Abstract
The ability of scoring functions to correctly select and rank docking poses of small molecules in protein binding sites is highly target dependent, which presents a challenge for structure-based drug discovery. Here we describe a virtual screening method that combines an energy-based docking scoring function with a molecular interaction fingerprint (IFP) to identify new ligands based on G protein-coupled receptor (GPCR) crystal structures. The consensus scoring method is prospectively evaluated by: 1) the discovery of chemically novel, fragment-like, high affinity histamine H1 receptor (H1R) antagonists/inverse agonists, 2) the selective structure-based identification of ß2-adrenoceptor (ß2R) agonists and 3) the experimental validation and comparison of the combined and individual scoring approaches. Systematic retrospective virtual screening simulations allowed the definition of scoring cut-offs for the identification of H1R and ß2R ligands and the selection of an optimal ß-adrenoceptor crystal structure for the discrimination between ß2R agonists and antagonists. The consensus approach resulted in the experimental validation of 53% of the ß2R and 73% of the H1R virtual screening hits with up to nanomolar affinities and potencies. The selective identification of ß2R agonists shows the possibilities of structure-based prediction of GPCR ligand function by integrating protein-ligand binding mode information.
Similar content being viewed by others
Introduction
In the past decade, there has been remarkable progress in the structural elucidation of G protein-coupled receptors (GPCRs), the largest family of transmembrane proteins in the human genome that plays an essential role in a plethora of cell signalling processes and has high potential as drug targets1,2. Currently, over 140 GPCR crystal structures have been published, covering 33 different GPCRs and 72 unique small molecule GPCR ligands with different functional effects on receptor signalling and with distinct binding modes in the receptor1,2. The increased amount of high resolution structural information on GPCRs has opened up new opportunities for the identification of novel GPCR ligands by structure-based virtual screening (SBVS)3,4,5,6. There are, however, still several hurdles for structure-based approaches for GPCRs, such as the efficient identification of chemically novel fragment-like ligands with high hit rates4,7 (i.e. the portion of experimentally validated hits) and the structure-based prediction of GPCR ligand function5,6,8. The last-mentioned hurdle has proven to be challenging as the functional effect of a ligand is inherently linked to the binding mode(s) it can adopt and the receptor conformations it can stabilize that are associated with specific receptor activation states9,10. The development of structure-based, ligand-function specific virtual screening methods is hampered by the fact that for most crystallized GPCRs only one or few structures are available as well as the limited diversity of the functional effect and binding modes of the co-crystallized ligands. Whereas the efficient identification of fragment-like compounds was shown to be hampered primarily by the (target-dependent) inability of docking scoring functions to accurately rank and score the different binding modes with an estimation of their binding energies11. These challenges are, for example, illustrated by a recent structure-based virtual screening study by Rodríguez et al. in which high-affinity adenosine antagonists were discovered while screening against an active-state agonist-bound X-rays of the A2A receptor in order to obtain A2A agonists12. In another virtual screening study against the ß2-adrenoceptor (ß2R), Weiss et al. were able to identify 1 fragment-like and 5 lead-like ß2-adrenoceptor (ß2R) agonists by making use of both an active-state agonist-bound and an inactive-state inverse-agonist bound structure and selecting compounds that were only scored high in the agonist structure13. In order to overcome issues with scoring and ranking docking poses consensus approaches have been devised14. These consensus scoring approaches have been applied retrospectively14,15 and prospectively16,17,18 in several studies. In those cases where the consensus approach has been prospectively applied the individual approaches have, however, never been experimentally validated. Only the prospective application, experimental validation and comparison of the individual and combined scoring methods would allow the assessment of the added value of consensus scoring in virtual screening.
In the current study we address both hurdles in virtual screening simultaneously by applying a novel docking scoring approach for the identification of novel fragment-like GPCR ligands and the prediction of their functional effect using GPCR crystal structures. This docking scoring approach combines a conventional docking scoring function (ChemPLP) using PLANTS19 docking with the molecular interaction fingerprint (IFP) rescoring approach20,21. This combination is based on the “complementary” hypothesis in which it is assumed that the combination of two (fundamentally) different scoring functions can result in increased performance by combining the strengths of each scoring function14. ChemPLP is an empirical energy-based scoring function whereas IFP compares the interaction pattern between the docking pose and the protein to a reference binding mode, most often the co-crystallized pose of a known ligand. By combining these methods we aimed to use the strengths of PLANTS to identify compounds with energetically favorable docking poses with the strength of IFP to select the most probable binding modes by selecting those with an interaction profile closely resembling the reference IFP20 (in this case the IFP of the co-crystallized ligand with the receptor). Moreover, in this study we have validated the performance of the consensus scoring approach in a truly prospective manner by also experimentally validating the results of the individual scoring approaches. We applied and systematically compared the combined and individual IFP and PLANTS scoring approaches with respect to their ability to: 1) identify chemically novel, fragment-like, high affinity histamine H1 receptor (H1R) antagonists/inverse agonists and 2) selectively retrieve ß2R agonists. The ß2R represents a rewarding additional target as there are many crystal structures available and we aim to selectively identify agonists, thereby representing a complementary case study to H1R. Building from our successful prospective H1R VS study21 and retrospective ß2R VS study22 reported earlier, we have in the current study for the first time explored and compared the virtual screening performances of the different scoring approaches and combinations for H1R and ß2R in a prospective manner.
The integration of protein-ligand interaction energy and interaction pattern similarity scores resulted in a better performance than each of the individual methods, although in all cases high hit-rates were obtained. Retrospective virtual screening studies based on multiple different ß-adrenoceptor crystal structures allowed us to select an optimal combination of reference interaction fingerprint and protein conformation for the selective retrieval of novel, fragment-like ß2R agonists. These results demonstrate the potential of structure-based prediction of GPCR ligand function by the integration of protein-ligand binding mode information.
Results and Discussion
Discovery new fragment-like H1R ligands
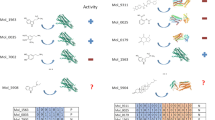
In order to analyse to what extent the combined scoring approach was responsible for the high hit-rate of our previously reported virtual screening on the doxepin-bound (1) H1R crystal structure (PDB-code 3RZE23) we experimentally validated the compound selections for each of the individual scoring approaches (Fig. 1). For both the PLANTS as well as the IFP approach the top 500 scoring compounds were selected and subsequently the compounds matching the combined approach were removed. The remaining compounds were processed in the same fashion as the compounds from the combined approach. We excluded hits that are similar to any known H1R ligand (ECFP-424 Tanimoto score of ≥0.4), visually clustered the compounds based on scaffold similarity and discarded compounds with buried polar groups that were placed in hydrophobic parts of the H1R binding site by visual inspection. In total 74 compounds were purchased and tested for their H1R affinity (including the 26 compounds reported in the previous article21). Table 1 gives an overview of the results and Fig. 2a illustrates the robustness of the assays performed.
Workflow of the virtual-screening approaches performed on both the H1R and the β2R.
The indicated PLANTS and IFP cut-offs for the top 500/750 compounds are indicative, only compounds within the top 500/750 compounds were selected for further processing. Notes: (a) The definition of these cut-offs has been described. (b) An ECFP-4 similarity cut-off of 0.4 and 0.5 was used for the H1R and ß2R selections, respectively. (c) The number before the brackets indicate the number of compounds that are unique to this selection, the number between brackets include compounds from the combined selection that match the criteria of the individual approach (see Fig. 3).
Representative radioligand displacement (H1R, reference compounds 1 and 2) and functional-response (ß2R, reference compound 46) curves of the reference ligands and selected compounds identified using the structure-based virtual screening on H1R (a) and ß2R (b). Curves for selected compounds from each of the scoring approaches curves are presented in Supplementary Fig. S2.
Discovery new β2R agonists
For the selection of the optimal β-adrenoceptor crystal structure we performed a retrospective virtual screening study (in line with the study performed by De Graaf et al.25) on 6 β1R and 7 β2R ligand-bound crystal structures (PDB accessed May 2011)26,27,28,29,30,31,32,33,34. β1R structures were also taken into account because of the high ligand and binding site similarity with β2R35. This retrospective VS analysis highlighted that the (first) active-state β2R crystal structure with the Gs-mimicking nanobody and the full agonist BI-167107 (47) (PDB-code 3P0G, Figs 1 and 3) was found to have one of the highest retrieval rate for agonists (EF1% 76.9) while maintaining a low retrieval rate for antagonists/inverse agonists (EF1% 7.7) when using the IFP scoring approach (see Supplementary Fig. S1). It should be noted that meanwhile, 35 β-adrenoceptor crystal structures have been deposited in the PDB (see Table S1). Systematic retrospective virtual screening studies against 31 of the β-adrenoceptor crystal structures indicated that docking and IFP scoring in 3P0G still gives one of the highest and most selective enrichments for the discrimination of partial/full agonists versus decoy molecules and inverse agonists/antagonists22. We performed a virtual screening on the selected active-state β2R structure using the same approach and fragment library as used for the H1R virtual screening. Based on the cut-offs used for the initial H1R virtual screening a set of only 318 compounds was selected for the combined scoring approach (Table S2). The consistency filter was not applied due to the low number of remaining compounds. A less strict novelty filter than on the H1R screening was applied (ECFP-4 Tanimoto score of ≤0.5 compared to any known ß2R ligand) as the many known ß2R ligands show limited chemical diversity13,25. The compounds for each of the individual scoring methods were selected based on a similar procedure as for the H1R, after which the top 750 scoring compounds for each scoring approach were selected. The final compounds were selected after MACCS and visual clustering of the compounds based on scaffold similarity for each of the scoring methods. During the final selection process priority was given to compounds with a lower ECFP-4 score. It should be noted that despite our efforts to perform this selection process systematically, the difference in size between the individual scoring selections (750 compounds) versus the combined scoring selection (318 compounds) might have introduced a positive bias for the individual approaches during the final compound selection as there was a larger pool of compounds to select from.
In total 63 compounds (Fig. 1) were purchased and experimentally tested for their ability to increase GPCR signalling measured by a cAMP response element (CRE) controlled luciferase reporter gene assay in β2R-transfected HEK293T cells. Table 2 gives an overview of the results and Fig. 2b illustrates the robustness of the assays performed.
Dissecting the contribution for each of the scoring methods
It has been frequently claimed that the combination of two virtual screening approaches resulted in increased performance in retrospective evaluations14,15 and prospective applications16,17,18. However, in these prospective applications the performance of the individual virtual screening approaches has, to the best of our knowledge, never been experimentally validated to allow true comparison of combined and individual virtual screening methods. Here we have validated both our combined PLANTS-IFP scoring approach as well as the individual IFP and PLANTS scoring functions on two different test cases in order to assess if the combination actually performs better than the individual approaches and is not the result of overfitting by retrospective optimization. The systematic evaluation of virtual screening strategies has enabled us to: i) assess the impact of the consensus and single scoring methods on chemical diversity and novelty of the hits, ii) to estimate the target-dependent performance of the approaches and iii) to dissect the contributions of the individual scoring methods to the consensus method.
The experimental validation of the compounds (Fig. 2 and Supplementary Fig. S2) showed that, although the combined-scoring approach was the most successful, the individual approaches resulted in high hit-rates as well. For the H1R VS hit rates of 73%, 61% and 45% were obtained for the combined, IFP and PLANTS approach respectively (Tables 1 and 2, Fig. 3a,c). For the β2R VS the hit rates were 53%, 44% and 39% for the combined, IFP and PLANTS approach respectively (Table 2, Fig. 3b,c). In short, both virtual screens show the same trends: the combined IFP and PLANTS scoring approach is most successful, followed initially by IFP-score ranking and finally by PLANTS-score ranking.
The compound sets for each of the individual scores overlap partially with the compound sets of the combined scoring approach as is illustrated by the Venn diagrams in Fig. 3d. By breaking the compounds sets (and thus hit-rates) down into unique subsets, the contribution of each approach individually can be derived more accurately (Fig. 3c,d) as the differences in hit-rates between the individual approaches are subtle (specifically for the β2R screening). Interestingly, the majority of the β2R hits that were identified in the individual IFP and PLANTS selection were also identified by the combined approach as well (11 of 16 and 8 of 11 hits respectively). Although similar results were observed in the H1R screening, they were less pronounced (7 out of the 20 hits and 4 of 15 hits respectively). This can mainly be ascribed to the fact that the distribution of the IFP and PLANTS scores for the docked compounds differ greatly between the H1R and β2R screening, as can be derived from the IFP versus PLANTS scoring scatterplots in Fig. 1. Overall the PLANTS scores are higher for the H1R than for the β2R and the IFP scores are more widely spread for the H1R (Fig. 1, Table S2). For the IFP scores this can be explained by the difference in size of the reference ligand of both targets. Compound 1 (Fig. 4a) is a fragment-like ligand with only 21 heavy atoms, thereby matching the size of the compounds in the screening library. Compound 47 on the other hand contains 27 heavy atoms thereby making more contacts and interactions than the fragments in the screening library are able to make based on their size. This discrepancy in size logically results in a reduced IFP-score for the compounds in the fragment-library. In the future this effect could be addressed by e.g. taking a different reference ligand using a fragment of the original reference ligand or by using a different similarity coefficient. The PLANTS scoring differences represent the differences in the shape and composition of the binding site36, thereby clearly demonstrating that each protein target benefits from a customized SBVS approach even if two targets are closely related like these two aminergic GPCRs. Thorough retrospective validation and optimization of a virtual screening method is therefore essential to maximize the potency of a method during a prospective application. It must, however, also be recognized that customization of a VS approach often introduces a subjective element (as do the visual clustering and visual inspection steps).
Proposed binding modes of a H1R hit from each of the scoring approaches compared to the X-ray structure.
Binding modes in H1R of (a) the co-crystallized 1 (magenta carbon atoms), (b) combined PLANTS-IFP-scoring hit 3 (orange carbon atoms), (c) PLANTS-scoring hit 35 (green carbon atoms) and (d) an IFP-scoring hit 22 (gold carbon atoms). (e) The interaction fingerprints of the compounds with each of the depicted residues.
The percentages of hits versus inactives for sets matching the combined scoring criteria are consistently higher (Fig. 3c). Moreover, of the tested H1R compounds that are unique to the combined scoring function (section 1 in Fig. 3d) 84.6% were found to be active. On the other hand, for the β2R screening most of the hits (16 of the 26 identified hits) were in the individual IFP selection (sections 3 and 4 in Fig. 3d) of which 69% also intersected with the combined scoring approach. This emphasizes once more that the presence of active compounds compared to inactive compounds is higher in the compound selection based on the combined scoring, as was hypothesized based on the retrospective H1R virtual screening study21. Other successful, non-GPCR, examples of prospective IFP-driven virtual screening studies such as for the Trypanosoma Brucei phosphodiesterase B137 and ligand-gated ion channel GABAA38 also demonstrate the added value of using IFP for the hit identification. Moreover, another advantage from a medicinal chemistry point of view is that the hits obtained through the (combined) use of IFP have reliable predicted binding modes as they are similar to previously observed and experimentally validated binding modes. These binding modes can serve as starting points for further ligand optimization efforts and can even be effectively combined with interaction-annotated chemogenomics databases (such as KLIFS39 for kinases and PDEStrIAn40 for phosphodiesterases) to drive optimization programs37 in order to target specific subpockets or to obtain interaction patterns associated with a specific functional effect22.
A comparative analysis of 29 GPCR structure-based virtual screening studies6,17,41,42,43,44,45,46,47,48 shows that the hit rate of the combined PLANTS-IFP H1R virtual screening study (73%) is the highest reported and the percentage of submicromolar-affinity hits (27%) is amongst the highest reported (the average submicromolar hit rate was 7%), together with the SBVS studies on the 5-HT1B serotonin receptor41 (36%), the α1A-adrenoceptor48 (30%) and the D3 dopamine receptor17 (40%).
Diverse scaffolds with conserved binding modes
Both virtual screening studies have resulted in the identification of novel and (relatively) potent fragment-like small molecules. The ligands identified with the combined or the IFP approach have a high IFP score indicating that they are expected to have similar interactions within the binding pocket, which translates to comparable binding modes obtained with diverse scaffolds. For H1R for example, hits 3, 22 and 1 (as shown in Fig. 4) occupy almost the same space in the binding site and make comparable interactions with the pocket residues despite their different molecular structure. However, many of the H1R hits that are unique for the PLANTS scoring method, e.g. 35 (Fig. 4c), 37, 40 and 45, are linear ligands (shape-wise) that therefore also address part of the minor pocket (between TM 2, 3 and 7) according to their predicted binding mode, unlike 1. This might be the result of the docking scoring function as molecules with more protein-ligand contacts (artificially) obtain higher interaction scores19, whereas molecules selected with IFP are ligands that adopt a similar binding mode (and thus shape) as the reference ligands. It should also be noted that the use of IFP can compensate for the fact that the PLANTS docking was performed with a rigid receptor (except for the hydroxyl groups, which could freely rotate). The PLANTS score does therefore not take binding site flexibility into account that could potentially compensate for less optimal binding poses, while the use of IFP is able to select compounds that do meet the IFP requirements while it prevents potential over-fitting of the pocket to accommodate the ligand docking pose. Since the reference molecule for the β2R screening, 47, is larger than the reference molecule for the H1R screening 1 and already addresses both the major (between TM 3, 5, 6 and 7) and the minor pocket, this effect is not observed for the β2R hits.
Interestingly, an analogue of the most potent hit from our previous study21 (3, pKi = 8.20, Fig. 4b), fragment 42, was selected using PLANTS scoring. In this analogue the piperidine is substituted on the 3-position instead of the 4-position and the benzyl has shifted from the 2 to the 4 position of the phenoxy moiety, resulting in a 400-fold lower affinity. Another hit obtained using PLANTS scoring is the bulky compound 36, which fully occupies the aromatic region between TM5 and TM635 with its 9,10-dihydro-9,10-ethanoanthracene moiety. The most potent H1R hits that were identified are 3 (pKi = 8.20, Fig. 4b), 22 (pKi = 7.05, Fig. 4d) and 35 (pKi = 6.97, Fig. 4c) for the Combined, IFP and PLANTS scoring approach respectively.
For β2R the well-known ethanolamine scaffold is prevalent within the hits (e.g. 50 and 66), however, more than 50% of all identified agonists do not have this scaffold (e.g. 53 and 69, Fig. 5b,d). Interestingly, a few ligands contain groups that are able to form a similar H-bond interaction network with N3127.39 as the conserved ethanolamine alcohol moiety (e.g. 72, Fig. 5c). It should be noted that Christopher et al.49 also reported the identification (and subsequent crystallization with β2R) of fragment-like β2R ligands without an ethanolamine moiety using biophysicial fragment screening. It was suggested by the authors that these fragment-like ligands were expected to be antagonists49. Hit 53 forms an H-bond with S2035.42 via its indole moiety (Fig. 5b). Interestingly, although this indole moiety is also observed in known β2R ligands like pindolol, the indole of 53 is substituted at the 3-position instead of the 4-position (e.g. hit 54). More surprising is that most of the identified agonists do not contain hydrogen-bond donors or acceptors for interacting with the serines in TM5, which were previously deemed essential for activation8,25,32,34,50. However, some of the identified agonists do contain halogen-substituents (e.g. 50 and 52) that could allow for halogen bonding. Strikingly, compounds like 66 are able to increase cAMP formation but do not contain any H-bonding partners for the serines in TM5 but do have a short linker to the aromatic head-group that was previously proposed to play a role in inducing the active-state via aromatic stacking with aromatic residues in TM651. 61 has a 2-(methylamino)-1-phenylethan-1-ol scaffold similar to 66 and 58. Despite the different substituents on this scaffold for 66 (2,4,6-trimethylbenzyl), 61 (methyladamantane), 58 (benzonitrile) all have comparable potencies (pEC50 around 6). However, compound 66 has a significantly higher efficacy (Emax 85% ± 6) compared to the other compounds (Emax ~60%). Based on the binding modes of these ligands (data not shown) this indicates that not only substitutions near TM5 and TM6, but also between TM3 and TM7 can result in a gain in efficacy. Interestingly, 58 is similar to 49 without the ethanolamine moiety. Their binding modes (not shown), however, are very different as the benzonitrile of 49 interacts with S2035.42 whereas it interacts with W3137.40 for 58.
Proposed binding modes of representative β2R hits from each of the scoring approaches compared to the X-ray structure.
Binding modes in β2R of (a) the co-crystallized 47 (salmon carbon atoms), (b) combined PLANTS-IFP-scoring hit 53 (slate carbon atoms), (c) PLANTS-scoring hit 72 (grey carbon atoms) and (d) an IFP-scoring hit 69 (brown carbon atoms). (e) The interaction fingerprints of the compounds with each of the depicted residues.
The agonist with the highest potency of all identified β2R agonists (pEC50 = 7.42), 48, was identified using the combined approach. However, due to the relatively high similarity of 48 to the well-known β2R agonist salbutamol (ECFP-4 similarity = 0.48) this is not surprising. The more novel agonists 50 and 71 have the highest potencies (pEC50 = 6.73 and pEC50 = 6.81) of the hits from the IFP and PLANTS approach, respectively.
It should be noted that the potency of isoprenaline as observed in these assays is relatively low (pEC50 = 6.51), which might be due to oxidation of the catechol moiety. We tried to address this using ascorbic acid (as was previously proposed52) but this unfortunately interfered with the reporter gene assay. It is noteworthy that the potency of isoprenaline varies throughout scientific publications and seem to be highly assay dependent, as previously reported potencies (pEC50) for isoprenaline range from 6.6253 to 10.113.
The combined scoring approach did select the compound with the highest affinity for H1R and potency for β2R. Despite this, there does not seem to be a clear correlation between affinity (pKi), potency (pEC50) and efficacy (Emax) when analysing the hits from the three selection approaches (Fig. 3a,b). Based on the ECFP-4 similarity compound 28 is the most novel hit for H1R. Although the structure of compound 28 is different from most known H1R ligands, the typical H1R pharmacophore elements can still be recognized. Compound 43, the second most novel H1R hit, on the other hand, has a very different scaffold with its furan, tetrahydrofuran and chlorobenzene moieties. From the identified β2R hits compound 56 is the most novel hit according to its ECFP-4 similarity to known β2R ligands. Interestingly, 56 has a similar scaffold as the novel H1R ligand 43 but the tetrahydrofuran group has been replaced with a thiophene and the chlorine atom was removed.
When comparing the entire screening library to the co-crystallized reference ligand using 2D topological (ECFP-4) and 3D shape-based (ROCS) similarity searches we observe two very different distributions for the targets (Fig. 6). This can be ascribed to the differences in size, shape and pharmacophore features of the two reference ligands, 1 and 47. As 1 itself also matches the fragment-like criteria of the screening library the overall similarity is much higher (Fig. 6a). The higher heavy atom count and thus volume of 47 results in a generally lower ECFP-4 and ROCS scores for the compounds from the fragments library (Fig. 6b). For the H1R hits only 3 of the identified ligands were present in the top 500 as selected by ROCS and 0 as selected by ECFP-4 (Fig. 6a). For β2R 7 and 4 of the ligands were present in the top 750 based on ROCS and ECFP-4 scoring, respectively, albeit with low similarity scores (Fig. 6b).
2D/3D ligand-based virtual screening on the fragment library using (a) the H1R reference ligand 1 and (b) the β2R reference ligand 47. The experimentally validated hits (blue) as well as all screened fragments (grey) are scattered based on their ECFP-424 and ROCS similarity to the doxepin reference ligand. The horizontal dotted line indicates the cut-off21 of 1.4 for the ROCS ComboScore. The vertical dotted lines indicate the cut-offs21 of 0.26 and 0.40 for ECFP-4.
We furthermore assessed the novelty of the identified ligands by performing SEA54 predictions (functional and binding) and the ChEMBL target prediction models in myChEMBL55 (using both the 10 μM and 1 μM models). The ChEMBL models predicted 1 of the 26 β2R SBVS hits as a human β2R ligand, another 2 hits as human β1R ligands and 4 of the 43 H1R SBVS hits as human H1R ligands. SEA (binding and functional combined) predicted 1 of the 26 β2R SBVS hits as a human β2R ligand, 2 as human β1R ligands and another 2 as animal β2R ligands and none of the 43 H1R SBVS hits as H1R ligand.
Predicting both ligand binding affinity and functional activity
The β2R screening tries to combine the identification of new compounds with affinity for β2R as well as predicting the functional activity by only searching for β2R agonists. Only limited studies with a similar goal have been performed on GPCRs of which most were retrospective analyses8,25,50. However, as described in the introduction, more recently a prospective virtual screening for β2 as well as D2 receptor agonists was published using the same active-state β2R structure (PDB 3P0G) as used in this study. Weiss et al. performed a systematic prospective study to investigate the effect of receptor conformation on virtual screening13. In this study, a library of 2.7 million lead-like and 0.4 million fragment-like molecules from the ZINC database was screened against the active-state 47-bound β2R crystal structure (PDB 3P0G) and carazolol-bound crystal structure (PDB 2RH1)13. During the docking the dipole moment of S2035.42, S2045.43, or S2075.46 was increased to augment docking scores for poses in polar contact with these residues. Compounds ranking within the top 0.2% of the active-state structure and ranking at least 5000 positions higher for the active-state compared to the carazolol-bound structure were selected for further processing. Compounds that had at least a positive charge, an ionic interaction with D1133.32 and at least one H-bond with any of the three aforementioned serines were visually inspected. In total 5 fragment-like and 17 lead-like molecules were experimentally validated resulting in the identification of 1 fragment-like and 5 lead-like β2R agonists, of which 1 known β2R agonist and 3 compounds with an ECFP-4 similarity higher than 0.4. Moreover, based on a homology model of an active-state D2 receptor a similar virtual screening was performed with the aim to identify new D2 agonists. 15 compounds were selected for experimental validation of which 3 were found to be hits with marginal potencies (of which 2 with an ECFP-4 similarity higher than 0.4): 2 agonists and 1 antagonist.
Although most structure-based virtual screenings tend to identify ligands with the same functional effect as the co-crystallized ligand or the ligand(s) used to refine a homology model21,43,44,45,46, there have been multiple prospective virtual screenings based on homology models of GPCRs that have resulted in the identification of ligands with a different functional effect13,17,42,47.
Despite the difficulties that come with these types of predictions, as illustrated by the examples above, we were able to selectively identify 26 β2R agonists. It should be noted that the β2R was also the ideal receptor for such an analysis, as much information is available for this receptor including (and most importantly) multiple crystal structures in different activation states and in complex with both agonists and antagonists/inverse agonists5,9,26,27,28,29,30,31,32,33,34. Generally, aminergic GPCRs have a deep and well-defined binding site without a large solvent-exposed area which make them suitable for docking simulations as demonstrated by other successful prospective structure-based virtual screening studies against aminergic GPCR crystal structures6,17,41,44,45,46,48. Other GPCRs such as, for example, the chemokine receptors CXCR456 and CCR557, have a larger and more open pocket, which provide challenges with respect to conformational sampling in molecular docking simulations and subsequent scoring of docking poses and structure-based virtual screening against chemokine receptors crystal structures have so far resulted in lower hit rates of larger hits with lower affinity6,43,58. Moreover, for many GPCRs structure-function relationships are not as well defined as for beta-adrenergic receptors59, thereby preventing the training of predictive models such as the one presented in the current study. On the other hand, the growing amount of GPCR crystal structures in complex with different ligands will provide structural information to guide and optimize ligand function-specific structure-based virtual screening studies against more and more GPCR targets. At this point in time, function-selective structure-based screenings such as described here are already in reach for the A2A60, M261, P2Y1262 and μ-opioid63 receptor for which both small molecule agonist and antagonist/inverse agonist bound crystal structures are available. As the insights for the different signalling pathways that ligands can block or induce is growing64 (together with the aforementioned availability of GPCR crystal structures) this could also open up new opportunities65 to finding ligands portraying specific biased signalling profiles4,66.
Conclusions
Conventional docking approaches are hampered by challenges in the prediction of the right binding mode and the correct ranking of those binding modes. To overcome these hurdles we have devised a novel docking scoring approach that combines a conventional energy-based scoring function with an interaction-profile-based rescoring approach. This approach was successfully applied by prospectively screening a fragment-like compound library on two GPCRs for which crystal structures are available. For both the H1 and β2 receptor high hit-rates were obtained. Moreover, besides the consensus approach also the individual approaches were experimentally validated in order to evaluate if the combination indeed resulted in an increased performance (hit-rates) in a prospective manner. Although the individual scoring approaches were effective as well, the combined approach did result in increased hit-rates and the retrieval of ligands with up to nanomolar affinities and potencies. It should also be noted that despite the fact that the combined scoring approach was proven to be effective for multiple protein targets, the scoring distributions for the targets did highlight a highly-target-specific score distribution due to the pocket composition and the impact of the reference ligand. It is therefore recommended to optimize this combined approach for each targeted protein using careful retrospective validation to e.g. optimize the docking scoring function and IFP scoring cut-offs. Accordingly, if multiple crystal structures are available, the selection of the right structure (or combination of structures13) will influence the outcome of the virtual screening in terms of retrieval rates but also functional effect of the obtained hits. For the β2R we were able to selectively retrieve novel fragment-like ligands with the desired functional effect. Moreover, this approach led to the identification of novel scaffolds for β2R agonists. Surprisingly, some of these hit compounds did not contain hydrogen bond acceptors or donors that would be able to interact with serines S2035.42, S2045.43, or S2075.46 in TM5, previously thought to be crucial for β2R activation8,25,32,34,50. The results show that the advances in GPCR crystallography open up new opportunities to selectively discover new GPCR ligands with the desired functional effect. These advances could ultimately lead to the prediction and structure-based optimization of ligands with designed (biased) signalling profiles4,65,66.
Experimental Procedures
Residue numbering and nomenclature
The Ballesteros–Weinstein residue numbering scheme67 was used throughout this manuscript. For residues in specific receptors, the UniProt residue number is given before the Ballesteros–Weinstein residue number in superscript (e.g. D1073.32 in H1R).
Preparation of prospective virtual screening database
The compound libraries of 15 vendors were obtained from the ZINC database in SMILES format totaling ~13 million unique compounds. Openeye’s filter (version 2.1.1) was used to only select fragment-like compounds were resulting in a subset of 757.728 compounds. Tauthor (version 1.4.90) and Blabber (version 1.4.90) from MolDiscovery’s MoKa package were used to compute plausible tautomers and protonation states. Subsequently, another filter was applied to remove all compounds without a positive formal charge to ensure only compounds, which could potentially form an ionic bond with key residue D3.32 were selected. These steps resulted in a final prospective virtual screening library comprising 108 790 compounds.
Automated docking and IFP post-processing
The dockings were performed using PLANTS19 and the resulting H1R and β-adrenergic docking poses were post-processed and ranked with IFP20,21. Using PLANTS 25 docking poses for each compound were calculated (speed setting 2) and scored using the ChemPLP scoring function. The docking site of β2R and H1R was defined by the coordinates of the center of cocrystallized ligand (BI-167107 and doxepin, respectively) and a radius around it based on the maximum distance from this center to the edge of the crystallized ligand +5 Å. All other options of PLANTS were left at their default setting. PLANTS employs an ant-colony-optimization algorithm for the prediction of binding poses of small molecules in a protein structure and an empirical scoring function, ChemPLP, for the scoring of the resulting binding poses. IFP evaluates a (predicted) binding mode of a compound in a protein structure by annotating the absence or presence of different types of interactions (hydrophobic, aromatic, H-bond, ionic) between each pocket residue and the molecule based on a set of rules20. This results in a molecular interaction fingerprint representing all interactions between the molecule and the protein in bit-string, allowing for the easy comparison and scoring (using the Tanimoto coefficient) of the similarity of multiple IFPs. The H1R and β-adrenoceptor pockets were defined by 33 pocket residues based on the consensus pocket definition by de Graaf et al.25: L/M1.35, L/M1.39, I1.42, T/I1.46, V2.57, M/V2.58, N/G2.61, L/I2.65, W3.28, L/T3.29, D3.32, Y/V3.33, S/V3.36, T3.37, I3.40, W/T4.56, I/P4.60, F/Y5.38, K/A5.39, T/S5.42, A/S5.43, N/S5.46, F5.47, F6.44, W6.48, Y/F6.51, F6.52, F/N6.55, H/Y7.35, I/N7.39, W7.40, Y7.43, N7.45 21. In the β1R/β2R retrospective validation the binding mode of the co-crystallized compound for each respective crystal structure was used for the calculation of the reference IFP. These reference IFPs were subsequently used to score the docking poses.
Retrospective virtual screening databases and analysis
The H1R retrospective validation was performed by docking a compound library comprising 543 known H1R ligands from ChEMBL, 59 CNS active drugs acting as inverse agonists on H1R and 7 088 physicochemically similar decoy molecules into the H1R crystal structure. Subsequently all docking poses were postprocessed using IFP and only docking poses in which the compounds made an ionic interaction with D1073.32 were analyzed. For each compound the best PLANTS and the best IFP score were selected and used for defining the score cut-offs for prospective application. With a PLANTS score cut-off of ≤−90 and an IFP score cut-off ≥0.75 high enrichment factors for the ligands over the decoys were obtained (EF1% of 39.3 and 57.6 for the ChEMBL and CNS H1R ligands respectively)21. The test set by de Graaf et al.25 was used for the retrospective virtual screening study on all 6 β1R and 7 β2R crystal structures available at that time (PDB accessed May 2011). This test set exists of 13 agonists, 13 antagonist/inverse agonists and 980 physicochemically-similar decoys and was extended with 7 agonists and 8 antagonists/inverse agonists from Baker et al.68. From each β-adrenergic crystal structure one chain was selected and used for the retrospective validation. The test set was docked into all selected chains and scored using PLANTS and IFP. Subsequently, the enrichment factors at a 1% false positive rate for the f/pAGO and ANT/iAGO over decoys were determined for each of the crystal structures (Supplementary Fig. S1).
Prospective virtual screening
The PLANTS and IFP scoring cut-offs (as described in the previous paragraph) were used for prospective virtual screening of a library of 108 790 fragment-like basic molecules21 against both H1R and β2R targets. For the combined scoring approach the filtering was performed by applying the D3.32 interaction filter, the PLANTS and IFP cut offs, the consistency cut off in case of H1R (only compounds with an IFP score of ≥0.7 according to the best PLANTS pose as well as a PLANTS score of ≤−75 according to the best IFP pose were kept) and a novelty filter (ECFP-4 ≤ 0.4 for H1R and ECFP-4 ≤ 0.5 for β2R). H1R and β2R hit selection consisted of a visual clustering of the remaining compounds, after which from each cluster the fragment with the highest IFP and/or PLANTS score was selected and fragments with docking poses in which polar atoms were buried in hydrophobic parts of the binding site were discarded21. For the individual PLANTS and IFP scoring approaches the top 500 (H1R) and top 750 (β2R) scoring compounds were selected and compounds matching the combined approach were removed. Hits that were similar to any known ligand of the respective H1R (ECFP-4 Tanimoto score of 0.4) or β2R (ECFP-4 Tanimoto score of ≥0.5) receptors were removed, compounds were visually clustered based on scaffold similarity and compounds with buried polar groups that were placed in hydrophobic parts of the receptor binding site were discarded by visual inspection.
ROCS 3D similarity search
The conformer database was generated using standard settings OMEGA (version 2.3.2; OpenEye Scientific Software: Santa Fe, NM.) and searched with ROCS (version 2.3.1; OpenEye Scientific Software: Santa Fe, NM.) using standard settings as well. The conformation of 1 found in the H1R X-ray structure (PDB-code 3RZE23) and the conformation of 47 in the β2R X-ray structure (PDB-code 3P0G) were used as query molecules for independent ROCS runs. The compounds from the screening library were ranked by decreasing Comboscore (combination of shape Tanimoto and the normalized colour score in this optimized overlay).
ECFP-4 2D similarity search
Two-dimensional similarity searches were carried out using ECFP-4 (extended connectivity fingerprints24) descriptors available in Pipeline Pilot (version 6.1.5; Accelrys Software Inc.: 5005 Waterridge Vista Dr, San Diego, CA 92121, United States.) and compared using the Tanimoto coefficient.
Compounds selected by virtual screening
The compounds selected by virtual screening were purchased from available screening collections of six vendors: Asinex (www.asinex.com), Chembridge (www.Hit2Lead.com), Enamine (www.enamine.com), IBScreen (www.ibscreen.com), Matrix Scientific (www.matrixscientific.com), Vitas-M (www.vitasmlab.com). Suppliers and supplier identifiers for each of the virtual screening hits are given in Table S3. The purity of all compounds was verified by liquid chromatography-mass spectrometry (LC-MS), all experimentally validated hits had a purity of 90% or higher (see Table S4), except compounds 40, 26 and 44, which in our hands had a purity of 75%, 76% and 88% respectively (reported to be at least 90% pure according to the suppliers).
Materials
Human H1R cDNA was kindly provided by Dr. H Fukui (Japan). The cDNA clone for Human β2R in pcDNA3.1+ was obtained from Missouri S&T cDNA Resource Center (www.cdna.org). Cell culture media were purchased from PAA (Pasching, Austria). Isoproterenol was bought from Sigma-Aldrich (St. Louis, MO, USA). Compounds used in the assays were obtained from different suppliers (see Supplementary Table S4). The cDNA clone for Human β2R in pcDNA3.1+ was obtained from Missouri S&T cDNA Resource Center (www.cdna.org).
Cell culture and transfection
The cell culture and transfection for H1R was performed as previously described21. HEK293T cells were cultured in Dulbecco’s modified Eagle medium (DMEM) supplemented with 10% fetal bovine serum, 50 IU/ml penicillin and 50 μg/ml streptomycin at 37 °C and 5% CO2. Approximately 2 × 106 million cells were seeded per 10-cm dish 1 day prior to transfection. Approximately 4 × 106 cells were transiently transfected with 5 μg of cDNA using 25 the polyethylenimine (PEI) method. Briefly, 10 ng β2R cDNA, 2,5 μg CRE-luc plasmid and 2.490 μg empty pcDNA3.1 were mixed with 20 μg of 25 kDa linear PEI (Polysciences, Warrington, PA, U.S.) in 500 μL of 150 mM NaCl. The transfection mix was incubated at 22 °C for 30 min. Meanwhile, medium in the 10 cm dish was replaced with fresh culture medium and transfection mix was subsequently added drop-wise to the cells. The next day, cells were collected and transferred to white-bottomed 96-well plates (50,000 cells/well).
Radioligand displacement assay
The radioligand displacement assay for H1R was performed as previously described21.
CRE (cyclic AMP response element) luciferase reporter gene assay
Two days after transfection, the medium was removed and the cells were stimulated for 6 h with ligands in serum-free DMEM supplemented with 1 mM thiourea to slow the oxidation of compounds, at 37 °C, 5% CO2. After 6 h, the medium was aspirated and 25 μl of luciferase assay reagent (LAR, 0.83 mM ATP, 0.83 mM D-luciferine, 18.7 mM MgCl2, 0.78 μM Na2HPO4, 38.9 mM Tris-H3PO4 (pH 7.8), 0.39% glycerol, 0.03% Triton X-100 and 2.6 μM dithiotreitol) was added to each well. Luminescence (1 s per well) was measured in a Victor3 1420 multi-label reader (Perkin Elmer Life and Analytical Sciences) after 30 min of incubation at 37 °C, 5% CO2. Non-linear curve-fitting and statistical analysis were performed using GraphPad Prism 6. Results are shown from pooled data (mean ± SEM) from at least three independent experiments performed in triplicate.
Additional Information
How to cite this article: Kooistra, A. J. et al. Function-specific virtual screening for GPCR ligands using a combined scoring method. Sci. Rep. 6, 28288; doi: 10.1038/srep28288 (2016).
References
Katritch, V., Cherezov, V. & Stevens, R. C. Structure-function of the G protein-coupled receptor superfamily. Annu. Rev. Pharmacol. Toxicol. 53, 531–556 (2013).
Piscitelli, C. L., Kean, J., de Graaf, C. & Deupi, X. A Molecular Pharmacologist’s Guide to G Protein-Coupled Receptor Crystallography. Mol. Pharmacol. 88, 536–551 (2015).
Congreve, M., Dias, J. M. & Marshall, F. H. Structure-based drug design for g protein-coupled receptors. Prog. Med. Chem. 53, 1–63 (2014).
Rodriguez, D., Ranganathan, A. & Carlsson, J. Discovery of GPCR Ligands by Molecular Docking Screening: Novel Opportunities Provided by Crystal Structures. Curr. Top. Med. Chem. 15, 2484–2503 (2015).
Shoichet, B. K. & Kobilka, B. K. Structure-based drug screening for G-protein-coupled receptors. Trends Pharmacol. Sci. 33, 268–272 (2012).
Kooistra, A. J., Leurs, R., de Esch, I. J. & de Graaf, C. From three-dimensional GPCR structure to rational ligand discovery. Adv. Exp. Med. Biol. 796, 129–157 (2014).
Vass, M., Schmidt, E., Horti, F. & Keseru, G. M. Virtual fragment screening on GPCRs: a case study on dopamine D3 and histamine H4 receptors. Eur. J. Med. Chem. 77, 38–46 (2014).
Katritch, V. et al. Analysis of full and partial agonists binding to beta2-adrenergic receptor suggests a role of transmembrane helix V in agonist-specific conformational changes. J. Mol. Recognit. 22, 307–318 (2009).
Lamichhane, R. et al. Single-molecule view of basal activity and activation mechanisms of the G protein-coupled receptor beta2AR. Proc. Natl. Acad. Sci. USA 112, 14254–14259 (2015).
Tehan, B. G., Bortolato, A., Blaney, F. E., Weir, M. P. & Mason, J. S. Unifying Family A GPCR Theories of Activation. Pharmacol. Ther. 143, 51–60 (2014).
Verdonk, M. L. et al. Docking performance of fragments and druglike compounds. J. Med. Chem. 54, 5422–5431 (2011).
Rodriguez, D., Gao, Z. G., Moss, S. M., Jacobson, K. A. & Carlsson, J. Molecular docking screening using agonist-bound GPCR structures: Probing the A2A adenosine receptor. J. Chem. Inf. Model. 55, 550–563 (2015).
Weiss, D. R. et al. Conformation guides molecular efficacy in docking screens of activated beta-2 adrenergic G protein coupled receptor. ACS Chem. Biol. 8, 1018–1026 (2013).
O’Boyle, N. M., Liebeschuetz, J. W. & Cole, J. C. Testing assumptions and hypotheses for rescoring success in protein-ligand docking. J. Chem. Inf. Model. 49, 1871–1878 (2009).
Svensson, F., Karlen, A. & Skold, C. Virtual screening data fusion using both structure- and ligand-based methods. J. Chem. Inf. Model. 52, 225–232 (2012).
Sanders, M. P. et al. A prospective cross-screening study on G-protein-coupled receptors: lessons learned in virtual compound library design. J. Med. Chem. 55, 5311–5325 (2012).
Varady, J. et al. Molecular modeling of the three-dimensional structure of dopamine 3 (D3) subtype receptor: discovery of novel and potent D3 ligands through a hybrid pharmacophore- and structure-based database searching approach. J. Med. Chem. 46, 4377–4392 (2003).
Park, H., Eom, J. W. & Kim, Y. H. Consensus scoring approach to identify the inhibitors of AMP-activated protein kinase alpha2 with virtual screening. J. Chem. Inf. Model. 54, 2139–2146 (2014).
Korb, O., Stutzle, T. & Exner, T. E. Empirical scoring functions for advanced protein-ligand docking with PLANTS. J. Chem. Inf. Model. 49, 84–96 (2009).
Marcou, G. & Rognan, D. Optimizing fragment and scaffold docking by use of molecular interaction fingerprints. J. Chem. Inf. Model. 47, 195–207 (2007).
de Graaf, C. et al. Crystal structure-based virtual screening for fragment-like ligands of the human histamine H(1) receptor. J. Med. Chem. 54, 8195–8206 (2011).
Kooistra, A. J., Leurs, R., de Esch, I. J. & de Graaf, C. Structure-Based Prediction of G-Protein-Coupled Receptor Ligand Function: A β-Adrenoceptor Case Study. J. Chem. Inf. Model. 55, 1045–1061 (2015).
Shimamura, T. et al. Structure of the human histamine H1 receptor complex with doxepin. Nature 475, 65–70 (2011).
Rogers, D. & Hahn, M. Extended-connectivity fingerprints. J. Chem. Inf. Model. 50, 742–754 (2010).
de Graaf, C. & Rognan, D. Selective structure-based virtual screening for full and partial agonists of the beta2 adrenergic receptor. J. Med. Chem. 51, 4978–4985 (2008).
Cherezov, V. et al. High-resolution crystal structure of an engineered human beta2-adrenergic G protein-coupled receptor. Science 318, 1258–1265 (2007).
Rasmussen, S. G. et al. Crystal structure of the human beta2 adrenergic G-protein-coupled receptor. Nature 450, 383–387 (2007).
Hanson, M. A. et al. A specific cholesterol binding site is established by the 2.8 A structure of the human beta2-adrenergic receptor. Structure 16, 897–905 (2008).
Warne, T. et al. Structure of a beta1-adrenergic G-protein-coupled receptor. Nature 454, 486–491 (2008).
Bokoch, M. P. et al. Ligand-specific regulation of the extracellular surface of a G-protein-coupled receptor. Nature 463, 108–112 (2010).
Wacker, D. et al. Conserved binding mode of human beta2 adrenergic receptor inverse agonists and antagonist revealed by X-ray crystallography. J. Am. Chem. Soc. 132, 11443–11445 (2010).
Rasmussen, S. G. et al. Structure of a nanobody-stabilized active state of the beta(2) adrenoceptor. Nature 469, 175–180 (2011).
Rosenbaum, D. M. et al. Structure and function of an irreversible agonist-beta(2) adrenoceptor complex. Nature 469, 236–240 (2011).
Warne, T. et al. The structural basis for agonist and partial agonist action on a beta(1)-adrenergic receptor. Nature 469, 241–244 (2011).
Kooistra, A. J., Kuhne, S., de Esch, I. J., Leurs, R. & de Graaf, C. A structural chemogenomics analysis of aminergic GPCRs: lessons for histamine receptor ligand design. Br. J. Pharmacol. 170, 101–126 (2013).
Kellenberger, E., Rodrigo, J., Muller, P. & Rognan, D. Comparative evaluation of eight docking tools for docking and virtual screening accuracy. Proteins 57, 225–242 (2004).
Jansen, C. et al. Discovery of novel Trypanosoma brucei phosphodiesterase B1 inhibitors by virtual screening against the unliganded TbrPDEB1 crystal structure. J. Med. Chem. 56, 2087–2096 (2013).
Richter, L. et al. Diazepam-bound GABAA receptor models identify new benzodiazepine binding-site ligands. Nat. Chem. Biol. 8, 455–464 (2012).
Kooistra, A. J. et al. KLIFS: a structural kinase-ligand interaction database. Nucleic Acids Res. 44, D365–371 (2016).
Jansen, C. et al. PDEStrIAn: A Phosphodiesterase Structure and Ligand Interaction Annotated Database As a Tool for Structure-Based Drug Design. J. Med. Chem. 10.1021/acs.jmedchem.5b01813 (2016).
Rodriguez, D., Brea, J., Loza, M. I. & Carlsson, J. Structure-Based Discovery of Selective Serotonin 5-HT1B Receptor Ligands. Structure 22, 1140–1151 (2014).
Tikhonova, I. G. et al. Discovery of novel agonists and antagonists of the free fatty acid receptor 1 (FFAR1) using virtual screening. J. Med. Chem. 51, 625–633 (2008).
Mysinger, M. M. et al. Structure-based ligand discovery for the protein-protein interface of chemokine receptor CXCR4. Proc. Natl. Acad. Sci. USA 109, 5517–5522 (2012).
Carlsson, J. et al. Ligand discovery from a dopamine D3 receptor homology model and crystal structure. Nat. Chem. Biol. 7, 769–778 (2011).
Katritch, V. et al. Structure-based discovery of novel chemotypes for adenosine A(2A) receptor antagonists. J. Med. Chem. 53, 1799–1809 (2010).
Kolb, P. et al. Structure-based discovery of beta2-adrenergic receptor ligands. Proc. Natl. Acad. Sci. USA 106, 6843–6848 (2009).
Kellenberger, E. et al. Identification of nonpeptide CCR5 receptor agonists by structure-based virtual screening. J. Med. Chem. 50, 1294–1303 (2007).
Evers, A. & Klabunde, T. Structure-based drug discovery using GPCR homology modeling: successful virtual screening for antagonists of the alpha1A adrenergic receptor. J. Med. Chem. 48, 1088–1097 (2005).
Christopher, J. A. et al. Biophysical fragment screening of the beta1-adrenergic receptor: identification of high affinity arylpiperazine leads using structure-based drug design. J. Med. Chem. 56, 3446–3455 (2013).
Reynolds, K. A., Katritch, V. & Abagyan, R. Identifying conformational changes of the beta(2) adrenoceptor that enable accurate prediction of ligand/receptor interactions and screening for GPCR modulators. J. Comput. Aided Mol. Des. 23, 273–288 (2009).
Liu, J. J., Horst, R., Katritch, V., Stevens, R. C. & Wuthrich, K. Biased signaling pathways in beta2-adrenergic receptor characterized by 19F-NMR. Science 335, 1106–1110 (2012).
Barber, R., Kelly, L. A., McGuire, R. F. & Butcher, R. W. Distortion of cyclic AMP responses to catecholamine due to destruction of the hormone. J. Cyclic Nucleotide Res. 3, 249–261 (1977).
Soriano-Ursua, M. A. et al. Cell-based and in-silico studies on the high intrinsic activity of two boron-containing salbutamol derivatives at the human beta(2)-adrenoceptor. Bioorg. Med. Chem. 20, 933–941 (2012).
Keiser, M. J. et al. Relating protein pharmacology by ligand chemistry. Nat. Biotechnol. 25, 197–206 (2007).
Ochoa, R., Davies, M., Papadatos, G., Atkinson, F. & Overington, J. P. myChEMBL: a virtual machine implementation of open data and cheminformatics tools. Bioinformatics 30, 298–300 (2014).
Wu, B. et al. Structures of the CXCR4 chemokine GPCR with small-molecule and cyclic peptide antagonists. Science 330, 1066–1071 (2010).
Tan, Q. et al. Structure of the CCR5 chemokine receptor-HIV entry inhibitor maraviroc complex. Science 341, 1387–1390 (2013).
Schmidt, D., Bernat, V., Brox, R., Tschammer, N. & Kolb, P. Identifying modulators of CXC receptors 3 and 4 with tailored selectivity using multi-target docking. ACS Chem. Biol. 10, 715–724 (2015).
Huang, X. P. et al. Allosteric ligands for the pharmacologically dark receptors GPR68 and GPR65. Nature 527, 477–483 (2015).
Xu, F. et al. Structure of an agonist-bound human A2A adenosine receptor. Science 332, 322–327 (2011).
Kruse, A. C. et al. Activation and allosteric modulation of a muscarinic acetylcholine receptor. Nature 504, 101–106 (2013).
Zhang, J. et al. Agonist-bound structure of the human P2Y12 receptor. Nature 509, 119–122 (2014).
Huang, W. et al. Structural insights into micro-opioid receptor activation. Nature 524, 315–321 (2015).
Kahsai, A. W. et al. Multiple ligand-specific conformations of the beta2-adrenergic receptor. Nat. Chem. Biol. 7, 692–700 (2011).
Marti-Solano, M., Schmidt, D., Kolb, P. & Selent, J. Drugging specific conformational states of GPCRs: challenges and opportunities for computational chemistry. Drug Discov. Today 21, 625–631 (2016).
van der Westhuizen, E. T., Breton, B., Christopoulos, A. & Bouvier, M. Quantification of ligand bias for clinically relevant beta2-adrenergic receptor ligands: implications for drug taxonomy. Mol. Pharmacol. 85, 492–509 (2014).
Ballesteros, J. A. & Weinstein, H. Integrated methods for the construction of three-dimensional models and computational probing of structure-function relations of G protein-coupled receptors. Methods Neurosci. 25, 366–428 (1995).
Baker, J. G. The selectivity of beta-adrenoceptor agonists at human beta1-, beta2- and beta3-adrenoceptors. Br. J. Pharmacol. 160, 1048–1061 (2010).
Grant, J. A., Gallardo, M. A. & Pickup, B. T. A fast method of molecular shape comparison: A simple application of a Gaussian description of molecular shape. J. Comput. Chem. 17, 1653–1666 (1996).
Acknowledgements
We thank Martien Kuijer and Mitchell Han for assistance with radioligand displacement and reporter gene assays and Niels J. Hauwert and Eric Karssen for performing the quality control of the compounds. This research was financially supported by The Netherlands Organization for Scientific Research (NWO VENI grant 700.59.408 to C.d.G. and NWO CW TOP-PUNT grant 718.014.002, 7 ways to 7TMR modulation (7-to-7), to R.L.) and the Netherlands eScience Center (NLeSC)/NWO (Enabling Technologies project: 3D-e-Chem, grant 027.014.201). A.J.K., H.F.V., D.M., I.J.P.d.E., R.L. and C.d.G. participate in the European Cooperation in Science and Technology Action CM1207 [GPCR-Ligand Interactions, Structures and Transmembrane Signalling: A European Research Network (GLISTEN)].
Author information
Authors and Affiliations
Contributions
A.J.K. and C.d.G. designed the research project. A.J.K. and C.d.G. performed the in silico experiments, H.F.V. and D.M.-F. performed the in vitro experiments. A.J.K., H.F.V., D.M.-F. and C.d.G. analyzed the data. H.F.V., I.J.P.d.E., R.L. and C.d.G. provided expertise regarding the experiments and interpretation of the results. A.J.K., I.J.P.d.E., R.L. and C.d.G. wrote the article.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Kooistra, A., Vischer, H., McNaught-Flores, D. et al. Function-specific virtual screening for GPCR ligands using a combined scoring method. Sci Rep 6, 28288 (2016). https://doi.org/10.1038/srep28288
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep28288
This article is cited by
-
Consensus scoring evaluated using the GPCR-Bench dataset: Reconsidering the role of MM/GBSA
Journal of Computer-Aided Molecular Design (2022)
-
Comparison of structure- and ligand-based scoring functions for deep generative models: a GPCR case study
Journal of Cheminformatics (2021)
-
New machine learning and physics-based scoring functions for drug discovery
Scientific Reports (2021)
-
G protein-coupled receptors: structure- and function-based drug discovery
Signal Transduction and Targeted Therapy (2021)
-
Ultra-large library docking for discovering new chemotypes
Nature (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.