Abstract
A common phenotypic difference among domestic animals is variation in coat color. Six-white-point is a pigmentation pattern observed in varying pig breeds, which seems to have evolved through several different mechanistic pathways. Herein, we re-sequenced whole genomes of 31 Diannan small-ear pigs from China and found that the six-white-point coat color in Diannan small-ear pigs is likely regulated by polygenic loci, rather than by the MC1R locus. Strong associations were observed at three loci (EDNRB, CNTLN and PINK1), which explain about 20 percent of the total coat color variance in the Diannan small-ear pigs. We found a mutation that is highly differentiated between six-white-point and black Diannan small-ear pigs, which is located in a conserved noncoding sequence upstream of the EDNRB gene and is a putative binding site of the CEBPB protein. This study advances our understanding of coat color evolution in Diannan small-ear pigs and expands our traditional knowledge of coat color being a monogenic trait.
Similar content being viewed by others
Introduction
Domestication of the pig has been accompanied with retention of a tremendous amount of phenotypic variation regarding growth, behavior and coat color. The diverse domestic pig breeds thus provide valuable animal models for genetic studies on phenotypic variation1,2,3.
Coat color was potentially exposed to artificial selection at an ancient time, dating back to about 5,000 years ago4. Variation in coat color may have important implications for pig breed development and conservation. Distinct coat colors have evolved in different pig breeds and may even have been used as visual characteristic to distinguish between breeds5. The most common coat colors found in domestic pigs include black, brown, white, red, spotted, belted and two-end-black, which are distinct from the camouflage coat color of wild boars4,6,7,8,9. The genetic mechanisms underlying variation in coat color is an interesting topic.
Six-white-point (SWP) is a common kind of coat color in pigs that and is characterized by a white coat on the four feet, the head and the end of tail but with the remaining coat being black. The SWP phenotype is observed in many well-known European and East Asian domestic pigs, including the Berkshire pigs from Europe10, as well as Diannan small-ear (DSE), Tibetan, Longlin and Anqing six-white-point pigs in China11.
The SWP coat color in the Berkshire pigs was found to be influenced by the melanocortin 1 receptor encoding (MC1R) gene, with this phenotype regarded as a special presentation of spotted color, as it is usually observed with black spots on white or red backgrounds5. In Berkshire pigs, two mutations in the MC1R gene are likely involved in formation of the SWP coat color pattern. The D124N MC1R allele probably causes a complete black body color4. SWP in the Berkshire is derived from the MC1R D124N allele through a 2-bp insertion in the coding region that leads to a coding frameshift and a premature stop codon. The background skin expresses the 2-bp inserted MC1R premature transcript, while the black-spotted area expresses a transcript without the 2-bp insertion, possibly due to somatic reversion events10. Additional to the MC1R locus, further evidence indicated that variation in the KITLG gene may also influence the coat color of Berkshire pigs12.
Due to the independent domestication of pigs in Europe and East Asia13,14, black coat color has putatively evolved through different genetic mutations in the coding sequence of the MC1R gene in East Asia (L102P) and Europe (D124N)4,15. Thus how the SWP phenotype evolved in East Asian domestic pigs, where these animals have a MC1R allele containing the L102P substitution remains mysterious.
In this study, we aimed to screen for genomic variants underlying the SWP phenotype using whole genome resequencing of DSE pigs from Yunnan province, China. DSE pigs have varying coat colors, including whole body black (denoted as “black” below) and SWP. Here we show that the SWP phenotype in DSE pigs is not attributed to variation in the MC1R gene, but rather it is associated with variations in multiple genes with differing phenotypic contributions.
Results
Coat colors in DSE pigs
To obtain coat color phenotype data on the DSE pig, we collected pig samples and classified their coat color as either SWP or black. A total of 1,977 DSE pig samples were collected with coat color phenotype information with 753 (38%) of the DSE pigs being black and 1,224 (62%) being SWP. A comparison of whole black body and SWP phenotypes of DSE pigs is shown in Fig. 1. The white point on the head of the DSE pigs is located on the forehead. The SWP phenotype in the examined DSE individuals is highly variable, with 1,171 of the 1,224 SWP having a partial phenotype with the absence of the white coat at one or more of the six points. From our findings, black individuals were observed in about 78.6% of litters produced by SWP parents and about 33% were black in the total population produced by SWP parents. SWP individuals were observed in about 25.5% of litters from black parents and about 21.6% were SWP in the total population from black parents. About 47% SWP and 53% black offspring were observed in the cross between black and SWP DSE pigs.
Whole genome resequencing of DSE pigs
To study the SWP phenotype in the DSE pigs, we selected 31 individuals (17 SWP and 14 black) for whole genome re-sequencing. Among the 17 SWP individuals, 11 had the typical SWP phenotype. Genomes were sequenced at about 7× depth of coverage for 8 SWP individuals and about 3× for the remaining 23 individuals.
Genomic reads were quality filtered and mapped to the pig reference genome16. On average, the resequencing data for each individual covered about 75% of the reference genome (Supplementary Table S1). We screened for genomic variants in the 31 individuals and a total of 23.9 million SNPs were identified. Among these SNPs, only about 150K and 107K were located in protein coding and UTR sequences, respectively, while most of them were located in noncoding sequences (6M in introns and 17.5M in intergenic sequences). Further classification showed that 69,707 SNPs were located in conserved sequences, 2,717,284 in putative regulatory sequences and 347,597 in putative transcription factor DNA-binding motifs. Detailed information on the distribution of SNPs on each chromosome is shown in Supplementary Table S2.
Analysis of the MC1R gene
To determine whether the SWP in the DSE pigs is associated with the 2-bp insertion in the coding sequence of the MC1R gene, as previously shown in Berkshire pigs10, we searched for the presence of a 2-bp insertion in MC1R in SWP and black DSE pigs. The 2-bp insertion in MC1R was not detected in any of the re-sequenced DSE pigs and both the SWP and black DSE pigs were homozygous for the expected L102P MC1R allele (Supplementary Table S3). This observation implies that the SWP phenotype in the DSE pigs is not caused by a 2-bp insertion in the MC1R gene.
Black East Asian domestic pigs carry a L102P missense mutation in the MC1R gene10, as compared to a D124N mutation in some black domestic pigs from Europe4 and all of the re-sequenced DSE samples are homozygous for L102P. No mutations were identified at the D124 position (Supplementary Table S3), confirming that the MC1R allele in DSE pigs is of East Asian ancestry and not of European origin. In our comparative analysis, the MC1R locus does not show strong differentiation between the black and SWP DSE pigs (data not shown). These results further imply that the SWP coat color in DSE pigs likely is not be attributed by MC1R mutations.
Screening for genomic regions putatively associated with SWP phenotype
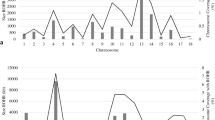
We first screened genomic regions that were highly differentiated between the SWP and black DSE pigs. In this analysis, 751 genes (Supplementary Table S4) were identified in genomic 10-kb sliding windows (containing at least 100 SNPs) with FST values above the chromosomal top 1% threshold (Fig. 2a, Supplementary Table S5). Some genes in the identified gene set are well known for roles in coat color regulation17, including the endothelin receptor type B (EDNRB) and dopachrome tautomerase (DCT) genes. EDNRB is important in the development of melanocytes18 and is involved in two-end-black phenotype regulation in pigs12,19,20. DCT null mice show diluted pigmentation with reduced melanin content in hair21.
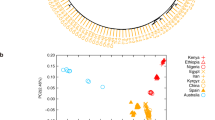
Whole genome screening of loci putatively associated with the SWP phenotype in DSE pigs.
(a) Autosomal plot of genomic differentiation (FST) between the SWP and black DSE pigs.(b) Autosomal XP-EHH plot in SWP DSE pigs with black DSE pigs as a background. (c) Association of IBD segments with the coat color phenotypes in the whole genome resequenced DSE pigs.
To determine whether some haplotypes are associated with the SWP phenotype in DSE pigs, we compared the extended haplotype homozygosity (EHH) between SWP and whole black DSE pigs with an XP-EHH analysis. In addition, as SWP DSE pigs may share some genomic sequences that were inherited from a recent common ancestor, we also examined the inheritance of genomic sequences by analyzing the sharing of identity-by-descent (IBD) segments in the DSE pigs.
The XP-EHH- and IBD-based screening approaches (Fig. 2b,c) identified several genomic regions (Supplementary Table S6) replicating the signals in the FST analysis at the PTEN induced putative kinase 1 (PINK1) locus on chromosome 6 and at the SLC19A2 gene on chromosome 4.
Association analysis of candidate genes in a large population of DSE pigs
To further analyze the association of SWP with candidate genes identified in our analysis, we focused on 18 SNPs, representing 18 different candidate genes (Supplementary Table S7). These 18 SNPs were chosen in genomic regions (<1Mb) showing a clustering of at least twenty 10-kb sliding windows with FST above the chromosomal top 10% threshold and containing at least three 10-kb sliding windows (each containing at least 100 SNPs) with window FST values above the chromosomal top 5% threshold. We selected the 18 SNPs with a rationale that each was from the top 10 SNPs having the highest FST values in each region, with a preference for a SNP covered by the largest number of resequenced individuals and avoiding SNPs located in repetitive sequences. A large population (n = 384) of DSE pigs with their coat color phenotype information was collected, where 215 were black and 169 were SWP pigs (37.87% of which had the typical SWP phenotype). The genotypes of all 18 SNPs were determined from all 384 individuals. Genotyping showed that the derived allele frequency difference between the SWP and black DSE populations were in concordance with that obtained in the whole genome re-sequenced population (Fig. 3). Of the 18 SNPs, 3 SNPs showed a statistically significant (P < 0.05; Pearson’s Chi-squared test) difference in their allele frequencies between the black and SWP individuals. However, none of the SNPs were completely fixed in the SWP pigs.
The association of the genotypes at the 18 SNPs and the SWP phenotype in the 384 phenotypic pigs was tested and unveiled an association with the SWP phenotype for 11 of these SNPs. Pronounced associations were observed at 3 SNPs from the genomic regions containing the EDNRB (P = 1.01 × 10−11; OR = 2.988), centlein, centrosomal protein (CNTLN) (P = 5.58 × 10−10; OR = 2.714) and PINK1 (P = 3.04 × 10−7; OR = 2.269) encoding genes. Additional associations were observed for eight SNPs linked to the Golgi apparatus membrane protein TVP23 homolog C (TVP23C) (P = 1.12 × 10−5; OR = 1.908), aryl hydrocarbon receptor nuclear translocator 2 (ARNT2) (P = 1.37 × 10−5; OR = 1.983), ring finger protein 19A, E3 ubiquitin protein ligase (RNF19) (P = 1.15 × 10−4; OR = 1.857), coiled-coil domain containing 182 (MSI2) (P = 1.80 × 10−4; OR = 1.851), Down Syndrome cell adhesion molecule (DSCAM) (P = 3.41 × 10−4; OR = 1.775), SRY-box 5 (SOX5) (P = 2.04 × 10−4; OR = 1.725), sentrin-specific protease 1 (SENP1) (P = 3.0 × 10−3; OR = 1.544) and optic atrophy 1 (OPA1) (P = 3.66 × 10−3; OR = 1.597) encoding genes.
In a least-squares linear regression analysis, we found that genetic variation in EDNRB, CNTLN and PINK1 genes individually explain about 12.68%, 9.36% and 7.14% coat color phenotypic variance in the DSE pigs respectively. However, the fraction of phenotypic variance explained by a single locus might be overestimated due to neglecting the fraction of variance explained by other loci, conditional on the genotypes at the focused locus. We performed a multiple regression analysis with the genetic variation combinations from these three loci, which showed that 18.52% of the coat color phenotypic variation was explained by the genetic co-variation from the EDNRB + CNTLN loci. Similarly, combinations of EDNRB + PINK1 and CNTLN + PINK1 explained 17.11% and 11.49% of the phenotypic variance, respectively. Genetic variation from the three loci explains 19.98% of the total phenotypic variance. The genetic variation from all 18 SNPs explains about 31.35% of the total phenotypic variance.
Potential regulatory sequence mutations in the EDNRB locus
We are interested in associations between the EDNRB gene and the SWP phenotype. Earlier studies revealed that EDNRB is an important gene in pigmentation regulation and is putatively involved in the two-end-black pigmentation in pigs12,19. In our analysis, the EDNRB locus showed the strongest association with the SWP phenotype and explained the largest fraction of the phenotypic variance.
We screened the SNPs at the EDNRB locus to identify mutations that are highly differentiated between the SWP and black DSE pigs. In the whole genome re-sequenced DSE population, a total of 452 SNPs were identified, of which 179 (Supplementary Table S8) were identified with differentiation between the SWP and black pigs above the chromosomal top 10% threshold. These 179 SNPs were all located in the intergenic sequence upstream of the EDRNB gene (Supplementary Figure S1).
In a 100-way cross-species whole genome alignment analysis, 12 of the 179 SNPs (Table 1) were located in conserved noncoding sequences (CNS). As CNSs are a class of noncoding sequences that have evolved under strong evolutionary constraint, they are considered to potentially influence gene expression22. The location of SNPs inside a CNS is indicative of functional changes to expressional regulation of the EDRNB gene.
Interestingly, a SNP (G>A; chr11: 54,769,424) is located in a sequence which is orthologous to the human counterpart that has been demonstrated to be a DNA-binding motif for the CCAAT/enhancer binding protein, beta (CEBPB), a b-ZIP transcription factor that binds to the consensus sequence (5′-T[TG]NNGNAA[TG]-3′) in promoters and upstream elements and stimulates gene expression23. A cross-species genome alignment showed that this CEBPB DNA-binding motif upstream of the EDNRB gene is highly conserved in mammals (Fig. 4a), indicating that it likely has an important biological function under strong evolutionary constraint. Strikingly, the G>A SNP position is centered at the binding motif in the CEBPB binding sequence and the mutation alters the DNA-binding sequence to 5′-T[TG]NNANAA[TG]-3′ (underlined “A” is the derived allele in the SWP DSE pigs), at a nucleotide that is highly conserved in the enhancers of a variety of genes23. This mutation may lead to a change in transactivation efficiency. In the whole genome resequencing, the G>A SNP was detected in reads from 12 SWP and 13 whole body black DSE pigs. Among the 12 SWP pigs, 10 were homozygous for the derived allele (adenine) and 2 were heterozygous, while in comparison, in the 13 black pigs, 6 were homozygous for the ancestral allele, 5 were heterozygous and only 2 were homozygous for the derived allele. However, this SNP is likely in linkage with, rather than being, a causative mutation, inferring from the distribution of mutant homozygotes in the black DSE pigs.
Cross-species alignment of genomic sequences enclosing two SNPs in the EDNRB and PINK1 loci.
(a) A SNP (chr11: 54,769,424; G>A) in the CEBPB DNA-binding motif upstream of the EDNRB gene. The position of the mutation position is indicated by a box. (b) A nonsynonymous mutation in the kinase domain of the PINK1. The arrow indicates the position of the mutation at a highly conserved residue from a variety of mammals.
Besides the SNP at the CEBPB recognition site, 11 additional SNPs (Table 1) were located in putative regulatory sequences. Among these, 3 SNPs (chr11: 54,769,314, 54,769,363 and 54,769,400) were found in the CEBPB binding sequence, with two SNPs (chr11: 54,770,500 and 54,770,533) identified in a sequence orthologous to a human sequence detected as a binding site by chromatin immunoprecipitation (ChIP) with the transcription factors STAT3, BACH1, FOS and MAFK. Two additional SNPs (chr11: 54,769,730 and 54,769,742) were located in sequences orthologous to a human BACH1 binding sequence and a DNase I hypersensitive site. The SNP data set provides important mutation candidates for the SWP phenotype study.
Mutation analysis of the PINK1 locus
A 280-kb genomic sequence on chromosome 6 showed FST values reaching the chromosomal top 5% threshold. A total of 2,140 SNPs were found in this region. Among them, 553 SNPs were identified with FST values above the chromosomal top 10%. Six protein coding genes, including CAMK2N1, MUL1, FAM43B, CDA, PINK1 and DDOST, were identified within this 280-kb genomic span. The peak of the FST values of the 10-kb sliding windows is located on the PINK1 gene.
Screening the 553 SNPs revealed two nonsynonymous and two synonymous mutations that were differentiated between the black and SWP DSE pigs. The two nonsynonymous mutations alter two residues (P195S and E396D) that are located in the kinase domain. The FST values are 0.42 and 0.24 at the P195S (chr6: 73,118,955; C>T) and E396D (chr6: 73,125,086; G>C) mutation sites. However, the SWP DSE pigs show an ancestral allele frequency (0.76) higher than the black DSE pigs (0.06) at the P195S coding site. In comparison, the derived allele frequency in the SWP DSE pigs (0.95) is much higher than in the black DSE pigs (0.40) at the E396D coding site. A multi-species sequence alignment showed that residue E396 is highly conserved in vertebrates (Fig. 4b), indicating a functional importance. Mutations at highly conserved residues in the kinase domain have been reported to impair or eliminate the PINK1 function24, thus the nonsynonymous mutation E396D potentially influences PINK1 function.
Discussion
Selection for coat color has occurred since the domestication of the pig. Substantial variation in coat color exists in domestic pigs and this phenotype has been a focus of evolutionary studies25. In this study, we examined the genetics underlying the SWP phenotype in DSE pigs, a well-known indigenous domestic pig from China.
The SWP phenotype is highly variable in the DSE pigs and with a complicated mode of inheritance. Our findings showed that black parents have SWP offspring while SWP parents black offspring. Therefore, we inferred from our observations that based on the occurrence of the SWP coat color in DSE pigs, this phenotype is likely to be under the regulation of polygenic loci. If the SWP was caused by a monogenic mutation, there should not be an occurrence of SWP parents with black offspring when SWP phenotype was inherited in a recessive mode, or alternatively, there should not be an occurrence of black parents with SWP offspring when SWP phenotype was inherited in a dominant mode. The inheritance of SWP phenotype is consistent with the multifactorial inheritance model of discrete traits proposed by Fisher26. However, we failed to identify an exact model for this phenotype inheritance. The higher proportion of black individuals observed as compared to the SWP individuals from a cross between black and SWP DSE pigs indicates a recessive mode of SWP inheritance at most of the associated loci. The association of the SWP phenotype with multiple loci is an indication of a complicated regulatory mechanism. Complete co-segregation of SWP with any mutation from a single locus in the DSE pigs could not be found, instead, many loci showed moderate levels of allele frequency differences in the SWP and black DSE populations. This suggests that different loci have varying contributions to the formation of the SWP phenotype. Furthermore, an increased fraction of the phenotypic variance can be explained by the combination of multiple loci compared to any individual locus. This is in sharp contrast to the black coat color that is likely under the regulation of missense mutations in the single MC1R gene in East Asian domestic pigs and wild boars4,15.
Our results support a role of EDNRB in the regulation of the SWP phenotype. EDNRB encodes a G-protein-coupled heptahelical receptor, for the ligand endothelin-3 (EDN3), which is an important signaling pathway for establishing melanocyte development18. Mutations in EDNRB have been widely reported to affect the pigmentation in a variety of organisms, including mice27,28,29, horse30,31, zebrafish32 and human33. It is interesting to observe the strong association of the EDNRB gene with the SWP phenotype in DSE pigs, as variation in EDNRB is also associated with the two-end-black pigmentation pattern seen in many different Chinese indigenous pig breeds34. In contrast, we found that the SWP phenotype is probably regulated by mutations in the upstream regulatory sequences of the EDNRB gene. The abundance of regulatory sequences inside and surrounding the EDNRB genes may partially explain the reason for the involvement of this locus in the regulation of several different coat color phenotypes. We infer that mutations in these putative EDNRB regulatory sequences might have altered tempospatial gene expression of downstream target genes. Consistently, EDNRB shows a temporal expression pattern18 and regulatory sequence variation results in altered pigmentation35.
The observed association of the PINK1 locus with SWP implies a role of melanocyte death in SWP regulation. PINK1 is widely accepted as a predisposition gene for early onset Parkinson disease36,37. Germline deletion of PINK1 in mice significantly impaired the mitochondrial function38. RNA interference-induced PINK1 deficiency causes mitochondrial accumulation of calcium, impaired mitochondrial respiration and ultimately cell death39. The early onset of Parkinson disease may be caused by PINK1-mediated selective death of nerve cells. We propose that the SWP phenotype in some DSE individuals is attributed to PINK1-mediated selective death of melanocytes. This speculation is supported by our observation that, in some SWP DSE pigs, the area of white coat on four legs can increase, but never decrease, as a SWP DSE pig ages (data not shown), a phenomenon that can be explained by an irreversible melanocyte death process.
Besides these well-known loci, our study identified several new candidate genes for further study of coat color phenotypic variation in pigs, although the underlying mechanism is not well understood. The putative role of these genes should be tested with functional experiments.
This study provided us an opportunity to detect genetic loci associated with the SWP phenotype in DSE pigs and enrich our understanding on pigmentation regulation in domestic pigs. The results expanded our knowledge on coat color regulation and provided additional insight on the well-addressed monogenic pigmentation regulation mechanisms and should have further implications on the conservation of the DSE pig.
Methods
All methods were performed in accordance with the guidelines approved by the Kunming Institute of Zoology, Chinese Academy of Sciences. All experimental protocols were approved by the Kunming Institute of Zoology, Chinese Academy of Sciences.
Sample and phenotype information collection
Samples and their phenotypes were collected from a purebred DSE pig population from our farm in Xishuangbanna, Yunnan Province, China. We collected DSE pig samples at day 1 after birth with a record of their coat color. All collected individuals were denoted as either black or SWP, irrespective of the moderate level of variations regarding the area, presence/absence of the white spots in some points in the SWP individuals. The occurrence of the SWP coat color was counted from 1,977 DSE pigs and 415 DSE pigs (186 SWP and 229 black) were used in this study.
Whole genome resequencing
17 SWP and 14 black DSE pigs were included in the whole genome resequencing. Genomic DNA was extracted using the phenol-chloroform method and precipitated by 75% alcohol. Genomic DNA resequencing libraries were constructed with an insert size of 500-bp, following the Illumina library protocols. 100-bp paired-end reads were generated from the resequencing libraries with a Hiseq2000 instrument (Illumina Inc.). 8 SWP samples were sequenced to a coverage depth of about 7× and the other samples to about 3×.
Variant calling and SNP annotation
The genomic reads were aligned to the pig reference genome (Sus Scrofa 10.2)16 with the BWA program40. Alignment outputs (BAM files) were sorted and PCR duplicates generated during genomic library construction were removed using SAMtools41. Genomic variants were called using the GATK42.
SNPs were annotated with the GTF file downloaded from the Ensembl website. In the annotation, a SNP was classified into protein coding (synonymous or nonsynonymous), untranslated region, intron, or intergenic according to their genomic position in the protein coding genes. To further annotate the SNPs in putative regulatory sequences, we downloaded human data from the Encyclopedia of DNA elements (ENCODE)43 project and identified putative regulatory sequences in the pig genome that are orthologous to the human counterpart with an ENCODE annotation (manuscript in preparation). Conserved noncoding sequences were identified from a 100-way whole genome alignment from 100 vertebrates that was downloaded from the UCSC genome browser (URL: http://genome.ucsc.edu). SNPs were analyzed according to their locations in the putative regulatory sequences and conserved noncoding sequences.
Identification of putative loci underlying SWP regulation
Genetic differentiation (FST) between SWP and black DSE pigs was calculated using VCFtools44, with 10-kb non-overlapping sliding windows for the autosomes. Genes in the sliding windows with FST above the chromosomal top 1% threshold were used as candidates for the SWP phenotype. Haplotypes were inferred using Beagle software45. XP-EHH was calculated to examine haplotypes that showed low levels of linkage decay in the SWP DSE pigs46,47,48. To examine inheritance from a common ancestor, identity-by-descendent (IBD) segments between any two individuals were identified using the GERMLINE program49 and the clustering of IBD segments was calculated using the DASH program50. The association between IBD clusters and the SWP phenotype was calculated using the PLINK software51.
Genotyping and association analysis
To further analyze the association of candidate loci with the SWP phenotypes in a large DSE population, the genotypes of 18 tagged SNPs was established from a total of 384 DSE pigs (215 black and 169 SWP) using the Sequenom MassArray platform (Agena Bioscience). The derived allele was deduced as alternative allele to that in the pig reference genome. The 18 SNPs were chosen as tags for 18 loci that were putatively associated with the SWP phenotypes. SNPs and primers are shown in Table 2. Associations were calculated with the PLINK software51. The fraction of phenotypic variance explained by each locus was calculated as the R2 value in a least-squares linear regression analysis with a model

where Y is the coat color phenotype, G is the fixed effects of genotyped SNPs and e is residual error.
Additional Information
How to cite this article: Lü, M.-D. et al. Genetic variations associated with six-white-point coat pigmentation in Diannan small-ear pigs. Sci. Rep. 6, 27534; doi: 10.1038/srep27534 (2016).
References
Andersson, L. & Georges, M. Domestic-animal genomics: deciphering the genetics of complex traits. Nat. Rev. Genet. 5, 202–212 (2004).
Rubin, C. J. et al. Strong signatures of selection in the domestic pig genome. Proc. Natl. Acad. Sci. USA 109, 19529–19536 (2012).
Ciobanu, D. C. et al. Genetic variation in two conserved local Romanian pig breeds using type 1 DNA markers. Genet. Sel. Evol. 33, 417–432 (2001).
Fang, M., Larson, G., Ribeiro, H. S., Li, N. & Andersson, L. Contrasting mode of evolution at a coat color locus in wild and domestic pigs. PLoS Genet. 5, e1000341 (2009).
Andersson, L. Melanocortin receptor variants with phenotypic effects in horse, pig and chicken. Ann. N. Y. Acad. Sci. 994, 313–318 (2003).
Johansson Moller, M. et al. Pigs with the dominant white coat color phenotype carry a duplication of the KIT gene encoding the mast/stem cell growth factor receptor. Mamm. Genome 7, 822–830 (1996).
Kijas, J. M. et al. Melanocortin receptor 1 (MC1R) mutations and coat color in pigs. Genetics 150, 1177–1185 (1998).
Bauer, G. L. et al. The role of MITF phosphorylation sites during coat color and eye development in mice analyzed by bacterial artificial chromosome transgene rescue. Genetics 183, 581–594 (2009).
Ren, J. et al. A 6-bp deletion in the TYRP1 gene causes the brown colouration phenotype in Chinese indigenous pigs. Heredity (Edinb) 106, 862–868 (2011).
Kijas, J. M., Moller, M., Plastow, G. & Andersson, L. A frameshift mutation in MC1R and a high frequency of somatic reversions cause black spotting in pigs. Genetics 158, 779–785 (2001).
China National Commission of Animal Genetic Resources. In Animal Genetic Resources In China: Pigs. 105–374 (China Agriculture Press, 2011).
Wilkinson, S. et al. Signatures of diversifying selection in European pig breeds. PLoS Genet. 9, e1003453 (2013).
Larson, G. et al. Worldwide phylogeography of wild boar reveals multiple centers of pig domestication. Science 307, 1618–1621 (2005).
Wu, G. S. et al. Population phylogenomic analysis of mitochondrial DNA in wild boars and domestic pigs revealed multiple domestication events in East Asia. Genome Biol. 8, R245 (2007).
Li, J. et al. Artificial selection of the melanocortin receptor 1 gene in Chinese domestic pigs during domestication. Heredity (Edinb) 105, 274–281 (2010).
Groenen, M. A. et al. Analyses of pig genomes provide insight into porcine demography and evolution. Nature 491, 393–398 (2012).
Montoliu, L., Oetting, W. S. & Bennett, D. C. Color Genes. European Society for Pigment Cell Research. (2011) Available at: http://www.espcr.org/micemut. (Accessed: 20th March 2016).
Shin, M. K., Levorse, J. M., Ingram, R. S. & Tilghman, S. M. The temporal requirement for endothelin receptor-B signalling during neural crest development. Nature 402, 496–501 (1999).
Okumura, N. et al. Sequencing, mapping and nucleotide variation of porcine coat colour genes EDNRB, MYO5A, KITLG, SLC45A2, RAB27A, SILV and MITF. Anim. Genet. 37, 80–82 (2006).
Ai, H., Huang, L. & Ren, J. Genetic diversity, linkage disequilibrium and selection signatures in chinese and Western pigs revealed by genome-wide SNP markers. PLoS One 8, e56001 (2013).
Guyonneau, L., Murisier, F., Rossier, A., Moulin, A. & Beermann, F. Melanocytes and pigmentation are affected in dopachrome tautomerase knockout mice. Mol. Cell. Biol. 24, 3396–3403 (2004).
Xie, H. B., Irwin, D. M. & Zhang, Y. P. Evolution of conserved secondary structures and their function in transcriptional regulation networks. BMC Genomics 9, 520 (2008).
Akira, S. et al. A nuclear factor for IL-6 expression (NF-IL6) is a member of a C/EBP family. Embo. J. 9, 1897–1906 (1990).
Kim, Y. et al. PINK1 controls mitochondrial localization of Parkin through direct phosphorylation. Biochem. Biophys. Res. Commun. 377, 975–980 (2008).
Spillman, W. J. Inheritance of color coat in swine. Science 24, 441–443 (1906).
Fisher, R. A. The correlation between relatives on the supposition of Mendelian inheritance. Philos. Trans. R. Soc. Edinb. 52, 399–433 (1918).
Matsushima, Y. et al. A mouse model of Waardenburg syndrome type 4 with a new spontaneous mutation of the endothelin-B receptor gene. Mamm. Genome 13, 30–35 (2002).
Gariepy, C. E., Cass, D. T. & Yanagisawa, M. Null mutation of endothelin receptor type B gene in spotting lethal rats causes aganglionic megacolon and white coat color. Proc. Natl. Acad. Sci. USA 93, 867–872 (1996).
Ceccherini, I. et al. Interstitial deletion of the endothelin-B receptor gene in the spotting lethal (sl) rat. Hum. Mol. Genet. 4, 2089–2096 (1995).
Yang, G. C. et al. A dinucleotide mutation in the endothelin-B receptor gene is associated with lethal white foal syndrome (LWFS); a horse variant of Hirschsprung disease. Hum. Mol. Genet. 7, 1047–1052 (1998).
Santschi, E. M. et al. Endothelin receptor B polymorphism associated with lethal white foal syndrome in horses. Mamm. Genome 9, 306–309 (1998).
Parichy, D. M., Rawls, J. F., Pratt, S. J., Whitfield, T. T. & Johnson, S. L. Zebrafish sparse corresponds to an orthologue of c-kit and is required for the morphogenesis of a subpopulation of melanocytes, but is not essential for hematopoiesis or primordial germ cell development. Development 126, 3425–3436 (1999).
Edery, P. et al. Mutation of the endothelin-3 gene in the Waardenburg-Hirschsprung disease (Shah-Waardenburg syndrome). Nat. Genet. 12, 442–444 (1996).
Wang, C. et al. Genome-wide analysis reveals artificial selection on coat colour and reproductive traits in Chinese domestic pigs. Mol. Ecol. Resour. 15, 414–424 (2015).
Yamada, T. et al. Reduced expression of the endothelin receptor type B gene in piebald mice caused by insertion of a retroposon-like element in intron 1. J. Biol. Chem. 281, 10799–10807 (2006).
Rohe, C. F. et al. Homozygous PINK1 C-terminus mutation causing early-onset parkinsonism. Ann. Neurol. 56, 427–431 (2004).
Chishti, M. A. et al. T313M PINK1 mutation in an extended highly consanguineous Saudi family with early-onset Parkinson disease. Arch. Neurol. 63, 1483–1485 (2006).
Gautier, C. A., Kitada, T. & Shen, J. Loss of PINK1 causes mitochondrial functional defects and increased sensitivity to oxidative stress. Proc. Natl. Acad. Sci. USA 105, 11364–11369 (2008).
Gandhi, S. et al. PINK1-associated Parkinson’s disease is caused by neuronal vulnerability to calcium-induced cell death. Mol. Cell 33, 627–638 (2009).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Li, H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27, 2987–2993 (2011).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
ENCODE Project Consortium. The ENCODE (ENCyclopedia Of DNA Elements) Project. Science 306, 636–640 (2004).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Browning, S. R. & Browning, B. L. Rapid and accurate haplotype phasing and missing-data inference for whole-genome association studies by use of localized haplotype clustering. Am. J. Hum. Genet. 81, 1084–1097 (2007).
Pickrell, J. K. et al. Signals of recent positive selection in a worldwide sample of human populations. Genome Res. 19, 826–837 (2009).
Coop, G. et al. The role of geography in human adaptation. PLoS Genet. 5, e1000500 (2009).
Pritchard, J. K., Pickrell, J. K. & Coop, G. The genetics of human adaptation: hard sweeps, soft sweeps and polygenic adaptation. Curr. Biol. 20, R208–215 (2010).
Gusev, A. et al. Whole population, genome-wide mapping of hidden relatedness. Genome Res. 19, 318–326 (2009).
Gusev, A. et al. DASH: a method for identical-by-descent haplotype mapping uncovers association with recent variation. Am. J. Hum. Genet. 88, 706–717 (2011).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (31472000), the National 973 Program of China (2013CB835203) and the Animal Branch of the Germplasm Bank of Wild Species, Chinese Academy of Sciences (the Large Research Infrastructure Funding).
Author information
Authors and Affiliations
Contributions
Y.P.Z. and H.B.X. supervised this work. Y.P.Z. and H.B.X. designed the research. M.D.L., X.M.H., Y.F.M. and H.B.X. performed data collection and data analysis. X.M.H., Y.G. and J.K.D. collected samples. H.B.X., M.D.L., X.M.H., D.M.I., A.C.A. and Y.P.Z. wrote and revised the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Lü, MD., Han, XM., Ma, YF. et al. Genetic variations associated with six-white-point coat pigmentation in Diannan small-ear pigs. Sci Rep 6, 27534 (2016). https://doi.org/10.1038/srep27534
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep27534
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.