Abstract
Like most other types of cancer cells, leukaemia cells undergo metabolic reprogramming to support rapid proliferation through enhancing biosynthetic processes. Pentose phosphate pathway (PPP) plays a pivotal role in meeting the anabolic demands for cancer cells. However, the molecular mechanism by which PPP contributes to leukaemia remains elusive. Here, we report that leukaemia cell proliferation is dependent on the oxidative branch of PPP, in particular the first and rate-limiting enzyme glucose-6-phosphate dehydrogenase (G6PD). Knockdown of G6PD reduces NADPH level in acute myeloid leukaemia (AML) cell lines. Exogenous lipid supplements partially restore the proliferation of G6PD-depleted cells. Deacetylase SIRT2 promotes NADPH production through deacetylating G6PD at lysine 403 (K403). Activation of G6PD by SIRT2 supports the proliferation and clonogenic activity of leukaemia cells. Chemical inhibitors against SIRT2 suppress G6PD activity, leading to reduced cell proliferation of leukaemia cells, but not normal hematopoietic stem and progenitor cells. Importantly, SIRT2 is overexpressed in clinical AML samples, while K403 acetylation is downregulated and G6PD catalytic activity is increased comparing to that of normal control. Together, our study reveals that acetylation regulation of G6PD is involved in the metabolic reprogramming of AML, and SIRT2 serves as a promising target for further therapeutic investigations.
Similar content being viewed by others
Introduction
Dysregulation of metabolic pathways, termed as metabolic reprogramming, is one of characteristic features of cancers1. Cancer cells take up and utilize more glucose for glycolysis than oxidative phosphorylation to support rapid cell proliferation (Warburg effect)2. The preference towards aerobic glycolysis is critical to provide tumour cells with not only energy, but also multiple building blocks for the biosynthesis of macromolecules, such as proteins, lipids, and nucleic acids etc3.
Acute myeloid leukaemia (AML) is the most common type of malignant myeloid disorder in adults4,5. Recent evidence revealed that leukaemia cells exhibited an increased dependence on aerobic glycolysis6,7. Mouse model studies further demonstrated that enhanced glucose uptake accelerated leukemogenesis in vivo8,9,10. Besides, glycolysis inhibition resulted in growth arrest or cell death of AML cells, and sensitized leukaemia cells to chemotherapeutic agents7,11. Notably, pentose phosphate pathway (PPP), a shunt for glycolysis, was also frequently altered in AML. Glucose turnover in leukaemia cells was possibly enhanced via PPP pathway rather than glycolysis12,13,14,15, indicating the implication of PPP pathway in metabolic reprogramming of leukaemia.
Glucose is converted into glucose-6-phosophate (G6P) upon entering into the cell. Afterward, G6P is either utilized in glycolysis for catabolism, or shunted towards pentose phosphate pathway. Based on the catalytic properties, PPP can be divided into oxidative and non-oxidative phases. In the oxidative branch, glucose-6-phosphate dehydrogenase (G6PD) and 6-phosphogluconate dehydrogenase (PGD) catalyse sequential dehydrogenation reactions of G6P, yielding five-carbon sugars (ribose) and nicotinamide adenine dinucleotide phosphate (NAPDH). In the non-oxidative phase, transketolase (TKT) and transaldolase (TALDO1) converted/recycled five-carbon sugars into glycolytic intermediates. Ribose is an essential precursor for nucleotide synthesis16, while NADPH functions as a reducing power for fatty acid synthesis and anti-oxidant processes. By producing theses two key intermediates (ribose and NADPH), PPP pathway plays a pivotal role in meeting the anabolic demands for cancer cells17,18,19,20,21.
G6PD is the first and rate-limiting enzyme of PPP. G6PD deficiency is one of the most common enzyme defects in humans, affecting 400 million people worldwide22. The lack of G6PD results in shortage of NADPH and deficiency of red blood cells (RBCs) in scavenging ROS, clinically characterized by hemolysis and anaemia23. Previously, we found that G6PD was post-translationally modified by lysine acetylation. SIRT2 deacetylated G6PD at lysine 403 (K403) and enhanced its activity in response to oxidative stress, protecting RBCs from oxidative damage24. In addition, recent studies demonstrated that G6PD was implicated in the development of multiple cancers, including breast cancer, liver cancer, leukaemia etc25. Leukaemia cells expressed high level of G6PD, even in G6PD-deficient subjects26. In chronic myelomonocytic leukaemia (CMML), overexpression of G6PD mRNA associated with poor diagnosis27. These observations indicate a potential link between G6PD deregulation and leukaemia development. However, the role of G6PD in metabolic reprogramming of leukaemia remains unclear. This study investigates the functional significance of PPP pathway, especially G6PD, in leukaemia development.
Results
Oxidative PPP is essential for the proliferation of leukaemia cells
PPP pathway sustains rapid cell growth by providing NADPH and pentose to biosynthetic processes (Fig. 1a). To dissect the contribution of PPP to leukaemia, we constructed a shRNA library targeting PPP enzymes and tested the dependence of leukaemia cell proliferation on these enzymes. Interestingly, depletion of enzymes in oxidative PPP, i.e. G6PD, PGLS (6-phosphogluconolactonase), and PGD, dramatically abrogated the proliferation of HL-60, a human promyelocytic leukaemia cell line (Fig. 1b–d and s1a). Contrarily, knockdown of non-oxidative PPP enzymes, including RPE (ribulose 5-phosphate 3-epimerase), RPI (ribulose 5-phosphate isomerase), TALDO1 (transaldolase), and TKT (transketolase), had negligible effects on cell proliferation (Fig. 1e–h and s1a). Accordingly, CCK-8 assay also demonstrated that oxidative PPP, but not non-oxidative PPP, is necessary for the proliferation of leukaemia cells (Fig. 1i). In support of these observations, cell growth of another two AML cell lines with different FAB subtypes (THP-1 and KG-1) was remarkably suppressed upon shRNA-induced G6PD knockdown (Supplementary Table 2 and Fig. 1j,k). Moreover, G6PD inhibitors, i.e. dehydroepiandrosterone (DHEA) and 6-aminonicotinamide (ANAD), significantly decreased the proliferation of HL-60, KG-1, and THP-1 cells in a dose-dependent manner (Fig. 1l,m). Together, these data demonstrate that leukaemia cell proliferation is dependent on the oxidative branch of PPP, in particular G6PD, across different subtypes.
G6PD is essential for the proliferation of leukaemia cells.
(a) Schematic overview of pentose phosphate pathway. Enzymes for individual chemical reactions are labelled as ovals and denoted next to the arrows connecting two metabolites. Metabolites and enzymes in oxidative PPP are colored in black, non-oxidative PPP in dark grey. G6P, glucose 6-phosphate; F6P, fructose 6-phosphate; F1,6BP, fructose 1,6-bisphosphate; DHAP, dihydroxyacetone phosphate; G3P, glyceraldehydes 3-phosphate; 6PGL, 6-phosphogluconolactone; 6PG, 6-phosphogluconate; R5P, ribulose 5-phosphate; X5P, xylulose 5-phosphate. (b–h) The proliferation curve of HL-60 cells expressing a control shRNA (shscr.) or shRNAs against G6PD (b), PGLS (c), PGD (d), RPI (e), RPE (f), TKT (g), or TALDO1 (h) was determined by cell counting. (i) HL-60 cells stably expressing control shRNA (scramble) or shRNAs targeting genes in PPP pathway as indicated were grown for 5 days, relative cell growth was determined by CCK8 assay. (j–k) The proliferation of KG-1 (j) and THP-1 (k) cells stably expressing control shRNA (shscr.) or shRNAs against G6PD were determined by cell counting. (l,m) HL-60, KG-1 and THP-1 cells were grown for 5 days with or without treatment of increasing concentrations of DHEA (l) or ANAD (m). Relative cell growth was determined by cell counting. Error bars represent mean ± SD from three replicates of each sample (*p < 0.05, **p < 0.01, n.s. = not significant for the indicated comparison).
G6PD maintains NADPH level in leukaemia cells
Next, we investigated metabolic alterations caused by G6PD knockdown. G6PD converts G6P and coenzyme NADP+ to 6PG and NADPH (Fig. 1a). Depletion of G6PD significantly reduced glucose consumption of HL-60, KG-1 and THP-1 cells (Fig. 2a–f). In accordance, knockdown of G6PD resulted in 1.4-fold accumulation of G6P (p = 0.015) and a 30% reduction of 6PG (p = 0.032) in HL-60 (Fig. 2g,h). Cellular NADPH/NADP+ ratio was significantly decreased by G6PD depletion in HL-60, KG-1 and THP-1 cells (Fig. 2i–k). These results suggest that G6PD is essential for cellular NADPH production in leukaemia cells.
G6PD maintains NADPH level in leukaemia cells.
(a–f) Knockdown efficiencies of shRNAs targeting G6PD in HL-60 (a), KG-1 (c), and THP-1 (e) cells was determined by western blotting. Relative glucose consumptions of HL-60 (b), KG-1 (d), and THP-1 (f) stable cells were determined. (g,h) Relative concentrations of G6P (glucose 6-phosphate) (g) and 6PG (6-phosphpogluconate) (h) in control or G6PD-knockdown HL-60 cells were determined. (i–k) Relative NADPH/NADP+ ratios in control or G6PD-knockdown HL-60 (i), KG-1 (j), and THP-1 (k) cells were determined. (l,m) Relative GSH/GSSG ratio (I) and ROS level (m) in control or G6PD-knockdown HL-60 cells were determined. Error bars represent mean ± SD from three replicates of each sample (*p < 0.05, **p < 0.01, n.s. = not significant for the indicated comparison).
NADPH can be utilized in the regeneration of reduced glutathione (GSH), which detoxifies reactive oxygen species (ROS). Interestingly, depletion of G6PD altered neither the ratio of reduced to oxidized glutathione (GSH/GSSG), nor cellular ROS level of HL-60 (Fig. 2l,m). These results indicate that ROS scavenging was not impaired by the decline of NADPH supply in G6PD-knockdown cells.
G6PD-deficient cells exhibit an increased demand for lipids
Alternatively, the NADPH produced by G6PD may support biosynthesis of macromolecules in leukaemia cells. If so, exogenous nutrient supplements may rescue the growth defect of G6PD-depleting cells. To identify potential biosynthetic process(es) affected by G6PD depletion, we added different supplements, including nucleosides and lipids, into the culture medium of G6PD-knockdown cells. In line with our earlier findings, reducing agents (GSH and N-acetyl cysteine) were incapable of restoring the proliferation of G6PD-depleting cells (Fig. 3a). Similarly, ribonucleosides or deoxyribonucleosides failed to restore the growth of G6PD-knockdown cells (Fig. 3a). However, supplementation of lipids, such as myristic acid (MA), palmitic acid (PA), or stearic acid (SA), partially rescued the growth defect of G6PD-knockdown cells (Fig. 3a). Of note, PA partially restored the proliferation of G6PD-knockdown cells, but not control HL-60 or THP-1 cells (Fig. 3b–d). These results indicate that the NADPH produced by G6PD is possibly utilized in lipogenic reactions of leukaemia cells, explaining increased lipid requirements of G6PD-deficient AML cells.
Supplementation of lipids rescues proliferation and colony formation of G6PD-knockdown cells.
(a) Control or G6PD-knockdown stable HL-60 cells were grown for 5 days. G6PD-depleted cells were either untreated or supplemented with ribonucleosides, deoxyribonucleosides, 5mM reducing agents (GSH or NAC), or 20 μM lipids (MA, myristic acid; PA, palmitic acid; SA, stearic acid) as indicated. Relative cell growth was determined by cell counting. (b) Control (shscr.) and G6PD-knockdown stable HL-60 cells were treated with or without palmitic acid (PA), relative cell proliferation was determined by cell counting. (c) Control or G6PD-knockdown stable HL-60 cells were treated with increasing concentrations of palmitic acid as indicated and grown for 5 days. Relative cell growth was determined by cell counting. (d) Control (shscr.) and G6PD-knockdown stable THP-1 cells were treated with or without palmitic acid (PA), relative cell proliferation was determined by cell counting. (e) Control and G6PD-knockdown stable HL-60 cells were treated with or without palmitic acid. Colony formation of was determined. Shown are representative images (Bar = 100 μm). (f) Control or G6PD-knockdown stable HL-60 cells were supplemented with increasing concentrations of palmitic acid as indicated. Relative colony formation was determined. (g) Control or G6PD-knockdown stable HL-60 cells were supplemented with or without different lipids (20 μM) as indicated. Relative colony formation was determined. Error bars represent mean ± SD from three replicates of each sample (*p < 0.05, **p < 0.01, n.s. = not significant for the indicated comparison).
We further investigated the effect of PA on clongenic ability of leukaemia cells. Knockdown of G6PD severely reduced the colony formation of HL-60 cells, which was partly rescued by PA supplementation in a dose dependent manner (Fig. 3e,f). Meanwhile, clongenic activity of control cells remained unaltered by PA treatment (Fig. 3e,f). Interestingly, MA and SA also partially restored the colony formation of G6PD-deficient cells (Fig. 3g). Together, G6PD activity is essential for both cell proliferation and colony formation of leukaemia cells, presumably through supporting de novo lipogenesis.
SIRT2-mediated deacetylation of G6PD promotes NADPH production
The dependence of leukaemia cell on G6PD implies that suppression of oxidative PPP, in particular G6PD, may serve as a promising strategy to inhibit leukaemia. However, targeting PPP remains challenging due to the lack of specific G6PD inhibitors. Previously, we found that G6PD was post-translationally modified by lysine acetylation. SIRT2 deacetylated G6PD at lysine 403 (K403) and activated its activity24. Thus, inhibition of SIRT2 would be an alternative approach to suppress G6PD and oxidative PPP. To this end, we examined the physiological significance of the interaction between SIRT2 and G6PD in leukaemia cells. Endogenous immunoprecipitation assay demonstrated that the physical interaction between G6PD and SIRT2 was readily detectable in HL-60, KG-1, and THP-1 cells (Fig. 4a). shRNA-induced knockdown of SIRT2 elevated G6PD K403 acetylation and remarkably decreased G6PD activity in HL-60 and THP-1 cells (Fig. 4b,c). Moreover, SIRT2-specific inhibitor AGK2 increased G6PD acetylation and reduced its catalytic activity in a dose-dependent manner (Fig. 4d). These data clearly indicate that G6PD is deacetylated and activated by SIRT2 in leukaemia cells.
Deacetylation of G6PD by SIRT2 promotes NADPH production.
(a) Endogenous G6PD protein was immunoprecipitated from human AML cell lines (HL-60, KG-1 and THP-1), followed by western blotting to detect SIRT2. Normal rabbit IgG was included as a negative control. (b,c) Endogenous G6PD protein was immunoprecipitated from control or SIRT2-knockdown stable HL-60 (b) or THP-1 (c) cells, followed by western blotting to detect K403 acetylation [acG6PD (K403)]. Catalytic activity of endogenous G6PD was determined. WCL denotes whole cell lysate (d) HL-60 cells were treated with increasing doses of AGK2 as indicated for 6hrs. K403 acetylation and catalytic activity of immunopurified endogenous G6PD was determined. (e) Relative NADPH/NADP+ ratio in HL-60 cells stably expressing shRNA against G6PD and/or SIRT2 was determined. (f,g) Control or G6PD-knockdown stable HL-60 (f) and THP-1 (g) cells were treated with or without 5 μM AGK2 for 6hrs. Relative NADPH/NADP+ ratio were determined. (h) Relative ROS level in HL-60 cells stably expressing shRNA against G6PD and/or SIRT2 was determined. (i) Control or G6PD-knockdown stable THP-1 cells were treated with or without 5 μM AGK2 for 6 hrs. Relative ROS level was determined. Error bars represent mean ± SD from three replicates of each sample (**p < 0.01, n.s. = not significant for the indicated comparison).
Next, we investigated the role of SIRT2-mediated deacetylation and activation of G6PD in glucose utilization and NADPH production. Glucose uptake of HL-60 was inhibited upon knockdown of SIRT2 or G6PD, while depletion of SIRT2 did not led to further reduction of glucose consumption in G6PD-knockdown cells (Fig. s2a,b). Additionally, NADPH/NADP+ ratio was decreased by two-fold in SIRT2-knockdown cells, while depleting SIRT2 exhibited no effect on NADPH level in G6PD-knockdown HL-60 cells (Fig. 4e). AGK2 treatment reduced the NADPH/NADP+ ratio of control cells, but not G6PD-knockdown HL-60 or THP-1 cells (Fig. 4f,g). Of note, inhibiting SIRT2 using shRNA or chemical inhibitor AGK2 displayed no effect on cellular redox state of HL-60 or THP-1 cells (Fig. 4h,i). These results suggest that SIRT2 maintains NADPH level in a manner dependent on G6PD.
Inhibition of SIRT2 reduces leukaemia cell proliferation
We next explored the functional role of acetylation regulation of G6PD by SIRT2 in leukaemia cell proliferation. Depletion of SIRT2 reduced both cell growth and clongenic ability of control HL-60 cells, but not G6PD-knockdown cells (Fig. 5a,b). Consistently, AGK2 treatment failed to suppress the proliferation and colony formation of G6PD-knockdown HL-60 and THP-1 cells (Fig. 5c–f). Interestingly, PA supplementation partially restored the proliferation and colony formation of SIRT2- and/or G6PD-knockdown cells, but not control HL-60 cells (Fig. 5g). Suppression of cell growth and colony formation by AGK2 was partly rescued by PA treatment (Fig. 5h). These data demonstrate that the contribution of SIRT2 to leukaemia cell proliferation and colony formation is dependent on G6PD.
Inhibiting the deacetylation of G6PD reduces leukaemia cell proliferation.
(a) Stable G6PD- and/or SIRT2-knockdown HL-60 cells as indicated were grown for 5 days, relative cell growth was determined by cell counting. (b) Colony formation of G6PD- and/or SIRT2-knockdown stable HL-60 cells was determined. (c,d) Control or G6PD-knockdown HL-60 (c) and THP-1 (d) cells were treated with or without AGK2. Cell proliferation was determined by cell counting. (e,f) Relative cell growth (e) and colony formation (f) of control or G6PD-knockdown HL-60 cells with indicated treatment was determined by CCK-8 assay and clonogenic assay, respectively. (g) G6PD- and/or SIRT2-knockdown stable HL-60 cells were supplemented with or without PA, relative cell growth was determined by cell counting after cells were grown for 5 days. (h) Control or G6PD-knockdown HL-60 cells were treated as indicated, colony formation was determined. Error bars represent mean ± SD from three replicates of each sample (*p < 0.05, **p < 0.01, n.s. = not significant for the indicated comparison).
Small molecule inhibitors of SIRT2 decreases G6PD activity and suppresses leukaemia cell proliferation
The observation that SIRT2-induced activation of G6PD contributes to leukaemia indicates that inhibiting SIRT2 with small molecule inhibitors may serve as a substitutive strategy to suppress G6PD. Recently, a spectrum of chemical inhibitors against SIRT2 has been developed, from which we utilized sirtinol, AGK2, and SirReal2 to evaluate their anti-proliferative effects (Supplementary Table 3). Interestingly, SIRT2 inhibitors dose-dependently decreased G6PD activity (Fig. 6a–c), and resulted in declined proliferation of HL-60 cells (Fig. 6d–f). SIRT2 inhibitors also suppressed G6PD activity (Fig. 6g–i) and cell proliferation (Fig. 6j–l) in KG-1 cells. To test the preference of SIRT2 inhibitors, we isolated normal hematopoietic stem and progenitor cells (HSPCs) and found that SIRT2-specific inhibitors (AGK2 and SirReal2) selectively decreased the proliferation of mouse leukaemia cell RAW264.7, but not HSPCs (Fig. 6m,n). These data suggest that SIRT2 inhibitors efficiently inactivate G6PD, and preferentially inhibit the proliferation of leukaemia cells.
Small molecule inhibitors of SIRT2 decrease G6PD activity and suppress leukaemia cells.
(a–c) HL-60 cells were treated with increasing doses of SIRT2 inhibitors, sirtinol (a), AGK2 (b) and SirReal2 (c), for 6 hours as indicated, G6PD activity was determined. (d–f) HL-60 cells were treated with increasing doses of SIRT2 inhibitors, sirtinol (d), AGK2 (e) and SirReal2 (f), relative cell growth was determined by cell counting after grown for 5 days. (g–i) KG-1 cells were treated with increasing doses of SIRT2 inhibitors, sirtinol (g), AGK2 (h) and SirReal2 (i), for 6 hours as indicated, G6PD activity was determined. (j–l) KG-1 cells were treated with increasing doses of SIRT2 inhibitors, sirtinol (j), AGK2 (k) and SirReal2 (l), relative cell growth was determined by cell counting after grown for 5 days. (m,n) Mouse hematopoietic stem and progenitor cells (HSPC) and RAW264.7 cells were treated with 5 μM AGK2 (m) or 150 nM SirReal2 (n) as indicated and grown for 5 days, cell proliferation curve was determined by cell counting. Error bars represent mean ± SD from three replicates of each sample (**p < 0.01, n.s. = not significant for the indicated comparison).
K403 acetylation of G6PD is downregulated in AML
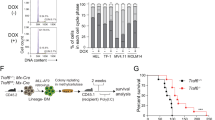
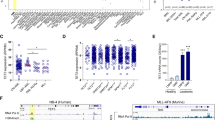
We further explored the pathological relevance of SIRT2-induced G6PD activation in clinical samples of AML. We collected mononuclear cells from bone marrows of 8 AML patients and 4 healthy donors (Supplementary Table 4). mRNA expression analysis showed that AML cells expressed higher level of SIRT2 than normal controls (p = 0.0024) (Fig. 7a), while no significant difference was observed in G6PD mRNA expression (Fig. 7b). Notably, AGK2 treatment strongly inhibited G6PD activity in AML cells (p = 0.0004), but not normal samples (Fig. 7c,d). Direct immunoblotting assay demonstrated that SIRT2 protein was significantly increased (p = 0.0107), while G6PD was mildly overexpressed in AML samples (Fig. 7e–g). Of note, G6PD enzymes in AML samples were more active than that from normal tissues (p = 0.0229) (Fig. 7h). Accordingly, K403 acetylation of G6PD was reduced in AML cells (p < 0.0001) (Fig. 7i). These data indicate that G6PD K403 acetylation is downregulated in AML. We further carried out a survival analysis by using TCGA AML cohort28. In line with our findings, high expression of G6PD, but not the other enzymes in PPP, significantly associated with poor survival rate (p = 0.0069) (Fig. 7j and Fig. s3a,b). Reduced survival rate was also observed in AML cases overexpressing SIRT2 (p = 0.0152) (Fig. 7k). These data strongly support that acetylation regulation of G6PD by SIRT2 is involved in the development of AML.
Acetylation at K403 of G6PD is downregulated in AML.
(a,b) Relative mRNA expression of SIRT2 (A) and G6PD (B) in control and AML samples was determined by qRT-PCR. (c,d) Leukaemia (c) or normal (d) cells from 4 healthy donors and 8 clinical AML patients were treated with or without 5 μM AGK2 for 6hrs, G6PD activity was determine. (e–i) Clinical AML samples and normal controls were lysed. Protein expression was determined by western blotting as indicated (e), SIRT2 (f) and G6PD (g) expression were quantified and normalized to β-actin (n.s. = not significant). G6PD activities were determined and normalized against G6PD protein (h). K403 acetylation level of immunopurified G6PD was quantified. Acetylation ratio was normalized against G6PD protein (i). (j,k) Kaplan-Meier survival curves for TCGA AML study. The patients (n = 173) were stratified by median G6PD (j) and SIRT2 (k) levels (‘high’ is greater than median, ‘low’ is less than or equal to median). *p < 0.05, **p < 0.01, n.s. = not significant for the indicated comparison.
Discussion
Cancer metabolism is reprogrammed to enhance the biosynthesis of diverse macromolecules, fulfilling requirements for rapid proliferation. In this study, we investigated the dysregulation of PPP pathway in AML. We demonstrated that proliferation of leukaemia cells was dependent on the oxidative branch of PPP, in particular G6PD. G6PD-knockdown cells had decreased NADPH/NADP+ ratio, and exhibited an enhanced demand for lipids, indicating that the NADPH produced by G6PD was necessary for de novo lipid synthesis. In addition, SIRT2 deacetylated and activated G6PD to maintain intracellular NADPH level. Chemical inhibitors against SIRT2 effectively impaired G6PD activity and suppressed the proliferation of leukaemia cells. Of note, we found that SIRT2 was overexpressed, with corresponding low K403 acetylation levels of G6PD, in clinical AML samples. High SIRT2 expression correlated with poor overall survival rate in AML patients. Taken together, our study reveals that deacetylation of G6PD by SIRT2 sustains NADPH production and promotes leukaemia cell proliferation (Fig. 8).
Although G6PD serves as a potential target to suppress leukaemia, ANAD is not amenable to in vivo use because of its toxic side effects29. DHEA is a naturally occurring hormone with pleiotropic effects that may be unrelated to G6PD inactivation30. Besides, DHEA may enhance G6PD mRNA expression, confounding its inhibitory effects31. The lack of G6PD-specific inhibitors prompted us to explore alternative strategies to target G6PD and oxidative PPP. Recent studies have identified multiple regulators of G6PD at transcription and post-translational levels. For example, tumour suppressor p53 directly bound to and inhibited G6PD17. TAp73, a p53 family member, activates G6PD gene transcription21. Phosphatase and tensin homologue (PTEN) suppresses PPP pathway through abrogating G6PD pre-mRNA splicing and preventing the formation of active G6PD dimers32. Upon exposure to irradiation, ATM kinase promotes G6PD activity to enhance nucleotide production33. Post-translationally, G6PD is modified by lysine acetylation, phosphorylation, ubiquitination and O-GlcNAcylation34,35,36,37. We previously found that G6PD is deacetylated and activated by SIRT238, which led us to explore the possibility of targeting SIRT2 to suppress G6PD and leukaemia39,40,41. SIRT2-mediated deacetylation of G6PD sustained NADPH production and promoted cell proliferation across different subtypes of AML. SIRT2-specific inhibitors effectively suppressed the proliferation of leukaemia cells, but not normal hematopoietic cells. The anti-proliferative effect of SIRT2 inhibitors merits further explorations in leukaemia mouse models and clinical trials. It is noteworthy that not every SIRT2 inhibitor achieved high efficacy of growth suppression in our cell proliferation assays. Development of more potent and specific inhibitors against SIRT2 is currently under active investigation42. Recently, it was proposed that SIRT2 inhibition may exhibit anti-cancer effect through promoting the degradation of Myc oncoprotein43. However, we found that shRNAs or chemical inhibitors against SIRT2 downregulated G6PD activity and suppressed leukaemia cell growth irrespective of Myc expression level (Fig. s1b). This observation indicates that the anti-cancer effect of SIRT2 inhibitors may be context-dependent.
Cancer cells consume large amounts of fatty acids for membrane production and lipid modification of proteins44. In the absence of G6PD, leukaemia cells exhibited enhanced demand for exogenous lipids, indicating that G6PD-knockdown cells are possibly deficient in de novo lipogenesis. Therefore, assimilating exogenous lipids would replenish lipid supplies and restore leukaemia cell proliferation. Supporting this notion, increased lipid mobilization had been observed in chronic lymphocytic leukaemia (CLL) patients as the lipolysis activity is significantly enhanced45,46.
G6PD depletion did not impair redox homeostasis of leukaemia cells, as GSH/GSSG ratio and ROS level remained unchanged in G6PD-knockdown cells. Ribose supplementation was incapable to rescue the growth defect of G6PD-knockdown cells. Yet it is possible that lack of G6PD affects ROS scavenging and ribose synthesis under stressed conditions, such as acute ROS challenge or DNA damage47,48.
Taken together, our study uncovers that leukaemia cells are dependent on oxidative PPP, in particular G6PD. SIRT2-mediated G6PD deacetylation and activation contributes to the development of AML and serves as a promising druggable target.
Methods
Generation of Stable Cell Pools
To generate stable knockdown cell pools in HL-60 cells, shRNA library containing two different shRNAs targeted towards each metabolic gene in pentose phosphate pathway were co-transfected with vectors expressing the gag and vsvg genes into HEK293T cells using a two-plasmid packaging system as previously described49,50. Retroviral supernatant was harvested 36 hrs after transfection, and mixed with 8 μg/mL polybrene to increase the infection efficiency. Cells were infected with the retrovirus and selected in 1 μg/ml puromycin for 1 week.
To generate stable G6PD- and/or SIRT2-knockdown cell pools in HL-60 and KG-1 cells, two different shRNAs targeting SIRT2 gene were constructed in pMKO.1-hyg vector as described previously51 (shSIRT2-#1: 5′-GCTCATCAACAAGGAGAAAGC-3′; shSIRT2-#2: 5′-GCATGGACTTTGACTCCAAGA-3′). The virus was produced as described above. Retroviral supernatant was harvested 36 hrs after transfection, and mixed with 8 μg/mL polybrene to increase the infection efficiency. G6PD-knockdown stable cells were infected with the retrovirus and selected in 200 μg/ml hygromycin B for 1 week.
Human Acute Myeloid Leukaemia Samples
The protocol of studies on human samples was approved by the Institutional Ethics Review Board of Southwest Hospital, Third Military Medical University. All methods were performed in accordance with the guidelines and regulations of Southwest Hospital, Third Military Medical University. Written informed consent was obtained from all participants. Leukaemia bone marrow samples were collected from AML patients. Control bone marrow samples were acquired from participants who have been excluded of haematological diseases. Mononuclear cells from these samples were prepared by red blood cell lyses method and were stored in liquid nitrogen until use. Bulk populations of normal or leukaemia cells were used for the analysis in this study.
Statistical Analysis
Statistical analyses were performed with a two-tailed unpaired Student’s t-test. All data shown represent the results obtained from triplicate independent experiments with standard errors of the mean (mean ± S.D.). Kaplan-Meier survival curves were prepared using tumours with mRNA data (RNA Seq V2) (n = 173) from TCGA AML study. The patients were stratified by median mRNA expression Z-score. ‘High’ indicates greater than median (n = 86), ‘low’ denotes less than or equal to median (n = 86). P-value was determined by the log-rank test. The values of p < 0.05 were considered statistically significant.
Additional Information
How to cite this article: Xu, S.-N. et al. SIRT2 activates G6PD to enhance NADPH production and promote leukemia cell proliferation. Sci. Rep. 6, 32734; doi: 10.1038/srep32734 (2016).
References
Ward, P. S. & Thompson, C. B. Metabolic reprogramming: a cancer hallmark even warburg did not anticipate. Cancer Cell 21, 297–308, doi: 10.1016/j.ccr.2012.02.014 (2012).
Singh, S. R., Tan, M. & Rameshwar, P. Cancer metabolism: targeting metabolic pathways in cancer therapy. Cancer Lett. 356, 147–148, doi: 10.1016/j.canlet.2014.06.002 (2015).
Yang, H. B. et al. Acetylation of MAT IIalpha represses tumour cell growth and is decreased in human hepatocellular cancer. Nat. Commun. 6, 6973, doi: 10.1038/ncomms7973 (2015).
Estey, E. & Dohner, H. Acute myeloid leukaemia. Lancet 368, 1894–1907, doi: 10.1016/S0140-6736(06)69780-8 (2006).
Wouters, B. J. & Delwel, R. Epigenetics and approaches to targeted epigenetic therapy in acute myeloid leukemia. Blood 127, 42–52, doi: 10.1182/blood-2015-07-604512 (2016).
Akers, L. J. et al. Targeting glycolysis in leukemia: a novel inhibitor 3-BrOP in combination with rapamycin. Leuk. Res. 35, 814–820, doi: 10.1016/j.leukres.2010.12.028 (2011).
Hulleman, E. et al. Inhibition of glycolysis modulates prednisolone resistance in acute lymphoblastic leukemia cells. Blood 113, 2014–2021, doi: 10.1182/blood-2008-05-157842 (2009).
Gottschalk, S., Anderson, N., Hainz, C., Eckhardt, S. G. & Serkova, N. J. Imatinib (STI571)-mediated changes in glucose metabolism in human leukemia BCR-ABL- positive cells. Clin. Cancer Res. 10, 6661–6668, doi: 10.1158/1078-0432.CCR-04-0039 (2004).
Karnauskas, R. et al. Bcl-x(L) and Akt cooperate to promote leukemogenesis in vivo. Oncogene 22, 688–698, doi: 10.1038/sj.onc.1206159 (2003).
Saito, Y., Chapple, R. H., Lin, A., Kitano, A. & Nakada, D. AMPK Protects Leukemia-Initiating Cells in Myeloid Leukemias from Metabolic Stress in the Bone Marrow. Cell Stem Cell 17, 585–596, doi: 10.1016/j.stem.2015.08.019 (2015).
Meynet, O. et al. Glycolysis inhibition targets Mcl-1 to restore sensitivity of lymphoma cells to ABT-737-induced apoptosis. Leukemia 26, 1145–1147, doi: 10.1038/leu.2011.327 (2012).
Chen, Y. et al. Inhibition of pentose phosphate pathway suppresses acute myelogenous leukemia. Tumour Biol. 37, 6027–6034, doi: 10.1007/s13277-015-4428-5 (2016).
Lowman, X. H. et al. The proapoptotic function of Noxa in human leukemia cells is regulated by the kinase Cdk5 and by glucose. Mol. Cell 40, 823–833, doi: 10.1016/j.molcel.2010.11.035 (2010).
Yamamoto, T. et al. Reduced methylation of PFKFB3 in cancer cells shunts glucose towards the pentose phosphate pathway. Nat. Commun. 5, 3480, doi: 10.1038/ncomms4480 (2014).
Shan, C. et al. Lysine acetylation activates 6-phosphogluconate dehydrogenase to promote tumor growth. Mol. Cell 55, 552–565, doi: 10.1016/j.molcel.2014.06.020 (2014).
Kuehne, A. et al. Acute Activation of Oxidative Pentose Phosphate Pathway as First-Line Response to Oxidative Stress in Human Skin Cells. Mol. Cell 59, 359–371, doi: 10.1016/j.molcel.2015.06.017 (2015).
Jiang, P. et al. p53 regulates biosynthesis through direct inactivation of glucose-6-phosphate dehydrogenase. Nat. Cell Biol. 13, 310–316, doi: 10.1038/ncb2172 (2011).
Jiang, P., Du, W. & Wu, M. Regulation of the pentose phosphate pathway in cancer. Protein Cell 5, 592–602, doi: 10.1007/s13238-014-0082-8 (2014).
Patra, K. C. & Hay, N. The pentose phosphate pathway and cancer. Trends Biochem. Sci. 39, 347–354, doi: 10.1016/j.tibs.2014.06.005 (2014).
Riganti, C., Gazzano, E., Polimeni, M., Aldieri, E. & Ghigo, D. The pentose phosphate pathway: an antioxidant defense and a crossroad in tumor cell fate. Free Radic. Biol. Med. 53, 421–436, doi: 10.1016/j.freeradbiomed.2012.05.006 (2012).
Du, W. et al. TAp73 enhances the pentose phosphate pathway and supports cell proliferation. Nat. Cell Biol. 15, 991–1000, doi: 10.1038/ncb2789 (2013).
Cappellini, M. D. & Fiorelli, G. Glucose-6-phosphate dehydrogenase deficiency. Lancet 371, 64–74, doi: 10.1016/S0140-6736(08)60073-2 (2008).
Leopold, J. A. et al. Aldosterone impairs vascular reactivity by decreasing glucose-6-phosphate dehydrogenase activity. Nat. Med. 13, 189–197, doi: nm1545 (2007).
Wang, Y. P. et al. Regulation of G6PD acetylation by SIRT2 and KAT9 modulates NADPH homeostasis and cell survival during oxidative stress. EMBO J. 33, 1304–1320, doi: 10.1002/embj.201387224 (2014).
Furuta, E., Okuda, H., Kobayashi, A. & Watabe, K. Metabolic genes in cancer: their roles in tumor progression and clinical implications. Biochim. Biophys. Acta. 1805, 141–152, doi: 10.1016/j.bbcan.2010.01.005 (2010).
Batetta, B. et al. G6PD activity and gene expression in leukemic cells from G6PD-deficient subjects. Cancer Lett. 140, 53–58 (1999).
Bou Samra, E. et al. New prognostic markers, determined using gene expression analyses, reveal two distinct subtypes of chronic myelomonocytic leukaemia patients. Br. J. Haematol. 157, 347–356, doi: 10.1111/j.1365-2141.2012.09069.x (2012).
Cancer Genome Atlas Research Network. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N. Engl.J. Med. 368, 2059–2074, doi: 10.1056/NEJMoa1301689 (2013).
Tian, W. N. et al. Importance of glucose-6-phosphate dehydrogenase activity for cell growth. J. Biol. Chem. 273, 10609–10617 (1998).
Di Monaco, M. et al. Role of glucose-6-phosphate dehydrogenase inhibition in the antiproliferative effects of dehydroepiandrosterone on human breast cancer cells. Br. J. Cancer 75, 589–592 (1997).
Hecker, P. A., Leopold, J. A., Gupte, S. A., Recchia, F. A. & Stanley, W. C. Impact of glucose-6-phosphate dehydrogenase deficiency on the pathophysiology of cardiovascular disease. Am. J. Physiol. Heart Circ. Physiol. 304, H491–H500, doi: 10.1152/ajpheart.00721.2012 (2013).
Hong, X. et al. PTEN antagonises Tcl1/hnRNPK-mediated G6PD pre-mRNA splicing which contributes to hepatocarcinogenesis. Gut 63, 1635–1647, doi: 10.1136/gutjnl-2013-305302 (2014).
Cosentino, C., Grieco, D. & Costanzo, V. ATM activates the pentose phosphate pathway promoting anti-oxidant defence and DNA repair. EMBO J. 30, 546–555, doi: 10.1038/emboj.2010.330 (2011).
Rao, X. et al. O-GlcNAcylation of G6PD promotes the pentose phosphate pathway and tumor growth. Nat. Commun. 6, 8468, doi: 10.1038/ncomms9468 (2015).
Xu, Y., Osborne, B. W. & Stanton, R. C. Diabetes causes inhibition of glucose-6-phosphate dehydrogenase via activation of PKA, which contributes to oxidative stress in rat kidney cortex. Am. J. Physiol. enal Physiol. 289, F1040–F1047, doi: 00076.2005 (2005).
Zhang, Z., Apse, K., Pang, J. & Stanton, R. C. High glucose inhibits glucose-6-phosphate dehydrogenase via cAMP in aortic endothelial cells. J. Biol. Chem. 275, 40042–40047, doi: 10.1074/jbc.M007505200 (2000).
Dong, L. H. et al. TRAF6-mediated SM22alpha K21 ubiquitination promotes G6PD activation and NADPH production, contributing to GSH homeostasis and VSMC survival in vitro and in vivo. Circ. Res. 117, 684–694, doi: 10.1161/CIRCRESAHA.115.306233 (2015).
Wu, L. E. & Sinclair, D. A. SIRT2 controls the pentose phosphate switch. EMBO J. 33, 1287–1288, doi: 10.15252/embj.201488713 (2014).
Gomes, P., Outeiro, T. F. & Cavadas, C. Emerging Role of Sirtuin 2 in the Regulation of Mammalian Metabolism. Trends Pharmacol. Sci. 36, 756–768, doi: 10.1016/j.tips.2015.08.001 (2015).
Kleszcz, R., Paluszczak, J. & Baer-Dubowska, W. Targeting aberrant cancer metabolism - The role of sirtuins. Pharmacol. Rep. 67, 1068–1080, doi: 10.1016/j.pharep.2015.03.021 (2015).
Chalkiadaki, A. & Guarente, L. The multifaceted functions of sirtuins in cancer. Nat. Rev. Cancer 15, 608–624, doi: 10.1038/nrc3985 (2015).
Rumpf, T. et al. Selective Sirt2 inhibition by ligand-induced rearrangement of the active site. Nat. Commun. 6, 6263, doi: 10.1038/ncomms7263 (2015).
Jing, H. et al. A SIRT2-selective inhibitor promotes c-Myc oncoprotein degradation and exhibits broad anticancer activity. Cancer Cell 29, 297–310, doi: 10.1016/j.ccell.2016.02.007 (2016).
Gao, X. et al. Acetate functions as an epigenetic metabolite to promote lipid synthesis under hypoxia. Nat. Commun. 7, 11960, doi: 10.1038/ncomms11960 (2016).
Pallasch, C. P. et al. Targeting lipid metabolism by the lipoprotein lipase inhibitor orlistat results in apoptosis of B-cell chronic lymphocytic leukemia cells. Leukemia 22, 585–592, doi: 2405058 (2008).
Southam, A. D. et al. Drug redeployment to kill leukemia and lymphoma cells by disrupting SCD1-mediated synthesis of monounsaturated fatty acids. Cancer Res. 75, 2530–2540, doi: 10.1158/0008-5472.CAN-15-0202 (2015).
Polimeni, M. et al. Modulation of doxorubicin resistance by the glucose-6-phosphate dehydrogenase activity. Biochem J. 439, 141–149, doi: 10.1042/BJ20102016 (2011).
Zhou, X. et al. Targeted polyubiquitylation of RASSF1C by the Mule and SCFbeta-TrCP ligases in response to DNA damage. Biochem. J. 441, 227–236, doi: 10.1042/BJ20111500 (2012).
Lei, Q. et al. NKX3.1 stabilizes p53, inhibits AKT activation, and blocks prostate cancer initiation caused by PTEN loss. Cancer Cell 9, 367–378, doi: S1535-6108(06)00118-8 (2006).
Lv, L. et al. Acetylation targets the M2 isoform of pyruvate kinase for degradation through chaperone-mediated autophagy and promotes tumor growth. Mol. Cell 42, 719–730, doi: 10.1016/j.molcel.2011.04.025 (2011).
Wang, Y. et al. WT1 recruits TET2 to regulate its target gene expression and suppress leukemia cell proliferation. Mol. Cell 57, 662–673, doi: 10.1016/j.molcel.2014.12.023 (2015).
Acknowledgements
This work was supported by Natural Science Foundation of China (NSFC) grant (No. 81502379 to Y.-P.W.), the Youth Cultivation Project of Medical Science and Technology of Chinese PLA (No. 14QNP089 to S.-N.X.), and “Chenguang Program” of Shanghai Education Development Foundation and Shanghai Municipal Education Commission (No. 14CG15 to Y.-P.W.)
Author information
Authors and Affiliations
Contributions
Y.-P.W. conceived the general framework of this study. T.-S.W. and Y.-P.W. designed experiments. S.-N.X., X.L. and T.-S.W. performed experiments. S.-N.X. and X.L. collected human samples. S.-N.X., T.-S.W. and Y.-P.W. prepared the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Xu, SN., Wang, TS., Li, X. et al. SIRT2 activates G6PD to enhance NADPH production and promote leukaemia cell proliferation. Sci Rep 6, 32734 (2016). https://doi.org/10.1038/srep32734
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep32734
This article is cited by
-
FBXO31 is upregulated by METTL3 to promote pancreatic cancer progression via regulating SIRT2 ubiquitination and degradation
Cell Death & Disease (2024)
-
Variation in pentose phosphate pathway-associated metabolism dictates cytotoxicity outcomes determined by tetrazolium reduction assays
Scientific Reports (2023)
-
NADPH homeostasis in cancer: functions, mechanisms and therapeutic implications
Signal Transduction and Targeted Therapy (2020)
-
IDH-1 deficiency induces growth defects and metabolic alterations in GSPD-1-deficient Caenorhabditis elegans
Journal of Molecular Medicine (2019)
-
Metabolic recoding of epigenetics in cancer
Cancer Communications (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.