Abstract
Chromosomes have a complex architecture in the cell nucleus, which serves vital functional purposes, yet its structure and folding mechanisms remain still incompletely understood. Here we show that genome-wide chromatin architecture data, as mapped by Hi-C methods across mammalian cell types and chromosomes, are well described by classical scaling concepts of polymer physics, from the sub-Mb to chromosomal scales. Chromatin is a complex mixture of different regions, folded in the conformational classes predicted by polymer thermodynamics. The contact matrix of the Sox9 locus, a region linked to severe human congenital diseases, is derived with high accuracy in mESCs and its molecular determinants identified by the theory; Sox9 self-assembles hierarchically in higher-order domains, involving abundant many-body contacts. Our approach is also applied to the Bmp7 locus. Finally, the model predictions on the effects of mutations on folding are tested against available data on a deletion in the Xist locus. Our results can help progressing new diagnostic tools for diseases linked to chromatin misfolding.
Similar content being viewed by others
Introduction
New technologies, such as Hi-C methods, are revealing that the genome of mammalian cells has a complex 3D architecture, comprising extensive long-range, functional interactions1,2,3,4,5. Chromosomes are folded into a sequence of 0.5–1.0 Mb long domains, known as TADs6,7, which are comparatively conserved between mice and humans. TADs, in turn, are thought to be one level of a hierarchy of higher-order domains (metaTADs), extending up to chromosomal scales8,9,10. Chromatin interactions have key functional roles, as for instance they control gene activity through the formation of physical loops between regulatory regions and target remote genes, disruptions being linked to human conditions11,12. Yet, the basic molecular mechanisms whereby chromosome 3D organisation is orchestrated in health and disease remain mysterious.
To make sense of genome-wide contact data and to expose the principles shaping chromosome 3D structure, models from polymer physics have been introduced13. Steric hindrance effects have been shown to play important roles in chromatin folding2,14,15,16, yet interacting polymer models have to be considered to describe FISH and Hi-C data in a more quantitative way17,18,19,20,21,22,23. In particular, we here we focus on the Strings&Binders (SBS) model of chromosome folding18 (Fig. 1a), a polymer model where the formation of chromatin loops is determined by specific interactions with DNA-binding molecules. The SBS model was the first to recapitulate within a single framework Hi-C and FISH data10,13,20.
A polymer model of chromatin.
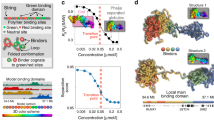
(a) The Strings&Binders (SBS) model is a Self-Avoiding (SAW) chain of beads interacting with molecular binders having a concentration, c and a binding affinity EInt. (b) Polymer physics dictates that the model different stable architectural classes correspond to the different phases of its phase diagram: at low EInt or c, the polymer is open and randomly folded in its coil phase; above its Θ-point transition, in the globule phase it is closed in more compact conformations as signalled by a drop in its gyration radius (Figure S1a); in the closed state, at higher values of EInt or c, its binders have a transition form a disordered to an ordered arrangement, as shown by their Structure Factor (Figure S1b).
Here we show that scaling concepts of classical polymer physics can explain the large scale behaviour of contact data over three orders of magnitudes in genomic separation, across different cell types and chromosomes. Chromatin is a complex mixture of different regions, folded in classes as predicted by polymer physics. The contact matrices of the Sox9 locus, a region associated to severe congenital diseases11 and of the Bmp7 locus, important in tissue development, are derived with high accuracy in mESC. Our models informed with genomic rearrangements can be used to anticipate their effects on the 3D architecture from only polymer physics. We test our in-silico predictions in the case of the ΔXTX deletion in the Xist locus where 5C data are available7. As our theory identifies the molecular determinants of folding, it can help progressing new diagnostic tools for diseases linked to chromatin misfolding, such as cancer and congenital disorders.
Results
In the SBS model, a chromatin filament is represented at a coarse-grained level as a Self-Avoiding (SAW) string of beads interacting with diffusing binders (Fig. 1a). We use the interaction potentials developed in classical studies of polymer physics24. The beads of the chain, each having a diameter σ, are connected by FENE bonds24 (see Supplementary Materials and Methods). The beads interact with the diffusing molecular binders through a Lennard-Jones attractive potential, having an energy scale, Eint. The binders have a concentration, c (and, for simplicity, have the same diameter σ); they can bridge the beads of the chain and, thus, fold the polymer. Each chromosome bead and each binder is subject to Brownian motion, here investigated by Molecular Dynamics (MD) computer simulations via LAMMPS25 in a box with periodic boundary conditions. To model a coarse-grained chromosome, we initially consider chains of N = 1000 beads; so, for instance, a 100 Mb long chromosome would have beads σ = 87 nm wide, each encompassing 100 k bases. In all cases, we work at scales much larger than the chromatin persistence length, where polymer scaling concepts26 are expected to apply.
Phase diagram and conformational classes
To characterize the thermodynamics features of the model, we first focus on the simple case of a homopolymer with identical beads, interacting with one type of DNA binders. In the space of the control parameters, Eint and c, the polymer has a known coil-globule folding transition13,26 signaled by a sharp drop in the value of its gyration radius at the Θ-point (Supplementary Figure S1a and Supplementary Materials and Methods). In the coil state, observed at small Eint and c, the binders do not succeed in establishing stable loops and the polymer has an open conformation falling in the universality class of the SAW (Fig. 1b). In the globular state, conversely, the binders fold the polymer in a closed conformation, occupying approximately 1% of the open state volume.
Here we discuss a new phase-transition, occurring in the polymer globular phase, wherby the binders, albeit not interacting directly with each other, undergo an order-disorder transition, which affect the finer structure of folding. At least two states exist: at low interaction energies or concentrations, the binders form a disordered lump around the chain, with a homogeneous density distribution and a flat structure function, S(k); at higher Eint or c, they form instead an ordered aggregate, revealed by sharp peaks in S(k) (Figure S1b and Materials and Methods). The phase diagram of the polymer-binders system is summarized in Fig. 1b: as dictated by polymer physics, the thermodynamics phases define the system stable conformational classes, which are expected to play an important role in the way chromatin folds. The phase transitions energies and corresponding concentrations fall in the weak biochemical energy range and in the nano- to micromole/litre range, as expected from biological considerations (Fig. 1b).
To characterize the folding state of our polymer model, we computed the average pairwise contact probability, Pc(s), of bead pairs at a given contour distance, s. Pc(s) only depends on the thermodynamics state of the system (Figure S2). In the coil state, Pc(s) decreases asymptotically as a power law with s, Pc(s) ~ s−α, with an exponent α ~ 2.1 in the SAW universality class. At the Θ-point, the exponent becomes α ~ 1.5, as known in polymer physics. In the globule state, Pc(s) depends on whether the system is in the disordered state, where after an initial decrease, a long plateau is found, or whether it is in the ordered state, where an exponent close to α ~ 1.0 is observed. Analogously, the mean square distance of site pairs, R2(s), which can be accessed by FISH measurements, depends on the system thermodynamics phase (Figure S3, Supplementary Materials and Methods).
The coil-globule and order-disorder transitions are general features of the physics of interacting polymers and more specific thermodynamics phases are known to exist depending on the molecular specificities of the system considered26. Even within our simplified context, the details of polymer conformations depend on other aspects, including the positioning of the binding sites along the bead chain, which could be unequally spaced and interspersed by ‘inert’ (non-interacting) sites, or local crowding and confinement. Different distributions of binding sites would produce, for instance, different types of ‘rosette-like’ globular conformations, with varying ranges of loop lengths. Furthermore, off-equilibrium, unstable conformations are also expected to be encountered in real chromosomal regions, in particular during changes in the folding state.
Chromatin is a mixture of different regions folded in their thermodynamics states
To compare our model against Hi-C data, we reasoned that a single chromosome is likely to be a mixture of a variety of different folded regions, including for instance eu- and heterochromatin domains, which can dynamically change from cell to cell according to functional purposes27. Yet, the stable spatial conformations of such regions must belong to one of folding classes determined by polymer physics (pure states), at least as a first approximation. To model such a scenario, we considered a mixture polymer system composed of different chain segments, each folded in one of the given thermodynamics states identified above (Fig. 2a).
Chromatin is a mixture of regions folded in different thermodynamics states.
(a) We model a chromatin filament as a mixture of differently folded regions, each belonging to one of the stable conformational classes envisaged by polymer physics (Fig. 1). In this view, the average pairwise contact probability is only determined by the relative abundances of the states in the mixture, as each state has a fixed, specific pairwise contact probability (Figure S2). (b) Genome-wide average contact frequencies across human cell types, from Hi-C and TCC technologies, can be fitted from the sub-Mb to chromosomal scales by such a mixture model. (c) Single chromosome data (here from IMR90 cells) can be similarly explained. (d) Different cell types (see colour scheme in panel b) have a different chromatin composition, with hESC (orange circle) more open than differentiated cells, such as IMR90 (blue circle). (e) Within a given cell type (here IMR90, as in c) distinct chromosomes have also a different composition, with chromosome X formed mostly of closed regions, whereas gene rich chromosomes, e.g., chr.19, are up to 70% open.
To test the biological significance of such a model of chromatin, we compared its predicted pairwise contact probability, P(s), with available Hi-C, TCC and in-situ Hi-C contact frequency data. In our mixture model, P(s) is just a linear combination of the contact probabilities of the pure states, independently derived above. It only depends on the relative abundances of the states in the mixture (and on a scale factor used to map bead sizes into genomic separations, see Supplementary Materials and Methods). We find that such a model can fit genome-wide averaged data (Fig. 2b, χ2 < 8 × 10−3) and single chromosome data (Fig. 2c, χ2 < 1.4 × 10−1) over approximately three orders of magnitude in length, from 0.5 Mb to chromosomal scales, across a variety of different cell types and experimental techniques.
Our approach also returns the mixture composition that best describes the given data (Fig. 2d,e, error bars are below 10% of signal, not shown for clarity). We find that different cell types have varying fractions of open state chromatin: embryonic stem cells6 (hESC) have the highest one, around 75%, while differentiated cells (e.g., IMR90) have values closer to 50%, in agreement with expectations. Different techniques (Hi-C v.s. TCC) give overall similar results in the same cell type: for instance, in GM12878 cells the fraction of closed ordered chromatin is 40% in both Hi-C and TCC data, yet the other states have a slightly different balance in the two cases. Different chromosomes can have very different compositions: in IMR90 cells, for instance, chromosome X is typically very compact (75%) with a prevalence of the closed states. In general, the shorter the chromosome the higher is its open fraction. For example, chromosome 1 is only 50% open; chromosome 11 or 12 are 40% in the closed-ordered conformation, with less than 5% in the disordered state; the gene rich chromosome 19 is one of the less compact (70% open), with the ordered and disordered closed states present in a 3/2 ratio. In brief, the mixture composition reflects the distribution of different folding domains along the chromosomes in the different cell types, across their thermodynamics states. Later, we will discuss the nature of the specific folding domains of important genomic loci, such as Sox9, Bmp7 and Xist.
Self-assembly of topological domains and higher-order structures
To investigate the patterns of Hi-C matrices beyond the average contact probability, we next studied the mechanisms underlying the self-assembly of topological domains. The average pairwise contact matrix of our homopolymer is different in the different thermodynamics phases and, as expected, the contact network is strongly enhanced in the closed states (Figure S4). We next considered the case of a block-copolymer with two types of beads (red/green) alternated in two pairs of blocks along the polymer chain, interacting each with a different type of binders (red/green, Fig. 3a, see Supplementary Materials and Methods). Each polymer block can fold in the conformational states discussed for the homopolymer. The random encounter of beads within the same block is favored by genomic proximity, resulting, above the Θ-point, in a lower level structure of ‘topological domains’ linked to single blocks. As similar beads in different blocks can also interact with each other, the long time contact matrices have a more complex, chessboard-like pattern (Fig. 3a), corresponding to a hierarchical organisation of higher-order structures deriving from intra- and inter-domain interactions. By those principles, the combinatorial action of a few different types of binders and corresponding binding sites can produce and regulate an exponential number of 3D conformations and patterns, as we discuss below. While the molecular nature of the blocks envisaged within our SBS framework is yet to be identified in general, a recent study in Drosophila has shown that the blocks could be also linked to local epigenetic patterns22. In our view, chromosomal structures discovered in Hi-C data, such as TADs6,7 and metaTADs10 and their differential re-wiring across tissues and cell types, emerge naturally by specialization of the involved molecular factors under general mechanisms of polymer physics.
Higher-order domains and symmetry breaking architectures.
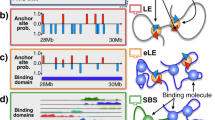
(a) We consider a block-copolymer composed of red and green binding domains which spontaneously folds, from an initially SAW conformation, in the ordered closed state of Fig. 1 (here c = 54 nmol/l and Eint = 4.1 kBT). The process is marked by a decrease of the gyration radius, Rg(t), in time (bottom panel) and by the formation of a hierarchy of higher-order domains, as reflected by the contact matrix pattern (top). (b) Pairs of sites with the same contour separation, differently positioned across a block boundary (see bottom panel), have the same average physical distances, r, in the open phase. Yet, in the closed states, the symmetry is broken by their different position relative to the boundary as the two pairs have a different physical distance, as seen from the corresponding distributions of r (from left to right Eint = 4.1, 3.1, 0 kBT and c = 54 nmol/l).
We also explored some additional, possibly functional consequences of the self-assembly of domains. As TAD boundaries have been associated to biological markers and, more specifically, to an insulating role, we focused on how they affect the physical distance of pairs of sites differently positioned relative to them. Within our toy block-copolymer model, we focused on pairs of sites with the same contour separation: we considered two cases where the pair is located symmetrically or asymmetrically with respect to a domain boundary (Fig. 3b). Interestingly, we found that the block boundary can have a simple symmetry-breaking effect: in the closed phases the sites of the symmetrically positioned pair have a larger physical distance than the asymmetrical pair (p-value = 0), whereas in the open phase no difference is recorded.
Finally, we investigated the occurrence of “many-body” contacts, i.e., co-localization events where multiple sites come simultaneously in physical proximity, beyond pairwise interactions. Within our toy homopolymer model, we measured the frequency of observing n sites in physical contact (Figure S5a, see also Fig. 4d and Supplementary Materials and Methods). As expected, in the closed states many-body contacts are exponentially more frequent than in the open state as n grows. The contact probability of bead triplets on a same polymer at different genomic separations, Pc(s1, s2), is given in Figure S5b. Multiple interactions are currently not detected by Hi-C methods, yet our model highlights that they are likely to be an abundant structural component of chromatin. That hints towards an overseen functional role of closed chromatin domains whereby multiple regulatory regions (enhancers) can loop simultaneously onto a given target (gene promoter) with a much higher probability than in open regions. Taken together our results support a view whereby basic mechanism of polymer folding could play key functional roles in the regulation of the genome by controlling the spatial organisation of chromatin.
Polymer physics captures the folding of the Sox9 locus.
(a) Top: the considered Sox9 locus in mESC-J1 cells, with a few marker genes. Bottom: the SBS polymer model that best explain the Hi-C contact map of the Sox9 region has the shown different types of binding sites, as seen in the zoom (different colors); their abundance is represented as an histogram over the genomic sequence. The bar at the bottom highlights three main regional areas to help 3D visualization. (b) The model derived pairwise contact frequency matrix (bottom) has a 95% Pearson correlation with Hi-C experimental data (top). (c) A snapshot of the Sox9 locus in its closed disordered state as derived by the polymer model, with the position of TSSs of some key genes highlighted. (d) In the locus, many-body contacts of n sites are exponentially more abundant than in random SAW conformations (the ratio of the average number is plotted v.s. n), which could help the simultaneous co-localization of multiple functional regulatory regions. (e) The Sox9 locus folding dynamics from an initially open conformation towards the closed disordered state is represented by the gyration radius, Rg(t). Chromatin domains self-assemble hierarchically in higher-order structures, in approx. 20s.
Folding of the Sox9 locus in mESC
Next, we asked whether our polymer models can also explain the details of folding of specific DNA regions. First, we considered a 6 Mbs sequence around the Sox9 gene (chr11:109000000-115000000, mm9), an important genomic locus linked to severe congenital diseases11, including gene rich areas as well as gene deserts (Fig. 4a). To capture the details of Sox9, we generalized our polymer model to accommodate different types of binding sites (colors) and molecular binders (Fig. 4a) and employed a chain with N = 2250 beads (σ = 26 nm) to increase resolution. With a Monte Carlo recursive optimization method (Supplementary Materials), we estimated the minimal arrangement and different types of binding sites that best reproduces the 40 kb Hi-C map available in mESC-J1 cells6. We informed our polymer model with the obtained arrangement of binding sites and run full-scale Molecular Dynamics simulations to derive the ensemble of 3D conformations of the locus (see Supplementary Materials and Methods). Figure 4a represents the histograms of the abundance of our inferred binding sites along the locus (different types in different colors), ordered left-to-right according to the location of the domain center of mass. Binding domains tend to overlap with the different TADs in the locus, but importantly they also overlap with each other producing higher-order interactions across TADs, i.e., metaTADs10, as visible in the original Hi-C data and in our 3D snapshots as well. In mESC-J1 cells, we find that the mixture best describing the locus is made 64% of the open and 36% of the closed disordered state (see Supplementary Materials and Methods).
The derived pairwise contact frequency matrix has a Pearson correlation with Hi-C data of 95% (Fig. 4b), supporting the view that our polymer model captures a relevant component of the molecular mechanisms determining the folding of Sox9. A snapshot of a single typical configuration, in the closed state, is shown in Fig. 4c, where the relative positioning of Sox9 and other genes in the locus, across its different higher-order domains, can be visualized. For instance, the transcription starting sites (TSSs) of the Sox9 and Kcnj2 genes have a genomic separation almost four times larger than the TSSs of Sox9 and Slc39a11 (1.72 Mb v.s. 0.46 Mb), but the average physical distances of the two pairs are proportionally closer (1.19 μm v.s. 0.59 μm) as the three genes belong to consecutive regional areas.
The Sox9 locus is marked by many-body contacts that are exponentially more abundant than expected in a randomly folded conformation (Fig. 4d, error bars within symbol size). The self-assembly of the locus architecture from initial open states proceeds hierarchically, with early formed local domains folding into larger and larger 3D structures encompassing the entire locus (Fig. 4e); the order of magnitude of the time-scale of the process is 20 sec. The variety of information on Sox9 and its folding mechanisms that can be inferred from polymer physics extends well beyond the Hi-C pair-wise contact data used to infer the model.
To check the general validity of our approach, we applied our polymer models to a different locus, a 2 Mb wide region around the Bmp7 gene (chr2:171090000-173430000) important in tissue development, where Hi-C data at 30 kb resolution are available in mESC-46C cells10. Also in the Bmp7 case, the inferred SBS binding domains produce a contact matrix having a pattern very similar to experimental data, with a 95% correlation (Figure S6).
Testing model predictions on the ΔXTX deletion in the Xist locus
Finally, to test our model, we considered the Xist locus, an important 1.3 Mb long region around the Xist gene (chrX:100298000-101373000), because data are also available for its ΔXTX deletion7. Starting from a 20 kb resolution 5C map in mESC-E14 cells7, we derived our polymer model describing the wild-type locus and after MD simulations we found a good agreement with 5C data (Fig. 5a,b; a 72% open and 28% closed disordered state mixture returns a 96% Pearson correlation of contact matrices). A 3D snapshot of the locus conformation and its two TADs is shown in Fig. 5c. Next, we implemented the ΔXTX deletion in our polymer model and calculated the contact map of the mutated system just from polymer physics, under no fitting or adjustable parameters. We find that the predicted interaction matrix has a pattern of extended ectopic interactions with respect to the wild-type case that is very similar to the one reported in 5C data on ΔXTX7, with an overall correlation of 91% (Fig. 5d). In the above calculation there are no adjustable parameters. In case the mixture composition is allowed to vary the best fit returns a similar mixture (80–20%), having a 91.4% correlation with the data. Such findings are in agreement with those of a similar interacting polymer model introduced for the Xist locus28. Our MD simulations show that, after the deletion, the regions flanking ΔXTX are spatially repositioned with respect to each other (3D snapshot in Fig. 5e) and contribute to establishing the observed ectopic interactions across regions sharing cognate binding sites. Summarizing, the binding domains identified in-silico by our model can describe the folding of Sox9 with high accuracy and predict the effects of genomic rearrangements based only on polymer physics.
Model predicted effects of the ΔXTX deletion in the Xist locus.
(a) The different binding domains are shown of the SBS polymer model of the Xist locus, inferred from 5C contact data7 in mESC-E14 cells. To help 3D visualization, the bars at the bottom highlight in cyan the wild-type region that is deleted in ΔXTX cells, its flanking sequences in yellow and the remaining regional areas overlapping the locus two TADs in violet and blue. (b) The model inferred contact matrix (bottom) has a 96% Pearson correlation with 5C experimental data (top). (c) A snapshot of the Xist locus in its closed disordered state. (d) The contact matrix predicted, under no adjustable parameters, by the very same model after the ΔXTX deletion (bottom) has a pattern of ectopic interactions (full line magenta box) w.r.t. wild-type data (panel b) very similar to the one seen in the corresponding experimental 5C data (top, correlation 91%). (e) Ectopic interactions extend across the yellow regions flanking the ΔXTX deletion, as visible in the snapshot of the deleted locus (closed disordered state).
Discussion
We have discussed a polymer model of chromatin where 3D conformations are shaped by the interactions of binding domains with their cognate binding molecular factors, such as DNA-binding molecules. Genome-wide and loci specific chromatin contact data can be explained by classical scaling concepts of polymer physics over orders of magnitude in genomic separation, up to chromosomal scales, across mammalian cell types. Consistent with our SBS picture, a model has been recently proposed focused, in particular, on the role of CTFC binding sites and cohesin mediated interactions to explain folding of loci at short scales, in the 10–500 kb range23,29. In facts, CTFC and cohesin molecules are examples of specific factors and DNA binders that contribute to chromatin folding. Contrary to previous approaches, our model does not require a previous knowledge of the molecular factors responsible for folding (e.g., location of CTCF sites or epigenetics features); conversely, it can be used to infer the nature of the key folding factors.
Pairwise contact data on the Sox9, Bmp7 and Xist loci are reproduced by the SBS model with an accuracy above 95%. Additionally, we found that the model predictions on the ectopic interactions induced by the ΔXTX deletion in the Xist locus are in a remarkably good agreement with available 5C data7,28. That supports the view that our model captures some of the key folding mechanisms of chromatin. Although current Hi-C technologies only provide information on pairwise contacts, we showed that ‘many-body’ interactions are an abundant, structural component of chromatin 3D organisation.
By combining polymer models and Hi-C data, a quantitative scenario emerges of the large-scale features of chromatin architecture where chromatin folding is determined by a complex system of binding domains and molecular factors, regulated by general mechanisms of polymer physics. As our polymer physics approach identifies the molecular determinants of folding and their mechanisms of action, it can help understanding the link between architecture and function and the design of novel approaches to personalized diagnosis and treatment of human diseases.
Materials and Methods
A detailed description of the materials and methods is provided in the Supplementary Materials. In particular, the SBS polymer model was investigated by Brownian Molecular Dynamics computer simulations implemented by LAMMPS. All the details about the model and computational parameters, as well as on the analyses performed are reported in the Supplementary Materials and Methods.
Additional Information
How to cite this article: Chiariello, A. M. et al. Polymer physics of chromosome large-scale 3D organisation. Sci. Rep. 6, 29775; doi: 10.1038/srep29775 (2016).
References
Misteli, T. Beyond the sequence: cellular organisation of genome function. Cell 128, 787–800 (2007).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–93 (2009).
Dekker, J., Marti-Renom, M. A. & Mirny, L. A. Exploring the three-dimensional organisation of genomes: interpreting chromatin interaction data. Nat. Rev. Genet. 14, 390–403 (2013).
Tanay, A. & Cavalli, G. Chromosomal domains: epigenetic contexts and functional implications of genomic compartmentalization. Curr. Op. Gen. & Dev. 23, 197–203 (2013).
Bickmore, W. A. & van Steensel, B. Genome architecture: domain organisation of interphase chromosomes. Cell 152, 1270–84 (2013).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–80 (2012).
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–5 (2012).
Sexton, T. et al. Three-dimensional folding and functional organisation principles of the Drosophila genome. Cell 148, 458–472 (2012).
Phillips-Cremins, J. E. et al. Architectural protein subclasses shape 3D organisation of genomes during lineage commitment. Cell 153, 1281–1295 (2013).
Fraser, J. et al. Hierarchical folding and reorganisation of chromosomes are linked to transcriptional changes during cellular differentiation. Mol. Sys. Bio. 11, 852 (2015).
Spielmann, M. & Mundlos, S. Structural variations, the regulatory landscape of the genome and their alteration in human disease. Bioessays 35, 533–543 (2013).
Lupianez, D. G. et al. Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions. Cell 161, 1012–1025 (2015).
Nicodemi, M. & Pombo, A. Models of chromosome structure. Curr. Op. Cell Bio. 28, 90–95 (2014).
Sachs, R. K. et al. A random-walk/giant-loop model for interphase chromosomes. Proc. Natl. Acad. Sci. USA 92, 2710–14 (1995).
Marenduzzo, D., Micheletti, C. & Cook, P. R. Entropy-driven genome organisation. Biophys. J. 90, 3712–3721 (2006).
Rosa, A. & Everaers, R. Structure and dynamics of interphase chromosomes. PLoS Comp. Bio. 4, e1000153 (2008).
Kreth, G. et al. Radial arrangement of chromosome territories in human cell nuclei: a computer model approach based on gene density indicates a probabilistic global positioning code. Biophys. J. 86, 2803–2812 (2004).
Nicodemi, M. & Prisco, A. Thermodynamic pathways to genome spatial organisation in the cell nucleus. Biophys. J. 96, 2168–2177 (2009).
Bohn, M. & Heermann, D. W. Diffusion-driven looping provides a consistent framework for chromatin organisation. PLoS ONE 5, e12218 (2010).
Barbieri, M. et al. Complexity of chromatin folding is captured by the Strings & Binders Switch model. Proc. Natl. Acad. Sci. USA 109, 16173–1678 (2012).
Brackley, C. A. et al. Nonspecific bridging-induced attraction drives clustering of DNA-binding proteins and and genome organisation. Proc Natl Acad Sci USA 110, E3605–11 (2013).
Jost, D., Carrivain, P., Cavalli, G. & Vaillant, C. Modeling epigenome folding: formation and dynamics of topologically associated chromatin domains. Nucleic Acids Res. 42, 9553–61 (2014).
Sanborn, A. L. et al. Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes. Proc. Natl. Acad. Sci. USA 112, E6456–65 (2015).
Kremer, K. & Grest, G. S. Dynamics of entangled linear polymer melts: A molecular‐dynamics simulation. J. Chern. Phys. 92, 5057–86 (1990).
Plimpton, S. Fast parallel algorithms for short-range molecular dynamics. J. Comput. Phys. 117, 1–19 (1995).
de Gennes, P. G. Scaling Concepts in Polymer Physics (Cornell Univ. Press, Ithaca, NY, 1979).
Nagano, T. et al. Single-cell Hi-C reveals cell-to-cell variability in chromosome structure. Nature 302, 59–64 (2013).
Giorgetti, L. et al. Predictive polymer modeling reveals coupled fluctuations in chromosome conformation and transcription. Cell 157, 950–63 (2014).
Dekker, J. & Mirny, L. The 3D Genome as Moderator of Chromosomal Communication. Cell 164, 1110–21 (2016).
Acknowledgements
We thank Ana Pombo for critical discussions and comments. This work was supported by grants to MN from the NIH ID 1U54DK107977-01 and CINECA ISCRA ID HP10CYFPS5. MN also acknowledges computer resources from INFN, CINECA and Scope at the University of Naples.
Author information
Authors and Affiliations
Contributions
M.N. designed the study; A.M.C., C.A., S.B. and M.N. developed the project; A.M.C., C.A., S.B. and A.E. run the computer simulations and performed the analyses; A.M.C., C.A., S.B. and M.N. prepared the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Chiariello, A., Annunziatella, C., Bianco, S. et al. Polymer physics of chromosome large-scale 3D organisation. Sci Rep 6, 29775 (2016). https://doi.org/10.1038/srep29775
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep29775
This article is cited by
-
Active transcription and epigenetic reactions synergistically regulate meso-scale genomic organization
Nature Communications (2024)
-
Interphase chromosomes of the Aedes aegypti mosquito are liquid crystalline and can sense mechanical cues
Nature Communications (2023)
-
Enhancer–promoter contact formation requires RNAPII and antagonizes loop extrusion
Nature Genetics (2023)
-
The dynamics of three-dimensional chromatin organization and phase separation in cell fate transitions and diseases
Cell Regeneration (2022)
-
Loop-extrusion and polymer phase-separation can co-exist at the single-molecule level to shape chromatin folding
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.