velocity spins a new weave for an old fabric. Current opinion in biotechnology 8, 14–24 (1997).
Philo, J. S. A method for directly fitting the time derivative of sedimentation velocity data and an alternative algorithm for calculating sedimentation coefficient distribution functions. Analytical biochemistry 279, 151–163, doi: 10.1006/abio.2000.4480 (2000).
Ormo, M. et al. Crystal structure of the Aequorea victoria green fluorescent protein. Science 273, 1392–1395 (1996).
Szabo, A., Horvath, G., Szollosi, J. & Nagy, P. Quantitative characterization of the large-scale association of ErbB1 and ErbB2 by flow cytometric homo-FRET measurements. Biophysical journal 95, 2086–2096, doi: 10.1529/biophysj.108.133371 (2008).
Weber, G. Dependence of polarization of the fluorescence on the concentration. Trans. Faraday Soc 50, 552–555 (1954).
Visscher, K., Brakenhoff, G. J. & Visser, T. D. Fluorescence Saturation in Confocal Microscopy. J Microsc-Oxford 175, 162–165 (1994).
Dross, N. Mobilität von eGFP-Oligomeren in lebenden Zellkernen PhD thesis, Ruprecht-Karls-Universität, Heidelberg (2009).
Genove, G., Glick, B. S. & Barth, A. L. Brighter reporter genes from multimerized fluorescent proteins. BioTechniques 39, 814, 816, 818 passim (2005).
Seibel, N. M., Eljouni, J., Nalaskowski, M. M. & Hampe, W. Nuclear localization of enhanced green fluorescent protein homomultimers. Analytical biochemistry 368, 95–99, doi: 10.1016/j.ab.2007.05.025 (2007).
Nalaskowski, M. M., Ehm, P., Giehler, S. & Mayr, G. W. A toolkit for graded expression of green fluorescent protein fusion proteins in mammalian cells. Analytical biochemistry 428, 24–27, doi: 10.1016/j.ab.2012.06.001 (2012).
Madl, J. et al. Resting state Orai1 diffuses as homotetramer in the plasma membrane of live mammalian cells. The Journal of biological chemistry 285, 41135–41142, doi: 10.1074/jbc.M110.177881 (2010).
Nagy, P., Claus, J., Jovin, T. M. & Arndt-Jovin, D. J. Distribution of resting and ligand-bound ErbB1 and ErbB2 receptor tyrosine kinases in living cells using number and brightness analysis. Proceedings of the National Academy of Sciences of the United States of America 107, 16524–16529, doi: 10.1073/pnas.1002642107 (2010).
Zhao, H., Casillas, E. Jr., Shroff, H., Patterson, G. H. & Schuck, P. Tools for the quantitative analysis of sedimentation boundaries detected by fluorescence optical analytical ultracentrifugation. PloS one 8, e77245, doi: 10.1371/journal.pone.0077245 (2013).
Petrasek, Z. & Schwille, P. Precise measurement of diffusion coefficients using scanning fluorescence correlation spectroscopy. Biophysical journal 94, 1437–1448, 10.1529/biophysj.107.108811 (2008).
Arrio-Dupont, M., Foucault, G., Vacher, M., Devaux, P. F. & Cribier, S. Translational diffusion of globular proteins in the cytoplasm of cultured muscle cells. Biophysical journal 78, 901–907 (2000).
Widengren, J. & Rigler, R. Photobleaching investigations of dyes using fluorescence correlation spectroscopy (FCS). Prog Biophys Mol Bio 65, Ph109–Ph109 (1996).
Shaner, N. C., Steinbach, P. A. & Tsien, R. Y. A guide to choosing fluorescent proteins. Nature methods 2, 905–909, doi: 10.1038/nmeth819 (2005).
Sednterp: Sedimentation Interpretation Program version 1.09. Alliance Protein Laboratories, Thousand Oaks, CA, USA. (1995).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta crystallographica. Section D, Biological crystallography 66, 486–501, doi: 10.1107/S0907444910007493 (2010).
Garcia De La Torre, J., Huertas, M. L. & Carrasco, B. Calculation of hydrodynamic properties of globular proteins from their atomic-level structure. Biophysical journal 78, 719–730, doi: 10.1016/S0006-3495(00)76630-6 (2000).
Philo, J. S. Improved methods for fitting sedimentation coefficient distributions derived by time-derivative techniques. Analytical biochemistry 354, 238–246, doi: 10.1016/j.ab.2006.04.053 (2006).
Stafford, W. F. 3rd . Boundary analysis in sedimentation transport experiments: a procedure for obtaining sedimentation coefficient distributions using the time derivative of the concentration profile. Analytical biochemistry 203, 295–301 (1992).
Krieger, J. W. & Langowski, J. QuickFit 3.0 (status: beta, compiled: Jan 5, 2015, SVN: 3695): A data evaluation application for biophysics, http://www.dkfz.de/Macromol/quickfit/v.3.0 (status: beta, compiled: Oct 2015) (2015).
Acknowledgements
We thank Dr. Dirk-Peter Herten (Bioquant, Heidelberg) for the use of the Picoquant fluorescence lifetime spectrometer. This work was supported by grants from the Hungarian Scientific Research Fund K103965 to G.V., the German Academic Exchange Service and the Tempus Public Foundation #73163 to K.T. and G.V.; the European Union and the State of Hungary, co-financed by the European Social Fund in the framework of TÁMOP-4.2.4.A/ 2-11/1-2012-0001 ‘National Excellence Program’ to G.V.
Author information
Authors and Affiliations
Contributions
G.M. prepared the proteins, U.C. and N.M. performed and analyzed AUC measurements, G.V. and K.T. performed and analyzed FCS measurements, J.W.K. made FCS simulations, K.T. carried out absorption and fluorescence spectroscopy experiments, G.V. set up the model, and all the authors edited the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Vámosi, G., Mücke, N., Müller, G. et al. EGFP oligomers as natural fluorescence and hydrodynamic standards. Sci Rep 6, 33022 (2016). https://doi.org/10.1038/srep33022
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep33022
This article is cited by
-
Current and future advances in fluorescence-based visualization of plant cell wall components and cell wall biosynthetic machineries
Biotechnology for Biofuels (2021)
-
Structural and mechanistic basis of capsule O-acetylation in Neisseria meningitidis serogroup A
Nature Communications (2020)
-
Optimal fluorescent protein tags for quantifying protein oligomerization in living cells
Scientific Reports (2018)
-
Monomerization of the photoconvertible fluorescent protein SAASoti by rational mutagenesis of single amino acids
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.