Abstract
Plant and cyanobacteria can perceive signals from soluble sugar and reactive oxygen species (ROS) and then coordinate gene expression under stress acclimation, but the underlying mechanism remains unclear. In this study, we found that the transcriptional factor PrqR (Slr0895) in Synechocystis can perceive signals from ROS generated after shifting from prolonged darkness with glucose into high-light. The deletion mutant (DprqR) showed increased growth rate and decreased ROS content, whereas the complementary strain (CprqR) restored the growth characteristics, phenotypes and ROS status of WT, thereby establishing PrqR as a negative regulator of ROS.LC/GC-MS-based metabolic profiling also showed active ROS mitigation in DprqR mutant. Further study by qRT-PCR, ChIP-PCR and deletion of both prqR and prqA (DprqR-DprqA mutant) revealed that PrqR exerts this negative regulation of ROS removal by controlling the expression of sodB and prqA (slr0896). Furthermore, PrqR also found to control glucose metabolism by regulating a positive regulator of glucose metabolism, sigE, and its regulons. Results suggest that PrqR was involved in perceiving signals from ROS under physiological condition, as well as in regulating stress removal and glucose metabolism.
Similar content being viewed by others
Introduction
Aquatic environments with regular light dark cycle are typical for cyanobacteria. However, cyanobacteria are also living under adverse ecological conditions with prolonged darkness; for example, cyanobacteria can be found self-shadowing in dense planktonic and benthic communities, lake sedimentation, and soil water or dense aquatic accumulation produced at surface1,2. Thus, cyanobacteria could be a good model to understand metabolism and signaling network in adverse condition. In addition, as an evolutionary ancestor of plant photosynthetic machineries, cyanobacteria can also be a model platform to understand metabolism under similar prolonged dark conditions, which occurs in non-green organs of plants, such as roots, stems, and flowers3. Cyanobacteria respond to adverse conditions by generating reactive oxygen species (ROS) that can modulate transcription of genes in a genetic network and connect an appropriate riposte to the conditions4,5. But, how do cyanobacteria and plants perceive signals from ROS, coordinate signals from different sources, and reprogram metabolism under adverse physiological conditions is largely unknown.
In photosynthetic organisms, glucose can cause an accelerated ROS production6,7,8 or mitigate by generating NADPH by using the OPP pathway6,9. The addition of excess sugar can force plant cells into auto-heterotrophic condition, thereby suppressing photosynthesis and the Calvin cycle6. Under such conditions, NADPH production diminishes, eventually favoring ROS production6,10,11. To avoid this situation, signals exerted by light, sugar, and/or oxidative stress must be connected to balance sugar metabolism and ROS generation, as well as photosynthetic activity12,13. Signaling network of this kind has been started to be revealed. In plants, stress signals have been found interacting with glucose signaling through the hexokinase independent pathway6. For instance, the psbA gene, which encodes the D1 protein of photosystem II, converges signals from light, ROS and sugar6. By contrast, cytosolic glucose-6-phosphate dehydrogenase (G6PDH) of the OPP pathway mediates signals from both light and sugar14. Moreover, genes involved in ROS defense in cyanobacteria, such as chalcone synthase (CHS), glutathione-S-transferase (GST), and biosynthetic genes of ascorbate and carotenoid are also responding signals from sugar and light6,15,16,17,18. For example, genes of the carotenoid biosynthetic pathway, such as phytoene synthase (crtB), phytoene desaturase (crtP), ζ-carotene desaturase (crtQ), and β-carotene hydroxylase (crtR), were upregulated under high light, which is the ROS generating condition, but downregulated in the dark19. The expression of these genes can also be restored in the dark by adding glucose suggesting that the genes can perceive signal from ROS and glucose20. Similarly, sigE, a type 2 sigma factor with its positive role in glucose metabolism21,22, can respond to ROS23. However, previous studies are lacked to find appropriate experimental conditions that can provide platform to understand the genetic network in ROS mitigation and its relation with glucose metabolism under stress condition.
The model cyanobacterium, Synechocystis sp. PCC 6803 (hereafter Synechocystis) contains transcriptional factor that sense signals of stress from the environment and transduce the signals to coordinate the gene expression accordingly24. In Synechocystis, the mutant (prq20) of the transcriptional factor PrqR was previously shown to be adaptive resistance to an externally added methyl viologen (Paraquat)25. Additionally, the photosynthetic apparatus was found less affected in the prq20 mutant than in the wild type (WT), as indicated by the damage generated by two strong-oxidative-stress producing compounds, namely, methyl viologen and benzyl viologen26. PrqR was found to contain a cis-acting regulatory sequence for auto-repression, and the frameshift mutation C134fs in the C-terminus DNA binding domain of PrqR inhibited this autoregulation25. Nevertheless, no information is available on the putative PrqR regulatory network under stress generating conditions.
With an initial goal to discover a response regulator that coordinate signals from both glucose and ROS, we established a unique nutritional condition, that is, GLC(+)DARK-LIGHT. In this condition, Synechocystis cultures were grown autotrophically (under illumination of 25 μmol m−2 s−2) until they reach exponential phase (OD730 ~ 0.40–0.45) in BG-11 media, with the addition of 5 mM glucose, samples were placed in the darkness for 72 h for GLC(+)DARK growth and then shifted back to grow under high light (80 μmol m−2 s−2) [GLC(+)DARK-LIGHT]. The results showed that oxidative stress was generated under GLC(+)DARK-LIGHT condition, and PrqR can be involved in perceiving ROS signals by repressing its downstream prqA gene, which encodes a putative multidrug-resistance protein. Furthermore, PrqR negatively regulated glucose metabolism by controlling the regulator of glucose metabolism. Overall, this study revealed that prqR plays a negative role in both glucose metabolism and ROS mitigation in Synechocystis under stress conditions.
Results
Phenotypic analysis of Synechocystis wild type and prqR mutants
A previous study found that the Synechocystis mutant Prq20 (L17Q in prqR transcription factor) was resistant to methyl viologen, a strong ROS-generating chemical26,27. In the current study, to further determine the roles of the prqR gene in ROS removal and glucose metabolism at the molecular level, a prqR-disrupted mutant (DprqR) was generated by inserting a kanamycin cassette under cpcB promoter into the Synechocystis (glucose-intolerant strain) chromosome. For complementation, a strain with prqR gene overexpressed under a psbA2 promoter along with the streptomycin cassette was also created in the DprqR mutant (CprqR) (the vectors and strains generated in this study are presented in Supplementary Fig. S1A). DprqR and CprqR were completely segregated on BG-11 plate supplemented with kanamycin and kanamycin + streptomycin (up to 100 μg/mL), respectively. The segregation was confirmed by PCR (Supplementary Figs S1B and S1C). WT, DprqR and CprqR showed normal growth in BG-11 liquid media in the presence of light at 25 μmol m−2 s−2, with shaking at 130 rpm and temperature of 30 °C (Fig. 1A). To assess glucose-dependent growth, we added 5 mM D-glucose into the culture when its OD730 was around 0.4 and then placed the culture into complete darkness [GLC(+)DARK] with shaking for different durations. Through the growth courses, the DprqR mutant grew almost similarly to WT and CprqR under GLC(+)DARK ; with subsequent placement of the cultures into high light, GLC(+)DARK-LIGHT(at 80 μmol m−2 s−2) led to approximately twofold increased growth of the DprqR mutant, whereas WT and CprqR showed significantly decreased growth as well as decolorization (Fig. 1B–D). To determine whether the phenotype attributed to the osmotic stress exerted by D-glucose, we repeated the experiments with D-sorbitol, which can produce an osmotic pressure similar to that D-glucose. However, all these strains grew equally normal (Supplementary Fig. S2), suggesting that the phenotype was not ascribed to osmotic stress. To determine whether the cells are dying under GLC(+)DARK and GLC(+)DARK-LIGHT conditions, we monitored the percentages of dead cells at two time points: at third day under GLC(+)DARK (72 h) condition and first day (24 h) under GLC(+)DARK-LIGHT condition. However, the results showed no significant changes in terms of cell death rates between WT and DprqR (Supplementary Fig. S3). Notably, the growth and chlorophyll content were found remarkably different in between DprqR and WT (and CprqR) after shifting from GLC(+)DARK to the GLC(+)DARK-LIGHT conditions (Fig. 1D,E).
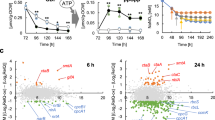
Culture of WT, DprqR and CprqR under normal (A); 72 h under GLC(+)DARK (B); and 48 h under GLC(+)DARK-LIGHT (C) conditions; (D), growth characteristics of WT, DprqR and CprqR under normal, GLC(+)DARK, and GLC(+)DARK-LIGHT cultural conditions in different durations; (E), Chlorophyll content of WT, DprqR and CprqR under normal, GLC(+)DARK, and GLC(+)DARK-LIGHT cultural conditions in different durations; total ROS content of WT, DprqR and CprqR corresponding at normal growth condition before adding glucose (F); 72 h under GLC(+)DARK (G); and 48 h under GLC(+)DARK-LIGHT (H) conditions. In (D,E), the number represent the incubation time in hours and N = 5 ± SEM. In (F–H), N = 5 ± SEM and “*” represents statistical significance as indicated by Student’s t-test with maximum p-value of P < 0.05.
ROS decreased in DprqR mutant
Assuming that a sudden change of metabolism from glucose-dependent [GLC(+)DARK] condition to both glucose- and light-dependent [GLC(+)DARK-LIGHT] conditions would generate ROS, we determined the total ROS contents at three time points: before placement under GLC(+)DARK condition (normal cells), after 72 h under GLC(+)DARK and after 48 h in GLC(+)DARK-LIGHT condition. The total ROS contents were measured using membrane-permeant fluorescence indicator 5-(and-6)-chloromethyl-2′, 7′-dichlorodihydro fluorescein diacetate, CM-H2DCFDA (Invitrogen, Life technology). The results showed that under normal and GLC(+)DARK conditions, the ROS contents of WT, DprqR and CprqR were at similar levels (Fig. 1F,G), although a slightly higher accumulation of ROS in CprqR under normal condition was found and may be ascribed to the stronger psbA2 promoter. However, significantly (more than twofold) higher ROS contents in WT and CprqR were found than those in DprqR under GLC(+)DARK-LIGHT condition after 48 h (Fig. 1H). In addition, no significant ROS accumulation difference was detected in WT, CprqR and DprqR in similar dark-light regime in the absence of glucose, that is, GLC(−)DARK and GLC(−)DARK-LIGHT (Supplementary Fig. S4). This finding suggests that GLC(+)DARK-LIGHT can cause ROS accumulation in Synechocystis cells, whereas the deletion of the prqR gene affected ROS mitigation.
Cyanobacteria exhibit diverse defense systems against oxidative stress, such as superoxide dismutase, which can remove superoxide radicals4. Synechocystis encompasses only one type of dismutase, that is, iron superoxide dismutase (FeSOD). Considering the role of FeSOD in ROS removal, we determined the expression order of sodB by qRT-PCR. The results showed that the sodB expression was upregulated in DprqR compared with WT at 24 and 48 h but equalized at 72 h under GLC(+)DARK condition. Subsequent shifting of the culture under GLC(+)DARK-LIGHT condition caused about 100-fold increase of sodB in DprqR strain compared with WT (Fig. 2A). To ensure that this fluctuation of the sodB expression in DprqR was ascribed to glucose rather than only high light, we maintained the GLC(−)DARK and GLC(−)DARK-LIGHT conditions but no significant upregulation or downregulation was observed (Fig. 2B).
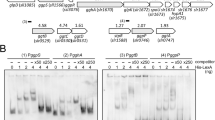
Expression of sodB at different time points of GLC(+)DARK and GLC(+)DARK-LIGHT (A); sodB expression in the absence of glucose, GLC(−)DARK (after 72 h) and GLC(−)DARK-LIGHT (after 24 h) (B,C), expression of prqA at GLC(+)DARK (after 72 h) and GLC(+)DARK-LIGHT (24 h); (D), expression of prqA in the absence of glucose, GLC(−)DARK (after 72 h) and GLC(−)DARK-LIGHT (after 24 h). In all the cases from (A–D), N = 5 ± SEM, and p-value P < 0.05, and were normalized by rnpB expression level. In (E), the designed primer sets (listed in Supplementary table S4) surrounded the predicted binding site of prqA (ATTTGTTTTTTCATA) promoter. ATG is the start codon of prqA; the bold “T” is the start site of transcription. The prqR-His was expressed in OEprqR strain. After ChIP, samples were assessed by quantitative PCR, and the relative enrichment was calculated against 1% input and IgG (F). For clarity, normal PCR with those primer sets were performed with 25 cycles (G). The picture in (G) was cropped from the gel run under similar condition in electrophoresis (400 V, 125 amps) with the sample from 25 cycle of PCR. The experiment was repeated twice and found similar results. The deletion of prqR and prqA from WT (DprqR-DprqA) restored the phenotype of WT and CprqR (H). The culture was taken in after 48 h under GLC(+)DARK-LIGHT condition.
PrqR binds to the promoter of prqA and negatively regulates the expression of prqA
In the Synechocystis chromosome, prqA located downstream of prqR encodes a hypothetical protein belonging to the multidrug resistance protein of the MATE family. Analysis of gene expression showed that prqA was upregulated in DprqR by threefold and fourfold than the WT under GLC(+)DARK and GLC(+)DARK-LIGHT conditions, respectively (Fig. 2C). However, these differential expressions was not obtained in the absence of glucose (Fig. 2D). To determine whether prqA was the regulatory target of PrqR, we overexpressed prqR with a His-tag at its C-terminus under cpcB promoter in the WT. The expression of prqR-6xHis was confirmed by Western blot (Supplementary Fig. S1E). Chromatin Immunoprecipitation (ChIP) was conducted in accordance with the manufacturer’s manual (Beyotime, China). Before primers were designed for PCR, the 1000 bp upstream of the start codon of prqA was subjected to analysis with BDGP online software and BPROM28 for the transcription start site and transcription factor binding sites (Fig. 2E). Primers were then designed spanning the predicted binding site for an amplicon of 150 bp to 200 bp. After ChIP, DNA was used for PCR and the relative enrichment of DNA was analyzed against 1% input and IgG. The results showed that the putative prqA promoter was able to enrich approximately 50 and 10 fold of DNA, relative to control IgG, by primers 2 and primer 3, correspondingly (Fig. 2F). Furthermore, semi-quantitative PCR also found that primers 2 and 3 were able to amplify more DNA in the sample with anti-His antibody compared with the IgG control (Fig. 2G). This finding demonstrates that PrqR can bind directly with the promoter of prqA, which is immediately downstream of prqR. Similar analysis was performed on gap2, sigE and sodB genes (data are not shown). However, no significant binding was observed, suggesting that these genes were not regulated directly by PrqR.
We further deleted both prqR and prqA genes, resulting in the DprqR-DprqA mutant (Supplementary Fig. S1D). The DprqR-DprqA mutant exhibited similar growth of WT and DprqR under normal condition but we found significant growth retardation under GLC(+)DARK-LIGHT condition (Fig. 2H), similar to that observed for WT. Thus, prqA may be negatively controlled by PrqR, which can be essential for the survival under high ROS conditions, such as GLC(+)DARK-LIGHT.
Glucose uptake and glycogen content varied in DprqR mutant
The mutant DprqR showed remarkable difference from WT and CprqR in both glucose uptake and glycogen content in the entire course of GLC(+)DARK and subsequent GLC(+)DARK-LIGHT condition. The glucose uptake in the first 24 h was found similar for WT, CprqR and DprqR mutants but declined more in WT and CprqR than in DprqR under GLC(+)DARK condition. After shifting from GLC(+)DARK to GLC(+)DARK-LIGHT both WT and CprqR showed a sharp increase in glucose uptake compared with DprqR (Fig. 3A) and after 48 h, DprqR consumed all the glucose added in the media, whereas WT stopped up-taking despite glucose remained in the media (Supplementary Fig. S5). This result suggests that DprqR continues glucose catabolism under the GLC(+)DARK condition, and more actively under GLC(+)DARK-LIGHT condition, until all the glucose is consumed. Glycogen aggregation was found to follow similar trends in the first 48 h under GLC(+)DARK but increased within 72 h in the WT, CprqR and DprqR mutants (Fig. 3B). The succeeding shift into the GLC(+)DARK-LIGHT condition led to a decreased glycogen content in the DprqR mutant, whereas the WT and CprqR maintained the same amount as those under the GLC(+)DARK condition.
(A) Glucose uptake ability of WT, DprqR and CprqR under GLC(+)DARK and GLC(+)DARK-LIGHT conditions; (B) Glycogen content of WT, DprqR and CprqR under GLC(+)DARK and GLC(+)DARK-LIGHT conditions. The expression levels of three common regulators, hik8 (sll0750), rre37 (sll1330) and hik31 (sll0790), of glucose metabolism under GLC(+)DARK (after 72 h) and GLC(+)DARK-LIGHT (after 24 h) conditions (C,D), represents the expression of sigE in different durations under GLC(+)DARK and GLC(+)DARK-LIGHT conditions, the number represents duration in hours; (E) is the expression of sigE in the absence of glucose, GLC(−)DARK and GLC(−)DARK-LIGHT conditions; (F) the expression level of regulons of sigE, two genes each in the glycolytic pathway (pfkB1 and gap1) and the OPP pathway (slr1843 and gnd) at GLC(+)DARK (after 72 h) and GLC(+)DARK-LIGHT (after 24 h) conditions; (G) gap2 expression in different durations under GLC(+)DARK and GLC(+)DARK-LIGHT conditions; and (H) expression of gap2 in the absence of glucose, GLC(−)DARK and GLC(−)DARK-LIGHT conditions. In (A,B), N = 3 ± SEM with three technical replications. “*” represents statistical significance, as shown by Student’s t-test with maximum p-value of < 0.05. From (C–H), N = 5 ± SEM, with maximum p-value < 0.05.
Dynamic expression of glucose metabolism regulators in DprqR mutant under GLC(+)DARK and GLC(+)DARK-LIGHT conditions
D-sorbitol did not affect cell growth under the D-SORBITOL(+)DARK-LIGHT condition (Supplementary Fig. S2). The possibility of osmotic stress was excluded. In addition, we washed the WT cells after 24 h under GLC(+)DARK-LIGHT. Afterward, we placed the cells into fresh media without glucose and found the cells resumed growth normally. These results led us to investigate the differential glucose metabolism abnormalities between WT and DprqR. Hence, we first targeted the following known regulators of the glucose metabolic pathway in Synechocystis through real-time qRT-PCR: a two-response regulatory protein Rre37 (sll1330) with HTH domain, which was previously found to control the expression of genes in glucose metabolic pathways in the presence of light and glucose29; Hik31 (sll0790) with regulatory role in glucose metabolism under heterotrophic growth30; a histidine kinase Hik8 (sll0750) involved in circadian rhythm and a regulator of glucose metabolism31; and a group 2 sigma factor SigE, which regulates the expression of several genes in glucose metabolic pathways under normal growth condition21,22. The results showed no significant difference among the expression of hik8, hik31 and rre37 between WT and DprqR under GLC(+)DARK condition (Fig. 3C). Under GLC(+)DARK-LIGHT condition, hik8 and hik31 exhibited a similar expression pattern in WT and DprqR, whereas rre37 displayed a relatively higher expression in DprqR than that in WT (Fig. 3C); this finding is consistent with a previous report29. Moreover, sigE had been found with differential expression in DprqR compared with WT (Fig. 3D) under GLC(+)DARK (24 and 48 h) and GLC(+)DARK-LIGHT (24 and 48 h) conditions. Interestingly, the expression levels of sigE in WT and DprqR were similar at 72 h under GLC(+)DARK but differed after shifting into GLC(+)DARK-LIGHT (Fig. 3D); this finding suggest that the above time point is crucial for the expression of sigE. To determine whether the change is attributed to dark-to-high light or from glucose-dark [i.e., GLC(+)DARK] to glucose-high light [i.e., GLC(+)DARK-LIGHT], we incubated both WT and DprqR in GLC(−)DARK for 72 h and then shifted into GLC(−)DARK-LIGHT condition. RNA was isolated from the samples after 72 h in GLC(−)DARK and 24 h in GLC(−)DARK-LIGHT and real-time qRT-PCR was performed for analysis. The results showed no significant change of sigE expression in DprqR at the two time points (Fig. 3E); this finding suggests that glucose not high light alone caused the elevated sigE expression in DprqR.
In a previous study, the sigE mutant was found defective in glucose catabolism with decreased expression of phosphofructokinase, pfkB1 (sll1196), glyceraldehyde-3-phosphate dehydrogenase (gap1) and pyruvate kinase (pyk1)21. To investigate role of sigE in terms of regulating genes of glucose metabolism under prqR-deficient background, we investigated the expression of three genes in the glycolytic pathway via real-time qRT-PCR, including two target genes of sigE (i.e., pfkB1 and gap1). Although the results showed no significant change between WT and DprqR under either GLC(+)DARK or GLC(+)DARK-LIGHT conditions for two sigE target genes, except a slight increase of gap1 expression in DprqR under GLC(+)DARK-LIGHT condition (Fig. 3F), the results showed upregulation of gap2, the gene encodes enzyme glyceraldyhyde-3-phosphate dehydrogenase in DprqR compared WT grown under GLC(+)DARK. In addition, we determined the expression pattern of gap2 at different durations of GLC(+)DARK and GLC(+)DARK-LIGHT conditions (Fig. 3G). The results showed that the gap2 expression in DprqR was more than 10 fold higher than that in WT throughout the GLC(+)DARK and GLC(+)DARK-LIGHT conditions. However, when we maintained a similar condition without glucose GLC(−)DARK for 72 h and then shifted to GLC(−)DARK-LIGHT for 24 h, no significant change of gap2 expression between DprqR and WT was found (Fig. 3H). This finding suggests that glucose caused the remarkable change of gap2 expression in D9prqR mutant under GLC(+)DARK and GLC(+)DARK-LIGHT conditions.
The OPP pathway is considered as the main route of carbon catabolism in Synechocystis and both zwf (slr1843 encoded enzyme glucose-6-phosphate-1 dehydrogenase, G6PDH) and gnd (sll0329 encoded enzyme 6-phosphogluconate dehydrogenase, 6PGD) play important roles in controlling the carbon flow regulated by sigE21,22. RT-PCR analysis showed that both zwf and gnd were slightly upregulated in DprqR under GLC(+)DARK condition compared with those in WT; a significant increase of zwf expression in DprqR was also observed under GLC(+)DARK-LIGHT condition (Fig. 3F), and this finding was consistent with the increased glucose and glycogen metabolism immediately after shifting from GLC(+)DARK to GLC(+)DARK-LIGHT.
Metabolite abundance of central metabolic pathways in WT and DprqR mutant under GLC(+)DARK and GLC(+)DARK-LIGHT conditions
Metabolites were analyzed by both LC-MS and GC-MS in accordance with previously established protocols32,33. LC-MS-based analysis detected 23 targeted metabolites, mostly in glucose metabolic pathways, pyrimidine nucleotides, amino acid biosynthesis and energy-containing adenylated phosphate. GC-MS-based non-targeted metabolomics identified around 600 peaks, among which 183 compounds were selected after reproducibility analysis through repeated experiments. The abundance of each metabolite at GLC(+)DARK (72 h in dark) and GLC(+)DARK-LIGHT (24 h) was compared between DprqR and WT, and the significance of the changes between WT and DprqR was statistically evaluated by t-test (p < 0.05) (Supplementary Table S1 and S2). DprqR was significantly different from WT, thereby suggesting different metabolite profiles between GLC(+)DARK and GLC(+)DARK-LIGHT conditions. In addition, PCA analyses based on 23 metabolites detected by LC-MS (Fig. 4A) and 183 metabolites detected by GC-MS (Fig. 4B) showed a distinct separation between WT and DprqR under both GLC(+)DARK and GLC(+)DARK-LIGHT, suggesting that the deletion of prqR reprogrammed cellular metabolism significantly.
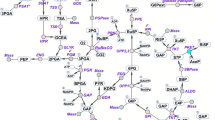
Significant differences between WT and DprqR were observed for metabolites of the glycolytic pathway, OPP pathway, and TCA cycle under GLC(+)DARK condition (Fig. 5A). For the glycolytic pathway, fructose-1,6-bisphosphate (FBP), glyceraldehyde-3-phosphate (GAP), 3-phosphoglycerate (3PG), and phosphoenolpyruvate (PEP), significantly increased in DprqR under GLC(+)DARK condition. Metabolites of gluconeogenesis (ADP-glucose), Calvin cycle [glyceraldehyde-3-phosphate (GAP), ribose-5-phosphate (R5P), and 3-phosphate glycerate (3PG)], TCA cycle (oxaloacetate) and oxidative pentose phosphate pathways [ribulose-5-phosphate (R5P)] were also found upregulated in DprqR compared with those in WT under GLC(+)DARK condition. The abundant changes of individual metabolites aligned in the metabolic pathway showed a sequential increase intermediate of the glycolytic pathway in DprqR, suggesting more glucose catabolism in GLC(+)DARK (Fig. 6A). Upon shifting from GLC(+)DARK to GLC(+)DARK-LIGHT, intermediates of the OPP pathway (R5P) (Figs 5B and 6B), as well as D-gluconic acid, a derivatization degradation product of 6-phosphogluconate, increased by twofold in DprqR (Supplementary Table S1); this finding suggests the active role of the OPP pathway in DprqR under the above mentioned condition. By contrast, the intermediates of the TCA cycle, namely, alpha-ketoglutarate and acetyl-CoA, decreased in DprqR.
Statistical comparison of metabolites detected by LC-MS under GLC(+)DARK (after 72 h) (A) and GLC(+)DARK-LIGHT (B) condition (after 24 h). F6P, fructose-6-phosphate; FBP, fructose-1,6-bisphosphte; DHP, dihydroxyacetone phosphate; GAP, glyceraldehyde-3-phosphate; 3PG, 3-phosphoglycerate; PEP, phosphoenolpyruvate; AcCoA, acetyl-CoA; ADP-GCS, ADP-glucose; R5P, Ribose-5-phosphate; AKG, alpha-ketoglutarate; OXA, oxaloacetate; and Glu, glutamate. The star (*) indicates the level of significance, as shown by Student t-tests. The p-value of < 0.05 is considered significant with four independent replications. The error bar represents ± SEM.
Changes of metabolites involved in oxidative stress mitigation
The metabolomics under oxidative stress in Synechocystis are not so available till now. Given that Cyanobacteria is evolutionary ancestor of plant and they share similar environmental habitat, we compared changed of metabolites in our work with some well-studied metabolic profile of plant under stress. Among many of such studies, Noctor et al.34 recently summarized a list of metabolites detected by GC/LC-MS that is more stably considered as marker to trace ROS status in Arabidopsis and Rice. The role and mechanism of many of these metabolites such as gluthathione, NADPH etc. in ROS acclimation are well-established. We compared the change of metabolites in our work with this information and listed. We found that under GLC(+)DARK condition, 2-monopalmitin, cadaverine, and L-rhamnose increased by more than twofold, whereas sorbitol and trehalose decreased more than twofold in DprqR than in WT (Supplementary Table S2). The most distinct change of metabolites involved in oxidative stress removal was observed under GLC(+)DARK-LIGHT condition. Myo-inositol, a sugar alcohol and signaling compound, decreased by approximately twofold in DprqR under GLC(+)DARK-LIGHT condition. Sugars such as lactose, maltose, L-sorbose and D-ribose were found to accumulate more than 1.5 fold in GLC(+)DARK-LIGHT condition in DprqR than in WT (Supplementary Table S1).
Glutathione (GSH) is considered as the most important defense metabolite against ROS damage in cells. GSH scavenges ROS and oxidize them to form glutathione disulfide (GSSG) by combining two GSH molecules with a sulfide bond. We were able to detect GSH by GC-TOF-MS, and the results showed that GSH was 1.75 fold higher in DprqR than in WT under GLC(+)DARK-LIGHT condition; however, no such difference was detected under GLC(+)DARK condition (Supplementary Table S1), and this finding was consistent with a previous report indicating that under the condition of oxidative stress, GSH declined and the redox state become more oxidized, thereby disrupting the cellular metabolic system35.
LC-MS based metabolite analyses revealed differential abundance of pyridine nucleotide and co-factors. Under GLC(+)DARK condition, ATP, AMP, NAD+, NADH, NADPH and CoA significantly decreased in DprqR mutant compared with those in WT. By contrast, ADP and NADP+ were remained unchanged (Fig. 7A). An almost similar trend was followed under GLC(+)DARK-LIGHT condition, in which ATP, AMP, NAD+, NADH, NADPH, Co-A, ADP and NADP+ decreased in DprqR mutant, compared with those in WT (Fig. 7B). The NADPH: NADP+ ratio decreased by ~42% and ~26% when shifted from GLC(+)DARK to GLC(+)DARK-LIGHT condition for WT and DprqR, respectively (Supplementary Table S5). Although the ratios of ATP:NADPH in WT and DprqR were similar under GLC(+)DARK condition, the ratio was relatively higher (1.90 unit) in the DprqR mutant than in WT under GLC(+)DARK-LIGHT condition (1.56 unit). This finding was consistent with a previous report indicating that changes in the ATP:NADPH ratio can lead to reponse in ROS generation4.
Series (A) represents the relative abundance of pyridine nucleotide detected by LC-MS under GLC(+)DARK (after 72 h) and (B) under GLC(+)DARK-LIGHT condition (24 h). The data were developed by t-test of means from four independent replications. The star (*) indicates the level of significance by Student t-tests. The p-value < 0.05 is considered significant with four independent replications. The error bar represents ± SEM.
Discussion
Studies have progressively elucidated that soluble sugars, such as glucose and sucrose, are strongly related to stress-induced ROS signaling in plants, cyanobacteria, Saccharomyces, and even mammals7,9,36,37. Unlike the known pro-oxidants and antioxidants which are only involved in ROS removal, soluble sugars such as glucose play a role in both ROS production and ROS-induced signal for scavenging. To clarify the mechanisms controlling ROS signaling pathways under environmental stress and their interplay with glucose metabolism, the focus should be shifted from studying signals under single stress to a combination of abiotic stresses in the presence of glucose to accurately represent the natural habitat. In contrast to previous studies, that focused on glucose metabolism under different trophic conditions, as well as cellular regulation of glucose uptake and metabolism21,22,38,39,40, the present study focused on GLC(+)DARK-LIGHT condition, which generates ROS (Fig. 1C,H), and provided a platform to investigate the complicated interaction of glucose and ROS as well as the signaling network.
Given that PrqR is a negative regulator of ROS26, the knockout mutant of prqR (DprqR mutant) differed from WT and CprqR for its lower ROS content was found with an increased glucose metabolism under GLC(+)DARK-LIGHT condition (Figs 1H and 3A,B, Supplementary Fig. S5); these findings reflect the ROS removal capacity is functional in DprqR under above-mentioned condition which can be related to glucose metabolism. In this study, we will discuss first how PrqR plays a role in ROS mitigation followed by glucose metabolism. PCA analysis of metabolites identified by LC-MS and GC-MS (Fig. 4A,B) showed reprogramming of metabolism in DprqR, whereas metabolic profiling revealed several key marker metabolite changes, which further indicated an increased ROS removal in DprqR. First, DprqR showed superiority of breaking down glucose following the glycolytic pathway under GLC(+)DARK condition (Figs 5A and 6A). This finding is in line with the ability of DprqR to enhance glucose uptake under GLC(+)DARK condition (Fig. 3A). In addition, major changes in shifting the glucose metabolism from the glycolytic pathway into the OPP pathway were observed under GLC(+)DARK-LIGHT condition (Fig. 6A,B). This switch is supposed to increase cellular NADPH given that the OPP pathway breaks down glucose to produce NADPH. On the contrary, we found decreased NADPH in DprqR cells under GLC(+)DARK-LIGHT condition (Fig. 7B). This finding explained that NADPH was used to reduce ROS-removing metabolites such as GSH. This NADPH-demand basis acceleration of the OPP pathway was also observed in photoheterotrophic condition in which NADPH production from photosynthesis was blocked and solely dependent on the OPP pathway41. Moreover, this re-routing of the central carbon metabolism, in favor of the OPP pathway to produce more NADPH, was previously reported in rice and yeast under oxidative stress removing cellular status42,43. Takahashi et al. found increased accumulation of glycolytic pathway intermediates such as fructose-6-phosphate, 3-phosphoglycerate, 2-phosphoglycerate, and phosphoenolpyruvate in mixotrophic condition in Synechocystis, similar to GLC(+)DARK-LIGHT in current study, except the prior extension of GLC(+)DARK incubation38. Overall, this study38 suggested an active mode of glycolytic pathway under mixotrophic condition which is sharply contrasting with our findings obtained under GLC(+)DARK-LIGHT condition. Thus, this particular information agrees with the understanding that glucose and ROS reprogrammed the central metabolic pathway favoring ROS removal in DprqR under GLC(+)DARK-LIGHT condition. Second, the reduced GSH increased in DprqR under GLC(+)DARK-LIGHT condition (Supplementary Table S1). This observation is similar to previous report indicating that the response of GSH to ROS acclimation is a time-series event and a successful acclimation accumulate reduced GSH with unsuccessful acclimation increased GSSG/GSH ratio35. Third, myo-inositol, a sugar alcohol which is a strong scavenger of ROS, was found with a sharply decreased concentration in DprqR mutant (Supplementary Table S1); this finding was consistent with a previous study in a Arabidopsis cat2 mutant, in which the ROS scavenger metabolites decreased in concentration during stress acclimation44,45. Finally, the results showed that the TCA cycle was perturbed in cells under GLC(+)DARK-LIGHT condition (Fig. 6B), which is within the expectation as a number of enzymes, including aconitase and 2-oxogluterate, are known to be affected during acclimation of oxidative stress34,46. Overall, metabolomics analysis demonstrated that the decreased ROS content and active ROS removal mechanism in DprqR mutant cell under GLC(+)DARK-LIGHT condition confirmed the negative regulation of ROS by PrqR (Fig. 8).
PrqR negatively regulates prqA directly, and sodB indirectly; the two genes which expression is crucial for ROS removal from cells under GLC(+)DARK-LIGHT condition. The elevated oxidative stress (ROS) may have been suppressing glucose catabolism and downregulating sigE and gap2 expression which together caused to develop the phenotype in WT and CprqR. The deleted prqR resulted upregulation of prqA as well as sodB, which helped to remove ROS. Thus, the cells with less ROS favored glucose metabolism through sigE and gap2 upregulation, which eventually helped DprqR to grow faster in GLC(+)DARK-LIGHT condition. The “solid black” arrow represents the direct relation in contrast to the “interrupted” arrow is not. Overall, the prqR acts as a regulator and control oxidative stress and glucose metabolism in GLC(+)DARK-LIGHT condition.
These properties of the DprqR mutant led to the assumption that PrqR functions as a regulatory protein for ROS removal. Therefore, a search for possible targets of PrqR ended up establishing PrqR as a negative regulator of prqA and sodB; qRT-PCR showed an upsurge of the prqA and sodB expression in DprqR under both GLC(+)DARK and GLC(+)DARK-LIGHT conditions (Fig. 2A,C). Although very limited information is available about sodB regulation by sugars in cyanobacteria, the participation of glucose in antioxidant induction has been reported20. Studies suggested that sugar-regulated genes involved in anti-oxidative systems are under control of sugar sensors14, leading to a hypothesis that both sodB and prqA may be regulated directly by PrqR. In contrast to this initial expectation, the ChIP results showed that PrqR bound only to the prqA but not sodB promoter (Fig. 2F,G); thus the result suggests that PrqR regulates prqA directly and sodB indirectly. PrqA is a putative multidrug resistance efflux protein previously shown to respond to methyl viologen along with PrqR47. Phenotype of DprqR-DprqA mutant under GLC(+)DARK-LIGHT condition (Fig. 2H), which was similar to WT, suggesting that prqA expression in DprqR mutant was crucial in ROS mitigation. However, the exact mechanism of how does prqA expression mitigate ROS is yet to be known. Altogether, the lower ROS content and elevated expression of ROS removing genes in DprqR implies that PrqR is a negative regulator of ROS under GLC(+)DARK-LIGHT condition (Fig. 8).
Instead of feeding the OPP pathway by glucose and replenishing NADPH that helps to scavenge ROS in plants under stress48,49, glucose was observed to generate ROS and impaired its metabolism under GLC(+)DARK-LIGHT condition in WT and CprqR (Fig. 1C,H). The enhanced glucose metabolism is understandable with overexpression of sigE, which is the positive regulator of glucose metabolism in Synechocystis21 (Fig. 3D), and its regulon in OPP pathway (i.e., zwf and gnd) (Fig. 3F)21,50. However, Chip-PCR measurement confirmed that PrqR does not regulate sigE directly (data not shown). Thus, an alternative explanation of glucose metabolism regulation in DprqR mutant through sigE expression under GLC(+)DARK-LIGHT condition implies that DprqR mutant mitigated ROS under GLC(+)DARK-LIGHT condition, thereby avoiding a negative consequence of ROS on sigE allowing sigE expression in the presence of glucose. Similarly, our result indicating that gap2 is overexpressed in DprqR under GLC(+)DARK-LIGHT condition (Fig. 3G,H), can be fit with the suggestion relevent to a previous study51. Together, this suggestion justifyes our hypothesis that DprqR helps to remove stress from cells grown under high light at GLC(+)DARK-LIGHT condition (Fig. 8). However, this suggestion cannot explain the increased expression of gap2 in DprqR at GLC(+)DARK condition, during which no significant accumulation of ROS occured.
In summary, the established GLC(+)DARK-LIGHT condition provides a platform to determine the ROS signaling and glucose metabolism in Synechocystis. Our results, as well as physiological analyses from a previous study26, demonstrated that the response regulator PrqR is involved in the regulation of glucose metabolism and oxidative stress acclimation in Synechocystis (Fig. 8).
Materials and Methods
Conditions of growth and maintenance for cyanobacteria
The Synechocystis sp. PCC 6803 strain used in this study was kindly provided by Prof. Weiwen Zhang (Tianjin University, China). Synechocystis was cultivated in BG-11 media (both solid and liquid)52. For normal growth, the culture was maintained in a shaker photo-incubator under continuous illumination at 25 μmol m−2 s−2, with shaking at 130 rpm and temperature of 30 °C. The mutant culture was supplemented with various antibiotics at different concentrations, as mentioned in the text (for DprqR mutant line, up to 100 μg/mL kanamycin; for CprqR, 100 μg/mL kanamycin + 100 mg/mL spectinomycin; for OEprqR, 100 μg/mL kanamycin; and for DprqR-DprqA, 100 μg/mL kanamycin). For heterotrophic [GLC(+)DARK] growth, 5 mM glucose was added into the culture which was then maintained in the dark by wrapping with aluminum foil while shaking at 130 rpm and temperature of 30 °C. In this study, we maintained a special condition called GLC(+)DARK-LIGHT in which cells were inoculated in liquid BG-11 media and kept for normal culture until the OD730 was around 0.4; we then added 5 mM glucose and placed the samples under GLC(+)DARK condition for 3 days with subsequent shifting into high light (at 80 μmol m−2 s−2) for 2 or 3 days. For plate culture, the solid BG-11 on plate was always maintained in photo-incubator with continuous illumination at 25 μmol m−2 s−2 and temperature of 30 °C. Antibiotics had been used during maintenance or selection of mutant strains but not when comparative study was conducted between WT and different mutants.
Cloning strategies, transformation and generation of stable strains
The constructed vectors and strains developed by transforming those vectors are listed in Fig. S1A. The vectors were constructed by combining the normal restriction-digestion-ligation system and fusion system, as described by Zhang et al.53. The two vector’s backbones, namely, pUC19 and pBluescript KS (+), were used to construct all other vectors. The primers used in these constructions are listed in Supplementary Table S3. The constructed vectors transformed into Synechocystis following the method described by Liu et al.54. Colonies that appeared on the plate were streaked several times (at least 10 times) to obtain complete segregation. The mutant was confirmed by PCR, and positive clones were set in liquid culture supplemented with appropriate antibiotics.
In DprqR mutant (“D” stands for deletion), 245 bp out of 645 bp of prqR gene was deleted and a kanamycin cassette was inserted. The CprqR strain (“C” represents complementation) was generated by expressing the prqR gene under psbA2 promoter and spectinomycin under cpcB promoter in DprqR mutant by inserting in a neutral site (NS) at slr1311. For ChIP assay, we attached a 6XHis Tag at the C-terminus of the prqR gene, which was expressed under psbA2 promoter in WT strain at NS site of slr1311. To delete the two neighboring gene prqR and prqA, vector was designed using an upstream fragment of prqR and a downstream fragment of prqA flanked with kanamycin cassette under cpcB promoter.
ChIP-PCR
Chip-PCR was conducted using all reagents and protocols provided by Beyotime (Shanghai, China). Briefly, 40 mL of cell culture was collected from the normal culture of OEprqR strain, crosslinked using 1% formaldehyde and stabilized using 125 mM (final concentration) glycine. Sonication was performed as follows: 1 s ON, 2 s OFF for 18 min with 25% amplitude, to achieve the DNA fragment length of 400 bp to 800 bp. After centrifugation, the supernatant was transferred (about 0.2 mL) into a 2 mL ice-bathed tube. ChIP was performed in accordance with the protocol given by Beyotime (Shanghai, China).
The real time q-PCR was set with a 25 μL system (12.5 μL of SYBR GreeN, 0.5 × 2 Primers, 1 μL of sample, 1 μL of 1% input, 9.5 μL of ddH2O). Three pairs of primers were used for each targeted set (Supplementary Table S4). The DNA sample eluted with normal unconjugated mouse IgG was used as control.
Semi-quantitative PCR was set using Taq 2X Master Mix Kit (New England Biolabs Inc). The reaction was set in a 25 μL system (12.5 Taq 2X Master Mix, 0.5 × 2 Primers, 1 μL each of sample and 1% input, and 10.5 ddH2O) for 25 cycles.
GC/LC-MS based metabolic profiling
For GC-MS, approximately 3 × 109 cells were collected from the culture under GLC(+)DARK and GLC(+)DARK-LIGHT conditions and placed into −80 °C for at least 2 min to stop further metabolic activity. Cells were collected by centrifugation, washed twice with ice-cold NaCl (0.9%) and placed in tubes in the presence of liquid nitrogen. Up to 1.5 mL of methanol-ddH2O (1:1 v/v) was added along with 10 μL of L-2-chlorophenylalanine (0.3 mg/mL in dH2O). Sonication was conducted as follows: 5 s ON, 10 s OFF for 15 min at 90% amplitude, and then repeated thrice. The supernatant was dried up completely by freeze drying with continuous flow of nitrogen. Samples were derivatized and run under GC-TOF-MS following the method reported by Liu et al.32. The data were acquired in full-scan mode with the m/z range of 30–550 at a rate of 20 spectra per second following a solvent delay of 360 s. The acquired data were processed and analyzed as previously described32,55. Metabolite identification was performed by Chroma TOF linked with spectral library databases of National Institute of Standards and Technology (NIST) (2005). Other original standards were used to further validate the data. The dataset extracted in MS files from GC/TOFMS analysis was exported into NetCDF format by ChromaTOF software (v3.30, Leco Co., CA). CDF files were processed using custom scripts (revised MATLAB toolbox, hierarchical multivariate curve resolution (H-MCR), developed by Jonsson et al.56,57 in MATLAB 7.0 (The Math Works). Principal component analysis (PCA) was performed with SIMCA-P_12.0 software package (Umetrics, Umeå, Sweden) and GraphPad Prism version 5.01.
For LC-MS analysis, samples with equal cells number were collected at two time points: 72 h under GLC(+)DARK and 24 h under GLC(+)DARK-LIGHT condition. The pellet was centrifuged and collected quickly, frozen in liquid nitrogen and then preserved it at −80 °C before further use. Metabolites were extracted by quenching following the protocol described by Su et. al.33. In quenching metabolites, the frozen pellet was dissolved in 500 μL of 100% LC-MS grade methanol and placed for thawing on dry ice. The supernatant was collected and stored at −80 °C. The LC-MS experiment was conducted using an Agilent 1260 series binary HPLC system (Agilent Technologies, Waldbronn, Germany) attached with an Agilent 6410 triple quadrupole mass analyzer equipped with an electronic spray ion (ESI) source. Aqueous 10 mM tributylamine (pH 4.95, adjusted with acetic acid) and 100% HPLC-grade methanol were used as Mobile Phase A (MPA) and Mobile Phase B (MPB), respectively. Agilent Mass Hunter workstation LC/QQQ acquisition software (version B.04.01) was applied for data processing and Agilent Mass Hunter workstation qualitative analysis software version (B.04.00) applied for statistical analysis. The data obtained were analyzed using SIMCA 12.0 software (for PCA analysis), MeV4_9_0 (for Heat Map), and GraphPad Prism version 5.01 (t-test and others).
Growth and chlorophyll content measurements
The growth models of WT, DprqR and CprqR were studied under normal, GLC(+)DARK and GLC(+)DARK-LIGHT conditions by detecting the optical density (OD) at different wavelengths by using a spectrophotometer (METASH). OD730 was used to determine growth curve. OD678, OD720 and OD750 were also measured, and the total chlorophyll content was quantified using following formula; 14.94 (OD678–OD750)−0.616 (OD720–OD750), described by58.
RNA isolation, cDNA synthesis, and qRT-PCR
Total RNA from 50 mL of culture had been isolated using TRIzol (Invitrogen) following the modified method described by Schwarzkopf et al.59. The collected RNA was then treated with DNase1 (RNase free), deactivated DNase1 to remove contaminated DNA following the provided protocol. Thus, the DNA-free RNA was subjected to convert cDNA applying kits, SuperScript® III First-Strand Synthesis System (Invitrogen). qRT-PCR was performed by applying FastStart Universal SYBER Green (ROX) (Roche) reagent in Applied Biosystems 7500 RealTime PCR System. The RNA subunit of ribonuclease P subunit B (rnpB) served as an internal control. The primers used to quantify gene expression are listed in Supplementary Table S4.
ROS measurement
We measured the total ROS by using membrane-permeant fluorescence indicator 5-(and-6)-chloromethyl-2′, 7′-dichlorodihydro fluorescein diacetate, CM-H2DCFDA (Invitrogen, Life technology), following the protocols described by Hakkila, et al.23 and provided by Life Technology. Samples from normal culture, GLC(+)DARK culture (72 h) and GLC(+)DARK-LIGHT (24 h) were taken to estimate the total ROS content. Up to 25 mM CM-H2DCFDA was added to 1 mL of culture in 2 mL tube and then mixed. About 1 mL of culture was also obtained without adding any CM-H2DCFDA and considered as control (auto-fluorescence). Samples were then incubated for 90 min. in complete dark at 32 °C. Cells were washed with BG-11 media and suspended with BG-11 media to reach the final volume of 0.5 mL. Up to 200 μL of cell suspension was pipetted to a white 96-well microtiter plate. Fluorescence from CM-H2DCFDA treated and untreated cell (auto-fluorescence) was measured using Synergy Multi-Mode Reader (BioTek). The data measured immediately after dark incubation were considered as “Initial Reading” and then incubated in the corresponding condition for 1 h and measured the fluorescence for “Final Reading.” The strategy was followed both for treated and untreated (auto-fluorescence) samples. The ROS content was then calculated following the equation, ROS content = (IRT-FRT)-(IRUT-FRUT) where IRT, Initial Reading of Treated sample; FRT, Final Reading of Treated sample; IRUT, Initial Reading of Un-treated sample; and FRUT, Final Reading of Un-treated sample.
The excitation and detection emission had been set to 485/20 nm and 535/20 nm respectively with sensitivity at 50, optic position in bottom and normal read speed. BioTek’s Gen5™ 1.10 Reader Control and Data Analysis Software were used to set and analyze the data.
Glucose (extracellular) and glycogen measurements
Both glycogen and glucose were measured described elsewhere60. For glycogen quantification, 3 mL sample was obtained and centrifuged; the pellet was collected and washed with sterile ddH2O and then freeze dried. The weight of the dried cell was recorded. The cell was then suspended in 30% w/v KOH, mixed and incubated at 97 °C for 2 h. Ice-cold ethanol was added to the final concentration of 70% to 75% and incubated on ice for at least 2 h. The mixture was centrifuged, the supernatant was removed, and the pellet was washed with 98% ethanol. The glycogen pellet was dried at 60 °C for 10 min and then dissolved in 100 mM sodium acetate (pH 4.5). The dissolved glycogen was hydrolyzed enzymatically, by treating with 2 mg/mL amyloglucosidase (Sigma) at 60 °C for 2 h, into glucose. The glucose obtained from glycogen and extracellular (in media added externally) was measured using a Glucose (GO) Assay kit (Sigma-Aldrich) in accordance with the manufacture’s protocol. For glycogen measurement, data were normalized by cell dry weight. To understand the glucose uptake ability of the cell, kinetics of the decrease of the glucose content in the growth medium under GLC(+)DARK and GLC(+)DARK-LIGHT conditions were calculated. The data was normalized by OD730 according to time duration.
Additional Information
How to cite this article: Khan, R. I. et al. Transcriptional regulator PrqR plays a negative role in glucose metabolism and oxidative stress acclimation in Synechocystis sp. PCC 6803. Sci. Rep. 6, 32507; doi: 10.1038/srep32507 (2016).
References
De Rosa, E. et al. [NiFe]-hydrogenase is essential for cyanobacterium Synechocystis sp. PCC 6803 aerobic growth in the dark. Sci Rep 5, 12424 (2015).
Stal, L. J. & Moezelaar, R. Fermentation in cyanobacteria. Fems Microbiology Reviews 21, 179–211 (1997).
Rosa, M. et al. Soluble sugars–metabolism, sensing and abiotic stress: a complex network in the life of plants. Plant Signal Behav 4, 388–93 (2009).
Latifi, A., Ruiz, M. & Zhang, C. C. Oxidative stress in cyanobacteria. FEMS Microbiol Rev 33, 258–78 (2009).
Singh, A. K. & Sherman, L. A. Reflections on the function of IsiA, a cyanobacterial stress-inducible, Chl-binding protein. Photosynth Res 93, 17–25 (2007).
Couee, I., Sulmon, C., Gouesbet, G. & El Amrani, A. Involvement of soluble sugars in reactive oxygen species balance and responses to oxidative stress in plants. J Exp Bot 57, 449–59 (2006).
Russell, J. W. et al. High glucose-induced oxidative stress and mitochondrial dysfunction in neurons. Faseb J 16, 1738–48 (2002).
Bonnefont-Rousselot, D. Glucose and reactive oxygen species. Curr Opin Clin Nutr Metab Care 5, 561–8 (2002).
Barros, M. H., Bandy, B., Tahara, E. B. & Kowaltowski, A. J. Higher respiratory activity decreases mitochondrial reactive oxygen release and increases life span in Saccharomyces cerevisiae. J Biol Chem 279, 49883–8 (2004).
Pego, J. V., Kortstee, A. J., Huijser, C. & Smeekens, S. C. Photosynthesis, sugars and the regulation of gene expression. J Exp Bot 51 Spec No, 407–16 (2000).
Rolland, F., Baena-Gonzalez, E. & Sheen, J. Sugar sensing and signaling in plants: conserved and novel mechanisms. Annu Rev Plant Biol 57, 675–709 (2006).
Baena-Gonzalez, E., Rolland, F., Thevelein, J. M. & Sheen, J. A central integrator of transcription networks in plant stress and energy signalling. Nature 448, 938–42 (2007).
Ramon, M., Rolland, F. & Sheen, J. Sugar sensing and signaling. Arabidopsis Book 6, e0117 (2008).
Hauschild, R. & von Schaewen, A. Differential regulation of glucose-6-phosphate dehydrogenase isoenzyme activities in potato. Plant Physiol 133, 47–62 (2003).
Price, J., Laxmi, A., St Martin, S. K. & Jang, J. C. Global transcription profiling reveals multiple sugar signal transduction mechanisms in Arabidopsis. Plant Cell 16, 2128–50 (2004).
Hellmann, H., Funck, D., Rentsch, D. & Frommer, W. B. Hypersensitivity of an Arabidopsis sugar signaling mutant toward exogenous proline application. Plant Physiol 123, 779–89 (2000).
Rossel, J. B., Wilson, I. W. & Pogson, B. J. Global changes in gene expression in response to high light in Arabidopsis. Plant Physiol 130, 1109–20 (2002).
Wagner, D. et al. The genetic basis of singlet oxygen-induced stress responses of Arabidopsis thaliana. Science 306, 1183–5 (2004).
Fernandez-Gonzalez, B., Martinez-Ferez, I. M. & Vioque, A. Characterization of two carotenoid gene promoters in the cyanobacterium Synechocystis sp. PCC 6803. Biochim Biophys Acta 1443, 343–51 (1998).
Ryu, J. Y. et al. Glucose-induced expression of carotenoid biosynthesis genes in the dark is mediated by cytosolic ph in the cyanobacterium Synechocystis sp. PCC 6803. J Biol Chem 279, 25320–5 (2004).
Osanai, T. et al. Positive regulation of sugar catabolic pathways in the cyanobacterium Synechocystis sp. PCC 6803 by the group 2 sigma factor sigE. J Biol Chem 280, 30653–9 (2005).
Osanai, T. et al. Genetic engineering of group 2 sigma factor SigE widely activates expressions of sugar catabolic genes in Synechocystis species PCC 6803. J Biol Chem 286, 30962–71 (2011).
Hakkila, K. et al. Oxidative stress and photoinhibition can be separated in the cyanobacterium Synechocystis sp. PCC 6803. Biochim Biophys Acta 1837, 217–25 (2014).
Marin, K. et al. Identification of histidine kinases that act as sensors in the perception of salt stress in Synechocystis sp. PCC 6803. Proc Natl Acad Sci USA 100, 9061–6 (2003).
Kirik, I. A., Zinchenko, V. V., Shestakov, S. V. & Babykin, M. M. [Trans- and cis-acting autorepressors of the prqR gene in Synechocystis cyanobacteria sp. PCC6803]. Mol Biol (Mosk) 37, 1035–44 (2003).
Kreslavskii, V. D. et al. [Effect of oxidative stress inductors on the photosynthetic apparatus in cyanobacterium Synechocystis sp. PCC 6803 Prq20 mutant resistant to methyl viologen]. Biofizika 52, 277–86 (2007).
Nefedova, L. N., Fantin Iu, S., Zinchenko, V. V. & Babykin, M. M. [The prqA and mvrA genes encoding carrier proteins control resistance to methyl viologen in the cyanobacterium Synechocystis sp. PCC6803]. Genetika 39, 336–40 (2003).
Solovyev, V. V., Shahmuradov, I. A. & Salamov, A. A. Identification of promoter regions and regulatory sites. Methods Mol Biol 674, 57–83 (2010).
Tabei, Y., Okada, K. & Tsuzuki, M. Sll1330 controls the expression of glycolytic genes in Synechocystis sp. PCC 6803. Biochem Biophys Res Commun 355, 1045–50 (2007).
Nagarajan, S., Sherman, D. M., Shaw, I. & Sherman, L. A. Functions of the duplicated hik31 operons in central metabolism and responses to light, dark, and carbon sources in Synechocystis sp. strain PCC 6803. J Bacteriol 194, 448–59 (2012).
Singh, A. K. & Sherman, L. A. Pleiotropic effect of a histidine kinase on carbohydrate metabolism in Synechocystis sp. strain PCC 6803 and its requirement for heterotrophic growth. J Bacteriol 187, 2368–76 (2005).
Liu, Y. et al. An ultrasonication-assisted extraction and derivatization protocol for GC/TOFMS-based metabolite profiling. Anal Bioanal Chem 400, 1405–17 (2011).
Su, Y. et al. Metabolomic and network analysis of astaxanthin-producing Haematococcus pluvialis under various stress conditions. Bioresour Technol 170, 522–9 (2014).
Noctor, G., Lelarge-Trouverie, C. & Mhamdi, A. The metabolomics of oxidative stress. Phytochemistry 112, 33–53 (2015).
Tausz, M., Sircelj, H. & Grill, D. The glutathione system as a stress marker in plant ecophysiology: is a stress-response concept valid? J Exp Bot 55, 1955–62 (2004).
Narainsamy, K., Cassier-Chauvat, C., Junot, C. & Chauvat, F. High performance analysis of the cyanobacterial metabolism via liquid chromatography coupled to a LTQ-Orbitrap mass spectrometer: evidence that glucose reprograms the whole carbon metabolism and triggers oxidative stress. Metabolomics 9, 21–32 (2013).
Ortega-Ramos, M. et al. Engineering Synechocystis PCC6803 for hydrogen production: influence on the tolerance to oxidative and sugar stresses. PLoS One 9, e89372 (2014).
Takahashi, H., Uchimiya, H. & Hihara, Y. Difference in metabolite levels between photoautotrophic and photomixotrophic cultures of Synechocystis sp. PCC 6803 examined by capillary electrophoresis electrospray ionization mass spectrometry. J Exp Bot 59, 3009–18 (2008).
Nakajima, T. et al. Integrated metabolic flux and omics analysis of Synechocystis sp. PCC 6803 under mixotrophic and photoheterotrophic conditions. Plant Cell Physiol 55, 1605–12 (2014).
Adebiyi, A. O., Jazmin, L. J. & Young, J. D. 13 C flux analysis of cyanobacterial metabolism. Photosynth Res 126, 19–32 (2015).
Yoshikawa, K. et al. Integrated transcriptomic and metabolomic analysis of the central metabolism of Synechocystis sp. PCC 6803 under different trophic conditions. Biotechnol J 8, 571–80 (2013).
Godon, C. et al. The H2O2 stimulon in Saccharomyces cerevisiae. J Biol Chem 273, 22480–9 (1998).
Ishikawa, T. et al. Metabolome analysis of response to oxidative stress in rice suspension cells overexpressing cell death suppressor Bax inhibitor-1. Plant Cell Physiol 51, 9–20 (2010).
Chaouch, S. & Noctor, G. Myo-inositol abolishes salicylic acid-dependent cell death and pathogen defence responses triggered by peroxisomal hydrogen peroxide. New Phytol 188, 711–8 (2010).
Chaouch, S. et al. Peroxisomal hydrogen peroxide is coupled to biotic defense responses by ISOCHORISMATE SYNTHASE1 in a daylength-related manner. Plant Physiol 153, 1692–705 (2010).
Baxter, C. J. et al. The metabolic response of heterotrophic Arabidopsis cells to oxidative stress. Plant Physiol 143, 312–25 (2007).
Babykin, M. M. et al. [On the involvement of the regulatory gene prqR in the development of resistance to methyl viologen in cyanobacterium Synechocystis sp. PCC6803]. Genetika 39, 25–32 (2003).
Debnam, P. M. et al. Altered activity of the P2 isoform of plastidic glucose 6-phosphate dehydrogenase in tobacco (Nicotiana tabacum cv. Samsun) causes changes in carbohydrate metabolism and response to oxidative stress in leaves. Plant J 38, 49–59 (2004).
Van den Ende, W. & Valluru, R. Sucrose, sucrosyl oligosaccharides, and oxidative stress: scavenging and salvaging? J Exp Bot 60, 9–18 (2009).
Osanai, T. et al. Pleiotropic effect of sigE over-expression on cell morphology, photosynthesis and hydrogen production in Synechocystis sp. PCC 6803. Plant J 76, 456–65 (2013).
Figge, R. M., Cassier-Chauvat, C., Chauvat, F. & Cerff, R. The carbon metabolism-controlled Synechocystis gap2 gene harbours a conserved enhancer element and a Gram-positive-like -16 promoter box retained in some chloroplast genes. Mol Microbiol 36, 44–54 (2000).
Stanier, R. Y., Kunisawa, R., Mandel, M. & Cohen-Bazire, G. Purification and properties of unicellular blue-green algae (order Chroococcales). Bacteriol Rev 35, 171–205 (1971).
Zhang, Y. et al. Identification of a transporter Slr0982 involved in ethanol tolerance in cyanobacterium Synechocystis sp. PCC 6803. Front Microbiol 6, 487 (2015).
Liu, X., Sheng, J. & Curtiss, R., 3rd . Fatty acid production in genetically modified cyanobacteria. Proc Natl Acad Sci USA 108, 6899–904 (2011).
Qiu, Y. et al. Serum metabolite profiling of human colorectal cancer using GC-TOFMS and UPLC-QTOFMS. J Proteome Res 8, 4844–50 (2009).
Jonsson, P. et al. A strategy for identifying differences in large series of metabolomic samples analyzed by GC/MS. Anal Chem 76, 1738–45 (2004).
Jonsson, P. et al. High-throughput data analysis for detecting and identifying differences between samples in GC/MS-based metabolomic analyses. Anal Chem 77, 5635–42 (2005).
Williams, J. G. K. [85] Construction of specific mutations in photosystem II photosynthetic reaction center by genetic engineering methods in Synechocystis 6803. 167, 766–778 (1988).
Schwarzkopf, M., Yoo, Y. C., Huckelhoven, R., Park, Y. M. & Proels, R. K. Cyanobacterial phytochrome2 regulates the heterotrophic metabolism and has a function in the heat and high-light stress response. Plant Physiol 164, 2157–66 (2014).
Grundel, M., Scheunemann, R., Lockau, W. & Zilliges, Y. Impaired glycogen synthesis causes metabolic overflow reactions and affects stress responses in the cyanobacterium Synechocystis sp. PCC 6803. Microbiology 158, 3032–43 (2012).
Acknowledgements
We thank Mr. Tao Yixin, Wang You from Bio-X Institute for their suggestions in experiment and constructive criticism on the manuscript. We are also thankful to CSC scholarship council for their support during this study. We would like to thank all participants and research staff of Bio-X Institute for their contribution to the study. This work was supported by the 973 Program (2011CBA00803), the National Key Technology R&D Program (2012BAI01B09), the National Natural Science Foundation of China (91127047 and 81421061), and the Natural Science Foundation of Shanghai (13ZR1421100).
Author information
Authors and Affiliations
Contributions
R.I.K., L.H. and G.M. conceived the project and designed the experiments. R.I.K., Y.W., S.A. and X.Z. co-performed the experiments. R.I.K., Y.W., S.A., B.W., L.C., Y.L., W.Z., L.H. and G.M. analyzed the data. Y.W., L.H. and G.M. contributed reagents/materials/analysis tools. R.I.K., W.Z. and G.M. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Khan, R., Wang, Y., Afrin, S. et al. Transcriptional regulator PrqR plays a negative role in glucose metabolism and oxidative stress acclimation in Synechocystis sp. PCC 6803. Sci Rep 6, 32507 (2016). https://doi.org/10.1038/srep32507
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep32507
This article is cited by
-
Integrative analysis of the salt stress response in cyanobacteria
Biology Direct (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.