Abstract
Long non-coding RNAs (lncRNAs) represent an emerging layer of cancer biology and have been implicated in the development and progression of cancers. However, the prognostic significance of lncRNAs in diffuse large-B-cell lymphoma (DLBCL) remains unclear and needs to be systematically investigated. In this study, we obtained and analyzed lncRNA expression profiles in three cohorts of 1043 DLBCL patients by repurposing the publicly available microarray datasets from the Gene Expression Omnibus (GEO) database. In the discovery series of 207 patients, we identified a set of six lncRNAs that was significantly associated with patients’ overall survival (OS) using univariate Cox regression analysis. The six prognostic lncRNAs were combined to form an expression-based six-lncRNA signature which classified patients of the discovery series into the high-risk group and low-risk group with significantly different survival outcome (HR = 2.31, 95% CI = 1.8 to 2.965, p < 0.001). The six-lncRNA signature was further confirmed in the internal testing series and two additional independent datasets with different array platform. Moreover, the prognostic value of the six-lncRNA signature is independent of conventional clinical factors. Functional analysis suggested that six-lncRNA signature may be involved with DLBCL through exerting their regulatory roles in known cancer-related pathways, immune system and signaling molecules interaction.
Similar content being viewed by others
Introduction
Diffuse large B-cell lymphoma (DLBCL) is the most common and aggressive subtype of non-Hodgkin lymphoma (NHL), constituting 30–58% of all diagnosed NHL cases1. The current standard treatment of DLBCL is a combination of Rituximab with traditional chemotherapy of cyclophosphamide-doxorubicin-vincristine-prednisone (R-CHOP). Although R-CHOP has been proven to be an effective treatment for this disease, nearly 40% of DLBCL patients still faced the failure of standard therapy and ultimately died from their disease2. Therefore, it is important to make risk stratification for DLBCL patients to identify high-risk patients who are unlikely to be cured with standard therapy and would therefore benefit from more effective therapy.
Recent advances in genomic and transcriptomic analysis have accelerated the discovery and identification of various types of non-coding RNAs (ncRNAs). Long non-coding RNAs, a recently discovered major class of ncRNAs, were arbitrarily defined as RNA transcripts of more than 200 bp in length that lack or have little protein coding capacity3. Growing evidence indicated that lncRNAs can function as a critical player of genome regulatory network to participate in the process of gene regulation, post-transcriptional regulation and epigenetic regulation4,5. Dysregulated lncRNAs expression have been observed frequently in tumors when compared to normal adjacent tissues, implying their oncogenic and tumor suppressor roles in cancer development6,7. Recently, transcriptome sequencing analysis has revealed the aberrant expression of lncRNAs between DLBCL and normal B cells8. Another lncRNA, PEG10, was reported to be unregulated in DLBCL compared with normal tissues9, implying their perspective in diagnostics and prognosis as potential biomarkers in DLBCL. However, the prognostic significance of lncRNAs in DLBCL remains unclear and needs to be systematically investigated.
In order to study the prognostic significance of lncRNAs for risk stratification of DLBCL, we obtained and analyzed lncRNA expression profiles on a large number of DLBCL patients by repurposing the publicly available microarray datasets from the Gene Expression Omnibus (GEO) database. Our analysis identified six prognostic lncRNAs that were significantly associated with survival outcome of DLBCL patients from the discovery series by using Cox regression analysis. We then developed a six-lncRNA signature by the risk score model method based on the expression levels of these six prognostic lncRNAs which could distinguish patients with good and poor survival. Moreover, additional results using the internal testing series and two independent non-overlapped patient datasets further confirmed the prognostic value of the six-lncRNA signature.
Results
Initial prognostic lncRNAs screening using lncRNA expression profiles and survival data in the discovery series
The 414 DLBCL patients from Lenz’s study10 (referred to as “Lenz dataset”) were divided randomly into a discovery series (n = 207) and an internal testing series (207) (see Supplementary Table S1). To identify prognosis-related lncRNAs, we first used univariate Cox proportional hazards regression model to evaluate the associations between the expression level of lncRNAs and overall survival (OS), and found that six lncRNAs were significantly associated with OS in the discovery series (adjusted p < 0.05 after Benjamini-Hochberg multiple testing correction, Table 1). To further investigate the expression pattern of these six lncRNAs, we performed an unsupervised hierarchical clustering for 207 DLBCL patients based on expression levels of these six lncRNAs in the discovery series. As shown in Fig. 1A, the resulting dendrogram showed two distinct patient clusters, which were highly correlated with patients’ survival status (p = 7.46e-03, Chi-square test). As previously described11,12, a panel of six-lncRNA signature was developed as a linear combination of the expression levels of these six lncRNAs and the estimated regression coefficients in the multivariate Cox regression analysis as the weight as follows: Risk Score = (SACS-AS1*0.4452) + (MME-AS1*−0.3143) + (CSMD2-AS1*−1.0086) + (RP11-360F5.1*−0.0827) + (RP11-25K19.1*−0.0181) + (CTC-467M3.1*−0.9599). We were able to calculate a lncRNA expression-based risk score (referred to as “LncRS”) for each patient in the discovery series and classified them into high-risk group (n = 104) or low-risk group (n = 103) using the median risk score of −1.2393 as the cutoff point. Patients were assigned to the high-risk group if their LncRSs were greater than or equal to the cutoff point, whereas low-risk group was composed of patients with LncRSs that were less than the cutoff point. As a result, patients in the high-risk group exhibited poor OS compared with those in the low-risk group (median OS 3.82 years vs. NA years, log rank p < 0.001) (Fig. 1B). There are 104 patients in the high-risk group and 103 patients in the low-risk group. The prognosis of patients in the discovery series showed 76 of dead (37%) and 131 of alive (63%). We performed odds ratio test to quantify the association between the real survival status of patients and the predicted risk group by the six-lncRNA signature. The odd ratio of the discovery series is 4.38 (95% CI = 2.37 to 8.11; p < 0.0001), suggesting that the high-risk group was more likely to have higher mortality than the low-risk group (53% vs. 20% for the discovery series) (see Supplementary Table S2).
The overall five- and ten-year relative survival rate of the high-risk group were 42.4% and 39.6%, respectively, whereas the corresponding rates in the low-risk group were 79.1% and 61.2%, respectively. Moreover, we found that the LncRS was significantly associated with OS (HR = 2.31, 95% CI = 1.8 to 2.965, p < 0.001) in a univariate analysis. Fig. 1C,D showed the LncRS distribution and expression pattern of six prognostic lncRNAs in patients of the discovery series (ranked according to increasing LncRS). As expected, five protective lncRNAs (MME-AS1, CSMD2-AS1, RP11-360F5.1, RP11-25K19.1 and CTC-467M3.1) tended to be expressed in patients with low LncRS, whereas one risky lncRNAs (SACS-AS1) was up-regulated in patients with high LncRS.
Prognostic value of the six-lncRNA signature for survival prediction in the internal testing series and entire Lenz dataset
The prognostic value of the six-lncRNA signature for survival prediction was evaluated using the internal testing series and entire Lenz dataset. With the same six-lncRNA signature score model and cutoff point derived from the discovery series, 207 patients of the internal testing series were divided into high-risk group (n = 110) and low-risk group (n = 97). As in the discovery series, OS in the high-risk group was significantly worse than that in the low-risk group (median OS 2.68 years vs. 10.62 years, log rank p = 0.015), and the proportions of OS in the high-risk group and low-risk group are 44.6% and 64.9% at five years, and 33.4% and 56.8% at ten years, respectively (Fig. 2A). The odd ratio of the internal testing series is 2.38 (95% CI = 1.35 to 4.19, p = 0.0028).
Similar results were observed in the entire Lenz dataset (i.e. combined discovery and internal testing series), which were comprised of 214 high-risk patients with median OS of 3.26 years and 200 low-risk patients with not reached median OS (log-rank p < 0.001) (Fig. 2B). The odd ratio of the entire Lenz dataset is 3.24 (95% CI = 2.14 to 4.90, p < 0.0001). In the univariate analysis, significant associations between the LncRS and OS also were found both in the testing series (HR = 1.362, 95% CI = 1.108 to 1.673, p = 0.003) and entire Lenz dataset (HR = 1.737, 95% CI = 1.484 to 2.034, p < 0.001). The LncRS distribution and expression pattern of six prognostic lncRNAs in patients of the testing series and entire Lenz dataset were shown in Supplementary Fig. S1, which were consistent with findings from the discovery series.
Confirmation of the six-lncRNA signature for survival prediction in additional independent dataset
We further evaluated the prognostic value of six-lncRNA signature for survival prediction in the additional independent patient dataset from Visco’s study13 (referred to as “Visco dataset”). The same six-lncRNA signature score model obtained from the discovery series was used to calculate the LncRS for each of 470 patients from the Visco dataset. With the same cutoff point derived from the discovery series, 470 patients of the Visco dataset were divided into the high-risk group (n = 383) and low-risk group (n = 87). As shown in Fig. 3A, patients with high-risk LncRS had significantly shorter survival than those belonging to low-risk group (median OS 6.27 years vs. NA years, log rank p < 0.001). The odd ratio of the Visco dataset is 3.26 (95% CI = 1.80 to 5.90, p = 0.0001). At four years and six years, the respective absolute differences in OS between the groups with high-risk and low-risk LncRS were 20.4% (61.7% vs. 82.1%) and 24.7% (53% vs. 77.7%), respectively. In univariate analysis of the Visco dataset, the LncRS were significantly correlated with OS (HR = 1.601, 95% CI = 1.315 to 1.95, p < 0.001) (Table 2). The distribution of LncRS and lncRNAs expression of patients in the Visco dataset is shown in Fig. 3B,C. Similar to the above findings, five protective lncRNAs were over-expressed and one risky lncRNA was down-regulated in the low-risk patients compared to the high-risk patients.
Further confirmation of prognostic lncRNAs using additional independent dataset with a different array platform
To further examine the predictive power and robustness of six prognostic lncRNAs, we performed a cross-platform test analysis of prognostic lncRNAs identified in the discovery series using another existing DLBCL patient dataset measured with Affymetrix HG-U133A array from Hummel’s study14 (referred to as “Hummel dataset”). We re-annotated the probes of Affymetrix HG-U133A array to lncRNAs as described in the Materials and Method, and found that only 3 lncRNAs from the six-lncRNA signature were covered on the Affymetrix HG-U133A array. Therefore, the predictive power of these three prognostic lncRNAs (MME-AS1, CSMD2-AS1 and CTC-467M3.1) was analyzed using the completely independent Hummel dataset of 159 DLBCL patients. The risk score of each patient in Hummel dataset was calculated based on the expression levels of these three lncRNAs according to the risk score model derived from the discovery series without re-estimating parameters. The median risk score obtained from Hummel dataset classified 159 DLBCL patients into the high-risk group (n = 77) and the low-risk group (n = 82). The Kaplan-Meier curves based on these three lncRNAs were markedly different (log rank p = 0.003), showing OS in 32.6% and 60.3% at five years, and 29% and 52.9% at ten years for patients with high-risk and low-risk LncRS, respectively (Fig. 4A). Furthermore, the univariate Cox regression analysis also showed that the risk scores were significantly associated with OS in DLBCL patients of the Hummel dataset (HR = 1.598, 95% CI = 1.098 to 2.327, p = 0.014). The results of LncRS distribution of patients and expression pattern of three prognostic lncRNAs also demonstrated their discriminatory power between patients with poor and good survival (Fig. 4B,C).
Survival prediction by the six-lncRNA signature is independent of conventional clinical factors
To assess whether the prognostic values of the six-lncRNA signature is independent of conventional clinical factors of DLBCL patients, we performed the multivariate Cox regression analyses using OS as the dependent variable and LncRS and other conventional clinical factors as explanatory variables, and found that the six-lncRNA signature still maintained an independent correlation with OS both in the Lenz and Visco datasets after adjustment for conventional clinical factors, including age, gender, stage, number of extranodal sites, lactate dehydrogenase (LDH) level, Eastern cooperative Oncology Group (ECOG) performance status and subtype (Table 2). However, we found that age, LDH level and ECOG performance status were also significant in the multivariate analysis. Therefore, data stratification analysis was performed to examine whether the six-lncRNA signature could provide prognostic value within the same clinical factors. For this, all 884 patients (combining Lenz and Visco datasets) were stratified into younger patient group (< = 60) and elder patient group (>60) according to age. With the same six-lncRNA signature and risk score cutoff point derived from the discovery series, all 388 patients with age < = 60 were divided into the high-risk group (n = 230) with poor survival or the low-risk group (n = 158) with good survival (log-rank p < 0.001) (Fig. 5A). Similar prognostic value of the six-lncRNA signature was observed for elder patients who were classified into the high-risk group (n = 366) with median OS of 3.94 years and low-risk group (n = 130) with median OS of 10.62 years (log rank p < 0.001) (Fig. 5B). Further analysis found that the six-lncRNA signature was able to separate patients with ECOG performance status score <2 into the high-risk group (n = 455) with median OS of 6.72 years and low-risk group (n = 215) with not reached median OS (log rank p < 0.001, Fig. 5C). Similarity, among those patients with poor general health status (ECOG performance status score of 2 or greater), the six-lncRNA signature also could distinguish between patients with significantly different survival (median OS 1.72 years vs. 6.08 years, log rank p = 0.045, Fig. 5D) Another important clinical factor, LDH level, stratified all 777 patients with LDH information into two subgroups with LDH level lower than 1*normal or greater than 1*normal. Within each LDH stratum, survival analysis also demonstrated significant differences in OS between the high-risk group and low-risk group (median OS 7.61 years vs. NA years, log-rank p < 0.001 for 317 patients with LDH < 1*normal, and median OS 4.06 years vs. 9.11 years, log-rank p = 0.001 for 460 patients with LDH> = 1*normal) (Fig. 5E,F). Taken together, these results suggested that the predictive capacity of the six-lncRNA signature is independent of conventional clinical factors for survival prediction of DLBCL patients.
(A) Kaplan–Meier survival curves for younger patients with DLBCL. (B) Kaplan–Meier survival curves for elder patients with DLBCL. (C) Kaplan–Meier survival curves for DLBCL patients with ECOG performance status score <2. (D) Kaplan–Meier survival curves for DLBCL patients with ECOG performance status score of 2 or greater. (E) Kaplan–Meier survival curves for DLBCL patients with LDH < 1*normal. (F) Kaplan–Meier survival curves for DLBCL patients with LDH> = 1*normal.
Functional implication of the six-lncRNA signature
We further investigated the functional implication of the six-lncRNA signature in the development of DLBCL by Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) functional enrichment analysis for PCGs co-expressed with six prognostic lncRNAs. We first measured the co-expressed relationships between the expression level of prognostic lncRNAs and that of protein-coding genes (PCGs), and considered those PCGs whose co-expressed correlations with lncRNAs were ranked in the top 1% of all correlations as lncRNAs-related PCGs. The results of functional enrichment analysis demonstrated that PCGs positively correlated with prognostic lncRNAs were enriched in three GO functional clusters (including immune system process, DNA repair and cell cycle) (Fig. 6A) and 12 KEGG pathways (Fig. 6B). The PCGs negatively co-expressed with prognostic lncRNAs clustered most significantly in cell death, cell adhesion, immune system process, and inflammatory response for GO biological process enrichment analysis (Fig. 6C), and in 12 biological pathways for KEGG enrichment analysis (Fig. 6D). Most of enriched functional catalogues have been found to be closely associated with the incidence and development of DLBCL15, which emphasized an implication of the six-lncRNA signature in DLBCL through exerting their regulatory roles on PCGs involved in these known DLBCL-related biological processes and pathways.
(A) The functional enrichment map of GO terms enriched by positively correlated protein-coding genes. (B) Significantly enriched KEGG pathways of positively correlated protein-coding genes. (C) The functional enrichment map of GO terms enriched by negatively correlated protein-coding genes. (D) Significantly enriched KEGG pathways of negatively correlated protein-coding genes.
Discussion
Recent advances in the transcriptomic analysis have demonstrated the pathological and molecular heterogeneity of DLBCL16. Distinctive molecular heterogeneity is closely associated with a wide range of clinical characteristics and outcome. For almost two decades, the International Prognostic Index (IPI) system has been widely used to guide risk stratification and predict the outcome of DLBCL patients17. The IPI takes into account a series of clinical criteria, including age, stage, LDH level, ECOG performance status and number of extranodal sites18. However, the fact that the IPI system does not account for factors underlying the molecular heterogeneity of DLBCL patients, and considerable differences in survival outcome have been observed even among patients with the same or similar IPI variables, leading to increasing attention for identifying additional molecular prognostic biomarkers18. Several expression-based prognostic models have been developed to meet this need at the mRNA level19,20,21. Recent studies have revealed the contribution of lncRNAs to cancer development, implying their potential as novel biomarkers in cancer diagnosis22,23,24,25,26,27,28,29.
In this study, we investigated the prognostic value of lncRNAs by analyzing lncRNAs expression profiles and clinical characteristics in a large cohort of 1043 DLBCL patients. By using the sample splitting method and Cox regression analysis, we identified six prognostic lncRNAs that were significantly associated with OS of DLBCL patients. Based on these six prognostic lncRNAs, we constructed a six-lncRNA signature which was able to classify DLBCL patients into the high-risk and low-risk groups with significantly different survival outcome. The predictive value of the six-lncRNA signature was successfully validated in a completely independent dataset of 470 DLBCL patients. Furthermore, we used another independent dataset with different array platform (HG-U133A array) to validate our findings. Even though only three out of six prognostic lncRNAs were covered in this array, expression levels of these three lncRNAs were again shown to have close association with OS of patients. These results with independent DLBCL patient datasets demonstrated the robustness and good reproducibility of the six-lncRNA signature in DLBCL. Further analysis revealed that the six-lncRNA signature is independent of conventional clinical factors, including age, gender, stage, number of extranodal sites, LDH level, ECOG performance status and subtype. Notably, the six-lncRNA signature was shown capable of predicting survival for patients with the same or similar IPI variables, suggesting that the six-lncRNA signature could provide additional prognostic information at the molecular level beyond the conventional IPI system.
To date, research has only just begun into the biological function of lncRNAs and only a handful of lncRNAs were functionally well-characterized. Recent study found that lncRNA CTC-467M3.1, one of the six prognostic lncRNAs, is involved in the cis-regulation of transcription factor MEF2C29 which is required for B cell proliferation and survival30. Increasing evidence has suggested that lncRNAs were involved in diverse biological processes by negatively or positively regulating gene expression at both the posttranscriptional and transcriptional levels31,32. Therefore, it is feasible to infer biological roles of lncRNAs by functional views of PCGs that are co-expressed with lncRNAs33,34,35. So we investigated the co-expression patterns of mRNAs and lncRNAs and performed functional enrichment analysis for co-expressed PCGs to predict biological function of prognostic lncRNAs in this signature. We found that PCGs whose expression value positively or negatively correlated with the six prognostic lncRNAs were enriched in three functional groups of known cancer-related pathways, immune system-related biological processes and signaling molecules interaction that are closely linked with the incidence and progression of DLBCL15. Thus, it is a plausible inference that the six-lncRNA signature may be involved with DLBCL through exerting their regulatory roles on PCGs in these known DLBCL-related biological processes and pathways.
In conclusion, our study investigated the prognostic potential of lncRNAs in DLBCL, and identified a potential panel of six-lncRNA signature as a composite biomarker for risk stratification of DLBCL patients at diagnosis. Moreover, the six-lncRNA signature was able to effectively predict the survival outcome of DLBCL patients with similar IPI variables. To our knowledge, this is the first report on efforts to identify lncRNA signature that predicts survival outcome of patients with DLBCL. With further confirmation, the six-lncRNA signature not only provides additional prognostic value beyond the conventional IPI system to identify high-risk patients who will benefit from more effective therapy, but also can improve our understanding about the molecular heterogeneity of DLBCL from the viewpoint of non-coding RNA. However, it should be noted that a large portion of known lncRNAs were missing in our study due to the intrinsic limitation of the microarray technique and probe repurposing method. Therefore, more research is needed to uncover novel diagnostic or prognostic lncRNAs candidates in DLBCL.
Materials and Methods
DLBCL patient datasets
Genome-wide gene expression profiles data generated from the Affymetrix platform (Affymetrix HG-U133 Plus 2.0 array and HG-U133A array) and corresponding clinical information of DLBCL patients were retrieved from the GEO database. After removing patients without available clinical information, a total of 1043 DLBCL patients were enrolled in this study, including 414 patients from Lenz’s study (the GEO accession number is GSE10846, Affymetrix HG-U133 Plus 2.0 array) (http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc = GSE10846)10, 470 patients from Visco’s study (the GEO accession number is GSE31312, Affymetrix HG-U133 Plus 2.0 array)(http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc = GSE31312)13 and 159 patients from Hummel’s study (the GEO accession number is GSE4475, Affymetrix HG-U133A array) (http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc = GSE4475)14. Detailed clinical information of DLBCL patients used in this study was shown in Supplementary Table S1.
Acquisition of lncRNA expression profiles of DLBCL patients
The raw array data (.CEL files) of 1043 DLBCL patients on the Affymetrix HG-U133 Plus 2.0 array or HG-U133A array were downloaded from the GEO database and were uniformly pre-processed using the Robust Multichip Average (RMA) algorithm for background correction, quantile normalization and log2-transformation36. To account for the heterogeneity in systematic measurement among multiple microarray datasets, each dataset was standardized independently by the Z-score transformation to scale expression intensities of each probe into having a mean of 0 and a standard deviation of 1.
The probe sequences of Affymetrix HG-U133 Plus 2.0 and HG-U133A arrays were retrieved from the Affymetrix website (http://www.affymetrix.com). We re-mapped those probes into the human genome (GRCh38) to obtain the chromosomal position of the probes using SeqMap tool37. Then the lncRNAs-specific probes were obtained by matching the chromosomal position of probes to the chromosomal position of lncRNAs based on the annotation from the GENCODE project (http://www.gencodegenes.org, release 22) as previously described22,23,38. The expression levels of lncRNAs were then obtained from the normalized intensity of lncRNAs-specific probes. Finally, 2330 lncRNAs for Affymetrix HG-U133 Plus 2.0 and 663 lncRNAs for Affymetrix HG-U133A were obtained for subsequent analysis.
Statistical analysis
The association between the expression level of each lncRNA and OS of DLBCL patients was evaluated using the univariate Cox regression analysis and Benjamini-Hochberg multiple testing correction. LncRNAs were considered as prognostic lncRNAs if their adjusted p-value were less than 0.05. Then a six-lncRNA expression signature was constructed using a linear combination of the expression levels of these six lncRNAs and the estimated regression coefficients in the multivariate Cox regression analysis as previously described11,12. With this six-lncRNA signature, patients in each dataset were classified into high-risk group and low-risk group by using the median LncRS of the discovery series as the cutoff point. Kaplan-Meier survival curves and log-rank test were used to assess the difference in OS between the two groups with high-risk and low-risk LncRS. Univariate and multivariate analyses with Cox proportional hazards regression for OS were performed on the individual conventional clinical variables with and without the six-lncRNA signature in each dataset. Hazard ratios (HR) and 95% confidence intervals (CI) were calculated. All statistical tests were two-sided and performed with R software.
Functional enrichment analysis
The Pearson correlation coefficients between the expression level of each lncRNAs and that of each PCG were calculated. The PCGs positively or negatively correlated with prognostic lncRNAs (ranked top 1%) were considered as lncRNAs-related PCGs. Functional enrichment analysis of lncRNAs-related PCGs for GO terms and KEGG pathway was performed using DAVID Bioinformatics Tool (https://david.ncifcrf.gov/, version 6.7)39. GO functional clusters limited to “Biological Process”(GOTERM-BP-FAT) with an enrichment score of >1.5 and KEGG pathway Functional Annotation with p-value of <0.05 using the whole human genome as background were considered as potential functional roles of prognostic RNAs. Significant GO terms with similar function were organized into an interaction network and visualized using the Enrichment Map40.
Additional Information
How to cite this article: Sun, J. et al. A potential panel of six-long non-coding RNA signature to improve survival prediction of diffuse large-B-cell lymphoma. Sci. Rep. 6, 27842; doi: 10.1038/srep27842 (2016).
Accession codes
References
Tilly, H. et al. Diffuse large B-cell lymphoma (DLBCL): ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann Oncol 26 Suppl 5, v116–125, doi: 10.1093/annonc/mdv304 (2015).
Younes, A. Prognostic Significance of Diffuse Large B-Cell Lymphoma Cell of Origin: Seeing the Forest and the Trees. J Clin Oncol 33, 2835–2836, doi: 10.1200/JCO.2015.61.9288 (2015).
Hangauer, M. J., Vaughn, I. W. & McManus, M. T. Pervasive transcription of the human genome produces thousands of previously unidentified long intergenic noncoding RNAs. PLoS Genet 9, e1003569, doi: 10.1371/journal.pgen.1003569 (2013).
Mercer, T. R., Dinger, M. E. & Mattick, J. S. Long non-coding RNAs: insights into functions. Nat Rev Genet 10, 155–159, doi: 10.1038/nrg2521 (2009).
Moran, V. A., Perera, R. J. & Khalil, A. M. Emerging functional and mechanistic paradigms of mammalian long non-coding RNAs. Nucleic Acids Res 40, 6391–6400, doi: 10.1093/nar/gks296 (2012).
Hauptman, N. & Glavac, D. Long non-coding RNA in cancer. Int J Mol Sci 14, 4655–4669, doi: 10.3390/ijms14034655 (2013).
Gibb, E. A., Brown, C. J. & Lam, W. L. The functional role of long non-coding RNA in human carcinomas. Mol Cancer 10, 38, doi: 10.1186/1476-4598-10-38 (2011).
Verma, A. et al. Transcriptome sequencing reveals thousands of novel long non-coding RNAs in B cell lymphoma. Genome Med 7, 110, doi: 10.1186/s13073-015-0230-7 (2015).
Peng, W., Fan, H., Wu, G., Wu, J. & Feng, J. Upregulation of long noncoding RNA PEG10 associates with poor prognosis in diffuse large B cell lymphoma with facilitating tumorigenicity. Clin Exp Med 16, 177–182, doi: 10.1007/s10238-015-0350-9 (2016).
Lenz, G. et al. Stromal gene signatures in large-B-cell lymphomas. N Engl J Med 359, 2313–2323, doi: 10.1056/NEJMoa0802885 (2008).
Lossos, I. S. et al. Prediction of survival in diffuse large-B-cell lymphoma based on the expression of six genes. N Engl J Med 350, 1828–1837, doi: 10.1056/NEJMoa032520 (2004).
Alizadeh, A. A. et al. Prediction of survival in diffuse large B-cell lymphoma based on the expression of 2 genes reflecting tumor and microenvironment. Blood 118, 1350–1358, doi: 10.1182/blood-2011-03-345272 (2011).
Visco, C. et al. Comprehensive gene expression profiling and immunohistochemical studies support application of immunophenotypic algorithm for molecular subtype classification in diffuse large B-cell lymphoma: a report from the International DLBCL Rituximab-CHOP Consortium Program Study. Leukemia 26, 2103–2113, doi: 10.1038/leu.2012.83 (2012).
Hummel, M. et al. A biologic definition of Burkitt’s lymphoma from transcriptional and genomic profiling. N Engl J Med 354, 2419–2430, doi: 10.1056/NEJMoa055351 (2006).
Zhang, Z. X. et al. Exploration of molecular mechanisms of diffuse large B-cell lymphoma development using a microarray. Asian Pac J Cancer Prev 14, 1731–1735 (2013).
Wright, G. et al. A gene expression-based method to diagnose clinically distinct subgroups of diffuse large B cell lymphoma. Proc Natl Acad Sci USA 100, 9991–9996, doi: 10.1073/pnas.1732008100 (2003).
Project, T. I. N.-H. s. L. P. F. A predictive model for aggressive non-Hodgkin’s lymphoma. The International Non-Hodgkin’s Lymphoma Prognostic Factors Project. N Engl J Med 329, 987–994, doi: 10.1056/NEJM199309303291402 (1993).
Sehn, L. H. & Gascoyne, R. D. Diffuse large B-cell lymphoma: optimizing outcome in the context of clinical and biologic heterogeneity. Blood 125, 22–32, doi: 10.1182/blood-2014-05-577189 (2015).
Bret, C., Klein, B. & Moreaux, J. Gene expression-based risk score in diffuse large B-cell lymphoma. Oncotarget 3, 1700–1710, doi: 10.18632/oncotarget.807 (2012).
Xu, Q. et al. Identification and validation of a two-gene expression index for subtype classification and prognosis in Diffuse Large B-Cell Lymphoma. Sci Rep 5, 10006, doi: 10.1038/srep10006 (2015).
Cheetham, S. W., Gruhl, F., Mattick, J. S. & Dinger, M. E. Long noncoding RNAs and the genetics of cancer. Br J Cancer 108, 2419–2425, doi: 10.1038/bjc.2013.233 (2013).
Zhou, M. et al. A potential signature of eight long non-coding RNAs predicts survival in patients with non-small cell lung cancer. J Transl Med 13, 231, doi: 10.1186/s12967-015-0556-3 (2015).
Zhou, M. et al. Identification and validation of potential prognostic lncRNA biomarkers for predicting survival in patients with multiple myeloma. J Exp Clin Cancer Res 34, 102, doi: 10.1186/s13046-015-0219-5 (2015).
Sun, J. et al. A potential prognostic long non-coding RNA signature to predict metastasis-free survival of breast cancer patients. Sci Rep 5, 16553, doi: 10.1038/srep16553 (2015).
Hu, Y. et al. A long non-coding RNA signature to improve prognosis prediction of colorectal cancer. Oncotarget 5, 2230–2242, doi: 10.18632/oncotarget.1895 (2014).
Zhou, M. et al. Relapse-related long non-coding RNA signature to improve prognosis prediction of lung adenocarcinoma. Oncotarget doi: 10.18632/oncotarget.8825 (2016).
Crea, F. et al. Identification of a long non-coding RNA as a novel biomarker and potential therapeutic target for metastatic prostate cancer. Oncotarget 5, 764–774, doi: 10.18632/oncotarget.1769 (2014).
Zhou, M. et al. Comprehensive analysis of lncRNA expression profiles reveals a novel lncRNA signature to discriminate nonequivalent outcomes in patients with ovarian cancer. Oncotarget, doi: 10.18632/oncotarget.8653 (2016).
Gudenas, B. L. & Wang, L. Gene Coexpression Networks in Human Brain Developmental Transcriptomes Implicate the Association of Long Noncoding RNAs with Intellectual Disability. Bioinform Biol Insights 9, 21–27, doi: 10.4137/BBI.S29435 (2015).
Wilker, P. R. et al. Transcription factor Mef2c is required for B cell proliferation and survival after antigen receptor stimulation. Nat Immunol 9, 603–612, doi: 10.1038/ni.1609 (2008).
Kornienko, A. E., Guenzl, P. M., Barlow, D. P. & Pauler, F. M. Gene regulation by the act of long non-coding RNA transcription. BMC Biol 11, 59, doi: 10.1186/1741-7007-11-59 (2013).
Zhang, X. et al. RAID: a comprehensive resource for human RNA-associated (RNA-RNA/RNA-protein) interaction. RNA 20, 989–993, doi: 10.1261/rna.044776.114 (2014).
Hao, Y. et al. Prediction of long noncoding RNA functions with co-expression network in esophageal squamous cell carcinoma. BMC Cancer 15, 168, doi: 10.1186/s12885-015-1179-z (2015).
Ma, H. et al. Molecular mechanisms and function prediction of long noncoding RNA. ScientificWorldJournal 2012, 541786, doi: 10.1100/2012/541786 (2012).
Liao, Q. et al. Large-scale prediction of long non-coding RNA functions in a coding-non-coding gene co-expression network. Nucleic Acids Res 39, 3864–3878, doi: 10.1093/nar/gkq1348 (2011).
Irizarry, R. A. et al. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4, 249–264, doi: 10.1093/biostatistics/4.2.249 (2003).
Jiang, H. & Wong, W. H. SeqMap: mapping massive amount of oligonucleotides to the genome. Bioinformatics 24, 2395–2396, doi: 10.1093/bioinformatics/btn429 (2008).
Sorensen, K. P. et al. Long non-coding RNA expression profiles predict metastasis in lymph node-negative breast cancer independently of traditional prognostic markers. Breast Cancer Res 17, 55, doi: 10.1186/s13058-015-0557-4 (2015).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res 37, 1–13, doi: 10.1093/nar/gkn923 (2009).
Merico, D., Isserlin, R., Stueker, O., Emili, A. & Bader, G. D. Enrichment map: a network-based method for gene-set enrichment visualization and interpretation. PLoS One 5, e13984, doi: 10.1371/journal.pone.0013984 (2010).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant No. 61403111), China Postdoctoral Science Foundation (Grant No. 2014M551268) and Postdoctoral Foundation of Heilongjiang Province (Grant No. LBH-Z14212).
Author information
Authors and Affiliations
Contributions
J.S. and M.Z. conceived and designed the experiments. J.S., L.C., H.S., Z.Z., H.Z., Z.W. and M.Z. analyzed data. M.Z. and J.S. wrote this manuscript. All authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Sun, J., Cheng, L., Shi, H. et al. A potential panel of six-long non-coding RNA signature to improve survival prediction of diffuse large-B-cell lymphoma. Sci Rep 6, 27842 (2016). https://doi.org/10.1038/srep27842
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep27842
This article is cited by
-
Circulating RNA biomarkers in diffuse large B-cell lymphoma: a systematic review
Experimental Hematology & Oncology (2021)
-
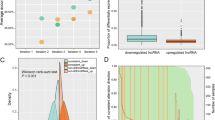
Circulating long non-coding RNAs HOTAIR, Linc-p21, GAS5 and XIST expression profiles in diffuse large B-cell lymphoma: association with R-CHOP responsiveness
Scientific Reports (2021)
-
Development of a Reproducible Prognostic Gene Signature to Predict the Clinical Outcome in Patients with Diffuse Large B-Cell Lymphoma
Scientific Reports (2019)
-
Discovery and validation of immune-associated long non-coding RNA biomarkers associated with clinically molecular subtype and prognosis in diffuse large B cell lymphoma
Molecular Cancer (2017)
-
Genome-scale analysis identifies NEK2, DLGAP5 and ECT2 as promising diagnostic and prognostic biomarkers in human lung cancer
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.