Abstract
Here we report two novel red emission fluorescent probes for the highly sensitive and selective detection of monoamine oxidase (MAO) with large Stokes shift (227 nm). Both of the probes possess solid state fluorescence and can accomplish the identification of MAO on test papers. The probe MAO-Red-1 exhibited a detection limit down to 1.2 μg mL−1 towards MAO-B. Moreover, the cleavage product was unequivocally conformedby HPLC and LCMS and the result was in accordance with the proposed oxidative deamination mechanism. The excellent photostability of MAO-Red-1 was proved both in vitro and in vivo through fluorescent kinetic experiment and laser exposure experiment of confocal microscopy, respectively. Intracellular experiments also confirmed the low cytotoxity and exceptional cell imaging abilities of MAO-Red-1. It was validated both in HeLa and HepG2 cells that MAO-Red-1 was capable of reporting MAO activity through the variation of fluorescence intensity.
Similar content being viewed by others
Introduction
Monoamine oxidase (MAO) is a flavin-dependent mitochondrial enzyme that catalyzes the oxidation of neuroactive and vasoactive amines such as dopamine, norepinephrine, adrenalin, and serotonin1,2. MAO-A and MAO-B are two isoforms of MAO, with 70% sequence identity and identical FAD cofactors3. Despite their resemblances, they also differ in a great many ways, including substrate and inhibitor specificities, and physiological functions4. MAO-A mainly metabolizes serotonin and epinephrine, the association between MAO-A and psychiatric disorders, such as social anxiety and depression, is well-documented. As a matter of fact, MAO-A selective inhibitors, take clorgyline and moclobemide for instance, are prescribed as clinical antidepressant agents5,6,7,8. MAO-B predominantly catalyzes the oxidative deamination of benzylamine and phenylethylamine and is selectively inhibited by pargyline and selegiline. MAO-B level is found to be associated with neurodegenerative disorders including Alzheimer’s disease and Parkinson’s disease and the selective inhibition of MAO-B is applied to the treatment of those two diseases9,10,11,12,13,14. Together, MAO-A and MAO-B play pivotal roles in maintaining the proper balance of neurotransmitter and other biogenic amines, which are essential for the healthy function of living organisms.
Due to their fundamental roles in sustaining the homeostasis of biogenic amines, MAO-A and MAO-B have drawn many attentions of researchers. Various technologies have been developed for monitoring real-time MAO levels, in an attempt to reveal MAO’s catalytic mechanism, better understand its roles in physiological processes, diagnose irregular MAO level caused diseases, and exploit MAO related drugs15,16,17,18,19,20,21. Among these technologies, fluorescence imaging22,23,24 as a powerful non-invasive method with high sensitivity and simplicity has attracted many attentions7,25,26,27,28. For example, Ahn et al. reported two coumarin based fluorophores that emits at 585 nm and achieved two-photon imaging of MAO29. Zhu and his colleagues designed a series of fluorescein probes, the maximum emission wavelength of which varies from 530 nm to 570 nm, and were successfully applied to live-cell imaging of MAO activites30,31. Last year, a group of activity-based MAO-B specific fluorescent probes with emission wavelength up to 530 nm were reported by Yao group32. Most of the previously developed MAO probes revealed good fluorescence imaging abilities and sensitivity towards MAO-B, whereas none of them had emission wavelength above 650 nm (Red emission). Fluorescent probes with emission wavelength above 650 nm have a great advantage in avoiding interference from biomolecular autofluorescence, minimizing photo-damage to living cells, and penetrating relatively deeper tissues33,34,35,36,37,38,39,40. Inspired by these merits, we designed the first red emission fluorescent turn-on probes for the detection of MAO with low detection limit, large Stokes shift, solid-state fluorescence and excellent imaging ability in monitoring MAO activities in living cells. The comprehensive improvement in the fluorescent properties will allow the probes to be widely adopted in biological applications.
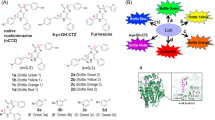
Our design strategy for the detection of MAO activities both in vitro and in vivo is based on the consecutive amine oxidation/β-elimination mechanism as shown in Fig. 1a. We chose (E)-2-(2-(4-hydroxystyryl)-4H-chromen-4-ylidene) malononitrile (DCPO) as the fluorescent reporter due to its long emission wavelength and biological compatibility33,40. Then, we installed two well-known MAO substrates, 3-amino-propoxy group and N, N-dimethylpropan-3-amino- propoxy group on the phenolic OH of DCPO to build MAO-Red-1 and MAO-Red-2 (Fig. 1b), respectively, which are not fluorescent at longer wavelength in the format of a phenolic ether (OFF state). We envisioned that with the oxidative deamination of MAO, the reactive site will be eliminated, leading to the ether bond cleavage and the release of DCPO. Subsequently, the fluorescence emission of DCPO at about 664 nm will be detected, and the OFF-ON fluorescence change will be observed.
Results
Spectra properties of MAO-Red-1 and MAO-Red-2
To explore the sensitivity and selectivity of MAO-Red-1 and MAO-Red-2, we first investigated their fluorescence properties in enzyme assay buffer (100 mM HEPES, pH 7.4 with 5% glycerol and 1% DMSO) at 37 °C. Prior to the treatment of MAO, the solution of MAO-Red-1 (10 μM) exhibited a fluorescence emission at around 550 nm. However, upon the addition of MAO (10 μg ml−1), the fluorescence intensity at 550 nm gradually decreased. Morever, a ~7.2-fold fluorescence increase at 664 nm were observed with MAO-B and ~3.8-fold augment with MAO-A. Additionally, since the maximum absorption wavelength was at 437 nm (Fig. S2), the Stokes shift was therefore calculated to be as large as 227 nm (Fig. 2a), which was highly conducive to minimizing background fluorescence. In addition to that, MAO-Red-1 displayed a detection limit of 1.2 μg mL−1 for MAO-B (Fig. S1), which was relatively low compared with previously reported MAO probes. Moreover, the results demonstrated that both of the probes were capable of detecting MAO activities, MAO-Red-1 exhibited better reactivity and was therefore further explored in all the subsequent studies. Fluorescence kinetic assays were later carried out between MAO and probes (Fig. 2a, inserted plot), it took less than 20 minutes for MAO-Red-1 to give a stable readout after the addition of MAO (Km value obtained for MAO-Red-1 with MAO-B was 270 μM), and about 40 minutes for MAO-Red-2 with MAO (Fig. S3). MAO-Red-1 exhibited the greatest signal-to-background ratio towards MAO-B (Fig. 2b), proving its capacity in selective detection of MAO and differentiate MAO-B from MAO-A with a selectivity ratio of about 2.2:1. Unfortunately, the excitation wavelength of the probes was in the UV-vis region, which will lead to the disadvantage in penetrating deep tissues compared with relatively longer excitation wavelength.
(a) Fluorescence intensity of MAO-Red-1 before and after reaction with MAO-A and MAO-B. Inserted plot: fluorescence kinetic assay of MAO-Red-1 (10 μM) after the addition of MAO (10 μg mL−1); (b) The signal-to-background ratio of MAO-Red-1 and MAO-Red-2 after reactions with MAO-A, MAO-B, subtilisin, BSA and inhibited MAO; [protein] = 0.01–0.15 mg mL−1. The data were recorded in enzyme assay buffer (100 mM HEPES, pH = 7.4 with 5% glycerol and 1% DMSO) at 37 °C (λex/λem = 420/664 nm); (c) Normalized fluorescent spectra of MAO-Red-1, MAO-Red-2 and DCPO in the solid state; (d) Paper imaging of MAO-Red-1, MAO-Red-2 and DCPO under 365 nm UV light.
To further validate the sensitivity of these probes towards MAO, MAO-Red-1 and MAO-Red-2 were respectively incubated with a group of different proteins, including subtilisin, bovine serum albumin (BSA), and pargyline inhibited MAO. No significant enhancement was detected upon addition of these proteins to our probes. We also noticed that MAO-Red-1, MAO-Red-2 and the theoretically product DCPO were all luminescent in the solid state. The maximum fluorescence emission of MAO-Red-1 and MAO-Red-2 were measured to be 592 nm and 599 nm, and the quantum yields were 10% and 5%, respectively. While DCPO exhibited a strong emission at about 633 nm (Fig. 2c), suggesting a red shift after the probes were incubated with MAO, which correlated well with the previous fluorescent results conducted in the aqueous solution. On this basis, we dipped filter papers into the methanol solution of MAO-Red-1, MAO-Red-2, and DCPO (1 mg mL−1), respectively. After drying up, DCPO showed obviously different colour from MAO-Red-1 and MAO-Red-2 (Fig. 2d) under 365 nm UV light. Given the advantages (i.e. high molar extinction coefficient, larger Stokes shifts, highly modifiable etc.) attributed to solid state fluorescent materials34 these novel probes might have the potential to be further employed in the application of MAO test papers.
Determination of the cleavage product through HPLC and LCMS tests
To further prove the proposed MAO catalytic mechanism as stated in Fig. 1, MAO-Red-1 (10 μM) was incubated with MAO-B (10 μg mL−1) in enzyme assay buffer (100 mM HEPES, pH 7.4 with 5% glycerol and 1% DMSO) at 37 °C for 1 hour. The reaction solution was then lyophilized, extracted with methanol, and tested on HPLC (Fig. 3). The cleavage product of MAO-Red-1 and DCPO displayed the same retention time, conforming they were of the same substance. We also verified the result using high resolution mass spectra, the characteristic peak at 625.1429 (m/z) in HRMS spectra was correspond to [DCPO + H]+ (Fig. S4). Thus, we could safely say that the cleavage product was DCPO.
Intracellular imaging of MAO
After demonstrating remarkable responsiveness and prominent selectivity towards MAO in vitro, we explored the possibility of applying our probes in vivo. Prior to imaging, it was essential for us to first validate the cytotoxicity of MAO-Red-1 using MTS reduction assay. HeLa cells were incubated with various concentrations of MAO-Red-1 (1, 1.25, 2, 2.5, 4, 5, 8, 10 μM) in 96-well plates for 24 h, the cell proliferation assay kit were then deployed according to standard procedure. Given the fact that above 80% of cells were still alive after treated with the maximum test concentration (10 μM) of MAO-Red-1 (Fig. S5), the toxicity of MAO-Red-1 is therefore neglectable.
Following that, we examined the photostability of MAO-Red-1 inside mammalian cells, which was a crucial factor for the successful imaging of MAO. Since MAO is a mitochondrial enzyme, we chose a commercially available Mito-Tracker Green as the comparative dye to co-stain HeLa cells with MAO-Red-1 for a total incubation time of 1 hour. The cells were then exposed to 405 nm (20 mW) and 588 nm (20 mW) channels of the solid state lasers, respectively. By the time of four minutes, the signal loss for Mito-tracker Green reached 91% while only 6% for MAO-Red-1 and 5% for DCPO (Fig. S6). Thus far, the photo stability of MAO-Red-1 is well proved both in vitro and in vivo.
Subsequently, we studied the cell imaging ability of MAO-Red-1. HeLa and HepG2 cells were chosen in this study on account of their positive expression of MAO. For the purpose of discussing the sensitivity of MAO-Red-1 in reporting MAO activities, both HeLa and HepG2 cells were divided into two groups (i.e. Group 1 and Group 2). Group 1 was incubated with 5 μM MAO-Red-1 for 1 h at 37 °C, while Group 2 was first treated with 100 μM MAO inhibitor pargyline for 2 h at 37 °C. Afterwards, MAO-Red-1 was added to Group 2 with the presence of pargyline, when the incubation time reached 1 h, the growth medium was removed and both groups were washed twice with PBS (PH = 7.4) to make sure no extracellular dye residue were left. The confocal imaging experiment was then carried out under dual-emission mode (λex = 405 nm), while no obvious change were observed in green channel, the red channel displayed distinct variation. The cells that were only treated with MAO-Red-1 showed significant red fluorescence, while the other cells that were pre-treated with MAO-B inhibitor pargyline exhibited very weak fluorescence (Fig. 4). This phenomenon was congruent with the previous carried out in vitro experiments, since the intensity at 550 nm (green channel) slightly decreased and a dramatic increase was observed at around 664 nm (red channel) after the addition of MAO.
(a,e,i,m) Bright field images of HeLa and HepG2 cells incubated with MAO-Red-1 (5 μM) for 1 h; (b,f,j,n) fluorescence images from Green channel (λex = 405 nm); (c,g,k,o) fluorescence images from red channel (λex = 405 nm); (e–h,m–p) fluorescence images of cells that was incubated with pargyline for 2 h followed by 1 h incubation of MAO-Red-1. The fluorescence images were collected at 420–520 nm (green channel) and 630–700 nm (red channel), respectively.
The successful imaging of MAO in living cells inspired us to further explore the intracellular staining abilities of MAO-Red-1 quantitatively via flow cytometry. As illustrated in Fig. S7, 90% cells that were pretreated with MAO-Red-1 displayed stronger fluorescence compared with untreated cells, while only 18% pargyline and MAO-Red-1 co-incubated cells exhibited the same increase. These results further demonstrated that MAO-Red-1 was adequate for intracellular detection of MAO activities.
Conclusion
In summary, we had designed and synthesized two red emission fluorescent probes with high sensitivity and selectivity towards MAO. In vitro, our probes successfully accomplished the identification of MAO on test papers and the detection limit demonstrated by MAO-Red-1 was relatively low compared with previously published MAO probes. The cleavage product was unequivocally conformed by HPLC and LCMS. In vivo, not only did the intracellular experiments validate the low cytotoxicity and high photostability of MAO-Red-1, but it was proven both in HeLa and HepG2 cells that MAO-Red-1 was capable of reporting MAO activities. Moreover, the CLSM experiments and flow cytometric studies jointly proved that MAO-Red-1could providea sensitive and accurate molecular tool for MAO imaging in complicated intracellular environment.
Methods
Chemicals and equipments
Unless otherwise stated, all reagents and solvents were obtained from commercial suppliers, and were used without further purification. Column chromatography was performed on silica gel (Qingdao haiyang) 300–400 mesh. All solvents used in spectra test systems were chromatographically pure. MAO-B inhibitor, Pargyline hydrochloride (P8013), human recombinant Monoamine Oxidase A (M7316) and B (M7441) (5 mg/ml) were purchased from Sigma Aldrich. Ultrapure water was used throughout.
1H NMR and 13C NMR spectra were recorded on a Bruker AMX-400 with chemical shifts expressed in parts per million (in deuteriochloroform or DMSO-d6, Me4Si as internal standard). UV−vis absorption spectra were recorded on a Hitachi PharmaSpec UV-1900 UV−visible spectrophotometer. Fluorescence spectra were determined using a HITAI F-7000 Spectro fluorophotometer. Absolute quantum yields were collected on a Horiba Jobin Yvon-Edison Fluoromax-4 fluorescence spectrometer with a calibrated integrating sphere system. To reduce the fluctuation in the excitation intensity, the xenon lamp was kept on for 1 hour prior to the experiment. The photostability experiments were performed on a Leica TCS SP8 confocal fluorescent microscope. The power percentages were both adjusted to 16.1%, the power on the focal plane were all 89 μW measured by a COHERENT LaserCheck power meter. The High-resolution mass spectra were obtained on a Finnigan LCQDECA and a Bruker Daltonics Bio TOF mass spectrometer. TLC analyses were performed on silica gel GF 254 using UV light as visualizing agent. The pH values were determined by a Leici pH3c (digital display) pH meter.
Synthetic procedures
The synthetic procedures are shown in Fig. 1b. All of the NMR and LCMS spectra for the corresponding products are shown in the supplementary information.
1-tert-Butoxycarbonylamino-3-bromopropane (S1)
A solution of 4-dimethylaminopyridine (244.3 mg, 2 mmol) and riethylamine (1.4 mL, 10 mmol) in dichloromethane (10 mL) was added dropwise to a suspension of 3-bromopropylamine hydrobromide (2.2 g, 10 mmol) and di-tert-butyl dicarbonate (2.3 g, 10.5 mmol) in dichloromethane (50 mL), then the mixture was stirred at room temperature for another 30 minutes to become a transparent solution. After that, the solution was washed successively with 0.5N HCl aq., 5 wt % of NaHCO3 aq., and brine, and then dried with NaSO4. After filtration, the solvent was removed under reduced pressure. S1 was obtained as colorless oil (2.3 g, 97% yield) and was then used without further purification.
(E)-2-(2-(4-methoxystyryl)-4H-chromen-4-ylidene) malononitrile (S2)
2-(2-methyl-4H-chromen-4-ylidene) malononitrile (1 g, 4.8 mmol) and 4-methoxybenzaldehyde (0.58 mL, 4.8 mmol) was dissolved in a mixture of ethanol and toluene (100 mL/40 mL), 20 drops of piperidine was added to the mixture. The solution was heated to reflux for 12 hours, and then the solvent was removed under reduced pressure. The crude product was then purified by chromatography on silica gel (petrol ether: dichloromethane, from 50:1 to 3:1, v/v) to afford S2 as a red solid. 1H NMR (400 MHz, CDCl3) δ 8.94 (d, J = 8.4 Hz, 1 H), 7.76 (m, J = 8.4 Hz, 1 H), 7.65–7.56 (m, 4 H), 7.47 (m, J = 8.2 Hz, 1 H), 6.99 (d, J = 8.6 Hz, 2 H), 6.85 (s, 1 H), 6.71 (d, J = 15.8 Hz, 1 H), 3.90 (s, 3 H);13C NMR (101 MHz, CDCl3) δ 161.7, 158.0, 152.8, 152.3, 138.7, 134.5, 129.7, 127.3, 125.8, 125.7, 118.5, 117.8, 116.9, 116.2, 115.9, 114.6, 106.1, 61.9, 55.4; HRMS (ESI) calcd.for C21H14N2O2, [M + H]+: 327.1134; found: 327.1109.
(E)-2-(2-(4-hydroxystyryl)-4H-chromen-4-ylidene) malononitrile (S3)
BBr3 (1.86 mL, 20 mmol) was diluted with anhydrous dichloromethane (30 mL), then was added dropwise over a period of 3 hours at 0 °C to the solution of S2 (1.3 g, 4 mmol) in 100 mL anhydrous dichloromethane. The mixture was allowed to warm up to room temperature and stirred overnight under nitrogen protection, after that the reaction was quenched with 10 mL water, the generated solid was filtered and washed with ethyl acetate. The filtrate was extracted with EtOAC (100 mL × 3); the organic layer was dried over NaSO4. The solvent was removed by rotary evaporation and the crude product was then purified by chromatography on silica gel (petrol ether: dichloromethane, from 10:1 to 0:1, v/v) furnished S3 as a red solid. 1H NMR (400 MHz, d6-DMSO) δ 10.14 (s, 1 H), 8.68 (dd, J = 8.4, 1.2 Hz, 1 H), 7.88 (m, 1 H), 7.75–7.72 (m, 1 H), 7.65–7.54 (m, 4 H), 7.21 (d, J = 15.9 Hz, 1 H), 6.89–6.83 (t, 3 H); 13C NMR (101 MHz, CDCl3) δ 165.2, 164.0, 157.9, 157.1, 144.4, 140.4, 135.5, 131.3, 131.2, 129.7, 124.1, 122.5, 122.2, 121.2, 121.0, 110.8, 64.3; HRMS (ESI) calcd. for C20H12N2O2, [M—H]−: 311.0826; found: 311.0827.
(E)-tert-butyl (3-(4-(2-(4-(dicyanomethylene)-4H-chromen-2-yl) vinyl) phenoxy) propyl) carbamate (S4)
To a stirred suspension of S3 (200 mg, 0.64 mmol), and K2CO3 (138.2 mg, 1 mmol) in DMF (10 mL), S1 (166 mg, 0.7 mmol) was added. After stirred at room temperature for 10 to 12 hours, the reaction was quenched by water (100 mL), aqueous phase was extracted with EtOAC (100 mL × 3), the organic layer was dried over NaSO4. The solvent was removed by rotary evaporation and the crude product was then purified by chromatography on silica gel (petrol ether: dichloromethane, from 10:1 to 0:1, v/v) to afford S4 as a red solid. 1H NMR (400 MHz, CDCl3) δ 8.91 (dd, J = 8.4, 1.2 Hz, 1 H), 7.73 (m, 1 H), 7.60–7.52 (m, 4 H), 7.44 (m, 1 H), 6.95 (d, J = 8.8 Hz, 2 H), 6.83 (s, 1 H), 6.68 (d, J = 15.8 Hz, 1 H), 4.74 (s, 1 H), 4.08 (t, J = 6.0 Hz, 2 H), 3.35 (q, 2 H), 2.05–1.99 (m, 2 H), 1.45 (s, 9 H); 13C NMR (101 MHz, CDCl3) δ 160.9, 157.9, 156.0, 152.8, 152.3, 138.7, 134.5, 129.7, 127.4, 125.8, 125.7, 118.5, 117.8, 116.9, 116.2, 115.1, 106.2, 79.3, 65.9, 62.0, 37.8, 29.5, 28.4; HRMS (ESI) calcd. for C28H27N3O4, [M + Na]+: 492.1899; found: 492.1868.
(E)-2-(2-(4-(3-aminopropoxy)styryl)-4H-chromen-4-ylidene) malononitrile (MAO-Red-1)
S4 (249 mg, 0.53 mmol) was dissolved in 5 mL dichloromethane, and was then added dropwise to saturated hydrogen chloride methanol solution. After stirred at room temperature for 10 to 12 hours, the mixture was concentrated under reduced pressure giving MAO-Red-1 (192 mg, 98%) as orange solid. 1H NMR (400 MHz, d6-DMSO) δ 8.68 (d, J = 8.2 Hz, 1 H), 8.04 (s, 3 H), 7.88 (t, J = 7.9 Hz, 1 H), 7.77–7.62 (m, 4 H), 7.56 (t, J = 7.6 Hz, 1 H), 7.31 (d, J = 15.8 Hz, 1 H), 7.00 (d, J = 8.4 Hz, 2 H), 6.92 (s, 1 H), 4.11 (t, J = 5.8 Hz, 2 H), 2.92 (t, J = 6.8 Hz, 2 H), 2.02 (m, 2 H); 13C NMR (101 MHz, d6-DMSO) δ 160.6, 158.9, 153.2, 152.3, 139.0, 135.7, 130.4, 128.1, 126.5, 125.0, 119.4, 117.6, 117.5, 117.4, 116.3, 115.5, 106.4, 65.3, 59.9, 36.5, 27.1; HRMS (ESI) calcd. for C23H19N3O2, [M + H]+: 370.1556; found: 370.1551.
(E)-2-(2-(4-(3-(dimethylamino) propoxy) styryl)-4H-chromen-4-ylidene) malononitrile (MAO-Red-2)
This compound was prepared according to the same process for the synthesis of S4, Yield: 61%.1H NMR (400 MHz, d6-DMSO) δ 9.29 (s, 1 H), 8.69 (d, J = 8.2 Hz, 1 H), 7.89 (t, J = 7.6 Hz, 1 H), 7.83–7.63 (m, 4 H), 7.58 (t, J = 7.6 Hz, 1 H), 7.34 (d, J = 16.0 Hz, 1 H), 7.02 (d, J = 8.5 Hz, 2 H), 6.95 (s, 1 H), 4.10 (t, J = 5.7 Hz, 2 H), 3.25–3.16 (m, 2 H), 2.80 (s, 6 H), 2.08 (m, 2 H);13C NMR (101 MHz, d6-DMSO) δ 160.5, 158.9, 153.2, 152.4, 138.9, 135.7, 130.4, 128.3, 126.5, 125.0, 119.4, 117.7, 117.6, 117.5, 116.3, 115.5, 106.5, 65.4, 59.9, 54.6, 42.8, 24.3; HRMS (ESI) calcd. for C25H23N3O2, [M + H]+: 398.1869; found: 398.1870.HeLa cells were cultured in Dulbecco’s modified Eagle medium (DMEM) containing 10% fetal bovine serum and 1% antibiotic–antimycotic at 37 °C in a 5% CO2/95% air incubator. Fluorescence imaging, cells (4 × 103 per well) were passed on confocal dishes and incubated for 24 h. Immediately before the staining experiments, the cells were washed twice with PBS (10 mM).
Parameters for the HPLC tests
HPLC measurements were carried out on Waters e2695 Separatins Module using Waters 2998 PDA detector equipped with a Symmetry C18 column (4.6 × 250 mm, 5 μm). Water (A) and methol (B) were used as eluents, the column was gradient eluted (A: B = 95-5%) for 20 minutes with a flow rate of 1 ml/min. 0.1%TFA was added to solvent A. 365 nm was used as the excitation wavelength.
Procedures for the cytotoxicity and confocal laser scanning microscopy (CLSM) experiments
HEK293 cells (human embryonic kidney) and HeLa cells were incubated in Dulbecco’s modified Eagle’s medium (DMEM) containing 10% fetal bovine serum (FBS) and 1% antibiotics (penicillin–streptomycin, 10,000 U mL−1) at 37 °C in a humidified atmosphere containing 5% CO2.
Toxicities of MAO-Red-1 toward HeLa cells were determined by using MTS reduction assay following literature procedures. About 1.0 × 104 cells/well was seeded into 96-well plates. After 24 h, various concentrations of MAO-Red-1 (1, 1.25, 2, 2.5, 4, 5, 8, 10 μM) were added to the cells. After another 24 h, 20 μL MTS and 100 μL PBS were added to each well and the plates were incubated at 37 °C for another 1 h. Then the absorbance of each sample was measured using an ELISA plate reader (model 680, BioRad) at a wavelength of 490 nm. The cell viability (%) was obtained according to the manufacturer’s instruction.
For the confocal laser scanning microscopy (CLSM) experiments, HEK 293 cells and HeLa cells were seeded at a density of 2.5 × 105 cells per well in 35 mm confocal dish (Φ = 15 mm), 24 h before the addition of MAO-Red-1 or pargyline followed by MAO-Red-1. The cells were observed with a ZEISS LSM 780 confocal laser scanning microscope.
Flow Cytometric Study
HeLa cells were seeded in 12-well plates (2.0 × 105 cells/well) and allowed to attach and grow for 24 h. Then the cells were treated with MAO-Red-1 (5 μM) or pargyline (100 μM, 2 h) followed by MAO-Red-1 (5 μM) in triplicate. Subsequently, the cells were washed with 1 × PBS, trypsinized, centrifuged and washed twice with 1 × PBS. The cell pellets were then resuspended. Mean fluorescence intensity was analyzed using the flow cytometer (BD Accuri C6) under the λex/λem mode (FL3 Red channel: filter 695 ± 40 nm, λex: 488 nm).
Additional Information
How to cite this article: Li, L.-L. et al. Red emission fluorescent probes for visualization of monoamine oxidase in living cells. Sci. Rep. 6, 31217; doi: 10.1038/srep31217 (2016).
References
Shih, J. C., Chen, K. & Ridd, M. J. Monoamine oxidase: from genes to behaviour. Annu. Rev. Neurosci. 22, 197–217 (1999).
Bortolato, M., Chen, K. & Shih, J. C. Monoamine oxidase inactivation: from pathophysiology to therapeutics. Adv. Drug Deliv. Rev. 60, 1527–1533 (2008).
Bach, A. W. J. et al. cDNA cloning of human liver monoamine oxidase A and B: Molecular basis of differences in enzymatic properties. Proc. Natl. Acad. Sci. USA 85, 4934–4938 (1988).
Belleau, B., Fang, M., Burba, J. & Moran, J. The absolute optical specificity of monoamine oxidase. J. Am. Chem. Soc. 82, 5752–5754 (1960).
Zhong, B. Y. & Silverman, R. B. Identification of the active site cysteine in bovineliver monoamine oxidase B. J. Am. Chem. Soc. 119, 6690–6691 (1997).
Li, L. et al. A sensitive two-photon probe to selectively detect monoamine oxidase B activity in Parkinson’s disease models. Nat. Commun. 5, 3276–3285 (2014).
Zhou, W. H. et al. New bioluminogenic substrates formonoamine oxidase assays. J. Am. Chem. Soc. 128, 3122–3123 (2006).
Ling, K. Q. & Sayre, L. M. Discovery of a sensitive, selective, and tightly binding fluorogenic substrate of bovine plasma amine oxidase. J. Org. Chem. 74, 339–350 (2009).
Johnson, J. P. Some observations upon a new inhibitor of monoamine oxidase in brain tissue. Biochem. Pharmacol. 17, 1285–1297 (1968).
Regina, L. et al. New pyrrole inhibitors of monoamine oxidase: synthesis, biological evaluation, and structural determinants of MAO-A and MAO-B selectivity. J. Med. Chem. 50, 922–931 (2007).
Chimenti, F. et al. Selective inhibitory activity against MAO and molecular modeling studies of 2-thiazolylhydrazone derivatives. J. Med. Chem. 50, 707–712 (2007).
Wu, J. B. et al. Monoamine oxidase A inhibitor-near-infrared dye conjugate reduces prostate tumor growth. J. Am. Chem. Soc. 137, 2366–2374 (2015).
Smith, D. M. et al. The separate and combined effects of monoamine oxidase inhibition and nicotine on P50 sensory gating. Psychopharmacology. 232, 1911–1920 (2015).
Huang, L. et al. Multitarget-directed benzylideneindanone derivatives: anti-beta-amyloid (Abeta) aggregation, antioxidant, metal chelation, and monoamine oxidase B (MAO-B) inhibition properties against Alzheimer’s disease. J. Med. Chem. 55, 8483–8492 (2012).
Repič, M. et al. Simulations of the hydride transfer step in the monoamine oxidase B catalyzed metabolism of dopamine. Proteins: Struct. Funct. Bioinform. 82, 3347–3355 (2014).
Torres, G. Z. et al. Revealing monoamine oxidase B catalytic mechanisms by means of the quantum chemical cluster approach. J. Chem. Inf. Model. 55, 1349–1360 (2015).
Chester, D. S. et al. Monoamine oxidase A (MAOA) genotype predicts greater aggression through impulsive reactivity to negative affect. Behav. Brain Res. 283, 97–101 (2015).
Guzel, D. M. & Iskra, E. B. The role of monoamine oxidase in humans and its metabolism. Psychiatric Annals. 44, 495–501 (2014).
Murray, A. T. et al. Catalytic amine oxidation under ambient aerobic conditions: mimicry of monoamine oxidase B. Angew.Chem. Int. Ed. 54, 8997–9000 (2015).
Silverman, R. B., Hoffman, S. J. & Catus, W. B. A mechanism for mitochondrial monoamine oxidase catalyzed amine oxidation. J. Am. Chem. Soc. 102, 7126–7128 (1980).
Yamaguchi, K. et al. Design of chemical shift-switching 19F magnetic resonance imaging probe for specific detection of human monoamine oxidase A. J. Am. Chem. Soc. 133, 14208–14211 (2011).
Le, T. T. et al. Aptamer-based biosensors for the rapid visual detection of flu viruses. Chem. Commun. 50, 15533–15536 (2014).
Liu, Y. et al. Fluorescence turn-on folding sensor to monitor proteome stress in live cells. J. Am. Chem. Soc. 137, 11303–11311 (2015).
Roth, A., Li, H., Anorma, C. & Chan, J. A reaction-based fluorescent probe for imaging of formaldehyde in living cells. J. Am. Chem. Soc. 137, 10890–10893 (2015).
Chen, G., Yee, D. J., Gubernator, N. G. & Sames, D. Design of optical switches as metabolic indicators: new fluorogenic probesfor monoamine oxidases (MAO A and B). J. Am. Chem. Soc. 127, 4544–4545 (2005).
Long, S. et al. An activity-based fluorogenic probe for sensitive and selective monoamine oxidase-B detection. Chem. Commun. 48, 7164–7166 (2012).
Albers, A. E., Rawls, K. A. & Chang, C. J. Activity-based fluorescent reporters for monoamine oxidases in living cells. Chem. Commun. 44, 4647–4649 (2007).
Krysiak, J. M. et al. Activity-based probes for studying the activity of flavin-dependent oxidases and for the protein target profiling of monoamine oxidase inhibitors. Angew. Chem. Int. Ed. 51, 7035–7040 (2012).
Kim, D. et al. Reaction-based two-photon probes for in vitro analysis and cellular imaging of monoamine oxidase activity. Chem. Commun. 48, 6833–6835 (2012).
Li, X. F. et al. Visualization of monoamine oxidases in living cells using “Turn-ON” fluorescence resonance energy transfer probes. Analyst. 139, 6092–6095 (2014).
Li, X. F. et al. Fluorescent probes for detecting monoamine oxidase activity and cell imaging. Org. Biomol. Chem. 12, 2033–2036 (2014).
Li, L. et al. A small-molecule probe for selective profiling and imaging of monoamine oxidase B activities in models of parkinson’s disease. Angew. Chem. Int. Ed. 54, 10821–10825 (2015).
Li, M. et al. A near-infrared colorimetric fluorescent chemodosimeter for the detection of glutathione in living cells. Chem. Commun. 50, 1751–1753 (2014).
Shao, A. D. et al. Far-red and near-IR AIE-active fluorescent organic nanoprobes with enhanced tumor-targeting efficacy: shape-specific effects. Angew. Chem. Int. Ed. 54, 7275–7280 (2015).
Weissleder, R. & Pittet, M. J. Imaging in the era of molecular oncology. Nature. 452, 580–589 (2008).
Liu, Y. et al. A cyanine-modified nanosystem for in vivo upconversion luminescence bioimaging of methylmercury. J. Am. Chem. Soc. 135, 9869–9876 (2013).
Wu, X. M. et al. In vivo and in situ tracking cancer chemotherapy by highly photostable NIR fluorescent theranostic prodrug. J. Am. Chem. Soc. 136, 3579–3588 (2014).
Atilgan, A. et al. Near-IR-triggered, remote-controlled release of metal ions: anovel strategy for caged ions. Angew. Chem. Int. Ed. 53, 10678–10681 (2014).
Guo, Z. Q., Park, S., Yoon, J. Y. & Shin, I. Recent progress in the development of near-infrared fluorescent probes for bioimaging applications. Chem. Soc. Rev. 43, 16–29 (2014).
Cao, J., Zhao, C. C. & Zhu, W. H. A near-infrared fluorescence chemodosimeter for fluoride via specific Si–O cleavage. Tetrahedron Lett. 53, 2107–2110 (2012).
Acknowledgements
This work was financially supported by the National Program on Key Basic Research Project of China (973 Program, 2012CB720603) and the National Science Foundation of China (nos 21232005, 21472131 and 21572147).
Author information
Authors and Affiliations
Contributions
K.L. and X.-Q.Y. designed the experiments, L.-L.L., Y.-H.L. and H.-R.X. conducted the experiments and prepared the figures. L.-L.L. and K.L. analyzed the results. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Li, LL., Li, K., Liu, YH. et al. Red emission fluorescent probes for visualization of monoamine oxidase in living cells. Sci Rep 6, 31217 (2016). https://doi.org/10.1038/srep31217
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep31217
This article is cited by
-
Kinetics, mechanism, and inhibition of monoamine oxidase
Journal of Neural Transmission (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.