Abstract
Mild cognitive impairment (MCI) is a precursor phase of Alzheimer’s disease (AD). As current treatments may be effective only at the early stages of AD, it is important to track MCI patients who will convert to AD. The aim of this study is to develop a high performance semi-mechanism based approach to predict the conversion from MCI to AD and improve our understanding of MCI-to-AD conversion mechanism. First, analysis of variance (ANOVA) test and lasso regression are employed to identify the markers related to the conversion. Then the Bayesian network based on selected markers is established to predict MCI-to-AD conversion. The structure of Bayesian network suggests that the conversion may start with fibrin clot formation, verbal memory impairment, eating pattern changing and hyperinsulinemia. The Bayesian network achieves a high 10-fold cross-validated prediction performance with 96% accuracy, 95% sensitivity, 65% specificity, area under the receiver operating characteristic curve of 0.82 on data from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) database. The semi-mechanism based approach provides not only high prediction performance but also clues of mechanism for MCI-to-AD conversion.
Similar content being viewed by others
Introduction
Alzheimer’s disease (AD), the most common form of dementia, is characterized by progressive neurodegenerative disorder1. 36 million people worldwide are affected by AD and the number is expected to almost triple by 20502. Many evidences indicate that AD has a years to decade preclinical period followed by a precursor phase termed as mild cognitive impairment (MCI)3. As new treatments are likely to be most effective at the early stages of AD, it is greatly urgent to track patients with MCI who will develop AD4,5.
Several sensitive imaging modalities such as structural magnetic resonance imaging (MRI) and positron emission tomography (PET) have been developed5. A number of previous researches have reported that MRI biomarkers can be used to predict the probability of conversion6,7,8. However, because some of structural changes may not be detected at visual inspection until MCI patients have converted to AD, predictions using MRI biomarkers only may not be accurate enough for application in the routine clinical setting or clinical drug trials3,5. Previous researches show that combined markers such as MRI and cerebrospinal fluid (CSF) biomarkers can improve the prediction accuracy5,9. But CSF sample collection requires lumbar puncture which is too invasive to be used as a routine clinical examination. As damage to the blood-brain barrier may occur in AD, this may increase movement of proteins between the brain and the blood10. It is therefore possible that AD and its precursor, MCI, may be associated with the variation of biomarkers detectable in plasma11. Recent work has demonstrated the possibility of predicting MCI-to-AD conversion based on plasma markers12. In addition, blood sample is more accessible and suitable for repeated collecting. These make plasma-based biomarkers promising for prediction of conversion from MCI to AD.
While the highly sensitive markers are beneficial on the conversion prediction, advanced machine learning methods can further improve the reliability of approaches. Machine learning is the study of algorithms and computational techniques that use previous examples in the form of multivariate datasets to help make future predictions13. A number of machine learning methods such as support vector machines (SVM) and logistic regression (LR) have been used to predict the conversion from MCI to AD5,8. Compared with the traditional data-driven machine learning methods, Bayesian network has unique advantages that it can quantify the causal relationships between the markers, visualize these relationships by the structure of network, and conduct the prediction task based on the causal relationships14. These attractive characteristics make Bayesian network a semi-mechanism method. On one hand, the semi-mechanism nature of Bayesian network can improve our understanding of conversion mechanism. On the other hand, because of the complex etiology and multiple pathogenesis of AD, the conversion from MCI to AD is affected by many uncertain factors which makes its prediction a complicated issue15. Bayesian network is especially well-suited to handle the intricacies of the prediction because it is designed for representing stochastic events and conducting prediction tasks under uncertainty16,17.
Lots of lectures based on data-driven methods, such as neural network with self-organizing maps (SOM), are focused on improving the classification performance and they have showed good performance in the diagnosis task. However, the contribution of these methods on improving our understanding of MCI-to-AD conversion mechanism is limited. As the semi-mechanism nature of Bayesian network can provide causal relationships of markers, this paper proposes a semi-mechanism method based on the combination of Bayesian network and lasso regression for not only the high performance of MCI-to-AD conversion prediction but also improving our understanding the mechanism of the conversion. The data from Alzheimer’s Disease Neuroimaging Initiative (ADNI) is used to develop the model. However, ADNI contains more than 500 biomarkers (including MRI markers and plasma markers), many of which may not relate to MCI-to-AD conversion. Irrelevant biomarkers may interfere the causal relationships identification and reduce the performance of prediction method. Therefore, biomarkers selection should be performed before the conversion prediction. In this study, lasso regression is proposed to conduct the markers selection, which combines variable selection with an efficient computational procedure18. Previous works have shown that lasso regression can enhance the prediction performance of models based on high dimension data sets19,20,21. As such, the combination of Bayesian network and lasso regression is proposed not only to conduct the prediction task but also to improve understanding of the AD-to-MCI conversion mechanism. Moreover, after the conversion probability is calculated, a subgroup analysis is performed for comparing the network disruption of high-risk patients and low-risk patients.
Results
Biomarkers selection
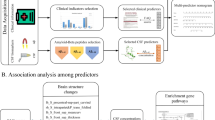
In this section, the process of biomarkers selection is described. The dataset used in this study contains 518 biomarkers (328 MRI markers and 190 plasma markers). 45 biomarkers (1 MRI marker and 44 plasma markers) are deleted during data checking due to too many missing entries. 75 biomarkers (57 MRI markers and 18 plasma markers) with significant difference between converters and non-converters are identified by ANOVA test. 34 biomarkers (25 MRI markers and 9 plasma markers) related to Alzheimer’s disease assessment scale (ADAS-cog) are selected by lasso regression. 7 biomarkers (5 MRI markers and 2 plasma markers) are eliminated during Bayesian network structure learning because they fail to connect to the Bayesian network. In addition, as 2 MRI markers are labeled as “unknown”, they are also eliminated. Finally, 25 biomarkers (18 MRI markers and 7 plasma markers) are selected for conversion prediction. The process of biomarkers identification is summarized in Fig. 1. The list of selected biomarkers is shown in Table 1.
Structure and performance of Bayesian network
In this section, we present the results of Bayesian structure learning and the performance of conversion prediction. The Bayesian network structure obtained by max-min hill-climbing (MMHC) is given in Fig. 2. It contains 26 nodes and 43 arcs.
It contains 26 nodes and 43 arcs. The nodes in order are: ST109TS, ST111CV, ST114TA, ST11SV, ST121TA, ST30SV, ST31TA, ST40CV, ST49TA, ST52CV, ST56CV, ST70SV, ST72CV, ST83CV, ST83TA, ST88SV, ST91CV, ST99CV, AGRP, C-peptide, CRP, FGF-4, Fibrinogen, Insulin, MMP-10, and “Whether patients converts to AD or not”.
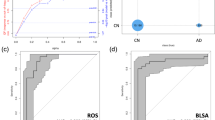
In order to evaluate the performance of Bayesian network, a 10-fold cross-validation is performed to estimate its accuracy, sensitivity and specificity. Furthermore, the performance of Bayesian network is compared to the performances of linear discriminant analysis (LDA) and SOM. The performances of all these methods are evaluated by 10-fold cross-validation. The results are given in Fig. 3. The Fig. 3A shows that the accuracy and sensitivity of Bayesian network are higher than those of LDA and SOM with markers selection. In Fig. 3B, the area under receiver operating characteristic curve (AUC-ROC) of Bayesian network is much higher than that of LDA and SOM with marker selection. Moreover, to evaluate the performance of markers selection, we apply SOM and Bayesian network with or without markers selection and compare their performances. With markers selection, the classification performances of both SOM and Bayesian network are improved.
(A) The receiver operating characteristic (ROC) curve of Linear discriminant analysis (LDA), self-organizing map (SOM) (with or without markers selection) and Bayesian network (with or without markers selection). (B) The performance of LDA, SOM (with or without markers selection) and Bayesian network (with or without selection) measured by three parameters: accuracy, sensitivity, specificity. All these parameters are evaluated by 10-fold cross-validation.
Network disruption profile
According to the result of Bayesian network, a group of highest conversion probability patients (high-risk group, n = 11) and a group of lowest conversion probability patients (low-risk group, n = 48) are drawn from the dataset. 11 biomarkers have significant difference (P < 0.05, ANOVA test) between high-risk group and low risk group. The mini network balance map Fig. 4A) shows that the high-risk group may suffer from more severe network disruption than the low risk group. The network disruption parameters coincide with the mini network balance map. Parameters U and  increase significantly in high risk group (P < 0.01, ANOVA test, shown in Fig. 4B) which may suggest that patients with greater U and
increase significantly in high risk group (P < 0.01, ANOVA test, shown in Fig. 4B) which may suggest that patients with greater U and  may have higher conversion risk.
may have higher conversion risk.
(A) Network disruption analysis of markers with significant difference between high-risk group and low-risk group. In normal state, the shape of radar graph is a regular polygon. With the shape deformation, the difference from normal state gets greater. (B) Box plot of parameters U, K, and  . If the value of disruption parameters U and
. If the value of disruption parameters U and  is beyond the horizontal lines in figures, the patient may have more conversion risk. *P < 0.05, **P < 0.01 vs low risk group.
is beyond the horizontal lines in figures, the patient may have more conversion risk. *P < 0.05, **P < 0.01 vs low risk group.
Discussion
In this study, we propose a semi-mechanism based Bayesian network to predict the conversion from MCI to AD. The proposed method has two contributions. Firstly, the proposed approach achieves relative high prediction performance. Secondly, as the Bayesian network can learn the causal relationships among biomarkers from the database, these causal relationships can provide some more insight into the mechanism of MCI-to-AD conversion.
The proposed model is compared to previous researches based on data-driven methods (Table 2). Comparing with LDA and SOM, Bayesian network has higher accuracy and sensitivity with markers selection. The high sensitivity of Bayesian network may lie in two points. On one hand, the semi-mechanism nature of Bayesian network may provide higher performance because it can learn causal relationships from data and combine these knowledge and data to conduct the prediction task22. On the other hand, plasma markers may be highly sensitive in conversion prediction12. Though the data-driven methods also achieved high performance, the Bayesian network still has its unique advantage. The structure of Bayesian network may contain the causal relationships of markers which makes it a semi-mechanism method and provide more information beyond the performance of classification.
In addition, with markers selection, the classification performance of Bayesian network is improved. It suggests that Bayesian network should work with an appropriate marker selection strategy. In another words, without markers selection, Bayesian network may produce false positive causal relationships which may not only decrease the performance but also mislead the MCI-to-AD conversion mechanism investigation. Therefore, combining Bayesian network and lasso marker selection strategy is very helpful in improving understanding the conversion mechanism and classification performance.
The semi-mechanism nature of Bayesian network is beneficial on investigating the mechanism of conversion. Structure of Bayesian network shows that 6 markers including volume of left middle temporal, cortical thickness average of right entorhinal, volume of right inferior temporal, AGRP, c-peptide, and fibrinogen may be related to the conversion directly. Our result, that destruction of entorhinal is associated with MCI-to-AD conversion, is consistent with previous research23. Previous researches had also reported the variations of temporal, c-peptide level, and fibrinogen level in AD patients19,24,25. However, our results suggest that these changes may have happened at MCI stage. It indicates that the conversion from MCI to AD may start with destruction of temporal, entorhnal, increased level of AGRP, c-peptide, and fibrin25.
Bayesian network identifies the variations of above six markers caused by MCI-to-AD conversion directly. But some of these changings may not be the key factors in the conversion. Therefore a reanalysis based on the results of Bayesian network is performed to identify the major factors. The subgroups network disruption profile suggests that the progress MCI patient may suffer from more severe network disruption than stable MCI patients. Network disruption may be related to the marker panel including 11 markers. The six markers identified by Bayesian network and marker panel related network disruption share three markers: Cortical Thickness of Entorhinal, Volume of Temporal and AGRP. These three markers may be the key factors in the conversion. In other word, they might be attributed to the warming signals of conversion. Previous researches showed that the destruction of entorhinal and temporal is associated with verbal memory impairment and verbal memory impairment might be the warming indicator of MCI-to-AD conversion26. In addition, clinical researches have reported that AD patients have greater preference for high-fat and sweet food than normal groups. However, our results suggested that such change in eating pattern may have happened at MCI stage27,28, as the elevated level of AGRP, an orexigenic peptide, in high-risk patients may increase the preference for a high fat diet29.
The crosstalk between cerebral destruction and plasma markers alteration revealed by Bayesian network can provide more clues for the mechanism of conversion. The crosstalk between C-peptide and cerebral destruction may play a vital role in the MCI-to-AD conversion. C-peptide is a measure of insulin secretion. Elevated C-peptide level represents high peripheral insulin secretion. It is reported that high peripheral insulin secretion can increase the risk of AD. Because high level peripheral insulin secretion impairs amyloid clearance by inhibiting brain insulin production which is a beneficial effect on amyloid clearance30. Bayesian network suggests that C-peptide may be related to the destruction of middle temporal, entorhinal, and inferior temporal. It suggests that amyloid may mainly aggregate in the above three regions at the MCI stage which may aggravate their damage. As all above three regions are involved in verbal memory, high level of C-peptide may impair to verbal memory which was confirmed by previous works26,31,32,33.
In summary, the analysis of Bayesian network shows that the conversion from MCI to AD may start with multiple pathological changes such as verbal memory impairment, vascular abnormalities, hyperinsulinemia and eating pattern change. In this study, a high performance semi-mechanism based approach is developed to predict the conversion from MCI to AD by combining MRI and plasma markers. The semi-mechanism based approach provides not only high performance prediction but also more insight into the mechanism of conversion from MCI to AD.
Subject and Method
Subject
Patients
In this study, the following criteria are used to select subjects for model developing:
-
Patients with baseline MRI scan records
-
Patients with baseline plasma-based biomarker data
-
Patients with baseline ADAS-cog scores
-
Patients with MCI due to Alzheimer’s disease
-
Patients with diagnosis records which can be used to determine whether they convert from MCI to AD in 18 months
Finally, a data set with complete imaging, plasma-based biomarkers, ADAS data is drawn from ADNI including 316 MCI patients (99 converters and 217 non-converters). The demographic information of subjects is given in Table 3.
Imaging biomarkers
Imaging data in this study is obtained from dataset UCSF—Cross-Sectional FreeSurfer (FreeSurfer Version 4.3). The dataset is available at https://ida.loni.usc.edu/pages/access/studyData.jsp. In this dataset, all scans were acquired on 1.5 T MRI scanners. The imaging data were processed and analyzed with FreeSurfer 4.3 by the UCSF team. The dataset includes 328 MRI biomarkers which can be grouped into 5 categories: average cortical thickness, standard deviation in cortical thickness, the volumes of cortical parcellations (based on regions of interest automatically segmented in the cortex), the volumes of specific white matter parcellations, and the total surface area of the cortex. Details of the analysis procedure are available at http://adni.loni.ucla.edu/research/mripost-processing/.
Plasma-based biomarkers
The plasma-based biomarker data is obtained from dataset Biomarkers Consortium Plasma Proteomics Project RBM multiplex data. The data is available at https://ida.loni.usc.edu/pages/access/studyData.jsp. The data was acquired by analyzing a subset of plasma samples from the ADNI cohort in a 190 analyte multiplex immunoassay panel. The panel, referred to as the human discovery map, was developed on the Luminex xMAP platform by Rules-Based Medicine (RBM) to contain proteins previously reported in the literature to be altered as a result of cancer, cardiovascular disease, metabolic disorders and inflammation. Details of the assay technology and validation has been described elsewhere (http://adni.loni.ucla.edu/wp-content/uploads/2010/11/BC_Plasma_Proteomics_Data_Primer.pdf).
Method
Considering that ADNI contains more than 500 biomarkers, it is essential to select the more predictive biomarkers to obtain a parsimonious model and avoid the classifier suffering overfitting. Then the Bayesian network is established based on the causal relationships among selected markers for predicting the AD-to-MCI conversion. Finally, a reanalysis of Bayesian network results is performed to profile the network disruption of the patients with highest probability of converting to AD and those with lowest probability. The framework is summarized in Fig. 5.
Biomarkers selection
Biomarkers selection includes two stages. At the first stage, ANOVA test is employed to screen biomarkers with significant difference (P < 0.05) between converters and non-converters. At the second stage, lasso regression is used to filter biomarkers related to ADAS-cog from the selected biomarkers at the first stage.
Lasso regression is a popular technique for feature selection which can continuously shrinks coefficients34. It drops biomarkers by shrinking some of coefficients to zero. In this study, a Least Angle Regression (LARS) algorithm is used to solve lasso35.
Bayesian network
Considering that the causal relationships among the selected markers may remain unknown, a Bayesian network structure learning algorithm termed as the max-min hill-climbing (MMHC) is employed to learn the causal relationships among the selected markers. MMHC algorithm is a hybrid method, using concepts and techniques from both constraint-based approaches and score-based approaches, which can achieve high quality in structure learning36. After the Bayesian network is learned from data, the most popular Bayesian network inference algorithm named junction tree is employed to acquire the conversion prediction37.
Model evaluation
In this study, the receiver operating characteristic (ROC) curve is used to evaluate the performance of Bayesian network. The ROC, which has become established as an important tool for classifier evaluation, is a graph of true positive rate (TPR) against false positive rate (FPR) at various operating points as a decision threshold38. The area under the ROC curve (AUC) is a measure of predictive ability39. Moreover, three parameters termed as accuracy (number of correctly classified samples divided by the total number of samples), sensitivity (the number of correctly classified converters divided by the total number of converters) and specificity (the number of correctly classified non-converters divided by the total number of non-converters) are calculated and evaluated by 10-fold cross-validation for a further measurement for the model performance40.
Network disruption analysis
To get more insight into the mechanism of the conversion, a reanalysis of Bayesian network results is performed using a mathematic method for evaluating the disruption of biology network which was proposed in our previous research41. In this study, subjects are divided into two subgroups high risk group and low risk group according to the results of Bayesian network and a mini network balance model is developed to evaluate the network disruption for both high-risk group and low-risk group. The network disruption comparison between these two subgroups may provide more insight into the mechanism of AD-to-MCI conversion.
The mini network balance model contains three parameters U, K, and  . U is response to both consistency variation and inconsistency variation comprehensively. K responds to multi-marker consistency variation.
. U is response to both consistency variation and inconsistency variation comprehensively. K responds to multi-marker consistency variation.  is response to the multi-marker inconsistency variation. These three parameters can be calculated as below:
is response to the multi-marker inconsistency variation. These three parameters can be calculated as below:



Let  be the state vector of patients with conversion risk and
be the state vector of patients with conversion risk and  be the state vector of normal control group.
be the state vector of normal control group.
Additional Information
How to cite this article: Liu, H. et al. A semi-mechanism approach based on MRI and proteomics for prediction of conversion from mild cognitive impairment to Alzheimer's disease. Sci. Rep. 6, 26712; doi: 10.1038/srep26712 (2016).
References
Trzepacz, P. T. et al. Comparison of neuroimaging modalities for the prediction of conversion from mild cognitive impairment to Alzheimer’s dementia. Neurobiology of aging 35, 143–151, doi: 10.1016/j.neurobiolaging.2013.06.018 (2014).
Barnes, D. E. & Yaffe, K. The projected effect of risk factor reduction on Alzheimer’s disease prevalence. The Lancet Neurology 10, 819–828, doi: http://dx.doi.org/10.1016/S1474-4422 (11)70072-2 (2011).
Petrella, J. R., Coleman, R. E. & Doraiswamy, P. M. Neuroimaging and early diagnosis of Alzheimer disease: A look to the future. Radiology 226, 315–336, doi: 10.1148/radiol.2262011600 (2003).
Drago, V. et al. Disease tracking markers for Alzheimer’s disease at the prodromal (MCI) stage. Journal of Alzheimer’s disease JAD 26 Suppl 3, 159–199, doi: 10.3233/JAD-2011-0043 (2011).
Shaffer, J. L. et al. Predicting cognitive decline in subjects at risk for Alzheimer disease by using combined cerebrospinal fluid, MR imaging, and PET biomarkers. Radiology 266, 583–591 (2013).
Cho, Y., Seong, J.-K., Jeong, Y. & Shin, S. Y. Individual subject classification for Alzheimer’s disease based on incremental learning using a spatial frequency representation of cortical thickness data. NeuroImage 59, 2217–2230, doi: http://dx.doi.org/10.1016/j.neuroimage.2011.09.085 (2012).
Coupé, P., Eskildsen, S. F., Manjón, J. V., Fonov, V. S. & Collins, D. L. Simultaneous segmentation and grading of anatomical structures for patient’s classification: Application to Alzheimer’s disease. NeuroImage 59, 3736–3747, doi: http://dx.doi.org/10.1016/j.neuroimage.2011.10.080 (2012).
Wolz, R. et al. Multi-method analysis of MRI images in early diagnostics of Alzheimer’s disease. PloS one 6, e25446 (2011).
Eckerstrom, C. et al. Multimodal Prediction of Dementia with up to 10 Years Follow Up: The Gothenburg MCI Study. Journal Of Alzheimers Disease 44, 205–214, doi: 10.3233/jad-141053 (2015).
Hye, A. et al. Proteome-based plasma biomarkers for Alzheimer’s disease. Brain 129, 3042–3050, doi: 10.1093/brain/awl279 (2006).
Jayasena, T. et al. Upregulation of glycolytic enzymes, mitochondrial dysfunction and increased cytotoxicity in glial cells treated with Alzheimer’s disease plasma. PLoS One 10, e0116092, doi: 10.1371/journal.pone.0116092 (2015).
Hye, A. et al. Plasma proteins predict conversion to dementia from prodromal disease. Alzheimer’s & Dementia 10, 799–807.e792, doi: 10.1016/j.jalz.2014.05.1749 (2014).
Challis, E. et al. Gaussian process classification of Alzheimer’s disease and mild cognitive impairment from resting-state fMRI. NeuroImage 112, 232–243, doi: 10.1016/j.neuroimage.2015.02.037 (2015).
Xu, B. G. Intelligent fault inference for rotating flexible rotors using Bayesian belief network. Expert Systems with Applications 39, 816–822, doi: 10.1016/j.eswa.2011.07.079 (2012).
Gomez-Ramirez, J. & Wu, J. Network-based biomarkers in Alzheimer’s disease: review and future directions. Frontiers in aging neuroscience 6, 12, doi: 10.3389/fnagi.2014.00012 (2014).
Bandyopadhyay, S. et al. Data mining for censored time-to-event data: a Bayesian network model for predicting cardiovascular risk from electronic health record data. Data Mining and Knowledge Discovery 29, 1033–1069, doi: 10.1007/s10618-014-0386-6 (2014).
Wang, K. J., Makond, B. & Wang, K. M. Modeling and predicting the occurrence of brain metastasis from lung cancer by Bayesian network: a case study of Taiwan. Computers in biology and medicine 47, 147–160, doi: 10.1016/j.compbiomed.2014.02.002 (2014).
Meinshausen, N. Relaxed Lasso. Computational Statistics & Data Analysis 52, 374–393, doi: 10.1016/j.csda.2006.12.019 (2007).
Zou, H. The adaptive lasso and its oracle properties. Journal of the American statistical association 101, 1418–1429 (2006).
Li, Z. & Sillanpaa, M. J. Overview of LASSO-related penalized regression methods for quantitative trait mapping and genomic selection. TAG. Theoretical and applied genetics. Theoretische und angewandte Genetik 125, 419–435, doi: 10.1007/s00122-012-1892-9 (2012).
Ogutu, J. O. & Piepho, H.-P. Regularized group regression methods for genomic prediction: Bridge, MCP, SCAD, group bridge, group lasso, sparse group lasso, group MCP and group SCAD. BMC Proceedings 8, 1–9, doi: 10.1186/1753-6561-8-s5-s7 (2014).
Needham, C. J., Bradford, J. R., Bulpitt, A. J., Care, M. A. & Westhead, D. R. Predicting the effect of missense mutations on protein function: analysis with Bayesian networks. BMC bioinformatics 7, doi: 10.1186/1471-2105-7-405 (2006).
Devanand, D. P. et al. MRI hippocampal and entorhinal cortex mapping in predicting conversion to Alzheimer’s disease. NeuroImage 60, 1622–1629, doi: 10.1016/j.neuroimage.2012.01.075 (2012).
Reiman, E. M. & Jagust, W. J. Brain imaging in the study of Alzheimer’s disease. NeuroImage 61, 505–516, doi: 10.1016/j.neuroimage.2011.11.075 (2012).
Cortes-Canteli, M., Mattei, L., Richards, A. T., Norris, E. H. & Strickland, S. Fibrin deposited in the Alzheimer’s disease brain promotes neuronal degeneration. Neurobiology of aging 36, 608–617, doi: 10.1016/j.neurobiolaging.2014.10.030 (2015).
Goto, M. et al. Entorhinal cortex volume measured with 3T MRI is positively correlated with the Wechsler Memory Scale-Revised logical/verbal memory score for healthy subjects. Neuroradiology 53, 617–622, doi: 10.1007/s00234-011-0863-1 (2011).
Mungas, D. et al. Dietary preference for sweet foods in patients with dementia. Journal of the American Geriatrics Society 38, 999–1007 (1990).
Cullen, P., Abid, F., Patel, A., Coope, B. & Ballard, C. Eating disorders in dementia. International journal of geriatric psychiatry 12, 559–562 (1997).
Barnes, M. J., Argyropoulos, G. & Bray, G. A. Preference for a high fat diet, but not hyperphagia following activation of mu opioid receptors is blocked in AgRP knockout mice. Brain research 1317, 100–107, doi: 10.1016/j.brainres.2009.12.051 (2010).
Luchsinger, J. A. & Gustafson, D. R. Adiposity, type 2 diabetes and Alzheimer’s disease. Journal of Alzheimer’s disease: JAD 16, 693 (2009).
Groussard, M. et al. Musical and verbal semantic memory: Two distinct neural networks ? NeuroImage 49, 2764–2773, doi: 10.1016/j.neuroimage.2009.10.039 (2010).
Shinoura, N. et al. Right temporal lobe plays a role in verbal memory. Neurological research 33, 734–738, doi: 10.1179/1743132811Y.0000000005 (2011).
Okereke, O. I. et al. Plasma C-peptide levels and rates of cognitive decline in older, community-dwelling women without diabetes. Psychoneuroendocrinology 33, 455–461, doi: 10.1016/j.psyneuen.2008.01.002 (2008).
Tibshirani, R. Regression shrinkage and selection via the lasso. Journal of the Royal Statistical Society. Series B (Methodological), 267–288 (1996).
Efron, B., Hastie, T., Johnstone, I. & Tibshirani, R. Least angle regression. The Annals of statistics 32, 407–499 (2004).
Tsamardinos, I., Brown, L. E. & Aliferis, C. F. The max-min hill-climbing Bayesian network structure learning algorithm. Machine Learning 65, 31–78, doi: 10.1007/s10994-006-6889-7 (2006).
Lauritzen, S. L. & Spiegelhalter, D. J. Local computations with probabilities on graphical structures and their application to expert systems. Journal of the Royal Statistical Society . Series B (Methodological), 157–224 (1988).
Bradley, A. P. ROC curve equivalence using the Kolmogorov–Smirnov test. Pattern Recognition Letters 34, 470–475, doi: 10.1016/j.patrec.2012.12.021 (2013).
Pinsky, P. F. Scaling of true and apparent ROC AUC with number of observations and number of variables. Communications In Statistics-Simulation And Computation 34, 771–781, doi: 10.1081/sac-200068366 (2005).
Moradi, E. et al. Machine learning framework for early MRI-based Alzheimer’s conversion prediction in MCI subjects. NeuroImage 104, 398–412, doi: 10.1016/j.neuroimage.2014.10.002 (2015).
Liu, H., Wei, C., He, H. & Liu, X. Evaluating Alzheimer’s disease progression by modeling crosstalk network disruption. Frontiers in Neuroscience 9, doi: 10.3389/fnins.2015.00523 (2015).
Casanova, R. et al. Alzheimer’s Disease Risk Assessment Using Large-Scale Machine Learning Methods. Plos One 8, doi: 10.1371/journal.pone.0077949 (2013).
Cheng, B., Liu, M., Zhang, D., Munsell, B. C. & Shen, D. Domain Transfer Learning for MCI Conversion Prediction. IEEE transactions on bio-medical engineering 62, 1805–1817, doi: 10.1109/TBME.2015.2404809 (2015).
Yu, G., Liu, Y., Thung, K. H. & Shen, D. Multi-task linear programming discriminant analysis for the identification of progressive MCI individuals. PLoS One 9, e96458, doi: 10.1371/journal.pone.0096458 (2014).
Acknowledgements
This study was supported by the National Natural Science Foundation of the People’s Republic of China. (Nos 81273588 and 81473274). Data collection and sharing for this project was funded by the Alzheimer’s Disease Neuroimaging Initiative (ADNI) (National Institutes of Health Grant U01 AG024904) and DOD ADNI (Department of Defense award number W81XWH-12-2-0012). ADNI is funded by the National Institute on Aging, the National Institute of Biomedical Imaging and Bioengineering, and through generous contributions from the following: Alzheimer’s Association; Alzheimer’s Drug Discovery Foundation; Araclon Biotech; BioClinica, Inc.; Biogen Idec Inc.; Bristol-Myers Squibb Company; Eisai Inc.; Elan Pharmaceuticals, Inc.; Eli Lilly and Company; EuroImmun; F. Hoffmann-La Roche Ltd and its affiliated company Genentech, Inc.; Fujirebio; GE Healthcare; IXICO Ltd.; Janssen Alzheimer Immunotherapy Research & Development, LLC.; Johnson & Johnson Pharmaceutical Research & Development LLC.; Medpace, Inc.; Merck & Co., Inc.; Meso Scale Diagnostics, LLC.; NeuroRx Research; Neurotrack Technologies; Novartis Pharmaceuticals Corporation; Pfizer Inc.; Piramal Imaging; Servier; Synarc Inc.; and Takeda Pharmaceutical Company. The Canadian Institutes of Rev December 5, 2013 Health Research is providing funds to support ADNI clinical sites in Canada. Private sector contributions are facilitated by the Foundation for the National Institutes of Health (www.fnih.org). The grantee organization is the Northern California Institute for Research and Education, and the study is coordinated by the Alzheimer’s Disease Cooperative Study at the University of California, San Diego. ADNI data are disseminated by the Laboratory for Neuro Imaging at the University of Southern California. A complete listing of ADNI investigators can be found at: http://adni.loni.usc.edu/wpcontent/uploads/how_to_apply/ADNI_Acknowledgement_List.pdf.
Author information
Authors and Affiliations
Consortia
Contributions
H.L., X.Z., H.J., H.H. and X.L. developed the model. H.L. do the computational work. H.L. wrote the manuscript. The data used in this manuscript is obtained from Alzheimer’s Disease Neuroimaging Initiative (ADNI). Database (adni.loni.usc.edu). As such, the investigators within the ADNI contributed to the design and implementation of ADNI and/or provided data but did not participate in analysis or writing of this report.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Liu, H., Zhou, X., Jiang, H. et al. A semi-mechanism approach based on MRI and proteomics for prediction of conversion from mild cognitive impairment to Alzheimer’s disease. Sci Rep 6, 26712 (2016). https://doi.org/10.1038/srep26712
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep26712
This article is cited by
-
Neuroimaging and analytical methods for studying the pathways from mild cognitive impairment to Alzheimer’s disease: protocol for a rapid systematic review
Systematic Reviews (2020)
-
Hybrid High-order Functional Connectivity Networks Using Resting-state Functional MRI for Mild Cognitive Impairment Diagnosis
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.